
Rice Science ›› 2024, Vol. 31 ›› Issue (1): 77-86.DOI: 10.1016/j.rsci.2023.11.007
收稿日期:2023-07-07
接受日期:2023-11-27
出版日期:2024-01-28
发布日期:2024-02-06
. [J]. Rice Science, 2024, 31(1): 77-86.
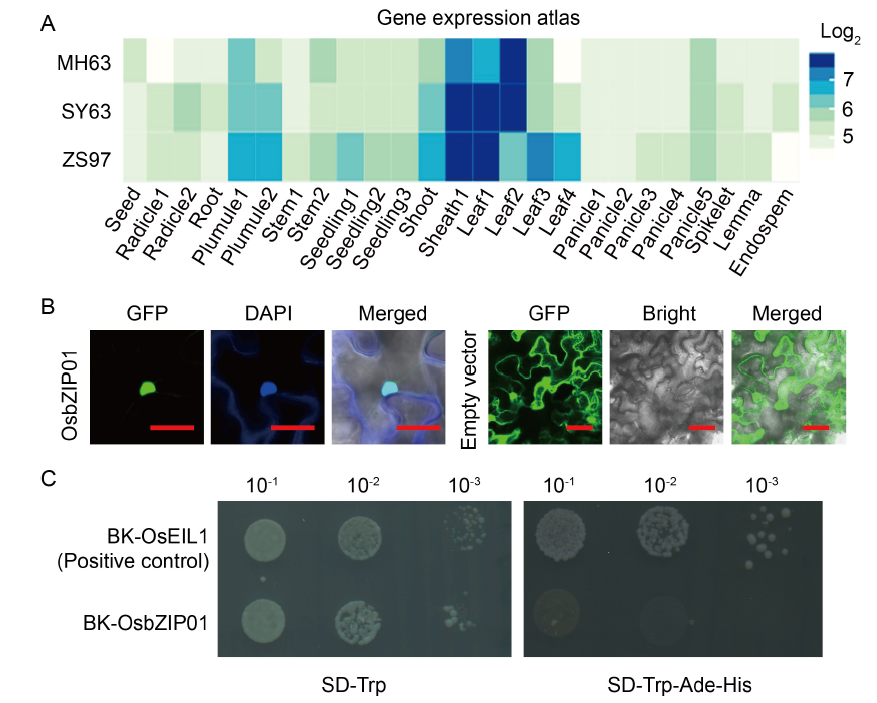
Fig. 1. Expression pattern of OsbZIP01 and its protein characterization. A, Gene expression atlas of OsbZIP01 in three representative indica varieties of China is from the CREP database (http://crep.ncpgr.cn/crep-cgi/home.pl). The expression value is log2-transformed. MH63, Minghui 63; SY63, Shanyou 63; ZS97, Zhenshan 97. B, Subcellular localization of OsbZIP01. DAPI, 4′,6-diamidino-2-phenylindole; GFP, Green fluorescent protein. Scale bars are 100 μm. C, Transactivation assay of OsbZIP01 in yeast cells. BK, pDEST-GBKT7; SD, Synthetic dropout medium; Trp, Tryptophan; Ade, Adenine; His, Histidine.
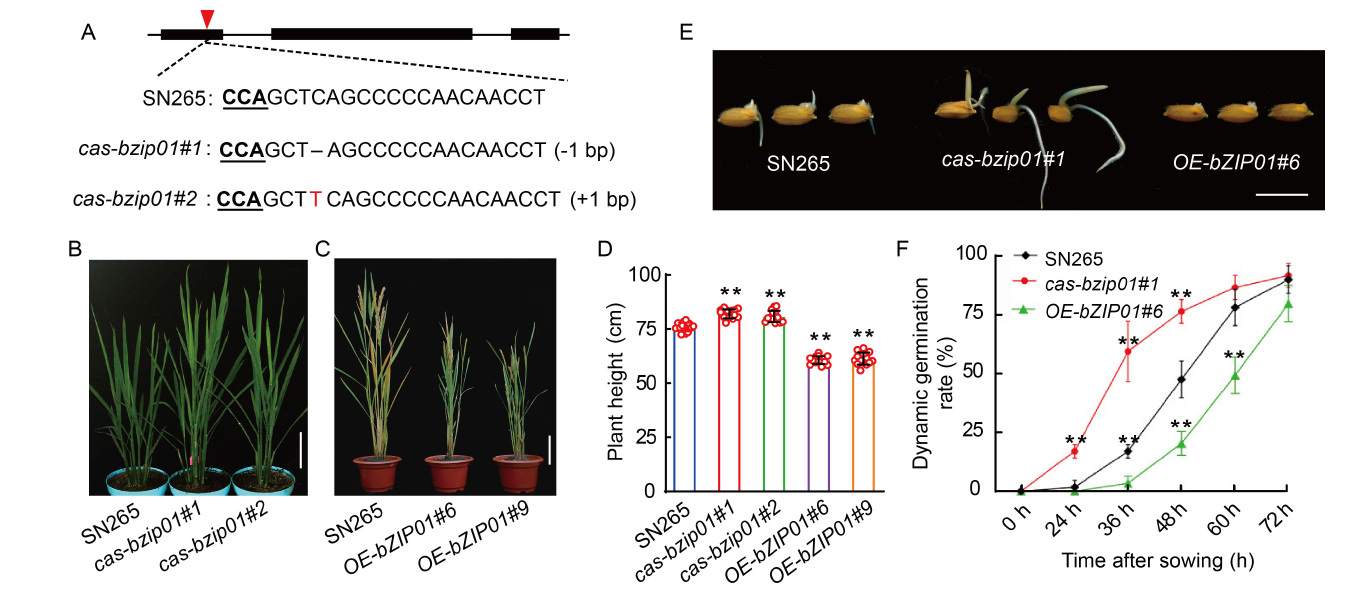
Fig. 2. Phenotypic characterization of Shennong 265 (SN265), cas-bzip01, and OE-bZIP01 transgenic plants. A, Schematic diagram of the sgRNA site of bZIP01 gene in SN265 and two mutants (cas-bzip01#1 and cas-bzip01#2) by CRISPR/Cas9 system. The red triangle indicates the sgRNA position. B, Phenotypes of SN265 and cas-bzip01 plants (cas-bzip01#1 and cas-bzip01#2) at the heading stage. Scale bar is 10 cm. C, Morphology of SN265 and overexpression lines (OE-bZIP01#6 and OE-bZIP01#9) at the maturity stage. Scale bar is 10 cm. D, Plant heights of SN265 and transgenic plants (cas-bzip01#1, cas-bzip01#2, OE-bZIP01#6, and OE-bZIP01#9) at the maturity stage. E, Seed germination phenotypes of SN265, cas-bzip01#1, and OE-bZIP01#6 plants. Sterile grains were sown on Murashige and Skoog plates, and photographs were taken at 48 h after germination at 30 ºC. Scale bar is 1 cm. F, Dynamic germination rates of SN265, cas-bzip01#1, and OE-bZIP01#6 after sown for 72 h. In D and F, Data are Mean ± SD of three independent experiments (n = 10 for each experiment). ** indicates the significant differences between transgenic plants and wild type (SN265) at the P < 0.01 by the Student’s t-test.
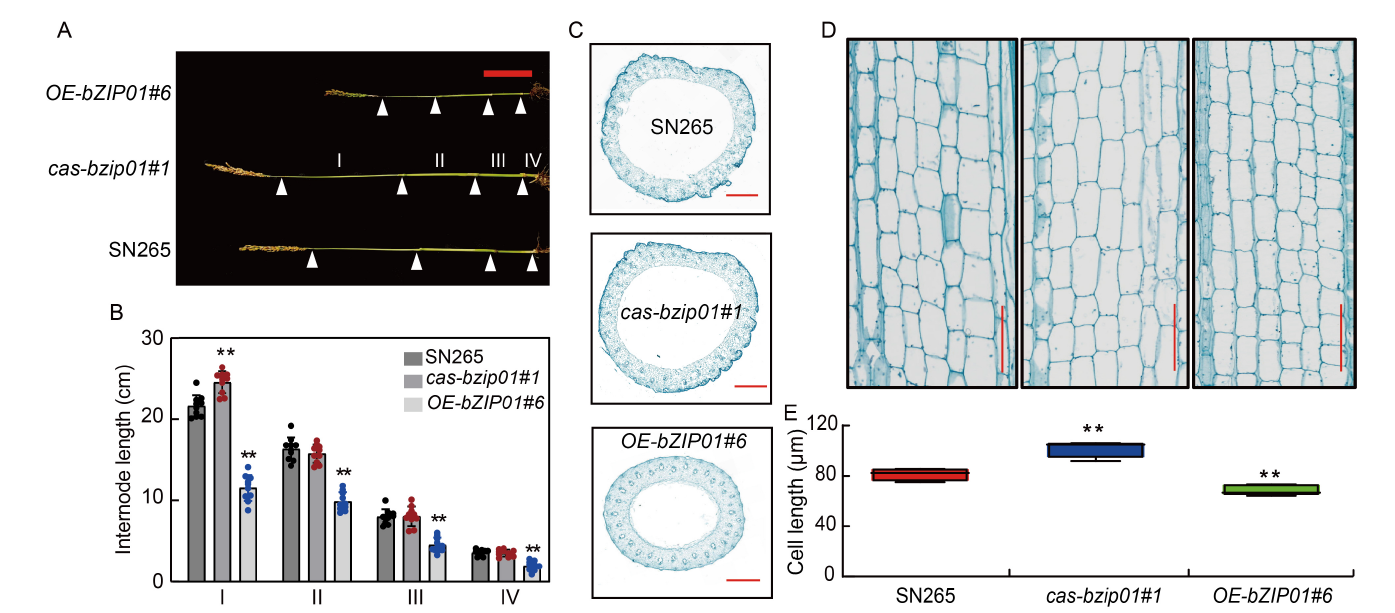
Fig. 3. Morphology comparison of cas-bzip01 and OE-bZIP01 transgenic plants. A and B, Comparison of internode lengths between wild type (Shennong 265, SN265), cas-bzip01#1, and OE-bZIP01#6 at the heading stage. Ⅰ, The first internode from the top; Ⅱ, The second internode from the top; Ⅲ, The third internode from the top; Ⅳ, The fourth internode from the top. Scale bar is 10 cm. C, Fast green-stained transverse sections of SN265, cas-bzip01#1, and OE-bZIP01#6 stems, showing differences in the diameter of the second internode from the top. Scale bars are 1 mm. D, Longitudinal sections of the first internode from the top of SN265, cas-bzip01#1, and OE-bZIP01#6. Scale bars are 120 μm. E, Comparison analysis of parenchyma cell lengths of the first internode from the top of SN265, cas-bzip01#1, and OE-bZIP01#6. In B and F, data are Mean ± SD of three independent experiments (n = 10 for each experiment). ** indicates the significant differences between transgenic plants and wild type at the P < 0.01 by the Student’s t-test.
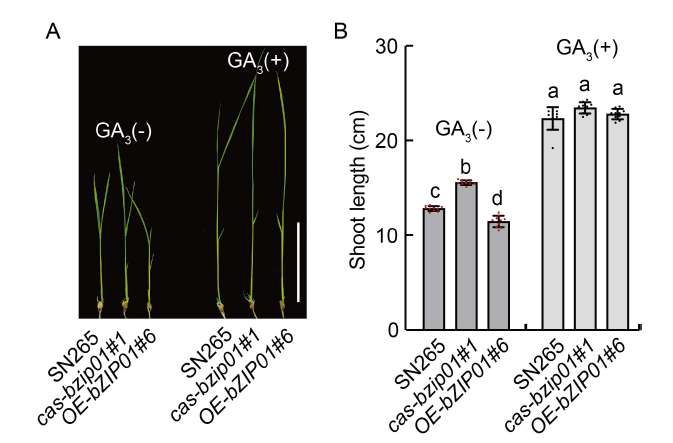
Fig. 4. Phenotype response of wild type (Shennong 265, SN265), cas-bzip01, and OE-OsbZIP01 to exogenous GA. A, Seedling phenotype of SN265 and transgenic lines (cas-bzip01#1 and OE-bZIP01#6), which were treated with exogenous 10 μmol/L GA3 for 10 d. GA3(-) represents the absence of GA3, and GA3(+) represents the application of GA3. Scale bar is 5 cm. B, Shoot lengths of SN265 and transgenic lines (cas-bzip01#1 and OE-bZIP01#6). Data are Mean ± SD of three independent experiments (n = 10 for each experiment). Different lowercase letters indicate the significant differences between transgenic plants and wild type (P < 0.01, Student’s t-test).
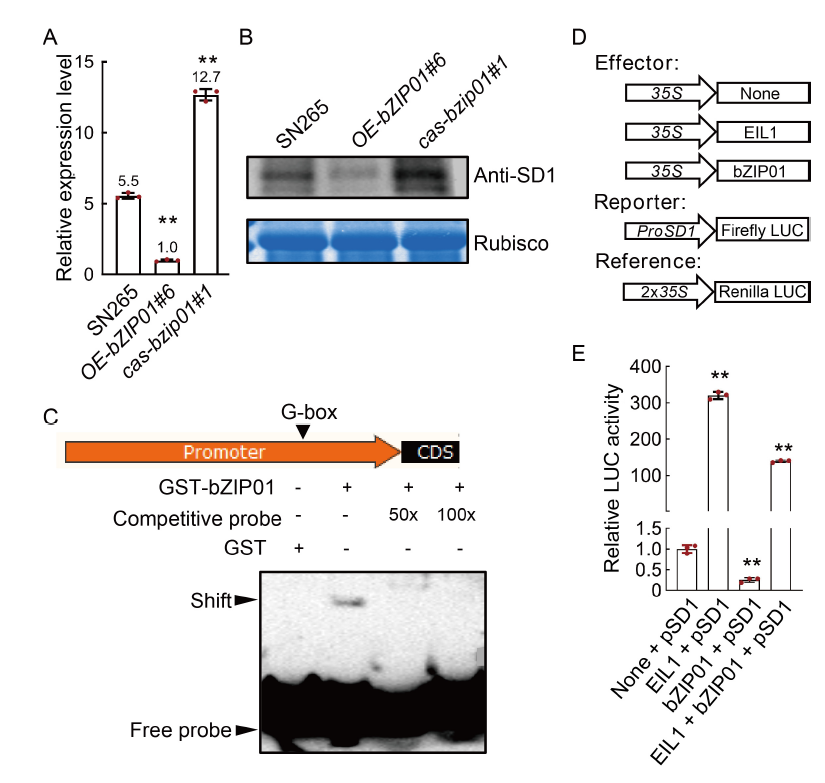
Fig. 5. OsbZIP01 directly binds to promoter region of SD1. A and B, Relative expression levels of mRNA (A) and protein (B) of SD1 in the stems of wild type (Shennong 265, SN265), cas-bzip01#1, and OE-bZIP01#6 transgenic plants. C, OsbZIP01 binds in vitro to G-box in the SD1 promoter by electrophoretic mobility shift assay. For the assay, the radiolabeled probes were incubated with OsbZIP01 protein. Competitive (unlabeled) probe (50× and 100×), G-box probe were used as indicated. GST, Glutathione S-transferase. CDS, Coding sequence. D, Schematic diagrams of the effector and reporter constructs used the dual-luciferase reporter assays. E, Transactivity assay in rice proplasts. EIL1 protein positively regulates SD1 expression, which is used as the positive control. LUC, Luciferase. Data in A and E are Mean ± SD of three independent experiments (n = 10 for each experiment). ** indicates the significant differences between transgenic plants and wild type at the P < 0.01 by the Student’s t-test.
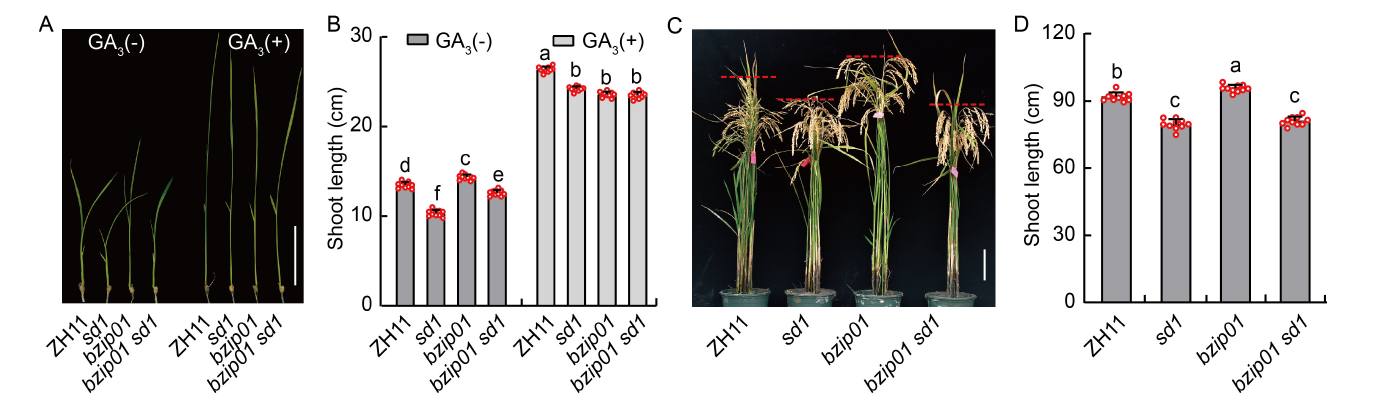
Fig. 6. Genetic relationship of OsbZIP01 and OsSD1. A and B, Shoot lengths of the single (sd1 and bzip01) and double (bzip01 sd1) mutant seedlings under Zhonghua 11 (ZH11) background with and without 10 μmol/L GA3 treatment [GA3(+) and GA3(-)] for 10 d. C and D, Phenotypic characterization (C) and shoot lengths (D) of the single (sd1 and bzip01) and double (bzip01 sd1) mutants at the maturity stage. In A and C, scale bars are 5 cm. In B and D, data are Mean ± SD of three independent experiments (n = 10 for each experiment), and different lowercase letters represent significant differences at the P < 0.05 by the Student’s t-test.
| [1] | Asano K, Yamasaki M, Takuno S, Miura K, Katagiri S, Ito T, Doi K, Wu J Z, Ebana K, Matsumoto T, Innan H, Kitano H, Ashikari M, Matsuoka M. 2011. Artificial selection for a green revolution gene during japonica rice domestication. Proc Natl Acad Sci USA, 108(27): 11034-11039. |
| [2] | Beyene G, Chauhan R D, Villmer J, Husic N, Wang N, Gebre E, Girma D, Chanyalew S, Assefa K, Tabor G, Gehan M, McGrone M, Yang M Z, Lenderts B, Schwartz C, Gao H R, Gordon- Kamm W, Taylor N J, MacKenzie D J. 2022. CRISPR/Cas9- mediated tetra-allelic mutation of the ‘Green Revolution’ SEMIDWARF-1 (SD-1) gene confers lodging resistance in tef (Eragrostis tef). Plant Biotechnol J, 20(9): 1716-1729. |
| [3] | Bhatnagar A, Burman N, Sharma E, Tyagi A, Khurana P, Khurana J P. 2023. Two splice forms of OsbZIP1, a homolog of AtHY5, function to regulate skotomorphogenesis and photomorphogenesis in rice. Plant Physiol, 193(1): 426-447. |
| [4] | Burman N, Bhatnagar A, Khurana J P. 2018. OsbZIP48, a HY5 transcription factor ortholog, exerts pleiotropic effects in light- regulated development. Plant Physiol, 176(2): 1262-1285. |
| [5] | Chen X, Lu S C, Wang Y F, Zhang X, Lv B, Luo L Q, Xi D D, Shen J B, Ma H, Ming F. 2015. OsNAC2 encoding a NAC transcription factor that affects plant height through mediating the gibberellic acid pathway in rice. Plant J, 82(2): 302-314. |
| [6] | Cho S H, Kang K, Lee S H, Lee I J, Paek N C. 2016. OsWOX3A is involved in negative feedback regulation of the gibberellic acid biosynthetic pathway in rice (Oryza sativa). J Exp Bot, 67(6): 1677-1687. |
| [7] | Chu Y L, Xu N, Wu Q, Yu B, Li X X, Chen R R, Huang J L. 2019. Rice transcription factor OsMADS57 regulates plant height by modulating gibberellin catabolism. Rice, 12(1): 38. |
| [8] | Gao S P, Chu C C. 2020. Gibberellin metabolism and signaling: Targets for improving agronomic performance of crops. Plant Cell Physiol, 61(11): 1902-1911. |
| [9] | Gao W W, Li M K, Yang S G, Gao C Z, Su Y, Zeng X, Jiao Z L, Xu W J, Zhang M Y, Xia K F. 2022. miR2105 and the kinase OsSAPK 10 co-regulate OsbZIP86 to mediate drought-induced ABA biosynthesis in rice. Plant Physiol, 189(2): 889-905. |
| [10] | Hasegawa T, Lucob-Agustin N, Yasufuku K, Kojima T, Nishiuchi S, Ogawa A, Takahashi-Nosaka M, Kano-Nakata M, Inari-Ikeda M, Sato M, Tsuji H, Wainaina C M, Yamauchi A, Inukai Y. 2021. Mutation of OUR1/OsbZIP1, which encodes a member of the basic leucine zipper transcription factor family, promotes root development in rice through repressing auxin signaling. Plant Sci, 306: 110861. |
| [11] | Hedden P. 2001. Gibberellin metabolism and its regulation. J Plant Growth Regul, 20(4): 317-318. |
| [12] | Hedden P, Phillips A L. 2000. Gibberellin metabolism: New insights revealed by the genes. Trends Plant Sci, 5(12): 523-530. |
| [13] | Im J H, Lee S G, Lee E, Park S R, Ahn I, Hwang D J. 2019. OsbZIP75 positively regulates plant defense against the bacterial leaf blight pathogen Xanthomonas oryzae pv. oryzae. Plant Biotechnol Rep, 13(6): 645-651. |
| [14] | Kuroha T, Nagai K, Gamuyao R, Wang D R, Furuta T, Nakamori M, Kitaoka T, Adachi K, Minami A, Mori Y, Mashiguchi K, Seto Y, Yamaguchi S, Kojima M, Sakakibara H, Wu J Z, Ebana K, Mitsuda N, Ohme-Takagi M, Yanagisawa S, Yamasaki M, Yokoyama R, Nishitani K, Mochizuki T, Tamiya G, McCouch S R, Ashikari M. 2018. Ethylene-gibberellin signaling underlies adaptation of rice to periodic flooding. Science, 361: 181-186. |
| [15] | Li J T, Zhao Y, Chu H W, Wang L K, Fu Y R, Liu P, Upadhyaya N, Chen C L, Mou T M, Feng Y Q, Kumar P, Xu J. 2015. SHOEBOX modulates root meristem size in rice through dose-dependent effects of gibberellins on cell elongation and proliferation. PLoS Genet, 11(8): e1005464. |
| [16] | Liu C, Zheng S, Gui J S, Fu C J, Yu H S, Song D L, Shen J H, Qin P, Liu X M, Han B, Yang Y Z, Li L G. 2018. Shortened basal internodes encodes a gibberellin 2-oxidase and contributes to lodging resistance in rice. Mol Plant, 11(2): 288-299. |
| [17] | Liu F, Wang P D, Zhang X B, Li X F, Yan X H, Fu D H, Wu G. 2018. The genetic and molecular basis of crop height based on a rice model. Planta, 247(1): 1-26. |
| [18] | Lu S J, Wei H, Wang Y, Wang H M, Yang R F, Zhang X B, Tu J M. 2012. Overexpression of a transcription factor OsMADS15 modifies plant architecture and flowering time in rice (Oryza sativa L.). Plant Mol Biol Rep, 30(6): 1461-1469. |
| [19] | Ma H Z, Liu C, Li ZX, Ran Q J, Xie G N, Wang B M, Fang S, Chu J F, Zhang J R. 2018. ZmbZIP4 contributes to stress resistance in maize by regulating ABA synthesis and root development. Plant Physiol, 178(2): 753-770. |
| [20] | Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M S, Sato M, Nasu S, Minobe Y. 2002. Positional cloning of rice semidwarfing gene, sd-1: Rice ‘green revolution gene’ encodes a mutant enzyme involved in gibberellin synthesis. DNA Res, 9(1): 11-17. |
| [21] | Nijhawan A, Jain M, Tyagi A K, Khurana J P. 2008. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol, 146(2): 323-324. |
| [22] | Oikawa T, Koshioka M, Kojima K, Yoshida H, Kawata M. 2004. A role of OsGA20ox1, encoding an isoform of gibberellin 20-oxidase, for regulation of plant stature in rice. Plant Mol Biol, 55: 687-700. |
| [23] | Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush G S, Kitano H, Matsuoka M. 2002. Green revolution: A mutant gibberellin-synthesis gene in rice. Nature, 416: 701-702. |
| [24] | Silverstone A L, Sun T P. 2000. Gibberellins and the green revolution. Trends Plant Sci, 5(1): 1-2. |
| [25] | Spielmeyer W, Ellis M H, Chandler P M. 2002. Semidwarf (sd-1), green revolution rice, contains a defective gibberellin 20-oxidase gene. Proc Natl Acad Sci USA, 99(13): 9043-9048. |
| [26] | Su S, Hong J, Chen X F, Zhang C Q, Chen M J, Luo Z J, Chang S W, Bai S X, Liang W Q, Liu Q Q, Zhang D B. 2021. Gibberellins orchestrate panicle architecture mediated by DELLA-KNOX signalling in rice. Plant Biotechnol J, 19(11): 2304-2318. |
| [27] | Tang L Q, Xu H Y, Wang Y F, Wang H M, Li Z Y, Liu X X, Shu Y Z, Li G, Liu W N, Ying J Z, Tong X H, Yao J L, Xiao W F, Tang S Q, Ni S, Zhang J. 2021. OsABF1 represses gibberellin biosynthesis to regulate plant height and seed germination in rice (Oryza sativa L.). Int J Mol Sci, 22(22): 12220. |
| [28] | Wang Q, Lin Q B, Wu T, Duan E C, Huang Y S, Yang C Y, Mou C L, Lan J, Zhou C L, Xie K, Liu X, Zhang X, Guo X P, Wang J, Jiang L, Wan J M. 2020. OsDOG1L-3 regulates seed dormancy through the abscisic acid pathway in rice. Plant Sci, 298: 110570. |
| [29] | Wang Y, Li J. 2008. Molecular basis of plant architecture. Annu Rev Plant Biol, 59: 253-279. |
| [30] | Yamaguchi S. 2008. Gibberellin metabolism and its regulation. Annu Rev Plant Biol, 59: 225-251. |
| [31] | Yang C, Ma Y M, Li J X. 2016. The rice YABBY4 gene regulates plant growth and development through modulating the gibberellin pathway. J Exp Bot, 67(18): 5545-5556. |
| [32] | Yang X C, Hwa C M. 2008. Genetic modification of plant architecture and variety improvement in rice. Heredity, 101(5): 396-404. |
| [33] | Ye H, Feng J H, Zhang L H, Zhang J F, Mispan M S, Cao Z Q, Beighley D H, Yang J C, Gu X Y. 2015. Map-based cloning of seed dormancy1-2 identified a gibberellin synthesis gene regulating the development of endosperm-imposed dormancy in rice. Plant Physiol, 169(3): 2152-2165. |
| [34] | Yin X M, Liu X, Xu B X, Lu P Y, Dong T, Yang D, Ye T T, Feng Y Q, Wu Y. 2019. OsMADS18, a membrane-bound MADS-box transcription factor, modulates plant architecture and the abscisic acid response in rice. J Exp Bot, 70(15): 3895-3909. |
| [35] | Zhang W, Cochet F, Ponnaiah M, Lebreton S, Matheron L, Pionneau C, Boudsocq M, Resentini F, Huguet S, Blázquez M Á, Bailly C, Puyaubert J, Baudouin E. 2019. The MPK8-TCP14 pathway promotes seed germination in Arabidopsis. Plant J, 100(4): 677-692. |
| [36] | Zhang Y, Su J B, Duan S, Ao Y, Dai J R, Liu J, Wang P, Li Y G, Liu B, Feng D R, Wang J F, Wang H B. 2011. A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods, 7(1): 30. |
| [37] | Zhou J P, Liu G Q, Zhao Y X, Zhang R, Tang X, Li L, Jia X Y, Guo Y C, Wu Y C, Han Y S, Bao Y, He Y, Han Q Q, Yang H, Zheng X L, Qi Y P, Zhang T, Zhang Y. 2023. An efficient CRISPR- Cas12a promoter editing system for crop improvement. Nat Plants, 9(4): 588-604. |
| [38] | Zhou W, Malabanan P B, Abrigo E. 2015. OsHox4 regulates GA signaling by interacting with DELLA-like genes and GA oxidase genes in rice. Euphytica, 201(1): 97-107. |
| [39] | Zong W, Tang N, Yang J, Peng L, Ma S Q, Xu Y, Li G L, Xiong L Z. 2016. Feedback regulation of ABA signaling and biosynthesis by a bZIP transcription factor targets drought-resistance-related genes. Plant Physiol, 171(4): 2810-2825. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||