
Rice Science ›› 2022, Vol. 29 ›› Issue (3): 247-256.DOI: 10.1016/j.rsci.2021.07.012
收稿日期:2021-06-17
接受日期:2021-07-12
出版日期:2022-05-28
发布日期:2022-03-10
. [J]. Rice Science, 2022, 29(3): 247-256.
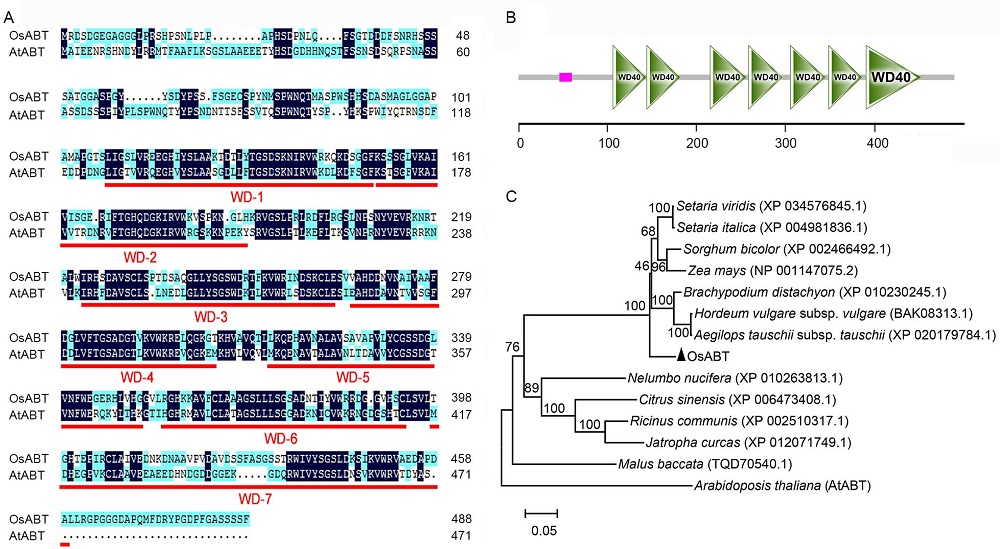
Fig. 1. Protein sequence analysis of OsABT and phylogenetic relationships of OsABT and its homologous proteins in plants. A, Multiple sequence alignment of OsABT and AtABT. WD-1 to WD-7 show the amino acid sequences of seven tryptophan-aspartate (WD) domains. B, Domain analysis of OsABT protein. The pink solid rectangle represents low compositional complexity, and the ruler represents the number of amino acids. C, Phylogenetic relationships of OsABT and its homologs in other plant species. The phylogenetic tree was generated in MEGA 6.0. Bootstrap values were 1 000 replications.
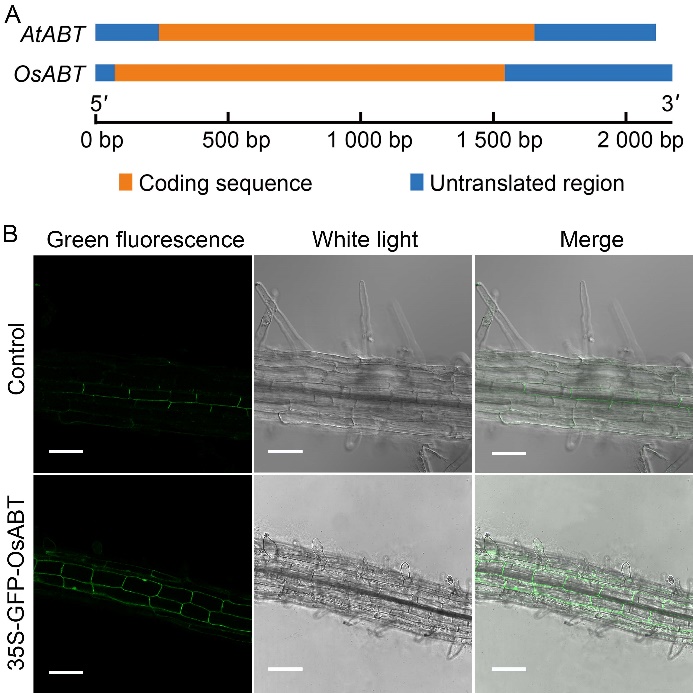

Fig. 2. Gene structure of OsABT and subcellular localization of OsABT protein. A, Comparison of gene structure between OsABT and AtABT. B, Protein location analysis of OsABT in Arabidoposis roots using confocal laser scanning microscopy. The three top images show free green fluorescence protein (GFP) signal, and the three subsequent images show GFP-OsABT fusion proteins. Scale bars, 20 μm.
| cis-element | Sequence | Position from ATG | Biological function |
|---|---|---|---|
| ABRELATERD1 | ACGTG | -1851, -1321 | ABA responsive element |
| ABRERATCAL | MACGYGB | -1694 | ABA responsive element |
| EBOXBNNAPA | CANNTG | -1264, -1089, -781, -762, -590 | ABA responsive element |
| DRE2COREZMRAB17 | ACCGAC | -232 | ABA responsive element, dehydration-responsive element |
| PROXBBNNAPA | CAAACACC | -846 | ABA responsive element |
| GT1CONSENSUS | GRWAAW | -515 | Salt responsive element |
| MYB1AT | WAACCA | -459, -401 | Dehydration-responsive element |
| MYB2AT | TAACTG | -353 | Dehydration-responsive element |
| MYB2CONSENSUSAT | YAACKG | -1669, -353, -169 | Dehydration-responsive element |
| MYCCONSENSUSAT | CANNTG | -1827, -1264, -1089, -781, -762, -590 | Dehydration-responsive element |
| DRECRTCOREAT | RCCGAC | -1581, -880, -232 | Dehydration-responsive element |
| CBFHV | RYCGAC | -1581, -880, -232 | Dehydration-responsive element |
| ACGTATERD1 | ACGT | -1858, -1852, -1322, -995 | Required for etiolation-induced expression of erd1 (early responsive to dehydration) |
| LTRECOREATCOR15 | CCGAC | -1581, -1566, -1035, -880, -840, -232 | Low-temperature-responsive element |
| LTRE1HVBLT49 | CCGAAA | -517 | Low-temperature-responsive element |
| CCAATBOX1 | CCAAT | -876, -377 | Heat stress response element |
| BIHD1OS | TGTCA | -1556, -653, -336 | Binding site of OsBIHD1 in disease resistance response |
Table 1. cis-elements that respond to abscisic acid (ABA) and stress in OsABT promoter.
| cis-element | Sequence | Position from ATG | Biological function |
|---|---|---|---|
| ABRELATERD1 | ACGTG | -1851, -1321 | ABA responsive element |
| ABRERATCAL | MACGYGB | -1694 | ABA responsive element |
| EBOXBNNAPA | CANNTG | -1264, -1089, -781, -762, -590 | ABA responsive element |
| DRE2COREZMRAB17 | ACCGAC | -232 | ABA responsive element, dehydration-responsive element |
| PROXBBNNAPA | CAAACACC | -846 | ABA responsive element |
| GT1CONSENSUS | GRWAAW | -515 | Salt responsive element |
| MYB1AT | WAACCA | -459, -401 | Dehydration-responsive element |
| MYB2AT | TAACTG | -353 | Dehydration-responsive element |
| MYB2CONSENSUSAT | YAACKG | -1669, -353, -169 | Dehydration-responsive element |
| MYCCONSENSUSAT | CANNTG | -1827, -1264, -1089, -781, -762, -590 | Dehydration-responsive element |
| DRECRTCOREAT | RCCGAC | -1581, -880, -232 | Dehydration-responsive element |
| CBFHV | RYCGAC | -1581, -880, -232 | Dehydration-responsive element |
| ACGTATERD1 | ACGT | -1858, -1852, -1322, -995 | Required for etiolation-induced expression of erd1 (early responsive to dehydration) |
| LTRECOREATCOR15 | CCGAC | -1581, -1566, -1035, -880, -840, -232 | Low-temperature-responsive element |
| LTRE1HVBLT49 | CCGAAA | -517 | Low-temperature-responsive element |
| CCAATBOX1 | CCAAT | -876, -377 | Heat stress response element |
| BIHD1OS | TGTCA | -1556, -653, -336 | Binding site of OsBIHD1 in disease resistance response |
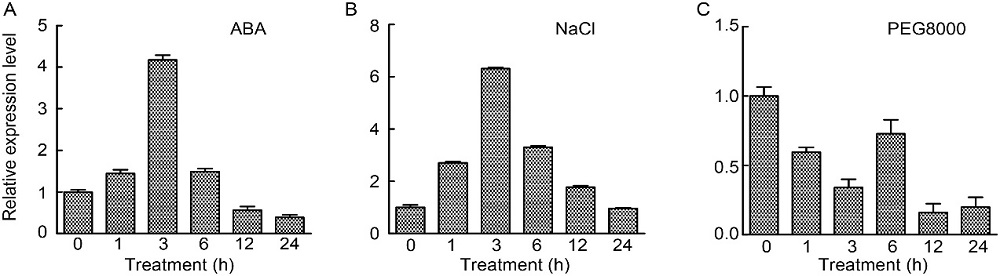
Fig. 3. Expression profiles of OsABT under stress. A?C, Expression patterns of OsABT in response to abscisic acid (ABA) (A), NaCl stress (B) and PEG8000 treatment (drought simulation) (C). RNA was extracted from rice roots at 0 (control), 1, 3, 6, 12 and 24 h after treatments with ABA (50 μmol/L), NaCl (150 mmol/L) and PEG8000 (10 g/mL). OseEF-1α was used as an internal reference. Data are Mean ± SD (n = 3).
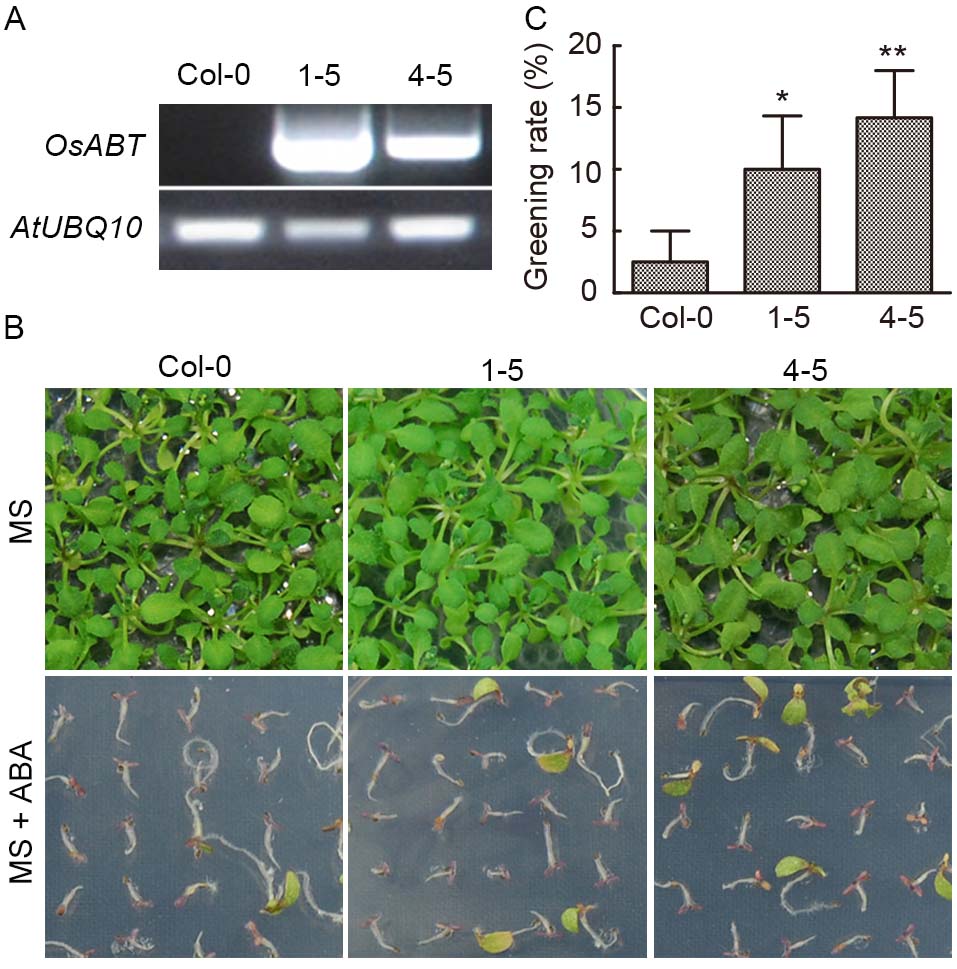
Fig. 4. Response of transgenic OsABT Arabidopsis lines (1-5 and 4-5) to abscisic acid (ABA). A, OsABT was detected in Col-0 and transgenic lines of Arabidopsis, with AtUBQ10 as an internal reference. B, Analysis of the greening rate of transgenic OsABT Arabidopsis on Murashige and Skoog (MS) medium and MS medium containing ABA (1 μmol/L). C, Statistic analysis of greening rate of transgenic lines at 14 d after germination. Data are Mean ± SD (n = 3). Asterisks represent significant differences between transgenic Arabidopsis lines and Col-0 plants (Student’s t-test, *, P < 0.05; **, P < 0.01).
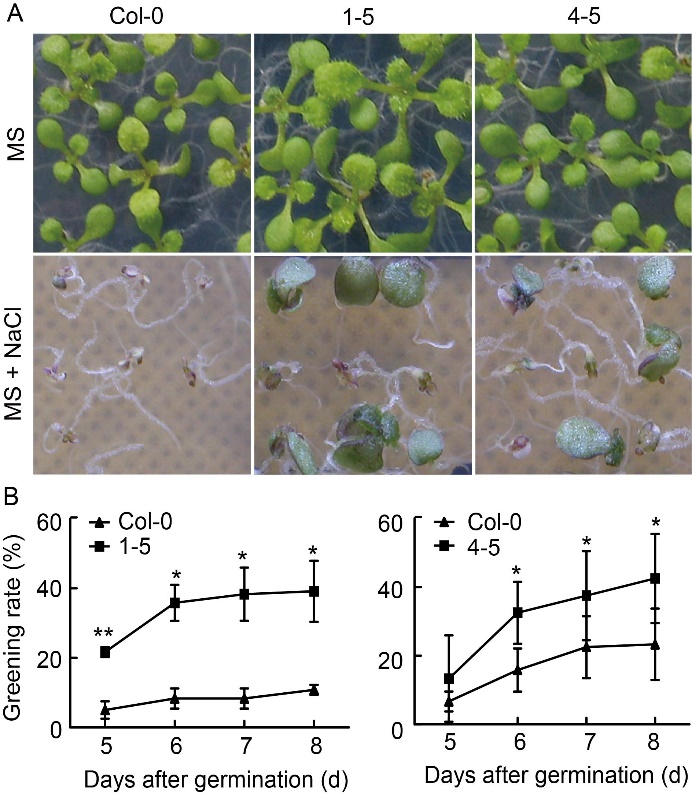
Fig. 5. Response of transgenic OsABT Arabidopsis to salt stress. A, Greening rate analysis of transgenic lines 1-5 and 4-5 on Murashige and Skoog (MS) medium and MS medium containing NaCl (125 mmol/L). B, Statistical analysis of greening rate of transgenic lines 1-5 and 4-5 at 5, 6, 7 and 8 d after germination, respectively. Data are Mean ± SD (n = 3). Significant differences compared with Col-0 (Student’s t-test): *, P < 0.05; **, P < 0.01.

Fig. 6. Response of transgenic Arabidopsis overexpression OsABT lines (1-5 and 4-5) to drought stress. A, Eleven-day-old seedlings of Col-0 and transgenic Arabidopsis overexpression lines were transplanted into different pots and grew for 14 d (before drought stress). Then, water was withheld for 2 weeks (after drought stress), after which the plants were rewatered for 1 week (after water recovery). The photos were taken at the time of before drought stress, after drought stress and after water recovery, respectively. B, Survival rates of transgenic lines 1-5 and 4-5 after drought stress compared with Col-0, respectively. Three independent biological replicates were applied. Data are Mean ± SD (n = 48, Student’s t-test, *, P < 0.05).
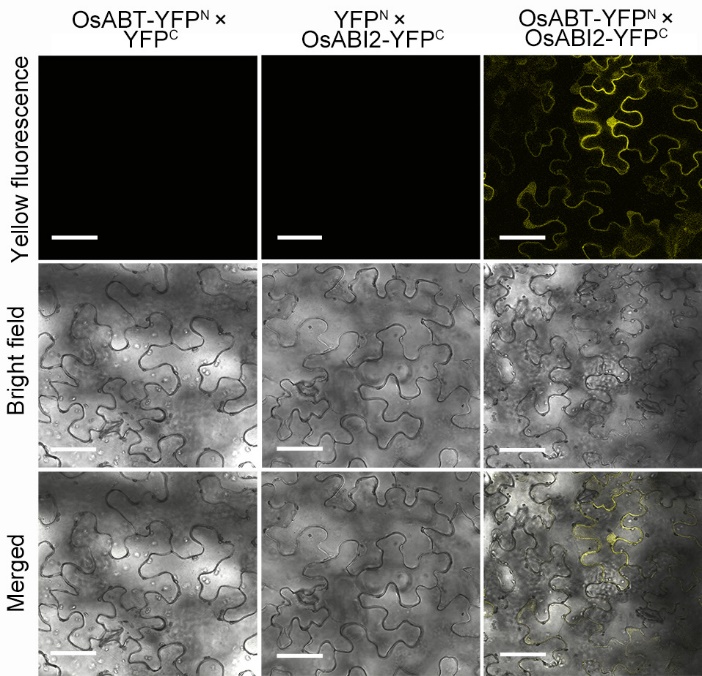
Fig. 7. Interaction between OsABT and OsABI2 as measured via bimolecular fluorescent complimentary assay. OsABT-YFPN and OsABI2-YFPC were co-localized in Nicotiana benthamiana leaf cells. Scale bars, 20 μm.
| [1] | Chen E Y, Zhang X Y, Yang Z E, Wang X Q, Yang Z R, Zhang C J, Wu Z X, Kong D P, Liu Z, Zhao G, Butt H I, Zhang X L, Li F G. 2017. Genome-wide analysis of the HD-ZIP IV transcription factor family in Gossypium arboreum and GaHDG11 involved in osmotic tolerance in transgenic Arabidopsis. Mol Genet Genomics, 292(3): 593-609. |
| [2] | Chinnusamy V, Zhu J H, Zhu J K. 2006. Salt stress signaling and mechanisms of plant salt tolerance. Genet Eng, 27: 141-177. |
| [3] | Chothia C, Hubbard T, Brenner S, Barns H, Murzin A. 1997. Protein folds in the all-beta and all-alpha classes. Annu Rev Biophys Biomol Struct, 26(1): 597-627. |
| [4] |
Clough S J, Bent A F. 1998. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J, 16(6): 735-743.
PMID |
| [5] | Cutler S R, Rodriguez P L, Finkelstein R R, Abrams S R. 2010. Abscisic acid: Emergence of a core signaling network. Annu Rev Plant Biol, 61(1): 651-679. |
| [6] | Dong H P, Yu H Q, Bao Z L, Guo X J, Peng J L, Yao Z, Chen G Y, Qu S P, Dong H S. 2005. The ABI2-dependent abscisic acid signalling controls HrpN-induced drought tolerance in Arabidopsis. Planta, 221(3): 313-327. |
| [7] |
Fülöp V, Böcskei Z, Polgár L. 1998. Prolyl oligopeptidase: An unusual beta-propeller domain regulates proteolysis. Cell, 94(2): 161-170.
PMID |
| [8] | Gong X Q, Zhang J Y, Hu J B, Wang W, Wu H, Zhang Q H, Liu J H. 2015. FcWRKY70, a WRKY protein of Fortunella crassifolia, functions in drought tolerance and modulates putrescine synthesis by regulating arginine decarboxylase gene. Plant Cell Environ, 38(11): 2248-2262. |
| [9] |
Gosti F, Beaudoin N, Serizet C, Webb A A, Vartanian N, Giraudat J. 1999. ABI1 protein phosphatase 2C is a negative regulator of abscisic acid signaling. Plant Cell, 11(10): 1897-1909.
PMID |
| [10] |
Hirayama T, Shinozaki K. 2007. Perception and transduction of abscisic acid signals: Keys to the function of the versatile plant hormone ABA. Trends Plant Sci, 12(8): 343-351.
PMID |
| [11] | Hoagland D R, Arnon D I. 1950. The Water-Culture Method for Growing Plants Without Soil. The College of Agriculture University of California, Berkeley, USA: 1-32. |
| [12] | Huang J, Wang M M, Bao Y M, Sun S J, Pan L J, Zhang H S. 2008. SRWD: A novel WD40 protein subfamily regulated by salt stress in rice (Oryza sativa L.). Gene, 424(1/2): 71-79. |
| [13] | Hubbard K E, Nishimura N, Hitomi K, Getzoff E D, Schroeder J I. 2010. Early abscisic acid signal transduction mechanisms: Newly discovered components and newly emerging questions. Genes Dev, 24(16): 1695-1708. |
| [14] |
Juhász T, Szeltner Z, Fülöp V, Polgár L. 2005. Unclosed beta- propellers display stable structures: Implications for substrate access to the active site of prolyl oligopeptidase. J Mol Biol, 346(3): 907-917.
PMID |
| [15] |
Kawasaki S, Borchert C, Deyholos M, Wang H, Brazille S, Kawai K, Galbraith D, Bohnert H J. 2001. Gene expression profiles during the initial phase of salt stress in rice. Plant Cell, 13(4): 889-905.
PMID |
| [16] | Kong D J, Li M J, Dong Z H, Ji H T, Li X. 2015. Identification of TaWD40D, a wheat WD40 repeat-containing protein that is associated with plant tolerance to abiotic stresses. Plant Cell Rep, 34(3): 395-410. |
| [17] | Kuhn J M, Boisson-Dernier A, Dizon M B, Maktabi M H, Schroeder J I. 2006. The protein phosphatase AtPP2CA negatively regulates abscisic acid signal transduction in Arabidopsis, and effects of abh1 on AtPP2CA mRNA. Plant Physiol, 140(1): 127-139. |
| [18] |
Leung J, Merlot S, Giraudat J. 1997. The Arabidopsis ABSCISIC ACID-INSENSITIVE2 (ABI2) and ABI1 genes encode homologous protein phosphatases 2C involved in abscisic acid signal transduction. Plant Cell, 9(5): 759-771.
PMID |
| [19] | Li Y J, Wang B, Dong R R, Hou B K. 2015. AtUGT76C2, an Arabidopsis cytokinin glycosyltransferase is involved in drought stress adaptation. Plant Sci, 236: 157-167. |
| [20] | Liu W C, Li Y H, Yuan H M, Zhang B L, Zhai S, Lu Y T. 2017. WD40-REPEAT 5a functions in drought stress tolerance by regulating nitric oxide accumulation in Arabidopsis. Plant Cell Environ, 40(4): 543-552. |
| [21] |
Miller G, Suzuki N, Rizhsky L, Hegie A, Koussevitzky S, Mittler R. 2007. Double mutants deficient in cytosolic and thylakoid ascorbate peroxidase reveal a complex mode of interaction between reactive oxygen species, plant development, and response to abiotic stresses. Plant Physiol, 144(4): 1777-1785.
PMID |
| [22] | Mishra A K, Puranik S, Bahadur R P, Prasad M. 2012. The DNA- binding activity of an AP2 protein is involved in transcriptional regulation of a stress-responsive gene, SiWD40, in foxtail millet. Genomics, 100(4): 252-263. |
| [23] | Neer E J, Schmidt C J, Nambudripad R, Smith T F. 1994. The ancient regulatory-protein family of WD-repeat proteins. Nature, 371: 297-300. |
| [24] | Ohta M, Guo Y, Halfter U, Zhu J K. 2003. A novel domain in the protein kinase SOS2 mediates interaction with the protein phosphatase 2C ABI2. Proc Natl Acad Sci USA, 100(20): 11771-11776. |
| [25] | Qiu Q S, Guo Y, Dietrich M A, Schumaker K S, Zhu J K. 2002. Regulation of SOS1, a plasma membrane Na+/H+ exchanger in Arabidopsis thaliana, by SOS2 and SOS3. Proc Natl Acad Sci USA, 99(12): 8436-8441. |
| [26] |
Rodriguez P L, Benning G, Grill E. 1998. ABI2, a second protein phosphatase 2C involved in abscisic acid signal transduction in Arabidopsis. FEBS Lett, 421(3): 185-190.
PMID |
| [27] | Sheen J. 1998. Mutational analysis of protein phosphatase 2C involved in abscisic acid signal transduction in higher plants. Proc Natl Acad Sci USA, 95(3): 975-980. |
| [28] |
Shinozaki K, Yamaguchi-Shinozaki K. 2007. Gene networks involved in drought stress response and tolerance. J Exp Bot, 58(2): 221-227.
PMID |
| [29] |
Smith T F, Gaitatzes C, Saxena K, Neer E J. 1999. The WD repeat: A common architecture for diverse functions. Trends Biochem Sci, 24(5): 181-185.
PMID |
| [30] | Sondek J, Bohm A, Lambright D G, Hamm H E, Sigler P B. 1996. Crystal structure of a G-protein beta gamma dimer at 2.1Å resolution. Nature, 379: 369-374. |
| [31] |
Stirnimann C U, Petsalaki E, Russell R B, Müller C W. 2010. WD40 proteins propel cellular networks. Trends Biochem Sci, 35(10): 565-574.
PMID |
| [32] | van Nocker S, Ludwig P. 2003. The WD-repeat protein superfamily in Arabidopsis: Conservation and divergence in structure and function. BMC Genomics, 4(1): 50. |
| [33] |
Wakasugi M, Kawashima A, Morioka H, Linn S, Sancar A, Mori T, Nikaido O, Matsunaga T. 2002. DDB accumulates at DNA damage sites immediately after UV irradiation and directly stimulates nucleotide excision repair. J Biol Chem, 277(3): 1637-1640.
PMID |
| [34] | Wang Z J, Ji H T, Yuan B J, Wang S F, Su C, Yao B, Zhao H T, Li X. 2015. ABA signalling is fine-tuned by antagonistic HAB1 variants. Nat Commun, 6: 8138. |
| [35] | Wang Z J, Ren Z Y, Cheng C H, Wang T, Ji H T, Zhao Y, Deng Z P, Zhi L Y, Lu J J, Wu X Y, Xu S M, Cao M M, Zhao H T, Liu L, Zhu J K, Li X. 2020. Counteraction of ABA-mediated inhibition of seed germination and seedling establishment by ABA signaling terminator in Arabidopsis. Mol Plant, 13: 1284-1297. |
| [36] | Xu C, Min J R. 2011. Structure and function of WD40 domain proteins. Protein Cell, 2(3): 202-214. |
| [37] | Zhang D P, Wang Y Z, Shen J Y, Yin J F, Li D H, Gao Y, Xu W F, Liang J S. 2018. OsRACK1A, encodes a circadian clock- regulated WD40 protein, negatively affect salt tolerance in rice. Rice, 11(1): 45. |
| [38] | Zhu J H, Jeong J C, Zhu Y M, Sokolchik I, Miyazaki S, Zhu J K, Hasegawa P M, Bohnert H J, Shi H Z, Yun D J, Bressan R A. 2008. Involvement of Arabidopsis HOS15 in histone deacetylation and cold tolerance. Proc Natl Acad Sci USA, 105(12): 4945-4950. |
| [39] |
Zhu J K. 2001. Plant salt tolerance. Trends Plant Sci, 6(2): 66-71.
PMID |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||