
Rice Science ›› 2021, Vol. 28 ›› Issue (4): 358-367.DOI: 10.1016/j.rsci.2021.05.006
收稿日期:2020-11-12
接受日期:2021-01-28
出版日期:2021-07-28
发布日期:2021-07-28
. [J]. Rice Science, 2021, 28(4): 358-367.
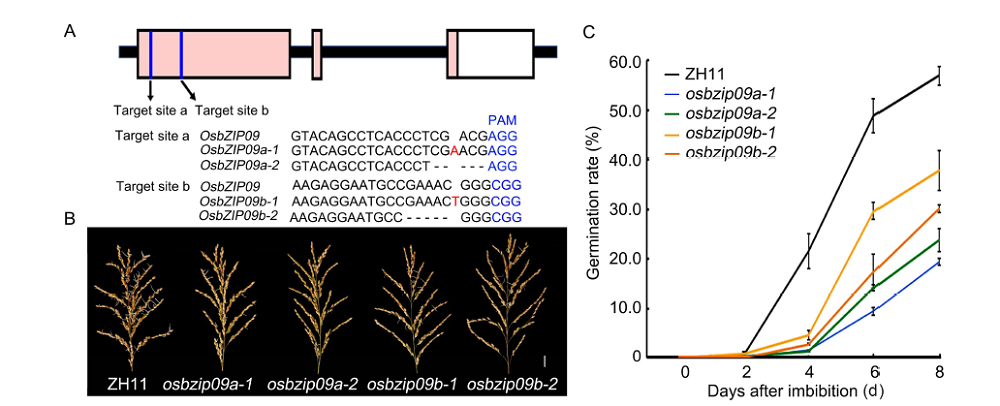
Fig. 1. Knock-out of OsbZIP09 alleviates rice pre-harvest sprouting (PHS).A, Schematic diagram of the two targets in OsbZIP09 gene and the mutation information of the related mutants. The protospacer adjacent motif (PAM) is highlighted in blue, insert nucleotide is highlighted in red, and deleted nucleotides are shown with minus sign ‘-’. B, Germination phenotype of mature rice panicles after 6 d imbibition in water. Scale bar, 2 cm. ZH11, Zhonghua 11.C, Time-course analysis of germination rates of seeds in the panicles. Data are Mean ± SD (n = 30).
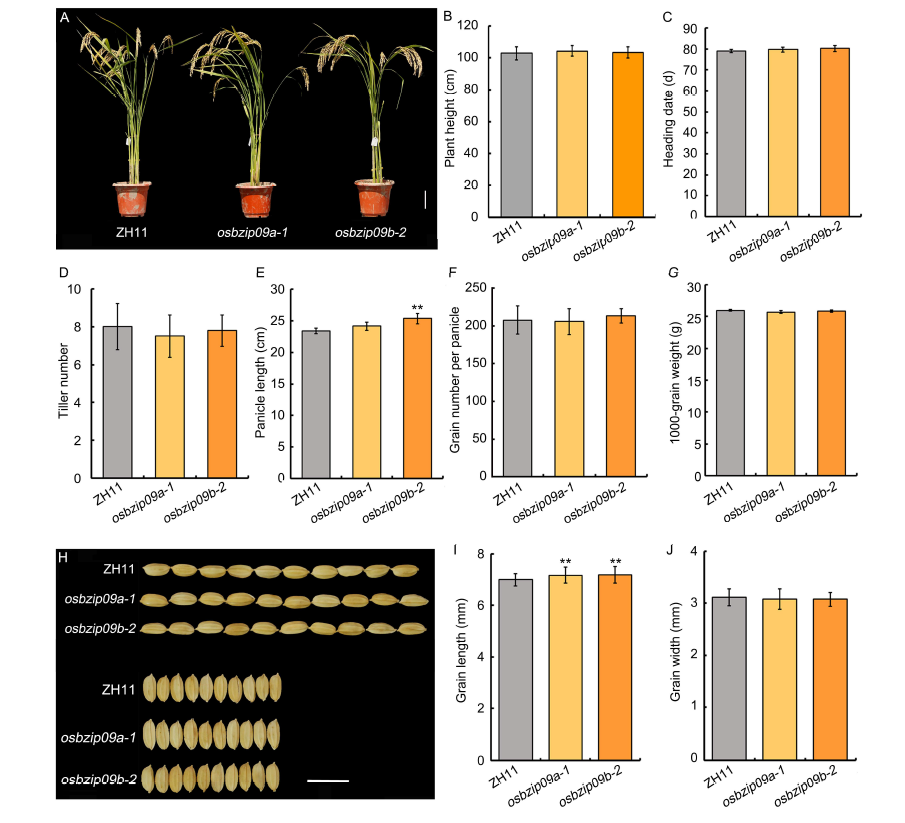
Fig. 2. Agronomic traits and normal seed germination assay of osbzip09 mutants.A, Morphology of osbzip09 mutants and its wild type Zhonghua 11 (ZH11). Scale bar, 10 cm. B?J, Plant height (B), heading date (C), tiller number per plant (D), panicle length (E), grain number per panicle (F), 1000-grain weight (G), grain shape (scale bar, 1 cm) (H), grain length (I) and grain width (J) of osbzip09 mutants and wild type. Data are Mean ± SD (n = 10 in B?G, and n = 30 in I and J). **, P < 0.01 (t-test).
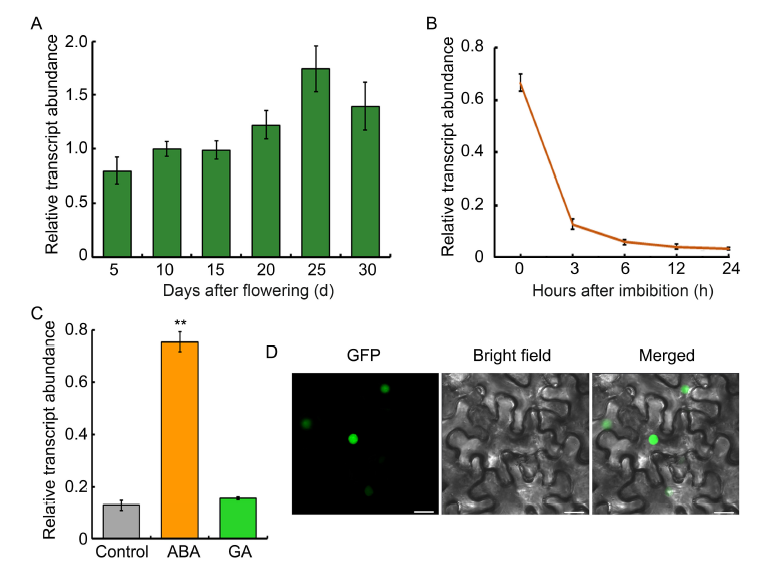
Fig. 3. Expression and sub-cellular localization analysis of OsbZIP09.A, Expression profiling of OsbZIP09 transcript abundance in developing seeds of rice. B, Time-course expression analysis of OsbZIP09 after imbibition. C, Expression analysis of OsbZIP09 in response to abscisic acid (ABA, 5 μmol/L) and gibberellins (GA, 10 μmol/L) treatments. D, Sub-cellular localization of OsbZIP09 (35S::OsbZIP09-GFP) in tobacco epidermal cells. Scale bars, 20 μm. Actin served as an internal reference gene for normalization in A?C. Data are Mean ± SD (n = 3). **, P < 0.01 (t-test).
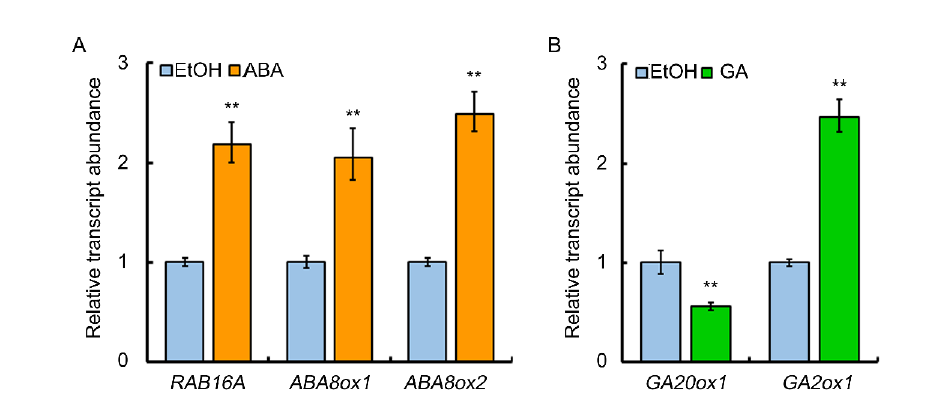
Fig. S2. Expression analysis of ABA or GA responsive marker genes.A, Expression analysis of RAB16A, ABA8ox1 and ABA8ox2 in response to ABA (5 μmol/L) treatment.B, Expression analysis of GA20ox1 and GA2ox1 in response to GA (10 μmol/L). Actin served as an internal reference gene for normalisation in A and B. Data are Mean ± SD (n = 3). ** indicates significant difference (P < 0.01, t-test).
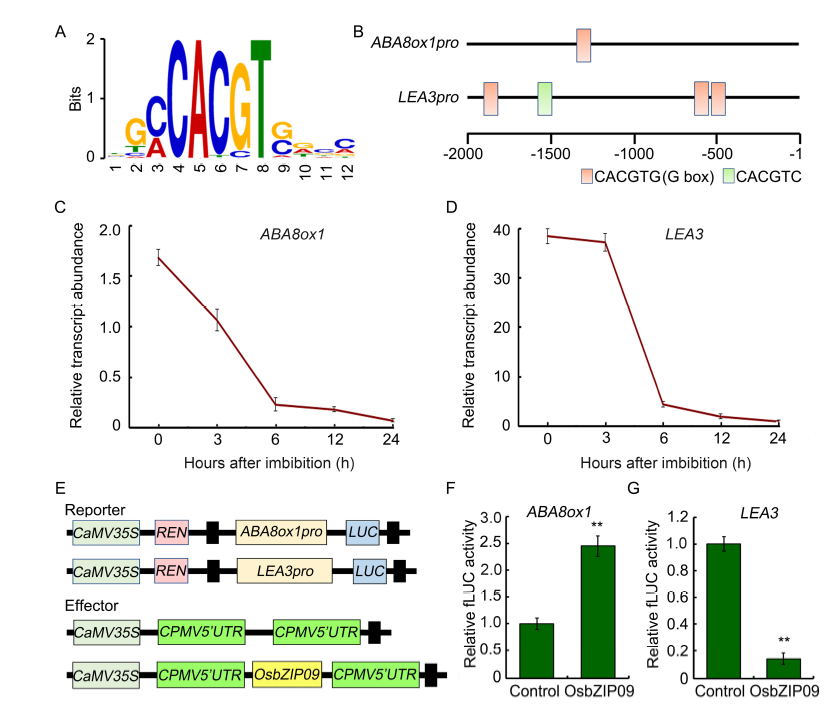
Fig. 4. Identification of OsbZIP09 binding motif and its targets involved in regulation of rice seed germination.A, Conserved motif identified from the DNA affinity purification sequencing (DAP-seq) data. B, Distribution of OsbZIP09 binding motifs in the 2-kb region preceding the initial codon (ATG) of ABA8ox1 and LEA3. C and D, Time-course expression analysis of ABA8ox1 and LEA3 after imbibition. Data are Mean ± SD (n = 3).E, Schematic program of the reporter and effector constructs used in the dual-luciferase reporter assay. F and G, OsbZIP09 activates the promoter activity of ABA8ox1 but suppresses the promoter activity of LEA3. Data are Mean ± SD (n = 5). ** indicates significant difference (P < 0.01, t-test).
| Chromosome | Start | End | -log10(P-value)' | Strand | Distance_ to_TSS | Category | Gene ID | Gene name |
|---|---|---|---|---|---|---|---|---|
| 5 | 26927280 | 26927615 | 7.60524 | + | 3 | Promoter | Os05g0542500 | OsLEA3; Similar to isoform 2 of late embryogenesis abundant protein 2C group 3. |
| 2 | 28994350 | 28994666 | 8.95194 | + | 1052 | Promoter | Os02g0703600 | OsABA8ox1; Similar to abscisic acid 8'-hydroxylase 1. (Os02t0703600-01) |
| 5 | 4663664 | 4663921 | 6.50438 | - | -1100 | Promoter | Os05g0178100 | OsGA3ox1;3beta-hydroxylase GA metabolism (Os05t0178100-01) |
Table S1. DAP-seq data related to ABA8ox1, LEA3 and GA3ox1.
| Chromosome | Start | End | -log10(P-value)' | Strand | Distance_ to_TSS | Category | Gene ID | Gene name |
|---|---|---|---|---|---|---|---|---|
| 5 | 26927280 | 26927615 | 7.60524 | + | 3 | Promoter | Os05g0542500 | OsLEA3; Similar to isoform 2 of late embryogenesis abundant protein 2C group 3. |
| 2 | 28994350 | 28994666 | 8.95194 | + | 1052 | Promoter | Os02g0703600 | OsABA8ox1; Similar to abscisic acid 8'-hydroxylase 1. (Os02t0703600-01) |
| 5 | 4663664 | 4663921 | 6.50438 | - | -1100 | Promoter | Os05g0178100 | OsGA3ox1;3beta-hydroxylase GA metabolism (Os05t0178100-01) |
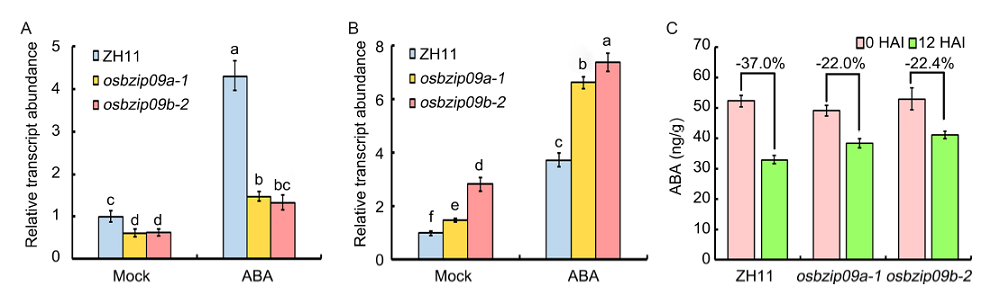
Fig. 5. Expression analysis of ABA8ox1 and LEA3 in rice seeds before and after imbibition of abscisic acid (ABA).A and B, Expression analysis of ABA8ox1 (A) and LEA3 (B) in osbzip09 mutants and wild type Zhonghua 11 (ZH11) with or without ABA treatment. Actin served as an internal reference gene for normalization. Different lowercase letters indicate significant differences at P < 0.05 level based on one-way analysis of variance. C, ABA content in osbzip09 mutants and ZH11 before or at 12 h after imbibition (HAI).
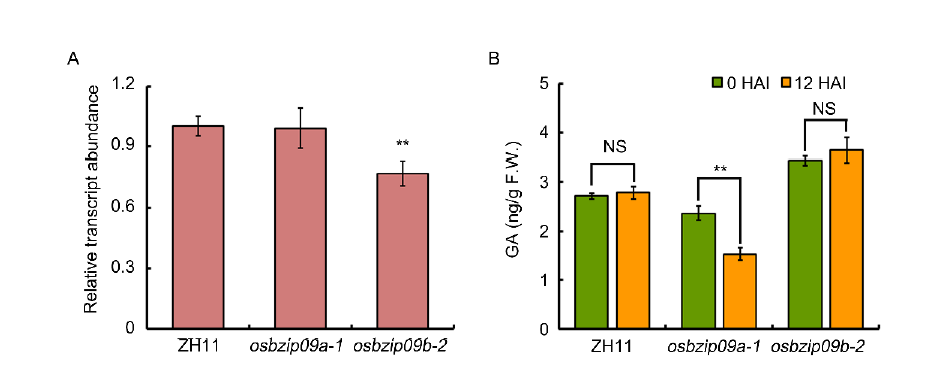
Fig. S3. Expression analysis of GA3ox1 and quantification of GA3 content in rice seeds before and after imbibition. A, Expression analysis of GA3ox1 in osbzip09 mutants and ZH11. Actin served as an internal reference gene for normalisation. B, GA3 content in osbzip09 mutants and ZH11 before or at 12 h after imbibition (HAI). Data are Mean ± SD (n = 3). ** indicates significant difference (P < 0.01, t-test). NS, Not significant.
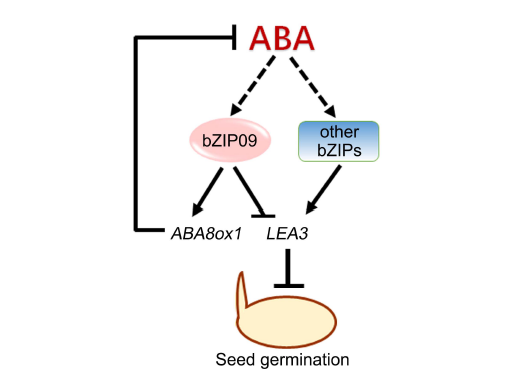
Fig. 6. Simplified model showing OsbZIP09 promotes rather than suppresses seed germination via reducing abscisic acid (ABA) accumulation and attenuating ABA-induction of downstream responsive genes in rice.
| Primer name | Sequence ( 5' → 3' ) | Usage |
|---|---|---|
| OsbZIP09a-SG-seq | GTACAGCCTCACCCTCGACG | SG sequence-1 for editing OsbZIP09 SG sequence-2 for editing OsbZIP09 |
| OsbZIP09b-SG-seq | AAGAGGAATGCCGAAACGGG | |
| OsbZIP09seq-F | ATCTCGTCTCGATCATGGGG | Primers for sequencing and confirming mutation of OsbZIP09 |
| OsbZIP09seq-R | CCCCAGCTTTGACCAGGAAA | |
| bZIP09qRT-F | GAGGCATGTCAGGGGATGTG | For qRT-PCR analysis of OsbZIP09 |
| bZIP09qRT-R | ACTTGGGTTCTGGTACAGGC | |
| RAB16AqRT-F | ATGAGGGAGGAGCACAAGAC | For qRT-PCR analysis of RAB16A |
| RAB16AqRT-R | TTGATCCCCTTCTTCCTCCT | |
| ABA8ox1qRT-F | AAGCTGGCAAAACCAACATC | For qRT-PCR analysis of ABA8ox1 |
| ABA8ox1qRT-R | TGGATTCCGTATTAGCACGG | |
| ABA8ox2qRT-F | GCGAGACGCTCCAGCTCT | For qRT-PCR analysis of ABA8ox2 |
| ABA8ox2qRT-R | GGGCACCCCAGCAGATT | |
| GA20ox1qRT-F | GCCACTACAGGGCCGACAT | For qRT-PCR analysis of GA20ox1 |
| GA20ox1qRT-R | TGGTTGCAGGTGACGATGAT | |
| GA2ox1qRT-F | TGACGATGATGACAGCGACAA | For qRT-PCR analysis of GA2ox1 |
| GA2ox1qRT-R | CCATAGGCATCGTCTGCAATT | |
| LEA3qRT-F | TCACTTCAAATTCGGTGCAA | For qRT-PCR analysis of LEA3 |
| LEA3qRT-R | CACACCCGTCAGAAATCCTC | |
| Actin1qRT-F | CCAAGGCCAATCGTGAGAAGA | Reference gene for qRT-PCR assay |
| Actin1qRT-R | AATCAGTGAGATCACGCCCAG | |
| pZIP09-Smal-F | TCCCCCGGGATGGCGTCGAAGGCCGGAG | For subcellular assay of OsbZIP09 |
| pZIP09-XbaI-R | GCTCTAGAGAAATCTGCGGAGCTTGTTC | |
| pZIP09-EcoRI-F | CGGAATTCATGGCGTCGAAGGCCGGAG | For construction of OsbZIP09-62-SK |
| pZIP09-KpnI-R | GGGGTACCTCAGAAATCTGCGGAGCTTG | |
| proLEA3Kpn-F | GGGGTACCGTCGGCCACCCTCCGCAATA | For construction of LEA3pro-pGreenII |
| proLEA3PstI-R | AACTGCAGACCTGCCCCGCCTTCTCCTG | |
| proABA8ox1Kpn-F | GGGGTACCGTAGCCTGTTGGTAGCTTGG | For construction of ABA8ox1pro-pGreenII |
| proABA8ox1PstI-R | AACTGCAGCGCCTGCTCACTTCACTTAT | |
| DAP-seq Adapter A | CACGACGCTCTTCCGATCT | For DAP-seq sampling |
| DAP-seq Adapter B | GATCGGAAGAGCACACGTCTG |
Table S2. Oligonucleotide primer sequences used.
| Primer name | Sequence ( 5' → 3' ) | Usage |
|---|---|---|
| OsbZIP09a-SG-seq | GTACAGCCTCACCCTCGACG | SG sequence-1 for editing OsbZIP09 SG sequence-2 for editing OsbZIP09 |
| OsbZIP09b-SG-seq | AAGAGGAATGCCGAAACGGG | |
| OsbZIP09seq-F | ATCTCGTCTCGATCATGGGG | Primers for sequencing and confirming mutation of OsbZIP09 |
| OsbZIP09seq-R | CCCCAGCTTTGACCAGGAAA | |
| bZIP09qRT-F | GAGGCATGTCAGGGGATGTG | For qRT-PCR analysis of OsbZIP09 |
| bZIP09qRT-R | ACTTGGGTTCTGGTACAGGC | |
| RAB16AqRT-F | ATGAGGGAGGAGCACAAGAC | For qRT-PCR analysis of RAB16A |
| RAB16AqRT-R | TTGATCCCCTTCTTCCTCCT | |
| ABA8ox1qRT-F | AAGCTGGCAAAACCAACATC | For qRT-PCR analysis of ABA8ox1 |
| ABA8ox1qRT-R | TGGATTCCGTATTAGCACGG | |
| ABA8ox2qRT-F | GCGAGACGCTCCAGCTCT | For qRT-PCR analysis of ABA8ox2 |
| ABA8ox2qRT-R | GGGCACCCCAGCAGATT | |
| GA20ox1qRT-F | GCCACTACAGGGCCGACAT | For qRT-PCR analysis of GA20ox1 |
| GA20ox1qRT-R | TGGTTGCAGGTGACGATGAT | |
| GA2ox1qRT-F | TGACGATGATGACAGCGACAA | For qRT-PCR analysis of GA2ox1 |
| GA2ox1qRT-R | CCATAGGCATCGTCTGCAATT | |
| LEA3qRT-F | TCACTTCAAATTCGGTGCAA | For qRT-PCR analysis of LEA3 |
| LEA3qRT-R | CACACCCGTCAGAAATCCTC | |
| Actin1qRT-F | CCAAGGCCAATCGTGAGAAGA | Reference gene for qRT-PCR assay |
| Actin1qRT-R | AATCAGTGAGATCACGCCCAG | |
| pZIP09-Smal-F | TCCCCCGGGATGGCGTCGAAGGCCGGAG | For subcellular assay of OsbZIP09 |
| pZIP09-XbaI-R | GCTCTAGAGAAATCTGCGGAGCTTGTTC | |
| pZIP09-EcoRI-F | CGGAATTCATGGCGTCGAAGGCCGGAG | For construction of OsbZIP09-62-SK |
| pZIP09-KpnI-R | GGGGTACCTCAGAAATCTGCGGAGCTTG | |
| proLEA3Kpn-F | GGGGTACCGTCGGCCACCCTCCGCAATA | For construction of LEA3pro-pGreenII |
| proLEA3PstI-R | AACTGCAGACCTGCCCCGCCTTCTCCTG | |
| proABA8ox1Kpn-F | GGGGTACCGTAGCCTGTTGGTAGCTTGG | For construction of ABA8ox1pro-pGreenII |
| proABA8ox1PstI-R | AACTGCAGCGCCTGCTCACTTCACTTAT | |
| DAP-seq Adapter A | CACGACGCTCTTCCGATCT | For DAP-seq sampling |
| DAP-seq Adapter B | GATCGGAAGAGCACACGTCTG |
| [1] | Alves M S, Dadalto S P, Gonçalves A B, De Souza G B, Barros V A, Fietto L G. 2013. Plant bZIP transcription factors responsive to pathogens: A review. Int J Mol Sci, 14(4): 7815-7828. |
| [2] | Blauwkamp T A, Chang M V, Cadigan K M. 2008. Novel TCF- binding sites specify transcriptional repression by Wnt signalling. EMBO J, 27: 1436-1446. |
| [3] | Chen K, Li G J, Bressan R A, Song C P, Zhu J K, Zhao Y. 2020. Abscisic acid dynamics, signaling, and functions in plants. J Integr Plant Biol, 62(1): 25-54. |
| [4] | Cheng X Y, Wu Y, Guo J P, Du B, Chen R Z, Zhu L L, He G C. 2013. A rice lectin receptor-like kinase that is involved in innate immune responses also contributes to seed germination. Plant J, 76(4): 687-698. |
| [5] | Dröge-Laser W, Snoek B L, Snel B, Weiste C. 2018. TheArabidopsis bZIP transcription factor family: An update. Curr Opin Plant Biol, 45: 36-49. |
| [6] | Dröge-Laser W, Weiste C. 2018. The C/S 1 bZIP network: A regulatory hub orchestrating plant energy homeostasis. Trends Plant Sci, 23(5): 422-433. |
| [7] | Finkelstein R, Reeves W, Ariizumi T, Steber C. 2008. Molecular aspects of seed dormancy. Annu Rev Plant Biol, 59: 387-415. |
| [8] | Hellens R P, Allan A C, Friel E N, Bolitho K, Grafton K, Templeton M D, Karunairetnam S, Gleave A P, Laing W A. 2005. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods, 1: 13. |
| [9] | Hossain M A, Cho J I, Han M, Ahn C H, Jeon J S, An G, Park P B. 2010. The ABRE-binding bZIP transcription factor OsABF2 is a positive regulator of abiotic stress and ABA signaling in rice. J Plant Physiol, 167(17): 1512-1520. |
| [10] | Jacobsen J V, Pearce D W, Poole A T, Pharis R P, Mander L N. 2002. Abscisic acid, phaseic acid and gibberellin contents associated with dormancy and germination in barley. Physiol Plant, 115(3): 428-441. |
| [11] | Joo J, Lee Y H, Song S I. 2019. OsbZIP42 is a positive regulator of ABA signaling and confers drought tolerance to rice. Planta, 249(5): 1521-1533. |
| [12] | Kawakatsu T, Takaiwa F. 2010. Differences in transcriptional regulatory mechanisms functioning for free lysine content and seed storage protein accumulation in rice grain. Plant Cell Physiol, 51(12): 1964-1974. |
| [13] | Kawakatsu T, Yamamoto M P, Touno S M, Yasuda H, Takaiwa F. 2009. Compensation and interaction between RISBZ1 and RPBF during grain filling in rice. Plant J, 59(6): 908-920. |
| [14] | Langmead B, Salzberg S L. 2012. Fast gapped-read alignment with Bowtie 2. Nat Methods, 9(4): 357-359. |
| [15] | Li Q F, Zhou Y, Xiong M, Ren X Y, Han L, Wang J D, Zhang C Q, Fan X L, Liu Q Q. 2020. Gibberellin recovers seed germination in rice with impaired brassinosteroid signalling. Plant Sci, 293: 110435. |
| [16] | Lu G J, Gao C X, Zheng X N, Han B. 2009. Identification of OsbZIP72 as a positive regulator of ABA response and drought tolerance in rice. Planta, 229(3): 605-615. |
| [17] | MacDonald B T, Tamai K, He X. 2009. Wnt/beta-catenin signaling: Components, mechanisms, and diseases. Dev Cell, 17: 9-26. |
| [18] | Machanick P, Bailey T L. 2011. MEME-ChIP: Motif analysis of large DNA datasets. Bioinformatics, 27(12): 1696-1697. |
| [19] | Martínez-Andújar C, Ordiz M I, Huang Z, Nonogaki M, Beachy R N, Nonogaki H. 2011. Induction of 9-cis-epoxycarotenoid dioxygenase in Arabidopsis thaliana seeds enhances seed dormancy. Proc Natl Acad Sci USA, 108(41): 17225-17229. |
| [20] | Miyakawa T, Fujita Y, Yamaguchi-Shinozaki K, Tanokura M. 2013. Structure and function of abscisic acid receptors. Trends Plant Sci, 18(5): 259-266. |
| [21] | Née G, Xiang Y, Soppe W J. 2017. The release of dormancy, a wake-up call for seeds to germinate. Curr Opin Plant Biol, 35: 8-14. |
| [22] | Nijhawan A, Jain M, Tyagi A K, Khurana J P. 2008. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol, 146(2): 333-350. |
| [23] | Nonogaki H. 2019. Seed germination and dormancy: The classic story, new puzzles, and evolution. J Integr Plant Biol, 61(5): 541-563. |
| [24] | Nonogaki M, Sall K, Nambara E, Nonogaki H. 2014. Amplification of ABA biosynthesis and signaling through a positive feedback mechanism in seeds. Plant J, 78(3): 527-539. |
| [25] | O’Malley R C, Huang S C, Song L, Lewsey M G, Bartlett A, Nery J R, Galli M, Gallavotti A, Ecker J R. 2016. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell, 165(5): 1280-1292. |
| [26] | Onodera Y, Suzuki A, Wu C Y, Washida H, Takaiwa F. 2001. A rice functional transcriptional activator, RISBZ1, responsible for endosperm-specific expression of storage protein genes through GCN4 motif. J Biol Chem, 276(17): 14139-14152. |
| [27] | Saika H, Okamoto M, Miyoshi K, Kushiro T, Shinoda S, Jikumaru Y, Fujimoto M, Arikawa T, Takahashi H, Ando M, Arimura S I, Miyao A, Hirochika H, Kamiya Y, Tsutsumi N, Nambara E, Nakazono M. 2007. Ethylene promotes submergence-induced expression of OsABA8ox1, a gene that encodes ABA 8′- hydroxylase in rice. Plant Cell Physiol, 48(2): 287-298. |
| [28] | Shu K, Liu X D, Xie Q, He Z H. 2016. Two faces of one seed: Hormonal regulation of dormancy and germination. Mol Plant, 9(1): 34-45. |
| [29] | Skubacz A, Daszkowska-Golec A, Szarejko I. 2016. The role and regulation of ABI5 (ABA-insensitive 5) in plant development, abiotic stress responses and phytohormone crosstalk. Front Plant Sci, 7: 1884. |
| [30] | Song S, Wang G F, Wu H, Fan X W, Liang L W, Zhao H, Li S L, Hu Y, Liu H Y, Ayaad M, Xing Y Z. 2020. OsMFT2 is involved in the regulation of ABA signaling mediated seed germination through interacting with OsbZIP23/66/72 in rice. Plant J, 103(2): 532-546. |
| [31] | Sornaraj P, Luang S, Lopato S, Hrmova M. 2016. Basic leucine zipper (bZIP) transcription factors involved in abiotic stresses: A molecular model of a wheat bZIP factor and implications of its structure in function. Biochim Biophy Acta: Genral Subjects, 1860: 46-56. |
| [32] | Sun Y, Fan X Y, Cao D M, Tang W, He K, Zhu J Y, He J X, Bai M Y, Zhu S, Oh E, Patil S, Kim T W, Ji H, Wong W H, Rhee S Y, Wang Z Y. 2010. Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev Cell, 19(5): 765-777. |
| [33] | Tong H N, Chu C C. 2018. Functional specificities of brassinosteroid and potential utilization for crop improvement. Trends Plant Sci, 23(11): 1016-1028. |
| [34] | Tong H N, Xiao Y H, Liu D P, Gao S P, Liu L C, Yin Y H, Jin Y, Qian Q, Chu C C. 2014. Brassinosteroid regulates cell elongation by modulating gibberellin metabolism in rice. Plant Cell, 26(11): 4376-4393. |
| [35] | Tuan P A, Kumar R, Rehal P K, Toora P K, Ayele B T. 2018. Molecular mechanisms underlying abscisic acid/gibberellin balance in the control of seed dormancy and germination in cereals. Front Plant Sci, 9: 668. |
| [36] | Wang C, Liu Q, Shen Y, Hua Y F, Wang J J, Lin J R, Wu M G, Sun T T, Cheng Z K, Mercier R, Wang K. 2019. Clonal seeds from hybrid rice by simultaneous genome engineering of meiosis and fertilization genes. Nat Biotechnol, 37(3): 283-286. |
| [37] | Wang Q, Lin Q B, Wu T, Duan E C, Huang Y S, Yang C Y, Mou C L, Lan J, Zhou C L, Xie K, Liu X, Guo X P, Wang J, Jiang L, Wan J M. 2020. OsDOG1L-3 regulates seed dormancy through the abscisic acid pathway in rice. Plant Sci, 298: 110570. |
| [38] | Wang Y F, Hou Y X, Qiu J H, Wang H M, Wang S, Tang L Q, Tong X H, Zhang J. 2020. Abscisic acid promotes jasmonic acid biosynthesis via a ‘SAPK10-bZIP72-AOC’ pathway to synergistically inhibit seed germination in rice (Oryza sativa). New Phytol, 228(4): 1336-1353. |
| [39] | Wu J H, Zhu C F, Pang J H, Zhang X R, Yang C L, Xia G X, Tian Y C, He C Z. 2014. OsLOL1, a C2C2-type zinc finger protein, interacts with OsbZIP58 to promote seed germination through the modulation of gibberellin biosynthesis in Oryza sativa. Plant J, 80(6): 1118-1130. |
| [40] | Xu H, Li X F, Zhang H, Wang L C, Zhu Z G, Gao J P, Li C S, Zhu Y. 2020. High temperature inhibits the accumulation of storage materials by inducing alternative splicing of OsbZIP58 during filling stage in rice. Plant Cell Environ, 43(8): 1879-1896. |
| [41] | Xu X, Wan W, Jiang G, Xi Y, Huang H, Cai J, Chang Y, Duan C G, Mangrauthia S K, Peng X, Zhu J K, Zhu G. 2019. Nucleocytoplasmic trafficking of theArabidopsis WD40 repeat protein XIW1 regulates ABI5 stability and abscisic acid responses. Mol Plant, 12(12): 1598-1611. |
| [42] | Yadukrishnan P, Datta S. 2020. Light and abscisic acid interplay in early seedling development. New Phytol, 229: 763-769. |
| [43] | Yamamoto M P, Onodera Y, Touno S M, Takaiwa F. 2006. Synergism between RPBF Dof and RISBZ1 bZIP activators in the regulation of rice seed expression genes. Plant Physiol, 141(4): 1694-1707. |
| [44] | Yan A, Chen Z. 2017. The pivotal role of abscisic acid signaling during transition from seed maturation to germination. Plant Cell Rep, 36(5): 689-703. |
| [45] | Yang J C, Zhang J H, Wang Z Q, Zhu Q S, Wang W. 2001. Hormonal changes in the grains of rice subjected to water stress during grain filling. Plant Physiol, 127(1): 315-323. |
| [46] | Yang W Q, Zhang W, Wang X X. 2017. Post-translational control of ABA signalling: The roles of protein phosphorylation and ubiquitination. Plant Biotechnol J, 15(1): 4-14. |
| [47] | Yang X, Yang Y N, Xue L J, Zou M J, Liu J Y, Chen F, Xue H W. 2011. Rice ABI5-like1 regulates abscisic acid and auxin responses by affecting the expression of ABRE-containing genes. Plant Physiol, 156(3): 1397-1409. |
| [48] | Yang Y M, Xu C N, Wang B M, Jia J Z. 2001. Effects of plant growth regulators on secondary wall thickening of cotton fibres. Plant Growth Regul, 35: 233-237. |
| [49] | Yu F F, Wu Y R, Xie Q. 2015. Precise protein post-translational modifications modulate ABI5 activity. Trends Plant Sci, 20(9): 569-575. |
| [50] | Zhang H Y, He H, Wang X C, Wang X F, Yang X Z, Li L, Deng X W. 2011. Genome-wide mapping of the HY5-mediated gene networks in Arabidopsis that involve both transcriptional and post-transcriptional regulation. Plant J, 65(3): 346-358. |
| [51] | Zhang Y, Liu T, Meyer C A, Eeckhoute J, Johnson D S, Bernstein B E, Nusbaum C, Myers R M, Brown M, Li W, Liu X S. 2008. Model- based analysis of ChIP-Seq (MACS). Genome Biol, 9(9): R137. |
| [52] | Zhao H Y, Nie K L, Zhou H P, Yan X J, Zhan Q D, Zheng Y, Song C P. 2020. ABI5 modulates seed germination via feedback regulation of the expression of the PYR/PYL/RCAR ABA receptor genes. New Phytol, 228(2): 596-608. |
| [53] | Zou M J, Guan Y C, Ren H B, Zhang F, Chen F. 2008. A bZIP transcription factor, OsABI5, is involved in rice fertility and stress tolerance. Plant Mol Biol, 66(6): 675-683. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||