
Rice Science ›› 2015, Vol. 22 ›› Issue (4): 180-188.DOI: 10.1016/S1672-6308(14)60293-6
收稿日期:2014-09-23
接受日期:2015-02-11
出版日期:2015-07-28
发布日期:2015-05-27
. [J]. Rice Science, 2015, 22(4): 180-188.
| Origin | Code for each material a | Year |
| Qingpu, Shanghai, China | Qp-1, Qp-2, Qp-3, Qp-4, Qp-5 | 2008 |
| Jiading, Shanghai, China | Jd-1, Jd-2, Jd-3, Jd-4, Jd-5, Jd-6 | 2007 |
| Nanhui, Shanghai, China | Nh-1, Nh-2, Nh-3, Nh-4 | 2010 |
| Pudong, Shanghai, China | Pd-1, Pd-2, Pd-3, Pd-4, Pd-5 | 2009 |
| Chongming, Shanghai, China | Cm-1, Cm-2, Cm-3, Cm-4, Cm-5 | 2008 |
| Fengxian, Shanghai, China | Fx-1, Fx-2, Fx-3, Fx-4, Fx-5 | 2009 |
| Taizhou, Jiangsu, China | Tz-1, Tz-2, Tz-3, Tz-4, Tz-5 | 2005 |
| Shenyang, Liaoning, China | Sy-1, Sy-2, Sy-3, Sy-4, Sy-5 | 2000 |
| Tieling, Liaoning, China | Tl-1, Tl-2, Tl-3, Tl-4, Tl-5 | 2004 |
| Haicheng, Liaoning, China | Hc-1, Hc-2, Hc-3, Hc-4, Hc-5 | 2003 |
| Dandong, Liaoning, China | Dd-1, Dd-2, Dd-3, Dd-4, Dd-5 | 2004 |
| Gongzhuling, Jilin, China | Gz-1, Gz-2, Gz-3, Gz-4, Gz-5 | 2006 |
| Jiamusi, Heilongjiang, China | Jm-1, Jm-2, Jm-3, Jm-4, Jm-5 | 2005 |
| Yesan, Chungcheong, Korea | Ys-1, Ys-2, Ys-3, Ys-4, Ys-5 | 2000 |
| Seosan, Chungcheong, Korea | Ss-1, Ss-2, Ss-3, Ss-4, Ss-5, Ss-6 | 2000 |
| Qp, Qingpu; Jd, Jiading; Nh, Nanhui; Pd, Pudong; Cm, Chongming; Fx, Fengxian; Tz, Taizhou; Sy, Shenyang; Tl, Tieling; Hc, Haicheng; Dd, Dandong; Gz, Gongzhuling; Jm, Jiamusi; Ys, Yesan; Ss, Seosan. a The numbers represent weedy rice accessions from each location. | ||
Table 1 Origins, codes and collected years of Northeast Asia weedy rice accessions used for SSR analysis.
| Origin | Code for each material a | Year |
| Qingpu, Shanghai, China | Qp-1, Qp-2, Qp-3, Qp-4, Qp-5 | 2008 |
| Jiading, Shanghai, China | Jd-1, Jd-2, Jd-3, Jd-4, Jd-5, Jd-6 | 2007 |
| Nanhui, Shanghai, China | Nh-1, Nh-2, Nh-3, Nh-4 | 2010 |
| Pudong, Shanghai, China | Pd-1, Pd-2, Pd-3, Pd-4, Pd-5 | 2009 |
| Chongming, Shanghai, China | Cm-1, Cm-2, Cm-3, Cm-4, Cm-5 | 2008 |
| Fengxian, Shanghai, China | Fx-1, Fx-2, Fx-3, Fx-4, Fx-5 | 2009 |
| Taizhou, Jiangsu, China | Tz-1, Tz-2, Tz-3, Tz-4, Tz-5 | 2005 |
| Shenyang, Liaoning, China | Sy-1, Sy-2, Sy-3, Sy-4, Sy-5 | 2000 |
| Tieling, Liaoning, China | Tl-1, Tl-2, Tl-3, Tl-4, Tl-5 | 2004 |
| Haicheng, Liaoning, China | Hc-1, Hc-2, Hc-3, Hc-4, Hc-5 | 2003 |
| Dandong, Liaoning, China | Dd-1, Dd-2, Dd-3, Dd-4, Dd-5 | 2004 |
| Gongzhuling, Jilin, China | Gz-1, Gz-2, Gz-3, Gz-4, Gz-5 | 2006 |
| Jiamusi, Heilongjiang, China | Jm-1, Jm-2, Jm-3, Jm-4, Jm-5 | 2005 |
| Yesan, Chungcheong, Korea | Ys-1, Ys-2, Ys-3, Ys-4, Ys-5 | 2000 |
| Seosan, Chungcheong, Korea | Ss-1, Ss-2, Ss-3, Ss-4, Ss-5, Ss-6 | 2000 |
| Qp, Qingpu; Jd, Jiading; Nh, Nanhui; Pd, Pudong; Cm, Chongming; Fx, Fengxian; Tz, Taizhou; Sy, Shenyang; Tl, Tieling; Hc, Haicheng; Dd, Dandong; Gz, Gongzhuling; Jm, Jiamusi; Ys, Yesan; Ss, Seosan. a The numbers represent weedy rice accessions from each location. | ||
| Origin | Code | Accession name |
| Shanghai, China | C-Sh | Jinfeng, Youfeng, Hanfeng, Qiufeng |
| Jiangsu, China | C-Js | Xiushui 7, Xiushui 9, Xiushui 61, Xiushui 110, Wuyunjing 7, Wuyunjing 3, Zhendao 88 |
| Liaoning, China | C-Ln | Liaojing 9, Liaojing 294, Liaojing 371, Liaoxing 15, Shendao 7 |
| Chungcheong, Korea | C-Cc | Ilpum, Singeumo, Gopum, Dongjin 2, Yeon 308 |
Table 2 List of cultivated rice collected in Northeast Asia used for SSR analysis.
| Origin | Code | Accession name |
| Shanghai, China | C-Sh | Jinfeng, Youfeng, Hanfeng, Qiufeng |
| Jiangsu, China | C-Js | Xiushui 7, Xiushui 9, Xiushui 61, Xiushui 110, Wuyunjing 7, Wuyunjing 3, Zhendao 88 |
| Liaoning, China | C-Ln | Liaojing 9, Liaojing 294, Liaojing 371, Liaoxing 15, Shendao 7 |
| Chungcheong, Korea | C-Cc | Ilpum, Singeumo, Gopum, Dongjin 2, Yeon 308 |
| Primer | Chr | SSR motif | Forward sequence | Reverse sequence | EPS | Na | Ne | I |
| RM1 | 1 | (GA)26 | GCGAAAACACAATGCAAAAA | GCGTTGGTTGGACCTGAC | 113 | 6 | 2.721 | 1.250 |
| RM9 | 1 | (GA)15GT(GA)2 | GGTGCCATTGTCGTCCTC | ACGGCCCTCATCACCTTC | 136 | 3 | 1.236 | 0.398 |
| RM14 | 1 | (GA)18 | CCGAGGAGAGGAGTTCGAC | GTGCCAATTTCCTCGAAAAA | 191 | 4 | 2.416 | 1.041 |
| RM23 | 1 | (GA)15 | CATTGGAGTGGAGGCTGG | GTCAGGCTTCTGCCATTCTC | 145 | 3 | 1.903 | 0.733 |
| RM212 | 1 | (CT)24 | CCACTTTCAGCTACTACCAG | CACCCATTTGTCTCTCATTATG | 136 | 2 | 1.403 | 0.462 |
| RM237 | 1 | (CT)18 | CAAATCCCGACTGCTGTCC | TGGGAAGAGAGCACTACAGC | 130 | 2 | 1.492 | 0.512 |
| RM110 | 2 | (GA)15 | TCGAAGCCATCCACCAACGAAG | TCCGTACGCCGACGAGGTCGAG | 156 | 2 | 1.280 | 0.377 |
| RM425 | 2 | (CGG)9 | CCAACGAAGATTCGAAGCTC | CAGCACCATGAAGTCGCC | 126 | 3 | 2.103 | 0.814 |
| RM525 | 2 | (AAG)12 | GGCCCGTCCAAGAAATATTG | CGGTGAGACAGAATCCTTACG | 131 | 4 | 3.034 | 1.179 |
| RM1178 | 2 | (AG)13 | CAGTGGGCGAGCATAGGAG | ATCCTTTTCTCCCTCTCTCG | 112 | 3 | 2.998 | 1.098 |
| RM156 | 3 | (CGG)8 | GCCGCACCCTCACTCCCTCCTC | TCTTGCCGGAGCGCTTGAGGTG | 160 | 3 | 2.090 | 0.879 |
| RM218 | 3 | (TC)24ACT5(GT)11 | TGGTCAAACCAAGGTCCTTC | GACATACATTCTACCCCCGG | 148 | 3 | 1.943 | 0.766 |
| RM489 | 3 | (ATA)8 | ACTTGAGACGATCGGACACC | TCACCCATGGATGTTGTCAG | 271 | 4 | 2.175 | 0.893 |
| RM8209 | 3 | (AG)15 | AGGAGAAGAGGAATCTTTGC | CGATCGAGAGCTACTATTGC | 188 | 3 | 1.043 | 0.117 |
| RM241 | 4 | (CT)31 | GAGCCAAATAAGATCGCTGA | TGCAAGCAGCAGATTTAGTG | 138 | 2 | 1.729 | 0.613 |
| RM470 | 4 | (CTT)14 | TCCTCATCGGCTTCTTCTTC | AGAACCCGTTCTACGTCACG | 83 | 4 | 3.406 | 1.297 |
| RM567 | 4 | (GA)21 | ATCAGGGAAATCCTGAAGGG | GGAAGGAGCAATCACCACTG | 261 | 1 | 1.000 | 0.000 |
| RM5473 | 4 | (TC)20 | ACACGGAGATCCGAGACACGAG | CGAGATTAACGTCGTCCTC | 105 | 5 | 4.117 | 1.493 |
| RM31 | 5 | (GA)15 | GATCACGATCCACTGGAGCT | AAGTCCATTACTCTCCTCCC | 140 | 6 | 2.737 | 1.266 |
| RM188 | 5 | (CA)8 | TCCGCCTCTCCTCTCGCTTCCC | GCAACGCACAACCGAACCGAGC | 210 | 3 | 2.283 | 0.924 |
| RM430 | 5 | (GA)25 | AAACAACGACGTCCCTGATC | GTGCCTCCGTGGTTATGAAC | 173 | 5 | 2.922 | 1.217 |
| RM193 | 6 | (GCT)5 | CGCCTCTTCTTCCTCGCCTCCG | CGGGTCCATCCCCCCTCTCCTC | 189 | 1 | 1.000 | 0.000 |
| RM276 | 6 | (AG)8A3(GA)33 | CTCAACGTTGACACCTCGTG | TCCTCCATCGAGCAGTATCA | 149 | 4 | 2.113 | 0.960 |
| RM402 | 6 | (ATA)7 | GAGCCATGGAAAGATGCATG | TCAGCTGGCCTATGACAATG | 133 | 3 | 1.321 | 0.487 |
| RM469 | 6 | (AG)15 | AGCTGAACAAGCCCTGAAAG | GACTTGGGCAGTGTGACATG | 105 | 2 | 1.465 | 0.498 |
| RM557 | 6 | (AC)13 | GTGGCGAGATCTATGTGGTG | GCTTTGTGTGTGTGTGTGTG | 211 | 1 | 1.000 | 0.000 |
| RM234 | 7 | (CT)25 | ACAGTATCCAAGGCCCTGG | CACGTGAGACAAAGACGGAG | 156 | 2 | 1.729 | 0.613 |
| RM346 | 7 | (CTT)18 | CGAGAGAGCCCATAACTACG | ACAAGACGACGAGGAGGGAC | 175 | 3 | 2.132 | 0.867 |
| RM429 | 7 | (TG)10 | TCCCTCCAGCAATGTCTTTC | CCTTCATCTTGCTTTCCACC | 159 | 3 | 1.876 | 0.744 |
| RM500 | 7 | (AAG)9 | GAGCTTGCCAGAGTGGAAAG | GTTACACCGAGAGCCAGCTC | 259 | 1 | 1.000 | 0.000 |
| RM256 | 8 | (CT)21 | GACAGGGAGTGATTGAAGGC | GTTGATTTCGCCAAGGGC | 127 | 2 | 1.021 | 0.058 |
| RM264 | 8 | GA)27 | GTTGCGTCCTACTGCTACTTC | GATCCGTGTCGATGATTAGC | 178 | 5 | 2.586 | 1.149 |
| RM281 | 8 | (GA)21 | ACCAAGCATCCAGTGACCAG | GTTCTTCATACAGTCCACATG | 138 | 3 | 2.073 | 0.799 |
| RM408 | 8 | (CT)13 | CAACGAGCTAACTTCCGTCC | ACTGCTACTTGGGTAGCTGACC | 128 | 3 | 2.014 | 0.783 |
| RM515 | 8 | (GA)11 | TAGGACGACCAAAGGGTGAG | TGGCCTGCTCTCTCTCTCTC | 211 | 4 | 2.419 | 1.005 |
| RM201 | 9 | (CT)17 | CTCGTTTATTACCTACAGTACC | CTACCTCCTTTCTAGACCGATA | 158 | 2 | 1.741 | 0.617 |
| RM242 | 9 | (CT)26 | GGCCAACGTGTGTATGTCTC | TATATGCCAAGACGGATGGG | 225 | 3 | 1.916 | 0.807 |
| RM257 | 9 | (CT)24 | CAGTTCCGAGCAAGAGTACTC | GGATCGGACGTGGCATATG | 147 | 4 | 3.504 | 1.314 |
| RM566 | 9 | (AG)15 | ACCCAACTACGATCAGCTCG | CTCCAGGAACACGCTCTTTC | 239 | 3 | 1.879 | 0.802 |
| RM216 | 10 | (CT)18 | GCATGGCCGATGGTAAAG | TGTATAAAACCACACGGCCA | 146 | 3 | 1.291 | 0.436 |
| RM228 | 10 | (CA)6(GA)36 | CTGGCCATTAGTCCTTGG | GCTTGCGGCTCTGCTTAC | 154 | 7 | 4.632 | 1.693 |
| RM484 | 10 | (AT)9 | TCTCCCTCCTCACCATTGTC | TGCTGCCCTCTCTCTCTCTC | 299 | 1 | 1.000 | 0.000 |
| RM224 | 11 | (AAG)8(AG)13 | ATCGATCGATCTTCACGAGG | TGCTATAAAAGGCATTCGGG | 157 | 4 | 2.390 | 1.070 |
| RM286 | 11 | (GA)16 | GGCTTCATCTTTGGCGAC | CCGGATTCACGAGATAAACTC | 110 | 4 | 2.484 | 1.066 |
| RM17 | 12 | (GA)21 | TGCCCTGTTATTTTCTTCTCTC | GGTGATCCTTTCCCATTTCA | 184 | 3 | 1.349 | 0.495 |
| RM101 | 12 | (CT)37 | GTGAATGGTCAAGTGACTTAGGTGGC | ACACAACATGTTCCCTCCCATGC | 324 | 3 | 2.563 | 1.001 |
| RM277 | 12 | (GA)11 | CGGTCAAATCATCACCTGAC | CAAGGCTTGCAAGGGAAG | 124 | 2 | 1.929 | 0.675 |
| RM1103 | 12 | (AG)12 | CAGCTGCTGCTACTACACCG | CTACTCCACGTCCATGCATG | 216 | 2 | 1.792 | 0.634 |
| Chr, Chromosome; EPS, Expected PCR product size; Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s information index. | ||||||||
Table 3 Repeat mofitis, sequences and genetic diversity of SSR primers.
| Primer | Chr | SSR motif | Forward sequence | Reverse sequence | EPS | Na | Ne | I |
| RM1 | 1 | (GA)26 | GCGAAAACACAATGCAAAAA | GCGTTGGTTGGACCTGAC | 113 | 6 | 2.721 | 1.250 |
| RM9 | 1 | (GA)15GT(GA)2 | GGTGCCATTGTCGTCCTC | ACGGCCCTCATCACCTTC | 136 | 3 | 1.236 | 0.398 |
| RM14 | 1 | (GA)18 | CCGAGGAGAGGAGTTCGAC | GTGCCAATTTCCTCGAAAAA | 191 | 4 | 2.416 | 1.041 |
| RM23 | 1 | (GA)15 | CATTGGAGTGGAGGCTGG | GTCAGGCTTCTGCCATTCTC | 145 | 3 | 1.903 | 0.733 |
| RM212 | 1 | (CT)24 | CCACTTTCAGCTACTACCAG | CACCCATTTGTCTCTCATTATG | 136 | 2 | 1.403 | 0.462 |
| RM237 | 1 | (CT)18 | CAAATCCCGACTGCTGTCC | TGGGAAGAGAGCACTACAGC | 130 | 2 | 1.492 | 0.512 |
| RM110 | 2 | (GA)15 | TCGAAGCCATCCACCAACGAAG | TCCGTACGCCGACGAGGTCGAG | 156 | 2 | 1.280 | 0.377 |
| RM425 | 2 | (CGG)9 | CCAACGAAGATTCGAAGCTC | CAGCACCATGAAGTCGCC | 126 | 3 | 2.103 | 0.814 |
| RM525 | 2 | (AAG)12 | GGCCCGTCCAAGAAATATTG | CGGTGAGACAGAATCCTTACG | 131 | 4 | 3.034 | 1.179 |
| RM1178 | 2 | (AG)13 | CAGTGGGCGAGCATAGGAG | ATCCTTTTCTCCCTCTCTCG | 112 | 3 | 2.998 | 1.098 |
| RM156 | 3 | (CGG)8 | GCCGCACCCTCACTCCCTCCTC | TCTTGCCGGAGCGCTTGAGGTG | 160 | 3 | 2.090 | 0.879 |
| RM218 | 3 | (TC)24ACT5(GT)11 | TGGTCAAACCAAGGTCCTTC | GACATACATTCTACCCCCGG | 148 | 3 | 1.943 | 0.766 |
| RM489 | 3 | (ATA)8 | ACTTGAGACGATCGGACACC | TCACCCATGGATGTTGTCAG | 271 | 4 | 2.175 | 0.893 |
| RM8209 | 3 | (AG)15 | AGGAGAAGAGGAATCTTTGC | CGATCGAGAGCTACTATTGC | 188 | 3 | 1.043 | 0.117 |
| RM241 | 4 | (CT)31 | GAGCCAAATAAGATCGCTGA | TGCAAGCAGCAGATTTAGTG | 138 | 2 | 1.729 | 0.613 |
| RM470 | 4 | (CTT)14 | TCCTCATCGGCTTCTTCTTC | AGAACCCGTTCTACGTCACG | 83 | 4 | 3.406 | 1.297 |
| RM567 | 4 | (GA)21 | ATCAGGGAAATCCTGAAGGG | GGAAGGAGCAATCACCACTG | 261 | 1 | 1.000 | 0.000 |
| RM5473 | 4 | (TC)20 | ACACGGAGATCCGAGACACGAG | CGAGATTAACGTCGTCCTC | 105 | 5 | 4.117 | 1.493 |
| RM31 | 5 | (GA)15 | GATCACGATCCACTGGAGCT | AAGTCCATTACTCTCCTCCC | 140 | 6 | 2.737 | 1.266 |
| RM188 | 5 | (CA)8 | TCCGCCTCTCCTCTCGCTTCCC | GCAACGCACAACCGAACCGAGC | 210 | 3 | 2.283 | 0.924 |
| RM430 | 5 | (GA)25 | AAACAACGACGTCCCTGATC | GTGCCTCCGTGGTTATGAAC | 173 | 5 | 2.922 | 1.217 |
| RM193 | 6 | (GCT)5 | CGCCTCTTCTTCCTCGCCTCCG | CGGGTCCATCCCCCCTCTCCTC | 189 | 1 | 1.000 | 0.000 |
| RM276 | 6 | (AG)8A3(GA)33 | CTCAACGTTGACACCTCGTG | TCCTCCATCGAGCAGTATCA | 149 | 4 | 2.113 | 0.960 |
| RM402 | 6 | (ATA)7 | GAGCCATGGAAAGATGCATG | TCAGCTGGCCTATGACAATG | 133 | 3 | 1.321 | 0.487 |
| RM469 | 6 | (AG)15 | AGCTGAACAAGCCCTGAAAG | GACTTGGGCAGTGTGACATG | 105 | 2 | 1.465 | 0.498 |
| RM557 | 6 | (AC)13 | GTGGCGAGATCTATGTGGTG | GCTTTGTGTGTGTGTGTGTG | 211 | 1 | 1.000 | 0.000 |
| RM234 | 7 | (CT)25 | ACAGTATCCAAGGCCCTGG | CACGTGAGACAAAGACGGAG | 156 | 2 | 1.729 | 0.613 |
| RM346 | 7 | (CTT)18 | CGAGAGAGCCCATAACTACG | ACAAGACGACGAGGAGGGAC | 175 | 3 | 2.132 | 0.867 |
| RM429 | 7 | (TG)10 | TCCCTCCAGCAATGTCTTTC | CCTTCATCTTGCTTTCCACC | 159 | 3 | 1.876 | 0.744 |
| RM500 | 7 | (AAG)9 | GAGCTTGCCAGAGTGGAAAG | GTTACACCGAGAGCCAGCTC | 259 | 1 | 1.000 | 0.000 |
| RM256 | 8 | (CT)21 | GACAGGGAGTGATTGAAGGC | GTTGATTTCGCCAAGGGC | 127 | 2 | 1.021 | 0.058 |
| RM264 | 8 | GA)27 | GTTGCGTCCTACTGCTACTTC | GATCCGTGTCGATGATTAGC | 178 | 5 | 2.586 | 1.149 |
| RM281 | 8 | (GA)21 | ACCAAGCATCCAGTGACCAG | GTTCTTCATACAGTCCACATG | 138 | 3 | 2.073 | 0.799 |
| RM408 | 8 | (CT)13 | CAACGAGCTAACTTCCGTCC | ACTGCTACTTGGGTAGCTGACC | 128 | 3 | 2.014 | 0.783 |
| RM515 | 8 | (GA)11 | TAGGACGACCAAAGGGTGAG | TGGCCTGCTCTCTCTCTCTC | 211 | 4 | 2.419 | 1.005 |
| RM201 | 9 | (CT)17 | CTCGTTTATTACCTACAGTACC | CTACCTCCTTTCTAGACCGATA | 158 | 2 | 1.741 | 0.617 |
| RM242 | 9 | (CT)26 | GGCCAACGTGTGTATGTCTC | TATATGCCAAGACGGATGGG | 225 | 3 | 1.916 | 0.807 |
| RM257 | 9 | (CT)24 | CAGTTCCGAGCAAGAGTACTC | GGATCGGACGTGGCATATG | 147 | 4 | 3.504 | 1.314 |
| RM566 | 9 | (AG)15 | ACCCAACTACGATCAGCTCG | CTCCAGGAACACGCTCTTTC | 239 | 3 | 1.879 | 0.802 |
| RM216 | 10 | (CT)18 | GCATGGCCGATGGTAAAG | TGTATAAAACCACACGGCCA | 146 | 3 | 1.291 | 0.436 |
| RM228 | 10 | (CA)6(GA)36 | CTGGCCATTAGTCCTTGG | GCTTGCGGCTCTGCTTAC | 154 | 7 | 4.632 | 1.693 |
| RM484 | 10 | (AT)9 | TCTCCCTCCTCACCATTGTC | TGCTGCCCTCTCTCTCTCTC | 299 | 1 | 1.000 | 0.000 |
| RM224 | 11 | (AAG)8(AG)13 | ATCGATCGATCTTCACGAGG | TGCTATAAAAGGCATTCGGG | 157 | 4 | 2.390 | 1.070 |
| RM286 | 11 | (GA)16 | GGCTTCATCTTTGGCGAC | CCGGATTCACGAGATAAACTC | 110 | 4 | 2.484 | 1.066 |
| RM17 | 12 | (GA)21 | TGCCCTGTTATTTTCTTCTCTC | GGTGATCCTTTCCCATTTCA | 184 | 3 | 1.349 | 0.495 |
| RM101 | 12 | (CT)37 | GTGAATGGTCAAGTGACTTAGGTGGC | ACACAACATGTTCCCTCCCATGC | 324 | 3 | 2.563 | 1.001 |
| RM277 | 12 | (GA)11 | CGGTCAAATCATCACCTGAC | CAAGGCTTGCAAGGGAAG | 124 | 2 | 1.929 | 0.675 |
| RM1103 | 12 | (AG)12 | CAGCTGCTGCTACTACACCG | CTACTCCACGTCCATGCATG | 216 | 2 | 1.792 | 0.634 |
| Chr, Chromosome; EPS, Expected PCR product size; Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s information index. | ||||||||
| Origin | Na | Ne | I | No | P (%) | Ho | He | Fis | Fit | Fst | t | |
| Qingpu (Qp), Shanghai, China | 1.833 | 1.560 | 0.422 | 30 | 62.50 | 0.034 | 0.295 | 0.855 | 0.883 | 0.192 | 0.062 | |
| Jiading (Jd), Shanghai, China | 1.458 | 1.300 | 0.228 | 15 | 31.25 | 0.000 | 0.155 | 1.000 | 1.000 | 0.230 | 0.000 | |
| Nanhui (Nh), Shanghai, China | 1.458 | 1.309 | 0.261 | 21 | 43.75 | 0.010 | 0.202 | 0.871 | 0.940 | 0.538 | 0.031 | |
| Pudong (Pd), Shanghai, China | 1.667 | 1.434 | 0.367 | 29 | 60.42 | 0.004 | 0.272 | 0.951 | 0.978 | 0.554 | 0.011 | |
| Chongming (Cm), Shanghai, China | 1.458 | 1.313 | 0.259 | 20 | 41.67 | 0.013 | 0.193 | 0.911 | 0.934 | 0.257 | 0.034 | |
| Fengxian (Fx), Shanghai, China | 1.708 | 1.416 | 0.365 | 29 | 60.42 | 0.050 | 0.262 | 0.708 | 0.785 | 0.263 | 0.120 | |
| Taizhou (Tz), Jiangsu, China | 1.958 | 1.685 | 0.489 | 31 | 64.58 | 0.067 | 0.337 | 0.728 | 0.769 | 0.149 | 0.131 | |
| Shenyang (Sy), Liaoning, China | 1.375 | 1.260 | 0.213 | 17 | 35.42 | 0.008 | 0.158 | 0.893 | 0.939 | 0.424 | 0.031 | |
| Tieling (Tl), Liaoning, China | 1.500 | 1.334 | 0.270 | 20 | 41.67 | 0.004 | 0.195 | 0.963 | 0.976 | 0.340 | 0.012 | |
| Haicheng (Hc), Liaoning, China | 1.500 | 1.308 | 0.266 | 21 | 43.75 | 0.004 | 0.193 | 0.972 | 0.980 | 0.282 | 0.010 | |
| Dandong (Dd), Liaoning, China | 1.458 | 1.349 | 0.269 | 20 | 41.67 | 0.004 | 0.202 | 0.975 | 0.981 | 0.235 | 0.010 | |
| Gongzhuling (Gz), Jilin, China | 1.292 | 1.218 | 0.168 | 12 | 25.00 | 0.000 | 0.124 | 1.000 | 1.000 | 0.241 | 0.000 | |
| Jiamusi (Jm), Heilongjiang, China | 1.438 | 1.337 | 0.258 | 19 | 38.58 | 0.000 | 0.193 | 1.000 | 1.000 | 0.468 | 0.000 | |
| Yesan (Ys), Chungcheong, Korea | 1.458 | 1.269 | 0.231 | 19 | 38.58 | 0.009 | 0.163 | 0.907 | 0.936 | 0.313 | 0.033 | |
| Seosan (Ss), Chungcheong, Korea | 1.542 | 1.27 | 0.267 | 25 | 52.08 | 0.007 | 0.186 | 0.947 | 0.959 | 0.234 | 0.021 | |
| Overall | 3.104 | 2.047 | 0.748 | 43 | 85.58 | 0.012 | 0.434 | 0.931 | 0.973 | 0.606 | 0.014 | |
| Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s diversity index; No, Number of polymorphic loci; P, Percentage of polymorphic loci; Ho, Observed heterozygosity; He, Expected heterozygosity; Fis, Fit and Fst, F-statistics of regional populations referred to | ||||||||||||
Table 4 Genetic characteristics of 76 weedy rice accessions in Northeast Asia by POPGENE according to 48 SSR markers.
| Origin | Na | Ne | I | No | P (%) | Ho | He | Fis | Fit | Fst | t | |
| Qingpu (Qp), Shanghai, China | 1.833 | 1.560 | 0.422 | 30 | 62.50 | 0.034 | 0.295 | 0.855 | 0.883 | 0.192 | 0.062 | |
| Jiading (Jd), Shanghai, China | 1.458 | 1.300 | 0.228 | 15 | 31.25 | 0.000 | 0.155 | 1.000 | 1.000 | 0.230 | 0.000 | |
| Nanhui (Nh), Shanghai, China | 1.458 | 1.309 | 0.261 | 21 | 43.75 | 0.010 | 0.202 | 0.871 | 0.940 | 0.538 | 0.031 | |
| Pudong (Pd), Shanghai, China | 1.667 | 1.434 | 0.367 | 29 | 60.42 | 0.004 | 0.272 | 0.951 | 0.978 | 0.554 | 0.011 | |
| Chongming (Cm), Shanghai, China | 1.458 | 1.313 | 0.259 | 20 | 41.67 | 0.013 | 0.193 | 0.911 | 0.934 | 0.257 | 0.034 | |
| Fengxian (Fx), Shanghai, China | 1.708 | 1.416 | 0.365 | 29 | 60.42 | 0.050 | 0.262 | 0.708 | 0.785 | 0.263 | 0.120 | |
| Taizhou (Tz), Jiangsu, China | 1.958 | 1.685 | 0.489 | 31 | 64.58 | 0.067 | 0.337 | 0.728 | 0.769 | 0.149 | 0.131 | |
| Shenyang (Sy), Liaoning, China | 1.375 | 1.260 | 0.213 | 17 | 35.42 | 0.008 | 0.158 | 0.893 | 0.939 | 0.424 | 0.031 | |
| Tieling (Tl), Liaoning, China | 1.500 | 1.334 | 0.270 | 20 | 41.67 | 0.004 | 0.195 | 0.963 | 0.976 | 0.340 | 0.012 | |
| Haicheng (Hc), Liaoning, China | 1.500 | 1.308 | 0.266 | 21 | 43.75 | 0.004 | 0.193 | 0.972 | 0.980 | 0.282 | 0.010 | |
| Dandong (Dd), Liaoning, China | 1.458 | 1.349 | 0.269 | 20 | 41.67 | 0.004 | 0.202 | 0.975 | 0.981 | 0.235 | 0.010 | |
| Gongzhuling (Gz), Jilin, China | 1.292 | 1.218 | 0.168 | 12 | 25.00 | 0.000 | 0.124 | 1.000 | 1.000 | 0.241 | 0.000 | |
| Jiamusi (Jm), Heilongjiang, China | 1.438 | 1.337 | 0.258 | 19 | 38.58 | 0.000 | 0.193 | 1.000 | 1.000 | 0.468 | 0.000 | |
| Yesan (Ys), Chungcheong, Korea | 1.458 | 1.269 | 0.231 | 19 | 38.58 | 0.009 | 0.163 | 0.907 | 0.936 | 0.313 | 0.033 | |
| Seosan (Ss), Chungcheong, Korea | 1.542 | 1.27 | 0.267 | 25 | 52.08 | 0.007 | 0.186 | 0.947 | 0.959 | 0.234 | 0.021 | |
| Overall | 3.104 | 2.047 | 0.748 | 43 | 85.58 | 0.012 | 0.434 | 0.931 | 0.973 | 0.606 | 0.014 | |
| Na, Observed number of alleles; Ne, Effective number of alleles; I, Shannon’s diversity index; No, Number of polymorphic loci; P, Percentage of polymorphic loci; Ho, Observed heterozygosity; He, Expected heterozygosity; Fis, Fit and Fst, F-statistics of regional populations referred to | ||||||||||||
| Origin | Qp | Jd | Nh | Pd | Cm | Fx | Tz | Sy | Tl | Hc | Dd | Gz | Jm | Ys |
| Jd | 0.078 | |||||||||||||
| Nh | 0.130 | 0.115 | ||||||||||||
| Pd | 0.093 | 0.145 | 0.112 | |||||||||||
| Cm | 0.076 | 0.143 | 0.173 | 0.174 | ||||||||||
| Fx | 0.446 | 0.579 | 0.543 | 0.410 | 0.527 | |||||||||
| Tz | 0.443 | 0.578 | 0.552 | 0.416 | 0.476 | 0.184 | ||||||||
| Sy | 0.738 | 0.889 | 0.832 | 0.622 | 0.777 | 0.254 | 0.145 | |||||||
| Tl | 0.715 | 0.850 | 0.902 | 0.652 | 0.689 | 0.266 | 0.228 | 0.085 | ||||||
| Hc | 0.698 | 0.854 | 0.941 | 0.689 | 0.721 | 0.211 | 0.162 | 0.114 | 0.068 | |||||
| Dd | 0.697 | 0.822 | 0.934 | 0.744 | 0.743 | 0.258 | 0.185 | 0.170 | 0.121 | 0.060 | ||||
| Gz | 0.747 | 0.859 | 0.930 | 0.737 | 0.821 | 0.311 | 0.199 | 0.202 | 0.201 | 0.103 | 0.087 | |||
| Jm | 0.665 | 0.771 | 0.904 | 0.668 | 0.760 | 0.246 | 0.220 | 0.188 | 0.208 | 0.115 | 0.107 | 0.098 | ||
| Ys | 0.721 | 0.849 | 0.863 | 0.687 | 0.828 | 0.228 | 0.220 | 0.157 | 0.180 | 0.115 | 0.138 | 0.106 | 0.053 | |
| Ss | 0.254 | 0.273 | 0.246 | 0.309 | 0.361 | 0.469 | 0.518 | 0.767 | 0.809 | 0.771 | 0.776 | 0.666 | 0.660 | 0.627 |
| Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea. | ||||||||||||||
Table 5 Genetic distance (Nei, 1978) among weedy rice groups in Northeast Asia.
| Origin | Qp | Jd | Nh | Pd | Cm | Fx | Tz | Sy | Tl | Hc | Dd | Gz | Jm | Ys |
| Jd | 0.078 | |||||||||||||
| Nh | 0.130 | 0.115 | ||||||||||||
| Pd | 0.093 | 0.145 | 0.112 | |||||||||||
| Cm | 0.076 | 0.143 | 0.173 | 0.174 | ||||||||||
| Fx | 0.446 | 0.579 | 0.543 | 0.410 | 0.527 | |||||||||
| Tz | 0.443 | 0.578 | 0.552 | 0.416 | 0.476 | 0.184 | ||||||||
| Sy | 0.738 | 0.889 | 0.832 | 0.622 | 0.777 | 0.254 | 0.145 | |||||||
| Tl | 0.715 | 0.850 | 0.902 | 0.652 | 0.689 | 0.266 | 0.228 | 0.085 | ||||||
| Hc | 0.698 | 0.854 | 0.941 | 0.689 | 0.721 | 0.211 | 0.162 | 0.114 | 0.068 | |||||
| Dd | 0.697 | 0.822 | 0.934 | 0.744 | 0.743 | 0.258 | 0.185 | 0.170 | 0.121 | 0.060 | ||||
| Gz | 0.747 | 0.859 | 0.930 | 0.737 | 0.821 | 0.311 | 0.199 | 0.202 | 0.201 | 0.103 | 0.087 | |||
| Jm | 0.665 | 0.771 | 0.904 | 0.668 | 0.760 | 0.246 | 0.220 | 0.188 | 0.208 | 0.115 | 0.107 | 0.098 | ||
| Ys | 0.721 | 0.849 | 0.863 | 0.687 | 0.828 | 0.228 | 0.220 | 0.157 | 0.180 | 0.115 | 0.138 | 0.106 | 0.053 | |
| Ss | 0.254 | 0.273 | 0.246 | 0.309 | 0.361 | 0.469 | 0.518 | 0.767 | 0.809 | 0.771 | 0.776 | 0.666 | 0.660 | 0.627 |
| Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea. | ||||||||||||||
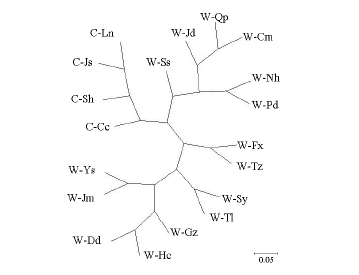
Fig. 2. Unrooted neighbor-joining tree of 15 weedy rice groups and 4 cultivated rice groups by POPGENE, based on a matrix of the inferred genetic diversity. W, Weedy rice; C, Cultivar rice; Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea.
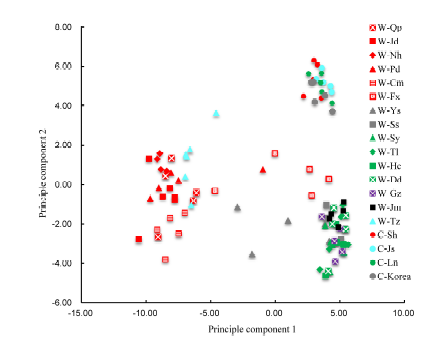
Fig. 4. Two-dimention principle analysis in 96 accessions for Northeast Asia. W, Weedy rice; C, Cultivar rice; Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea.
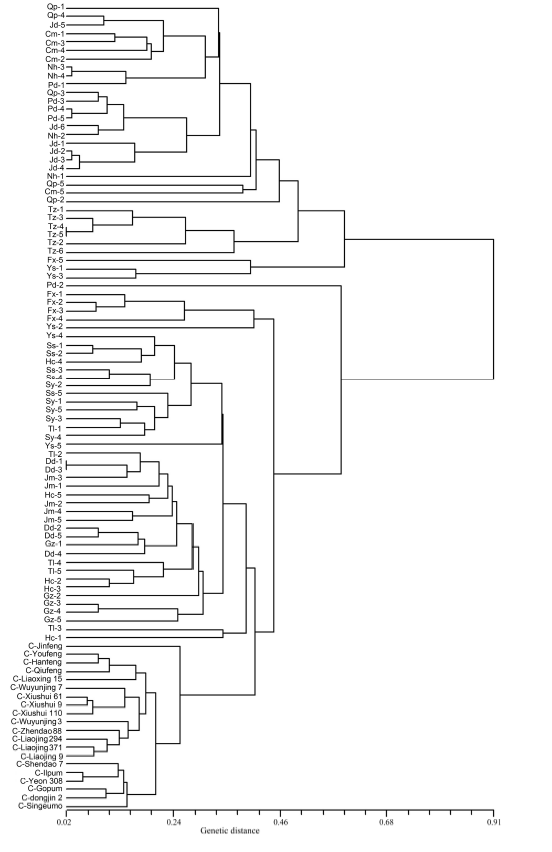

Fig. 3. Cluster diagram of 76 weedy rice and 20 rice accessions in Northeast Asia based on the genetic distance derived from 48 SSR markers, using Nei’s unbiased genetic distance coefficients (Nei, 1978). C, Cultivar rice; Qp, Qingpu, Shanghai, China; Jd, Jiading, Shanghai, China; Nh, Nanhui, Shanghai, China; Pd, Pudong, Shanghai, China; Cm, Chongming, Shanghai, China; Fx, Fengxian, Shanghai, China; Tz, Taizhou, Jiangsu, China; Sy, Shenyang, Liaoning, China; Tl, Tieling, Liaoning, China; Hc, Haicheng, Liaoning, China; Dd, Dandong, Liaoning, China; Gz, Gongzhuling, Jilin, China; Jm, Jiamusi, Heilongjiang, China; Ys, Yesan, Chungcheong, Korea; Ss, Seosan, Chungcheong, Korea.
| 1 | Baker H G.1974. The evolution of weeds.Annu Rev Ecol Syst, 5: 1-24. |
| 2 | Bres-Patry C, Lorieux M, Clément G, Bangratz M, Ghesquière A.2001. Heredity and genetic mapping of domestication-related traits in a temperate japonica weedy rice.Theor Appl Genet, 102(1): 118-126. |
| 3 | Burger J C, Lee S, Ellstrand N C.2006. Origin and genetic structure of feral rye in the western United States.Mol Ecol, 15(9): 2527-2539. |
| 4 | Cao Q J, Lu B R, Xia H, Rong J, Sala F, Spada A, Grassi F.2006. Genetic diversity and origin of weedy rice (Oryza sativa f. spontanea) populations found in north-eastern China revealed by simple sequence repeat (SSR) markers.Ann Bot, 98(6): 1241-1252. |
| 5 | Chen H Z, Xuan S N, Wang W X, Shao G S, Sun Z X.2004. Freezing tolerance and germination ability at low temperature of Dandong weedy rice.Chin J Rice Sci, 18(2): 109-112. (in Chinese with English abstract) |
| 6 | Cho Y C, Chung T Y, Suh H S.1995. Genetic characteristics of Korean weedy rice (Oryza sativa L.) by RFLP analysis.Euphytica, 86(2): 103-110. |
| 7 | Dekker J.1997. Weed diversity and weed management.Weed Sci, 45: 357-363. |
| 8 | Doyle J J, Doyle J L.1987. A rapid DNA isolation procedure for small quantities of flesh leaf tissue.Pytochem Bull, 19: 11-15. |
| 9 | Federici M T, Vaughan D, Tomooka N, Kaga A, Wang X W, Doi K, Francis M, Zorrilla G, Saldain N.2001. Analysis of Uruguayan weedy rice genetic diversity using AFLP molecular markers.Electron J Biotechnol, 4(3): 130-145. |
| 10 | Hartl D L, Clark A G.1989. Principles of Population Genetics. Sunderland, Massachusetts, USA: Sinauer Associate: 682. |
| 11 | Hossain M, Narciso J H.. |
| 12 | Ishikawa R, Toki N, Imai K, Sato Y I, Yamagishi H, Shimamoto Y, Ueno K, Morishima H, Sato T.2005. Origin of weedy rice grown in Bhutan and the force of genetic diversity.Genet Res Crop Evol, 52(4): 395-403. |
| 13 | Jiang H, Wu J L, Wang G L, Wang S.1985. Study on weedy rice from Lianyungang, China.Crop Res China, (2): 4-7. (in Chinese) |
| 14 | Li M B, Ma D R, Xu Z J, Chen W F.2006. Preliminary study on biological trait of Liaoning weedy rice.J Anhui Agric Sci, 34(20): 5224-5225. (in Chinese with English abstract) |
| 15 | Li M B, Ma D R, Xu Z J, Huang C, Piao Z Z, Lee J R, Chen W F.2008. Subspecies characteristics and their relationships with economic characters of weedy rice in Liaoning Province.Liaoning Agric Sci, (4): 1-5. (in Chinese with English abstract) |
| 16 | Li X Y, Qiang S, Song X L, Cai K, Dai W M.2014. Haplotype analysis of Rc gene for weedy rice in Jiangsu Province.Chin J Rice Sci, 28(3): 304-313. (in Chinese with English abstract) |
| 17 | Londo J P, Chiang Y C, Hung K H, Chiang T Y, Schaal B A.2006. Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa.Proc Natl Acad Sci USA, 103(25): 9578-9583. |
| 18 | Ma D R, Chen W F, Xu Z J, Zhang W Z.2005. Origin and control management of weedy rice in Liaoning.Chin Agric Sci Bull, 21(8): 358-360. (in Chinese with English abstract) |
| 19 | Ma D R, Li M B, Wang N, Xu Z J, Chen W F.2008. Genetic diversity and population differentiation of weedy rice in Liaoning Province of China.Acta Agron Sin, 34(3): 403-4l1. (in Chinese with English abstract) |
| 20 | Ma D R, Kong D X, Gao Q, Xu F, Zhao M H, Tang L, Xu Z J, Chen W F.2014. Effects of weedy rice competition on yield and paddy environment of transplanted rice in Northeast China.Chin J Rice Sci, 28(2): 211-216. (in Chinese with English abstract) |
| 21 | Nei M.1978. Estimation of average heterozygosity and genetic distance from a small number of individuals.Genetics, 89(3): 583-590. |
| 22 | Oka H I.1988. Origin of Cultivated Rice. Tokyo: Japan Science Society Press. |
| 23 | Panaud O, Chen X, McCouch S R.1996. Development of microsatellite markers and characterization of simple sequence length polymorphism (SSLP) in rice (Oryza sativa L.).Mol Gen Genet, 252(5): 597-607. |
| 24 | Pyšek P, Prach K.2003. Research into plant invasions in a cross-roads region: History and focus.Biol Invas, 5(4): 337-348. |
| 25 | Rohlf F J.1998. NTSYSpc: Numerical Taxonomy and Multivariate Analysis System, Version 2_02. Setauket, New York: Exeter Software. |
| 26 | Suh H S, Sato Y I, Morishima H.1997. Genetic characterization of weedy rice (Oryza sativa L.) based on morpho-physiology, isozymes and RAPD markers.Theor Appl Genet, 94(3/4): 316-321. |
| 27 | Wright S.1978. Evolution and the Genetics of Population: Variability Within and Among Natural Populations. Chicago: University of Chicago Press. |
| 28 | Xu C, Wu W C.1996. Ecological investigating and identifying of weedy rices in Hainan Island.Chin J Rice Sci, 10(4): 247-249. (in Chinese with English abstract) |
| 29 | Yang S R, Zhao J S.1959. A research on indica-japonica hybridization.Agric Sci, 10: 256-268. (in Chinese) |
| 30 | Yeh F C, Yang R C, Boyle T.1999. Microsoft Window-Based Freeware for Population Genetic Analysis (POPGENE), Version 1.31. |
| 31 | Yu G Q, Bao Y, Shi C H, Dong C Q, Ge S.2005. Genetic diversity and population differentiation of Liaoning weedy rice detected by RAPD and SSR markers.Biochem Genet, 43(5/6): 261-270. |
| 32 | Yu L Q, Mortimer A M, Xuan S N, Lu Y L, Zhou Y J.2005. Stress-resistance of weedy rice Luolijing (Oryza sativa).Chin J Appl Ecol, 16(4): 717-720. (in Chinese with English abstract) |
| 33 | Yuan Q H, Shi L, Wang F, Cao B, Qian Q, Lei X M, Liao Y L, Liu W G, Cheng L, Jia S R.2007. Investigation of rice transgene flow in compass sectors by using male sterile line as a pollen detector.Theor Appl Genet, 115(4): 549-560. |
| 34 | Zhang J, Burgos N R, Ma K, Zhou Y J, Geng R M, Yu L Q.2008. Genetic diversity and relationship of weedy rice in Taizhou City, Jiangsu Province, China.Chin J Rice Sci, 15(4): 295-302. (in Chinese with English abstract) |
| 35 | Zhang Z L, Tan X L, Deng A F.2002. Comprehensive evaluation of weedy rice germplasm.J Plant Genet Res, 3(4): 47-50. (in Chinese with English abstract) |
| 36 | (Managing Editor: Fang Hongmin) |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||