
Rice Science ›› 2023, Vol. 30 ›› Issue (5): 364-368.DOI: 10.1016/j.rsci.2023.04.001
收稿日期:2022-11-04
接受日期:2023-04-28
出版日期:2023-09-28
发布日期:2023-08-10
. [J]. Rice Science, 2023, 30(5): 364-368.
| Trait | Locus | Chr | Start (Mb) | End (Mb) | R2 (%) | P-value | |||
|---|---|---|---|---|---|---|---|---|---|
| Minimum | Maximum | Minimum | Maximum | ||||||
| CD2017, CR2017, TD2020 | locus1-1 | 1 | 7.6 | 8.8 | 4.9 | 7.2 | 6.67E-05 | 8.95E-04 | |
| CD2018, CD2020, CR2017, CR2018, CR2020, TD2020 | locus1-2 | 1 | 9.3 | 12.8 | 5.0 | 8.2 | 2.23E-05 | 8.96E-04 | |
| GL2017, GL2018 | locus1-3 | 1 | 24.7 | 24.8 | 5.2 | 5.9 | 3.13E-04 | 6.82E-04 | |
| CD2017, CD2018, CR2017, GW2020 | locus2-1 | 2 | 2.1 | 2.9 | 5.0 | 7.8 | 3.53E-05 | 8.44E-04 | |
| CD2017, GL2020 | locus2-2 | 2 | 9.8 | 11.6 | 5.1 | 3.8 | 7.64E-04 | 7.49E-04 | |
| CR2017 | locus2-3 | 2 | 15.3 | 15.3 | 5.5 | 5.5 | 4.85E-04 | 4.85E-04 | |
| CR2017, GW2017, GW2018, GW2020 | locus2-4 | 2 | 28.8 | 31.6 | 3.6 | 5.2 | 6.36E-04 | 9.46E-04 | |
| CD2017, GLW2020, TD2018 | locus2-5 | 2 | 33.4 | 35.7 | 4.8 | 6.1 | 2.20E-04 | 9.81E-04 | |
| GL2017, GL2018, GL2020, GLW2017, GLW2018, GLW2020 | locus3-1 | 3 | 15.3 | 17.7 | 4.8 | 27.7 | 1.02E-13 | 9.24E-04 | |
| GL2017, GL2018, GL2020, GLW2018, GLW2020, GW2020 | locus3-2 | 3 | 20.7 | 24.7 | 4.9 | 11.3 | 8.51E-07 | 9.21E-04 | |
| CD2018, CR2017, CR2018 | locus3-3 | 3 | 27.7 | 28.0 | 4.9 | 6.6 | 1.26E-04 | 9.93E-04 | |
| CD2017, CR2017, TD2017, TD2018, GL2018 | locus4-1 | 4 | 16.0 | 19.4 | 5.0 | 7.6 | 4.25E-05 | 9.03E-04 | |
| TD2017, TD2020 | locus4-2 | 4 | 21.4 | 21.6 | 3.8 | 5.2 | 1.03E-04 | 8.60E-04 | |
| CD2020 | locus4-3 | 4 | 34.7 | 34.7 | 5.0 | 5.0 | 1.16E-04 | 1.16E-04 | |
| GW2017, GW2018, GW2020 | locus5-1 | 5 | 4.3 | 4.8 | 5.0 | 5.3 | 4.90E-04 | 7.31E-04 | |
| TD2017, TD2018 | locus6-1 | 6 | 5.6 | 7.1 | 4.9 | 9.0 | 8.56E-06 | 9.19E-04 | |
| GW2020 | locus6-2 | 6 | 28.0 | 28.0 | 4.0 | 4.0 | 4.89E-04 | 4.89E-04 | |
| CD2018, CR2018, CR2020, GL2017, GL2018, GL2020, GLW2017, GLW2018, GLW2020, GW2017, GW2018, GW2020 | locus7-1 | 7 | 23.7 | 25.0 | 4.8 | 12.8 | 1.45E-07 | 9.94E-04 | |
| GLW2020, GW2017, GW2020 | locus8-1 | 8 | 0.6 | 1.7 | 4.9 | 4.0 | 4.85E-04 | 8.32E-04 | |
| CD2020, CR2020, TD2020 | locus8-2 | 8 | 4.9 | 5.7 | 3.6 | 4.1 | 4.97E-04 | 9.82E-04 | |
| GLW2020, GW2017, GW2018, GW2020 | locus8-3 | 8 | 26.3 | 27.9 | 3.6 | 8.4 | 1.45E-05 | 9.23E-04 | |
| CR2020 | locus9-1 | 9 | 6.3 | 6.9 | 3.7 | 4.7 | 1.81E-04 | 9.27E-04 | |
| TD2017, CD2018 | locus11-1 | 11 | 21.2 | 23.7 | 5.0 | 6.8 | 1.20E-04 | 8.15E-04 | |
| GW2020 | locus12-1 | 12 | 25.2 | 25.2 | 3.7 | 3.7 | 8.62E-04 | 8.62E-04 | |
Table 1. Loci for milled grain appearance traits in Bandillo indica multiparent advanced generation intercross population by genome-wide association study.
| Trait | Locus | Chr | Start (Mb) | End (Mb) | R2 (%) | P-value | |||
|---|---|---|---|---|---|---|---|---|---|
| Minimum | Maximum | Minimum | Maximum | ||||||
| CD2017, CR2017, TD2020 | locus1-1 | 1 | 7.6 | 8.8 | 4.9 | 7.2 | 6.67E-05 | 8.95E-04 | |
| CD2018, CD2020, CR2017, CR2018, CR2020, TD2020 | locus1-2 | 1 | 9.3 | 12.8 | 5.0 | 8.2 | 2.23E-05 | 8.96E-04 | |
| GL2017, GL2018 | locus1-3 | 1 | 24.7 | 24.8 | 5.2 | 5.9 | 3.13E-04 | 6.82E-04 | |
| CD2017, CD2018, CR2017, GW2020 | locus2-1 | 2 | 2.1 | 2.9 | 5.0 | 7.8 | 3.53E-05 | 8.44E-04 | |
| CD2017, GL2020 | locus2-2 | 2 | 9.8 | 11.6 | 5.1 | 3.8 | 7.64E-04 | 7.49E-04 | |
| CR2017 | locus2-3 | 2 | 15.3 | 15.3 | 5.5 | 5.5 | 4.85E-04 | 4.85E-04 | |
| CR2017, GW2017, GW2018, GW2020 | locus2-4 | 2 | 28.8 | 31.6 | 3.6 | 5.2 | 6.36E-04 | 9.46E-04 | |
| CD2017, GLW2020, TD2018 | locus2-5 | 2 | 33.4 | 35.7 | 4.8 | 6.1 | 2.20E-04 | 9.81E-04 | |
| GL2017, GL2018, GL2020, GLW2017, GLW2018, GLW2020 | locus3-1 | 3 | 15.3 | 17.7 | 4.8 | 27.7 | 1.02E-13 | 9.24E-04 | |
| GL2017, GL2018, GL2020, GLW2018, GLW2020, GW2020 | locus3-2 | 3 | 20.7 | 24.7 | 4.9 | 11.3 | 8.51E-07 | 9.21E-04 | |
| CD2018, CR2017, CR2018 | locus3-3 | 3 | 27.7 | 28.0 | 4.9 | 6.6 | 1.26E-04 | 9.93E-04 | |
| CD2017, CR2017, TD2017, TD2018, GL2018 | locus4-1 | 4 | 16.0 | 19.4 | 5.0 | 7.6 | 4.25E-05 | 9.03E-04 | |
| TD2017, TD2020 | locus4-2 | 4 | 21.4 | 21.6 | 3.8 | 5.2 | 1.03E-04 | 8.60E-04 | |
| CD2020 | locus4-3 | 4 | 34.7 | 34.7 | 5.0 | 5.0 | 1.16E-04 | 1.16E-04 | |
| GW2017, GW2018, GW2020 | locus5-1 | 5 | 4.3 | 4.8 | 5.0 | 5.3 | 4.90E-04 | 7.31E-04 | |
| TD2017, TD2018 | locus6-1 | 6 | 5.6 | 7.1 | 4.9 | 9.0 | 8.56E-06 | 9.19E-04 | |
| GW2020 | locus6-2 | 6 | 28.0 | 28.0 | 4.0 | 4.0 | 4.89E-04 | 4.89E-04 | |
| CD2018, CR2018, CR2020, GL2017, GL2018, GL2020, GLW2017, GLW2018, GLW2020, GW2017, GW2018, GW2020 | locus7-1 | 7 | 23.7 | 25.0 | 4.8 | 12.8 | 1.45E-07 | 9.94E-04 | |
| GLW2020, GW2017, GW2020 | locus8-1 | 8 | 0.6 | 1.7 | 4.9 | 4.0 | 4.85E-04 | 8.32E-04 | |
| CD2020, CR2020, TD2020 | locus8-2 | 8 | 4.9 | 5.7 | 3.6 | 4.1 | 4.97E-04 | 9.82E-04 | |
| GLW2020, GW2017, GW2018, GW2020 | locus8-3 | 8 | 26.3 | 27.9 | 3.6 | 8.4 | 1.45E-05 | 9.23E-04 | |
| CR2020 | locus9-1 | 9 | 6.3 | 6.9 | 3.7 | 4.7 | 1.81E-04 | 9.27E-04 | |
| TD2017, CD2018 | locus11-1 | 11 | 21.2 | 23.7 | 5.0 | 6.8 | 1.20E-04 | 8.15E-04 | |
| GW2020 | locus12-1 | 12 | 25.2 | 25.2 | 3.7 | 3.7 | 8.62E-04 | 8.62E-04 | |
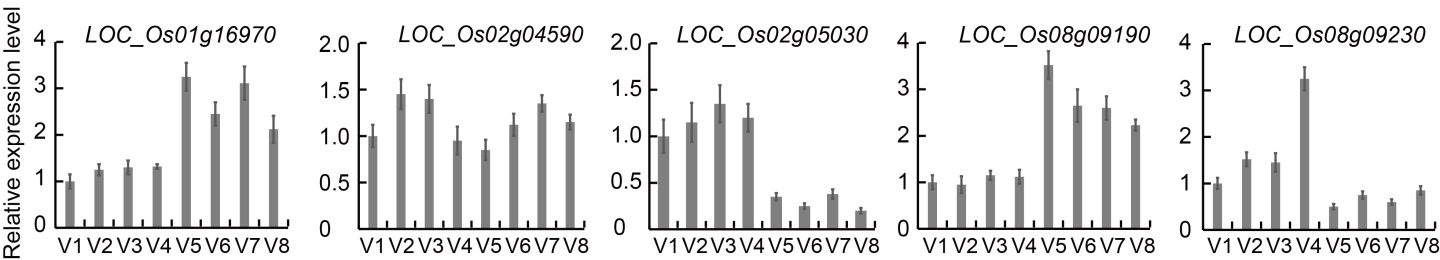
Fig. 1. Quantitative real-time PCR for candidate genes related to grain appearance traits in eight parents. V1, Fedearroz 50; V2, Shanhuangzhan 2; V3, IR64633-87-2-2-3-3; V4, IR4630-22-2-5-1-3; V5, IR45427-2B-2-2B-1-1; V6, IR84196-12-32; V7, IR77298-14-1-2-10; V8, IR77186-122-2-2-3.
| [1] | Ayaad M, Han Z M, Zheng K, Hu G, Abo-Yousef M, Sobeih S E S, Xing Y Z. 2020. Bin-based genome-wide association studies reveal superior alleles for improvement of appearance quality using a 4-way MAGIC population in rice. J Adv Res, 28: 183-194. |
| [2] |
Bai S, Yu H, Wang B, Li J. 2018. Retrospective and perspective of rice breeding in China. J Genet Genomics, 45(11): 603-612.
PMID |
| [3] |
Bandillo N, Raghavan C, Muyco P A, Sevilla M A L, Lobina I T, Dilla-Ermita C J, Tung C W, McCouch S, Thomson M, Mauleon R, Singh R K, Gregorio G, Redoña E, Leung H. 2013. Multi- parent advanced generation inter-cross (MAGIC) populations in rice: Progress and potential for genetics research and breeding. Rice, 6(1): 11.
PMID |
| [4] |
Barnaby J Y, Huggins T D, Lee H, McClung A M, Pinson S R M, Oh M, Bauchan G R, Tarpley L, Lee K J, Kim M, Edwards J D. 2020. Vis/NIR hyperspectral imaging distinguishes sub-population, production environment, and physicochemical grain properties in rice. Sci Rep, 10(1): 9284.
PMID |
| [5] | Choi B S, Kim Y J, Markkandan K, Koo Y J, Song J T, Seo H S. 2018. GW2 functions as an E3 ubiquitin ligase for rice expansin-like 1. Int J Mol Sci, 19(7): 1904. |
| [6] |
Descalsota G I L, Swamy B P M, Zaw H, Inabangan-Asilo M A, Amparado A, Mauleon R, Chadha-Mohanty P, Arocena E C, Raghavan C, Leung H, Hernandez J E, Lalusin A B, Mendioro M S, Diaz M G Q, Reinke R. 2018. Genome-wide association mapping in a rice MAGIC plus population detects QTLs and genes useful for biofortification. Front Plant Sci, 9: 1347.
PMID |
| [7] | Duan P G, Xu J S, Zeng D L, Zhang B L, Geng M F, Zhang G Z, Huang K, Huang L J, Xu R, Ge S, Qian Q, Li Y H. 2017. Natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol Plant, 10(5): 685-694. |
| [8] | Fan C, Xing Y, Mao H, Lu T, Han B, Xu C, Li X, Zhang Q. 2006. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrance protein. Theor Appl Genet, 112: 1164-1171. |
| [9] | Hou M M, Luo F F, Wu D X, Zhang X H, Lou M M, Shen D F, Yan M, Mao C Z, Fan X R, Xu G H, Zhang Y L. 2021. OsPIN9, an auxin efflux carrier, is required for the regulation of rice tiller bud outgrowth by ammonium. New Phytol, 229(2): 935-949. |
| [10] | Liu C, Song J L, Wang Y C, Huang X R, Zhang F, Wang W S, Xu J L, Zhang Y, Yu H X, Pang Y H, Bao J S. 2020. Rapid prediction of head rice yield and grain shape for genome-wide association study in indica rice. J Cereal Sci, 96: 103091. |
| [11] | Liu J D, He Z H, Rasheed A, Wen W E, Yan J, Zhang P Z, Wan Y X, Zhang Y, Xie C J, Xia X C. 2017. Genome-wide association mapping of black point reaction in common wheat (Triticum aestivum L.). BMC Plant Biol, 17(1): 220. |
| [12] | Ponce K S, Ye G Y, Zhao X Q. 2018. QTL identification for cooking and eating quality in indica rice using multi-parent advanced generation intercross (MAGIC) population. Front Plant Sci, 9: 868. |
| [13] | Ponce K S, Meng L J, Guo L B, Leng Y J, Ye G Y. 2021. Advances in sensing, response and regulation mechanism of salt tolerance in rice. Int J Mol Sci, 22(5): 2254. |
| [14] | Ruan B P, Shang L G, Zhang B, Hu J, Wang Y X, Lin H, Zhang A P, Liu C L, Peng Y L, Zhu L, Ren D Y, Shen L, Dong G J, Zhang G H, Zeng D L, Guo L B, Qian Q, Gao Z Y. 2020. Natural variation in the promoter of TGW2 determines grain width and weight in rice. New Phytol, 227(2): 629-640. |
| [15] | Shi C L, Dong N Q, Guo T, Ye W W, Shan J X, Lin H X. 2020. A quantitative trait locus GW6 controls rice grain size and yield through the gibberellin pathway. Plant J, 103(3): 1174-1188. |
| [16] |
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M. 2008. Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet, 40(8): 1023-1028.
PMID |
| [17] |
Song X J, Kuroha T, Ayano M, Furuta T, Nagai K, Komeda N, Segami S, Miura K, Ogawa D, Kamura T, Suzuki T, Higashiyama T, Yamasaki M, Mori H, Inukai Y, Wu J Z H, Kitano H, Sakakibara H, Jacobsen S E, Ashikari M. 2015. Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield, and plant biomass in rice. Proc Natl Acad Sci USA, 112(1): 76-81.
PMID |
| [18] | Tang Z B, Gao X Y, Zhan X Y, Fang N Y, Wang R Q, Zhan C F, Zhang J Q, Cai G, Cheng J P, Bao Y M, Zhang H S, Huang J. 2021. Natural variation in OsGASR7 regulates grain length in rice. Plant Biotechnol J, 19(1): 14-16. |
| [19] | Tatiana R, Julie D, Philippe L, Julien F, Isabelle R R, Noronirina V R, Tuong-Vi C, Kirsten V B, Alain R, Nourollah A, Louis-Marie R. 2021. Genome-wide association study of nitrogen use efficiency and agronomic traits in upland rice. Rice Sci, 28(4): 379-390. |
| [20] |
Wang D W, Sun W Q, Yuan Z Y, Sun Q, Fan K, Zhang C P, Yu S B. 2021. Identification of a novel QTL and candidate gene associated with grain size using chromosome segment substitution lines in rice. Sci Rep, 11(1): 189.
PMID |
| [21] | Wang S, Wu K, Yuan Q, Liu X, Liu Z, Lin X, Zeng R, Zhu H, Dong G, Qian Q, Zhang G, Fu X. 2012. Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet, 44(8): 950-954. |
| [22] |
Wu W G, Liu X Y, Wang M H, Meyer R S, Luo X J, Ndjiondjop M N, Tan L B, Zhang J W, Wu J Z, Cai H W, Sun C Q, Wang X K, Wing R A, Zhu Z F. 2017. A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication. Nat Plants, 3: 17064.
PMID |
| [23] | Ying J Z, Ma M, Bai C, Huang X H, Liu J L, Fan Y Y, Song X J. 2018. TGW3, a major QTL that negatively modulates grain length and weight in rice. Mol Plant, 11(5): 750-753. |
| [24] | You H, Zhang O L, Xu L, Liang C, Xiang X C. 2020. Effects of soluble starch synthase lla allelic variation on rice grain quality with different Waxy backgrounds. J Sci Food Agric, 100(15): 5344-5351. |
| [25] | Zhao D S, Li Q F, Zhang C Q, Zhang C, Yang Q Q, Pan L X, Ren X Y, Lu J, Gu M H, Liu Q Q. 2018. GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat Commun, 9(1): 1240. |
| [26] | Zhou H, Yun P, He Y Q. 2019. Rice appearance quality. In: Bao J S. Rice: Chemistry and Technology 4 edn. AACC International Press: 371-383. |
| [27] | Zhou Y, Miao J, Gu H Y, Peng X R, Leburu M, Yuan F H, Gu H W, Gao Y, Tao Y J, Zhu J Y, Gong Z Y, Yi C D, Gu M H, Yang Z F, Liang G H. 2015. Natural variations in SLG7 regulate grain shape in rice. Genetics, 201(4): 1591-1599. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||