
Rice Science ›› 2025, Vol. 32 ›› Issue (6): 766-771.DOI: 10.1016/j.rsci.2025.06.008
收稿日期:2025-03-02
接受日期:2025-06-30
出版日期:2025-11-28
发布日期:2025-12-04
. [J]. Rice Science, 2025, 32(6): 766-771.
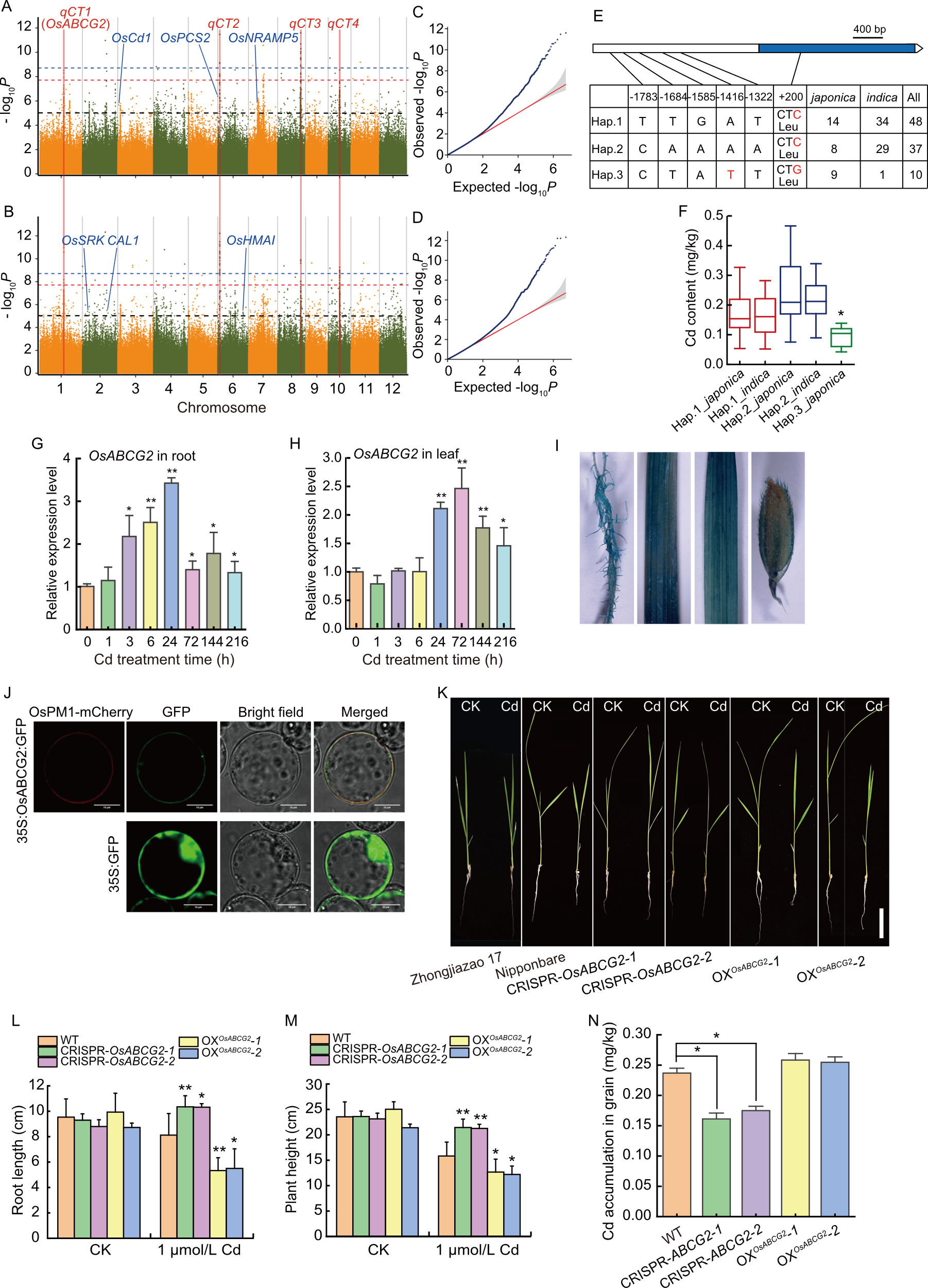
Fig. 1. Genome-wide association analysis of cadmium (Cd) accumulation in Ting’s core collection and functional analysis of OsABCG2. A and B, Manhattan plots for Cd accumulation in 2013 (A) and 2016 (B) using EMMAX. The blue and red fonts represent the cloned genes related to Cd and the novel QTL regions identified in the present study, respectively. Blue, red, and black horizontal dashed lines indicate the significant thresholds of 8.71 [-log10(1.94 × 10-9)], 7.71 [-log10(1.93 × 10-8)], and 5.09 [-log10(8.17 × 10-6)], respectively. C and D, Quantile-Quantile plots for Cd accumulation in 2013 (C) and 2016 (D). Red and blue symbols represent expected -log10(P-value) and observed -log10(P-value), respectively. E, Haplotype analysis of OsABCG2. Blue frame, white frame, and white triangle represent the exon, promoter, and 3′-untranslated region of OsABCG2, respectively. The numbers with minus and plus markers represent the positions to start codon of variations in promoter and coding sequence regions, respectively. F, Cd contents in grains of different haplotypes in Ting’s core collection. G and H, Relative expression of OsABCG2 in Nipponbare’s roots (G) and leaves (H) exposed to CdCl2 (20 μmol/L). I, β-Glucuronidase (GUS) staining in root, stem, leaf at the seedling stage, and grains driven by OsABCG2 promoter. J, Subcellular localization of OsABCG2. OsPM1-mCherry is the plasma membrane (PM)-localization marker. Scale bars, 10 µm. K, Performances of seedlings with (1 μmol/L) or without (CK) CdCl2 among wild type (WT, Nipponbare), OsABCG2 knockout lines (CRISPR-OsABCG2-1 and CRISPR-OsABCG2-2), and overexpression lines (OXOsABCG2-1 and OXOsABCG2-2). Scale bar, 4 cm. L and M, Root length (L) and plant height (M) of seedlings with (1 μmol/L) or without (CK) CdCl2 among WT, OsABCG2 knockout lines (CRISPR-OsABCG2-1 and CRISPR-OsABCG2-2), and overexpression lines (OXOsABCG2-1 and OXOsABCG2-2). Data are mean ± SD (n = 8-10). N, Comparison of Cd accumulation in grains among WT, OsABCG2 knockout lines (CRISPR-OsABCG2-1 and CRISPR-OsABCG2-2), and overexpression lines (OXOsABCG2-1 and OXOsABCG2-2). * and ** represent P < 0.05 and P < 0.01 levels by the Student’s t-test.
| [1] | Chen J, Huang X Y, Salt D E, et al. 2020. Mutation in OsCADT1 enhances cadmium tolerance and enriches selenium in rice grain. New Phytol, 226(3): 838-850. |
| [2] | Chen J G, Zou W L, Meng L J, et al. 2019. Advances in the uptake and transport mechanisms and QTLs mapping of cadmium in rice. Int J Mol Sci, 20(14): 3417. |
| [3] | Chen J M, Yang Y J, Huang Q N, et al. 2017. Effects of continuous flooding on cadmium absorption and its regulation mechanisms in rice. Sci Agric Sin, 50(17): 3300-3310. (in Chinese with English abstract) |
| [4] | Clemens S, Ma J F. 2016. Toxic heavy metal and metalloid accumulation in crop plants and foods. Annu Rev Plant Biol, 67: 489-512. |
| [5] | Clemens S, Aarts M G M, Thomine S, et al. 2013. Plant science: The key to preventing slow cadmium poisoning. Trends Plant Sci, 18(2): 92-99. |
| [6] | Das N, Bhattacharya S, Bhattacharyya S, et al. 2017. Identification of alternatively spliced transcripts of rice phytochelatin synthase 2 gene OsPCS2 involved in mitigation of cadmium and arsenic stresses. Plant Mol Biol, 94(1/2): 167-183. |
| [7] | Ding S L, Liu C L, Shang L G, et al. 2021. Identification of QTLs for cadmium tolerance during seedling stage and validation of qCDSL1 in rice. Rice Sci, 28(1): 81-88. |
| [8] | Ding Y F, Ye Y Y, Jiang Z H, et al. 2016. microRNA390 is involved in cadmium tolerance and accumulation in rice. Front Plant Sci, 7: 235. |
| [9] | Do T H T, Martinoia E, Lee Y, et al. 2021. 2021 update on ATP-binding cassette (ABC) transporters: How they meet the needs of plants. Plant Physiol, 187(4): 1876-1892. |
| [10] | Fu S, Lu Y S, Zhang X, et al. 2019. The ABC transporter ABCG36 is required for cadmium tolerance in rice. J Exp Bot, 70(20): 5909-5918. |
| [11] | Hao X H, Zeng M, Wang J, et al. 2018. A node-expressed transporter OsCCX2 is involved in grain cadmium accumulation of rice. Front Plant Sci, 9: 476. |
| [12] | Ishikawa S, Ishimaru Y, Igura M, et al. 2012. Ion-beam irradiation, gene identification, and marker-assisted breeding in the development of low-cadmium rice. Proc Natl Acad Sci USA, 109(47): 19166-19171. |
| [13] | Liang L, Wang C C, Chen T. 2024. Advances in understanding cadmium stress and breeding of cadmium-tolerant crops. Rice Sci, 31(5): 507-525. |
| [14] | Lu C N, Zhang L X, Tang Z, et al. 2019. Producing cadmium-free indica rice by overexpressing OsHMA3. Environ Int, 126: 619-626. |
| [15] | Luo J S, Huang J, Zeng D L, et al. 2018. A defensin-like protein drives cadmium efflux and allocation in rice. Nat Commun, 9(1): 645. |
| [16] | Miyadate H, Adachi S, Hiraizumi A, et al. 2011. OsHMA3, a P1B-type of ATPase affects root-to-shoot cadmium translocation in rice by mediating efflux into vacuoles. New Phytol, 189(1): 190-199. |
| [17] | Nguyen V N T, Moon S, Jung K H. 2014. Genome-wide expression analysis of rice ABC transporter family across spatio-temporal samples and in response to abiotic stresses. J Plant Physiol, 171(14): 1276-1288. |
| [18] | Oda K, Otani M, Uraguchi S, et al. 2011. Rice ABCG43 is Cd inducible and confers Cd tolerance on yeast. Biosci Biotechnol Biochem, 75(6): 1211-1213. |
| [19] | Sasaki A, Yamaji N, Yokosho K, et al. 2012. Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice. Plant Cell, 24(5): 2155-2167. |
| [20] | Seregin I V, Ivanov V B. 2001. Physiological aspects of cadmium and lead toxic effects on higher plants. Russ J Plant Physiol, 48: 523-544. |
| [21] | Song W Y, Lee H S, Jin S R, et al. 2015. Rice PCR1 influences grain weight and Zn accumulation in grains. Plant Cell Environ, 38(11): 2327-2339. |
| [22] | Sun L, Xu X X, Jiang Y R, et al. 2016. Genetic diversity, rather than cultivar type, determines relative grain Cd accumulation in hybrid rice. Front Plant Sci, 7: 1407. |
| [23] | Takahashi R, Ishimaru Y, Shimo H, et al. 2012. The OsHMA2 transporter is involved in root-to-shoot translocation of Zn and Cd in rice. Plant Cell Environ, 35(11): 1948-1957. |
| [24] | Tian S Q, Liang S, Qiao K, et al. 2019. Co-expression of multiple heavy metal transporters changes the translocation, accumulation, and potential oxidative stress of Cd and Zn in rice (Oryza sativa). J Hazard Mater, 380: 120853. |
| [25] | Ueno D, Yamaji N, Kono I, et al. 2010. Gene limiting cadmium accumulation in rice. Proc Natl Acad Sci USA, 107: 16500-16505. |
| [26] | Uraguchi S, Kamiya T, Sakamoto T, et al. 2011. Low-affinity cation transporter (OsLCT1) regulates cadmium transport into rice grains. Proc Natl Acad Sci USA, 108: 20959-20964. |
| [27] | Yamaji N, Xia J X, Mitani-Ueno N, et al. 2013. Preferential delivery of zinc to developing tissues in rice is mediated by P-type heavy metal ATPase OsHMA2. Plant Physiol, 162(2): 927-939. |
| [28] | Yan H L, Xu W X, Xie J Y, et al. 2019. Variation of a major facilitator superfamily gene contributes to differential cadmium accumulation between rice subspecies. Nat Commun, 10(1): 3301. |
| [29] | Zhang P, Zhong K Z, Zhong Z Z, et al. 2019a. Genome-wide association study of important agronomic traits within a core collection of rice (Oryza sativa L.). BMC Plant Biol, 19(1): 259. |
| [30] | Zhang P, Zhong K Z, Zhong Z Z, et al. 2019b. Mining candidate gene for rice aluminum tolerance through genome wide association study and transcriptomic analysis. BMC Plant Biol, 19(1): 490. |
| [31] | Zhao J L, Yang W, Zhang S H, et al. 2018. Genome-wide association study and candidate gene analysis of rice cadmium accumulation in grain in a diverse rice collection. Rice, 11(1): 61. |
| [32] | Zhong K Z, Xie L H, Hu S K, et al. 2022. Genome-wide association study of zinc toxicity tolerance within a rice core collection (Oryza sativa L.). Plants, 11(22): 3138. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||