
Rice Science ›› 2026, Vol. 33 ›› Issue (2): 221-231.DOI: 10.1016/j.rsci.2025.10.006
收稿日期:2025-07-07
接受日期:2025-08-18
出版日期:2026-03-28
发布日期:2026-04-01
. [J]. Rice Science, 2026, 33(2): 221-231.

Fig. 1. Phylogenetic tree and protein structure of JMJ30 in selected monocotyledons and Arabidopsis. A, Phylogenetic analysis of JMJ30 proteins from wheat (Triticum aestivum), barley (Hordeum vulgare), purple false brome (Brachypodium distachyon), rice (Oryza sativa), sorghum (Sorghum bicolor), foxtail millet (Setaria italica), and Arabidopsis (Arabidopsis thaliana). The phylogenetic tree was constructed using the neighbor-joining method based on JMJ30 protein sequences, with 1 000 bootstrap replicates in MEGA11. The values at the nodes indicate the percentage of the bootstrap confidence level. B, Protein domain architecture of JMJ30 in different species. Gray boxes represent variable regions, while blue boxes denote conserved JmjC domains.
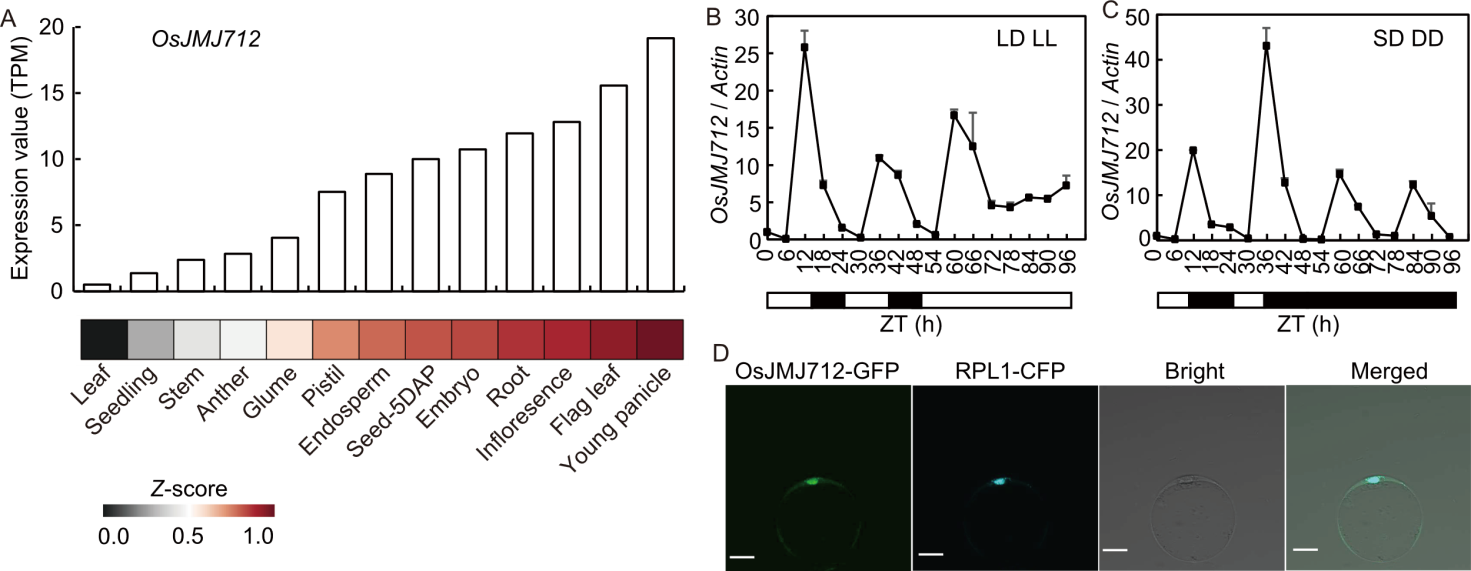
Fig. 2. Expression pattern and subcellular localization of OsJMJ712. A, Expression levels of OsJMJ712 in various rice tissues. The bar chart display expression values, and the heatmap shows gene expression as transcripts per million (TPM) values normalized by Z-score transformation. RNA-seq data were obtained from the Rice Genome Annotation Project database (RGAP, https://rice.uga.edu/expression.shtml). Transcript abundances were quantified as TPM. DAP, Days after pollination. B and C, Rhythmic expression patterns of OsJMJ712 under long-day (LD) transferred to constant light (LL) (B) and short-day (SD) transferred to constant dark (DD) (C) conditions. Fourteen-day-old seedlings grown under LD or SD conditions were transferred to LL or DD, respectively. Samples were collected every 6 h. The Actin gene was used as an internal control. Data are mean ± SD (n = 3). Black bars represent the dark period and light bars indicate the light period. ZT, Zeitgeber time; ZT0 is defined as the time of lights on. D, Subcellular localization of OsJMJ712-GFP. 35S:OsJMJ712-GFP was transformed into rice protoplasts and visualized by fluorescence microscopy. 35S:RPL1-CFP was used as a nuclear marker. Bright-field and merged images are shown. Scale bars, 10 μm. GFP, Green fluorescent protein; CFP, Cyan fluorescent protein.
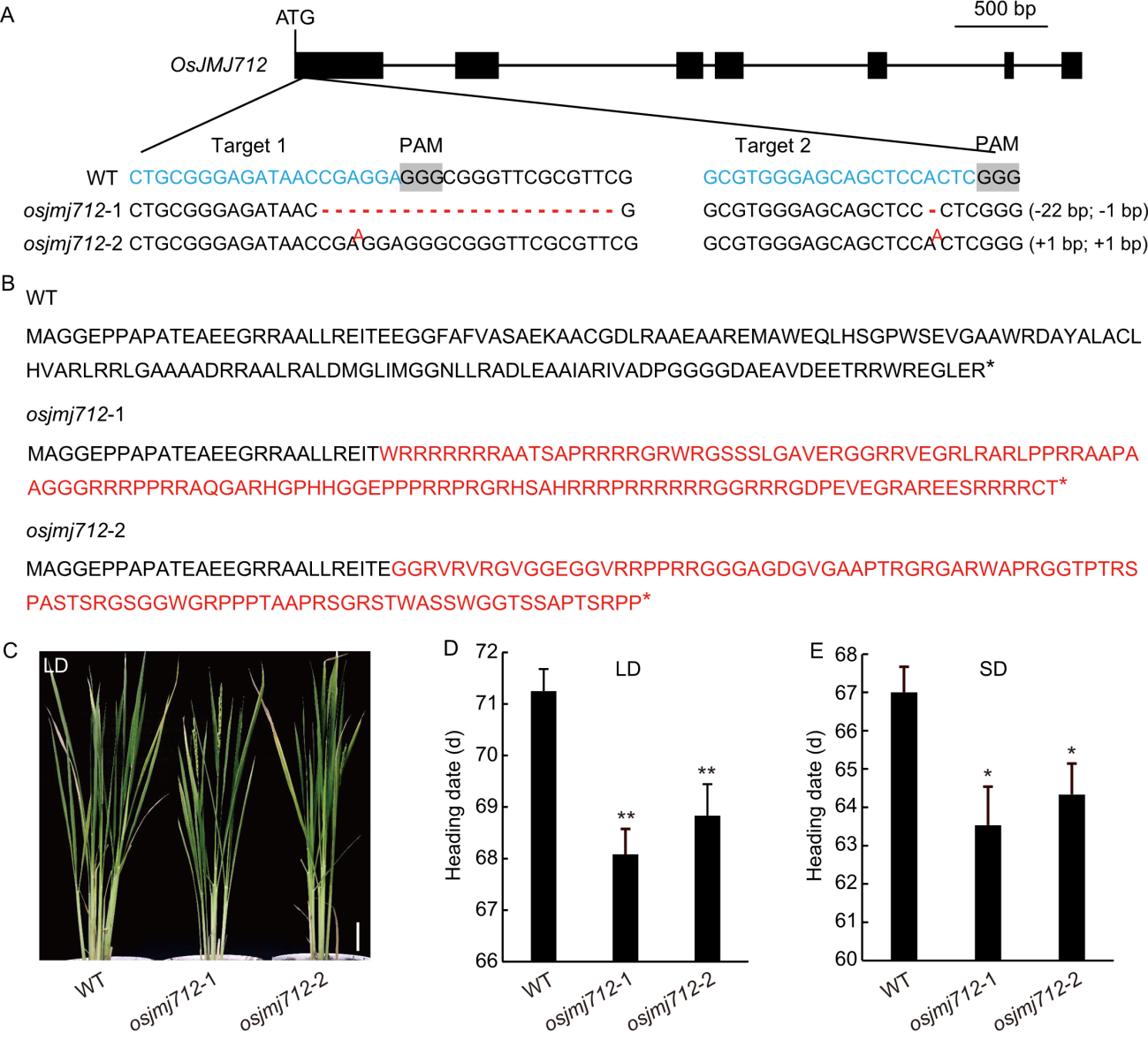
Fig. 3. Generation of OsJMJ712 gene-editing mutants and their heading date phenotypes under long-day (LD) and short-day (SD) conditions. A, Schematic of OsJMJ712 gene structure and CRISPR/Cas9-targeted mutation sites. sgRNA targets and protospacer adjacent motif (PAM) sequences are highlighted in blue and gray shading in the wild type (WT), respectively. Red nucleotides indicate insertions; red dashes indicate deletions. Sequence gap lengths are shown in parentheses. B, Partial amino acid sequences of WT and osjmj712 mutants. Red letters highlight predicted altered amino acids. Asterisks (*) indicate stop codons. C, Flowering phenotype of WT and osjmj712 mutants under LD conditions. Scale bar, 10 cm. D and E, Statistical analysis of heading date for WT and osjmj712 mutants under LD (D) and SD (E) conditions. Data are mean ± SD (n = 12). Significant differences were determined by a two-tailed unpaired Student’s t-test (*, P ≤ 0.05; **, P ≤ 0.01).
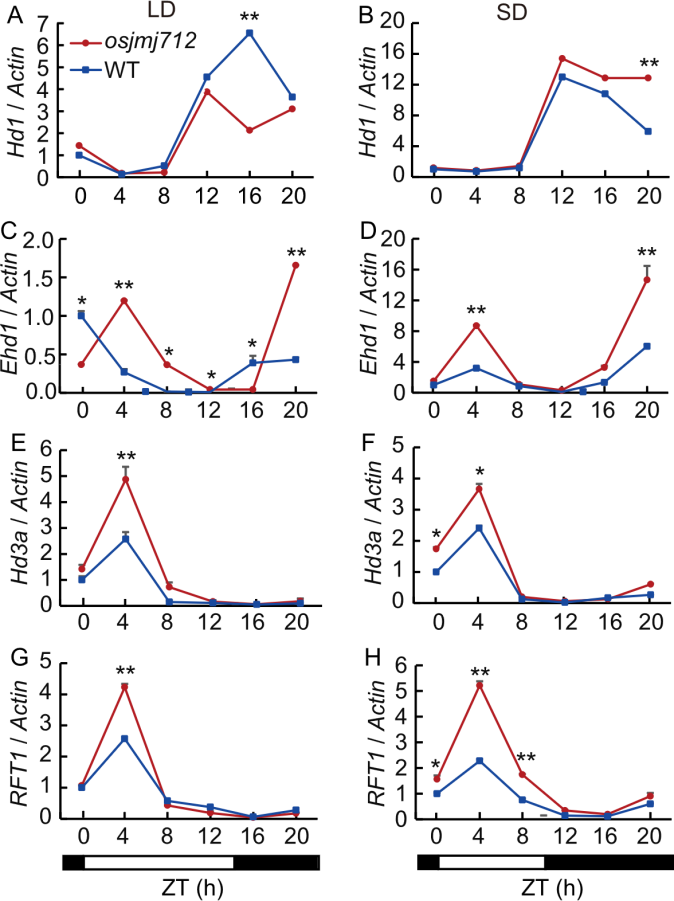
Fig. 4. Diurnal expression patterns of key flowering-related genes in wild type (WT) and osjmj712 mutants under long-day (LD) and short-day (SD) conditions. Black bars represent the dark period and light bars indicate the light period. ZT, Zeitgeber time; ZT0 is defined as the time of lights on. The Actin gene was used as an internal control. Data are mean ± SD (n = 3). Significant differences were determined by a two-tailed unpaired Student’s t-test (*, P ≤ 0.05; **, P ≤ 0.01).
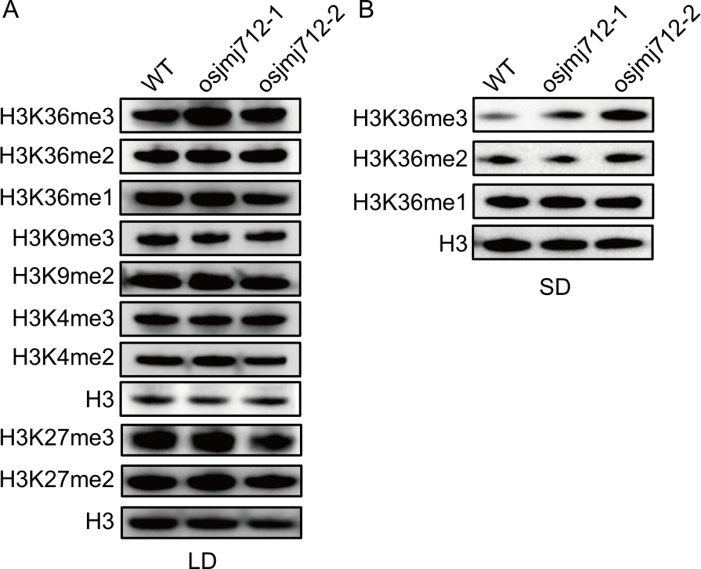

Fig. 5. OsJMJ712 is an H3K36me3 demethylase. Assay of histone H3 lysine methylation levels in vivo. Western blot analysis of histones isolated from wild type (WT) and osjmj712 mutants under long-day (LD, A) and short-day (SD, B) conditions using the indicated methylation-specific antibodies listed on the left. Anti-H3 was used as a loading control.
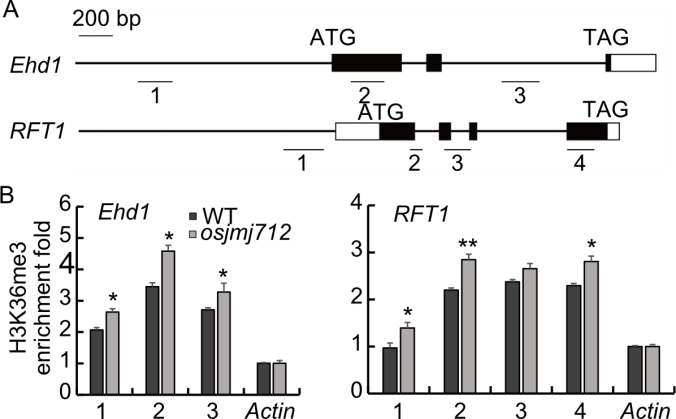
Fig. 6. H3K36me3 levels at Ehd1 and RFT1 loci in osjmj712 mutant and wild type (WT). A, Schematic diagrams of Ehd1 and RFT1 gene structures and the fragments detected by chromatin immunoprecipitation-quantitative PCR (ChIP-qPCR). Numbers 1‒4 represent the regions analyzed. B, ChIP-qPCR analysis of H3K36me3 enrichment at the Ehd1 and RFT1 loci. The Actin gene was used as an internal control for normalization. Numbers 1‒4 represent regions 1‒4 in A. IgG was used as a negative control. Data are mean ± SD (n = 3). Asterisks indicate significant differences between osjmj712 mutant and WT (Student’s t-test, *, P ≤ 0.05; **, P ≤ 0.01).
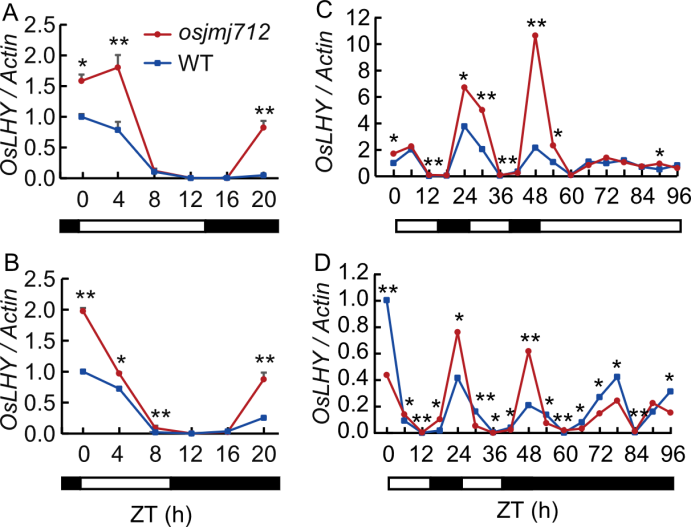
Fig. 7. Rhythmic expression of OsLHY in osjmj712 mutant and wild type (WT). A and B, Diurnal expression patterns of OsLHY in WT and osjmj712 mutant under long-day (LD, A) and short-day (SD, B) conditions. C and D, Free-running rhythms of OsLHY expression in WT and osjmj712 mutant under constant light (LL, C) and dark (DD, D) conditions. Black bars represent the dark period and light bars indicate the light period. ZT, Zeitgeber time; ZT0 is defined as the time of lights on. The Actin gene was used as an internal control. Data are mean ± SD (n = 3). Significant differences were determined by a two-tailed unpaired Student’s t-test (*, P ≤ 0.05; **, P ≤ 0.01).
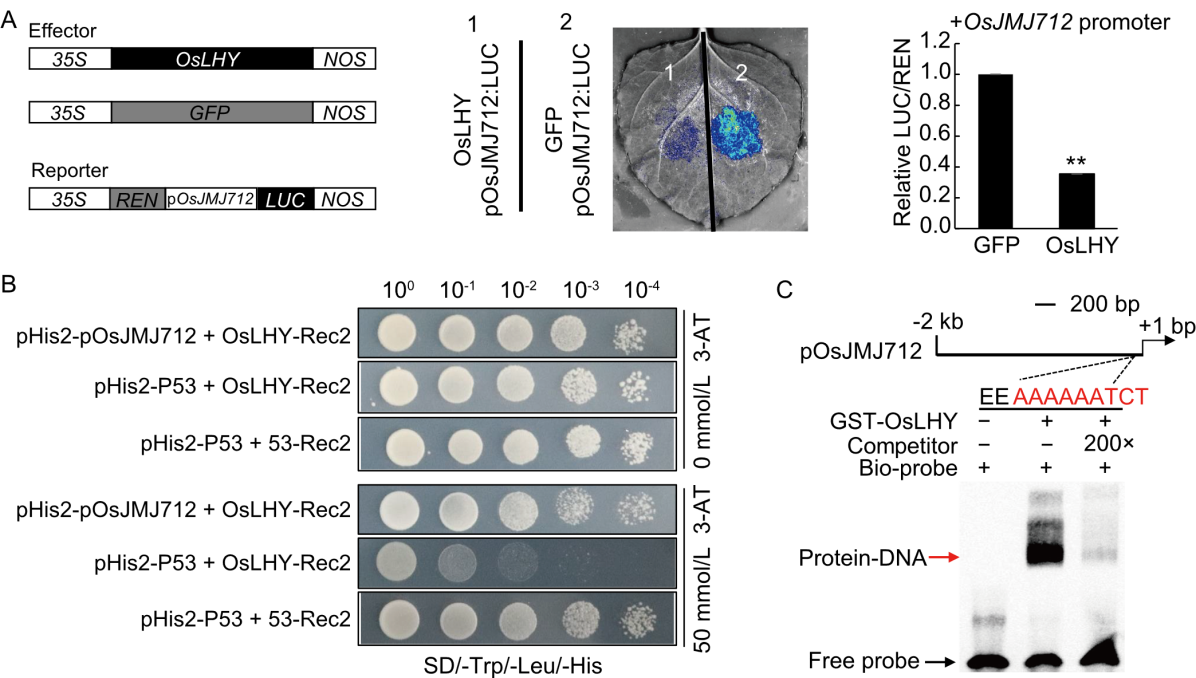
Fig. 8. OsLHY directly binds to OsJMJ712 promoter. A, Transcriptional activity of the OsJMJ712 promoter was assessed by dual-luciferase assays in Nicotiana benthamiana leaves. REN, Renilla luciferase; LUC, Firefly luciferase. Data are mean ± SD (n = 5). Significant differences were determined by Student’s t-test (**, P ≤ 0.01). B, Interaction between OsLHY and OsJMJ712 promoter detected by yeast one-hybrid (Y1H) assays. pHis2‐pOsJMJ712 and OsLHY‐Rec2 were co-transformed into yeast cells, and transformants were grown on SD/‐Trp/‐Leu/‐His (SD/-TLH) medium and SD/‐TLH medium supplemented with 50 mmol/L 3‐amino‐1,2,4‐triazole (3‐AT). C, Binding of OsLHY to evening element (EE) sequences in OsJMJ712 promoter detected by electrophoretic mobility shift assay (EMSA). EE fragments were 5ʹ-biotin-labelled as the Bio‐probe. Unlabelled probe was used as competitor. Purified GST‐OsLHY fusion protein was incubated with the designed probes. The red arrowhead indicates the protein‐DNA complex, and the black arrowhead indicates the free probe.
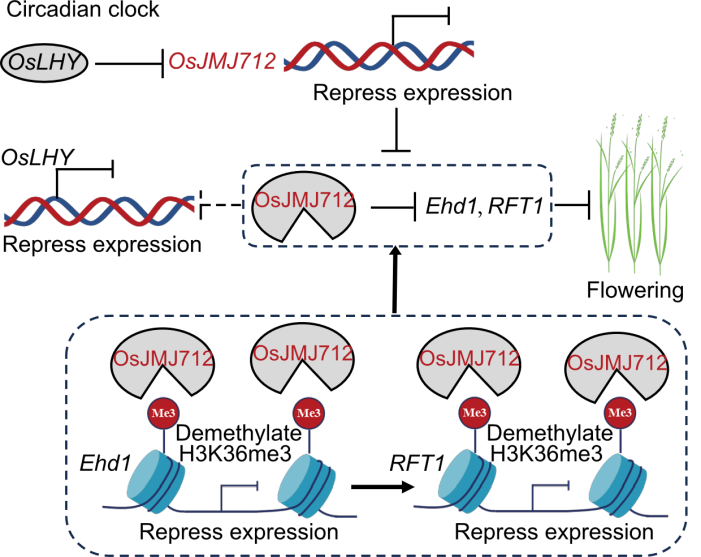
Fig. 9. Proposed model for the role of OsJMJ712 in regulating flowering in rice. OsJMJ712 is a component of the circadian clock, and its expression is directly suppressed by OsLHY. Loss of function of OsJMJ712 increases OsLHY expression, indicating a potential negative transcriptional feedback loop. Acting as an H3K36me3 demethylase, OsJMJ712 removes H3K36me3 marks at the Ehd1 and RFT1 loci, thereby repressing their transcription and ultimately delaying flowering.
| [1] | Bart R, Chern M, Park C J, et al. 2006. A novel system for gene silencing using siRNAs in rice leaf and stem-derived protoplasts. Plant Methods, 2: 13. |
| [2] | Cai M H, Zhu S S, Wu M M, et al. 2021. DHD4, a CONSTANS-like family transcription factor, delays heading date by affecting the formation of the FAC complex in rice. Mol Plant, 14(2): 330-343. |
| [3] | Cazzonelli C I, Millar T, Finnegan E J, et al. 2009. Promoting gene expression in plants by permissive histone lysine methylation. Plant Signal Behav, 4(6): 484-488. |
| [4] | Chai J X, Gu X Y, Song P Y, et al. 2024. Histone demethylase JMJ713 interaction with JMJ708 modulating H3K36me2, enhances rice heat tolerance through promoting hydrogen peroxide scavenging. Plant Physiol Biochem, 217: 109284. |
| [5] | Chen K, Du K X, Shi Y C, et al. 2021. H3K36 methyltransferase SDG708 enhances drought tolerance by promoting abscisic acid biosynthesis in rice. New Phytol, 230(5): 1967-1984. |
| [6] | Cheng S F, Tan F, Lu Y, et al. 2018. WOX11 recruits a histone H3K27me3 demethylase to promote gene expression during shoot development in rice. Nucleic Acids Res, 46(5): 2356-2369. |
| [7] | Doi K, Izawa T, Fuse T, et al. 2004. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev, 18(8): 926-936. |
| [8] | Gan E S, Xu Y F, Wong J Y, et al. 2014. Jumonji demethylases moderate precocious flowering at elevated temperature via regulation of FLC in Arabidopsis. Nat Commun, 5: 5098. |
| [9] | Geng L, Tan M, Deng Q, et al. 2024. Transcription factors WOX11 and LBD16 function with histone demethylase JMJ706 to control crown root development in rice. Plant Cell, 36(5): 1777-1790. |
| [10] | Hamilton J P, Li C, Buell C R. 2024. The rice genome annotation project: An updated database for mining the rice genome. Nucleic Acids Res, 53(D1): D1614-D1622. |
| [11] | Hayama R, Yokoi S, Tamaki S, et al. 2003. Adaptation of photoperiodic control pathways produces short-day flowering in rice. Nature, 422: 719-722. |
| [12] | He Y H, Michaels S D, Amasino R M. 2003. Regulation of flowering time by histone acetylation in Arabidopsis. Science, 302: 1751-1754. |
| [13] | Hung F Y, Lai Y C, Wang J H, et al. 2021. The Arabidopsis histone demethylase JMJ28 regulates CONSTANS by interacting with FBH transcription factors. Plant Cell, 33(4): 1196-1211. |
| [14] | Izawa T. 2007. Adaptation of flowering-time by natural and artificial selection in Arabidopsis and rice. J Exp Bot, 58(12): 3091-3097. |
| [15] | Jia J T, Luo Y J, Wu Z Y, et al. 2024. OsJMJ718, a histone demethylase gene, positively regulates seed germination in rice. Plant J, 118(1): 191-202. |
| [16] | Jones M A, Morohashi K, Grotewold E, et al. 2019. Arabidopsis JMJD5/JMJ30 acts independently of LUX ARRHYTHMO within the plant circadian clock to enable temperature compensation. Front Plant Sci, 10: 57. |
| [17] | Kouzarides T. 2007. Chromatin modifications and their function. Cell, 128(4): 693-705. |
| [18] | Lee S J, Kang K, Lim J H, et al. 2022. Natural alleles of CIRCADIAN CLOCK ASSOCIATED1 contribute to rice cultivation by fine-tuning flowering time. Plant Physiol, 190(1): 640-656. |
| [19] | Li C, Liu X J, Yan Y, et al. 2022. OsLHY is involved in regulating flowering through the Hd1- and Ehd1-mediated pathways in rice (Oryza sativa L.). Plant Sci, 315: 111145. |
| [20] | Li C, He Y Q, Yu J, et al. 2024. The rice LATE ELONGATED HYPOCOTYL enhances salt tolerance by regulating Na+/k+ homeostasis and ABA signalling. Plant Cell Environ, 47(5): 1625-1639. |
| [21] | Liu B, Wei G, Shi J L, et al. 2016. SET DOMAIN GROUP 708, a histone H3 lysine 36-specific methyltransferase, controls flowering time in rice (Oryza sativa). New Phytol, 210(2): 577-588. |
| [22] | Liu C Y, Lu F L, Cui X, et al. 2010. Histone methylation in higher plants. Annu Rev Plant Biol, 61: 395-420. |
| [23] | Lu F L, Li G L, Cui X, et al. 2008. Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J Integr Plant Biol, 50(7): 886-896. |
| [24] | Lu F L, Cui X, Zhang S B, et al. 2010. JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res, 20(3): 387-390. |
| [25] | Lu S X, Knowles S M, Webb C J, et al. 2011. The Jumonji C domain-containing protein JMJ30 regulates period length in the Arabidopsis circadian clock. Plant Physiol, 155(2): 906-915. |
| [26] | Ma X L, Zhang Q Y, Zhu Q L, et al. 2015. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant, 8(8): 1274-1284. |
| [27] | Nemoto Y, Nonoue Y, Yano M, et al. 2016. Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7. Plant J, 86(3): 221-233. |
| [28] | Noh B, Lee S H, Kim H J, et al. 2004. Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell, 16(10): 2601-2613. |
| [29] | Putterill J, Robson F, Lee K R, et al. 1995. The CONSTANS gene of Arabidopsis promotes flowering and encodes a protein showing similarities to zinc finger transcription factors. Cell, 80(6): 847-857. |
| [30] | Sui P F, Shi J L, Gao X Y, et al. 2013. H3K36 methylation is involved in promoting rice flowering. Mol Plant, 6(3): 975-977. |
| [31] | Sun C H, Fang J, Zhao T L, et al. 2012. The histone methyltransferase SDG724 mediates H3K36me2/3 deposition at MADS50 and RFT1 and promotes flowering in rice. Plant Cell, 24(8): 3235-3247. |
| [32] | Sun C H, Chen D, Fang J, et al. 2014. Understanding the genetic and epigenetic architecture in complex network of rice flowering pathways. Protein Cell, 5(12): 889-898. |
| [33] | Sun C H, Zhang K, Zhou Y, et al. 2021. Dual function of clock component OsLHY sets critical day length for photoperiodic flowering in rice. Plant Biotechnol J, 19(8): 1644-1657. |
| [34] | Sun Q W, Zhou D X. 2008. Rice jmjC domain-containing gene JMJ706 encodes H3K9 demethylase required for floral organ development. Proc Natl Acad Sci USA, 105: 13679-13684. |
| [35] | Turck F, Fornara F, Coupland G. 2008. Regulation and identity of florigen: FLOWERING LOCUS T moves center stage. Annu Rev Plant Biol, 59: 573-594. |
| [36] | Weng X Y, Zhou S L, Zong W, et al. 2018. ChIP assay in rice. Bio-101: e1010135. (in Chinese) |
| [37] | Yan Y Y, Shen L S, Chen Y, et al. 2014. A MYB-domain protein EFM mediates flowering responses to environmental cues in Arabidopsis. Dev Cell, 30(4): 437-448. |
| [38] | Yang H C, Han Z F, Cao Y, et al. 2012a. A companion cell-dominant and developmentally regulated H3K4 demethylase controls flowering time in Arabidopsis via the repression of FLC expression. PLoS Genet, 8(4): e1002664. |
| [39] | Yang H C, Mo H X, Fan D, et al. 2012b. Overexpression of a histone H3K4 demethylase, JMJ15, accelerates flowering time in Arabidopsis. Plant Cell Rep, 31(7): 1297-1308. |
| [40] | Yang W N, Jiang D H, Jiang J F, et al. 2010. A plant-specific histone H3 lysine 4 demethylase represses the floral transition in Arabidopsis. Plant J, 62(4): 663-673. |
| [41] | Yang Y, Peng Q, Chen G X, et al. 2013. OsELF3 is involved in circadian clock regulation for promoting flowering under long-day conditions in rice. Mol Plant, 6(1): 202-215. |
| [42] | Yano M, Katayose Y, Ashikari M, et al. 2000. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell, 12( 12): 2473-2484. |
| [43] | Yokoo T, Saito H, Yoshitake Y, et al. 2014. Se14, encoding a JmjC domain-containing protein, plays key roles in long-day suppression of rice flowering through the demethylation of H3K4me 3 of RFT1. PLoS One, 9(4): e96064. |
| [44] | Zhang C C, Yuan W Y, Zhang Q F. 2012. RPL1, a gene involved in epigenetic processes regulates phenotypic plasticity in rice. Mol Plant, 5: 482-493. |
| [45] | Zhao W J, Wang X Y, Zhang Q, et al. 2022. H3K36 demethylase JMJ710 negatively regulates drought tolerance by suppressing MYB48-1 expression in rice. Plant Physiol, 189(2): 1050-1064. |
| [46] | Zheng S Z, Hu H M, Ren H M, et al. 2019. The Arabidopsis H3K27me3 demethylase JUMONJI 13 is a temperature and photoperiod dependent flowering repressor. Nat Commun, 10(1): 1303. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||