
Rice Science ›› 2022, Vol. 29 ›› Issue (6): 535-544.DOI: 10.1016/j.rsci.2022.04.001
收稿日期:2022-01-07
接受日期:2022-04-24
出版日期:2022-11-28
发布日期:2022-09-09
. [J]. Rice Science, 2022, 29(6): 535-544.
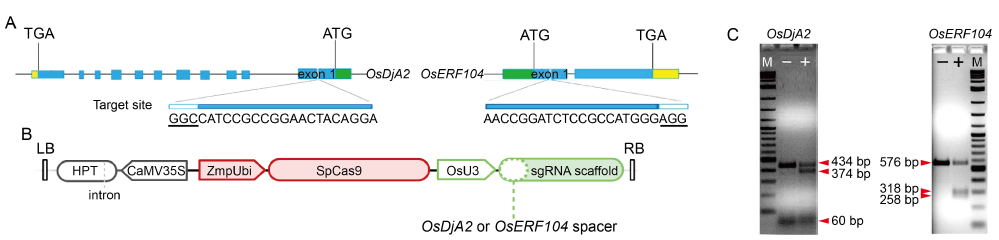
Fig. 1. CRISPR/Cas9 design and T7EI assay for sgRNA gene-editing activity. A, Schematic map of gRNA target sites on genomic regions of OsDjA2 and OsERF104. Exons are indicated as blue boxes, interspaced by introns shown as lines. Promoter and transcription termination sites are represented by green and yellow boxes, respectively. Protospacer adjacent motif is underlined and represented as white boxes. ATG and TGA represent start codon and stop codon, respectively. B, Simplified schematic representation of CRISPR/Cas9 T-DNA structure. LB and RB, T-DNA left and right borders, respectively; HPT, Hygromycin resistance gene; CaMV35S, Cauliflower mosaic virus 35S promoter; ZmpUbi, Maize ubiquitin promoter; SpCas9, Streptococcus pyogenes Cas9 gene; OsU3, Oryza sativa PolII U3 promoter sequence. C, Assessment of gRNA cleavage activity of rice protoplast genomic DNA via T7EI assay. ‘-’ means non-cleaved PCR product derived from wild type protoplast transformed with a control plasmid; ‘+’ means cleaved PCR product derived from protoplasts transformed with CRISPR/Cas9 final vector. M, Marker.
| Parameter | OsDjA2 (LOC_Os02g56040) | OsERF104 (LOC_Os08g36920) |
|---|---|---|
| No. of total T0 transgenic plants | 24 | 24 |
| No. of total T0 transgenic plants with T-DNA PCR positive | 23 | 23 |
| No. of total T0 transgenic plants harboring 1-2 copies of T-DNA | 15 | 17 |
| No. of total T0 transgenic plants harboring site-mutations (Proportion, %) | 14 (93.3%) | 12 (70.6%) |
| Biallelic mutation (Proportion, %) | 8 (57.1%) | 5 (41.6%) |
| Homozygous mutation (Proportion, %) | 5 (35.7%) | 6 (50.0%) |
| Heterozygous mutation (Proportion, %) | 1 (7.1%) | 1 (8.3%) |
Table 1. Efficiency of CRISPR/Cas9-mediated genome editing of target genes and ratios of mutant genotypes in T0 plants.
| Parameter | OsDjA2 (LOC_Os02g56040) | OsERF104 (LOC_Os08g36920) |
|---|---|---|
| No. of total T0 transgenic plants | 24 | 24 |
| No. of total T0 transgenic plants with T-DNA PCR positive | 23 | 23 |
| No. of total T0 transgenic plants harboring 1-2 copies of T-DNA | 15 | 17 |
| No. of total T0 transgenic plants harboring site-mutations (Proportion, %) | 14 (93.3%) | 12 (70.6%) |
| Biallelic mutation (Proportion, %) | 8 (57.1%) | 5 (41.6%) |
| Homozygous mutation (Proportion, %) | 5 (35.7%) | 6 (50.0%) |
| Heterozygous mutation (Proportion, %) | 1 (7.1%) | 1 (8.3%) |
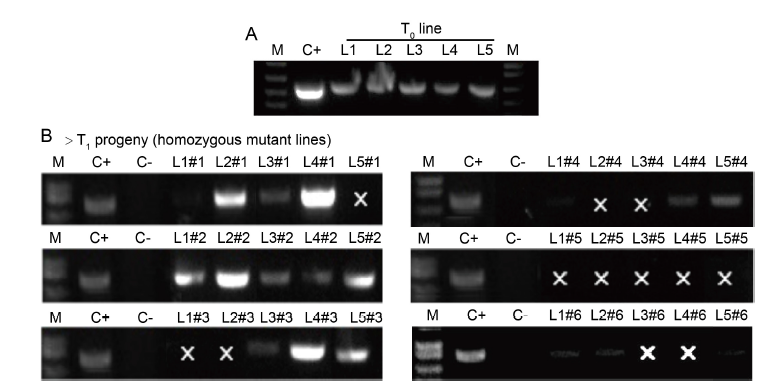
Fig. 2. PCR-based screening for presence of T-DNA in rice mutant plants. A, T0 homozygous primary transformants L1 (OsDjA2_20.1), L2 (OsDjA2_24.1), L3 (OsERF104_1.1), L4 (OsERF104_5.1) and L5 (OsERF104_6.1). B, T1 progeny plants (n = 6 of each independent mutant line as #1, #2, #3, #4, #5 and #6), using specific Cas9 primer pair. M, DNA molecular ladder; C+, CRISPR plasmid; C-, Genomic DNA of wild type Nipponbare; ‘×’ indicates PCR negative for T-DNA.
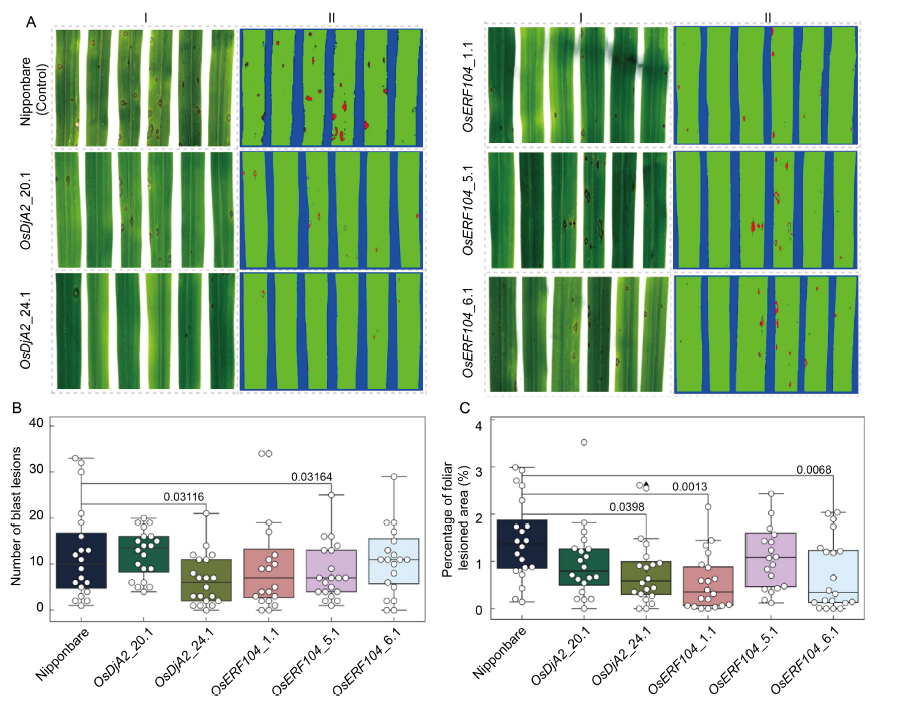
Fig. 3. Identification of blast resistance in CRISPR/Cas9-edited rice mutant plants. A, Phenotypes upon blast infection of wild type Nipponbare, and T0 homozygous mutant plants (OsDjA2_20.1 and OsDjA2_24.1; OsERF104_1.1, OsERF104_5.1 and OsERF104_6.1) of each target gene. The fourth leaves of each line were detached at 7 d post-infection, scanned and analyzed for the number of blast lesions (A-I) and the percentage of the lesioned foliar area (A-II) using the software Quant®. B and C, Boxplot merged with swarmplot data representation for the number of blast lesions (B) and the percentage of foliar lesioned area (C), respectively, observed on each of the 20 leaves. The numbers above the boxplot indicate statistical significance (P < 0.05, two-sample t-test).
| [1] | Abiri R, Shaharuddin N A, Maziah M, Yusof Z N B, Atabaki N, Sahebi M, Valdiani A, Kalhori N, Azizi P, Hanafi M M. 2017. Role of ethylene and the APETALA 2/ethylene response factor superfamily in rice under various abiotic and biotic stress conditions. Environ Exp Bot, 134: 33-44. |
| [2] | Ahn S W, Seshu D V. 1991. Blast reaction of durably resistance rice cultivar in multiplication trials. Phytopathology, 81(10): 1150. |
| [3] |
Bes M, Herbert L, Mounier T, Meunier A C, Durandet F, Guiderdoni E, Périn C. 2021. Efficient genome editing in rice protoplasts using CRISPR/CAS9 construct. Methods Mol Biol, 2238: 173-191.
PMID |
| [4] | Bevitori R, Sircar S, de Mello R N, Togawa R C, Côrtes M V C B, Oliveira T S, Grossi-de-Sá M F, Parekh N. 2020. Identification of co-expression gene networks controlling rice blast disease during an incompatible reaction. Genet Mol Res, 19(3): gmr18579. |
| [5] | Bonman J M, Khush G S, Nelson R J. 1992. Breeding rice for resistance to pests. Annu Rev Phytopathol, 30: 507-528. |
| [6] | Boston R S, Viitanen P V, Vierling E. 1996. Molecular chaperones and protein folding in plants. Plant Mol Biol, 32(1/2): 191-222. |
| [7] |
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, van Daelen R, van der Lee T, Diergaarde P, Groenendijk J, Töpsch S, Vos P, Salamini F, Schulze-Lefert P. 1997. The barley Mlo gene: A novel control element of plant pathogen resistance. Cell, 88(5): 695-705.
PMID |
| [8] | Choi Y, Sims G E, Murphy S, Miller J R, Chan A P. 2012. Predicting the functional effect of amino acid substitutions and indels. PLoS One, 7(10): e46688. |
| [9] | Concordet J P, Haeussler M. 2018. CRISPOR: Intuitive guide selection for CRISPR/Cas9 genome editing experiments and screens. Nucleic Acids Res, 46(W1): W242-W245. |
| [10] |
Dehairs J, Talebi A, Cherifi Y, Swinnen J V. 2016. CRISP-ID: Decoding CRISPR mediated indels by Sanger sequencing. Sci Rep, 6: 28973.
PMID |
| [11] | do Vale F X R, Fernandes Filho E I, Liberato J R. 2003. A Software for Plant Disease Severity Assessment. Christchurch, New Zealand: 8th International Congress of Plant Pathology: 105. |
| [12] |
Feng K, Hou X L, Xing G M, Liu J X, Duan A Q, Xu Z S, Li M Y, Zhuang J, Xiong A S. 2020. Advances in AP2/ERF super-family transcription factors in plant. Crit Rev Biotechnol, 40(6): 750-776.
PMID |
| [13] | Fukagawa N K, Ziska L H. 2019. Rice: Importance for global nutrition. J Nutr Sci Vitaminol, 65: S2-S3. |
| [14] | Gasteiger E, Gattiker A, Hooland C, Ivanyi I, Appel R D, Bairoch A. 2003. ExPASy: The proteomics server for in-death protein knowledge and analysis. Nucleic Acids Res, 31(13): 3784-3788. |
| [15] |
Hiei Y, Ohta S, Komari T, Kumashiro T. 1994. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J, 6(2): 271-282.
PMID |
| [16] | Hong Y B, Liu Q N, Cao Y R, Zhang Y, Chen D B, Lou X Y, Cheng S H, Cao L Y. 2019. The OsMPK15 negatively regulates Magnaporthe oryzae and Xoo disease resistance via SA and JA signaling pathway in rice. Front Plant Sci, 10: 752. |
| [17] | Jain P, Singh P K, Kapoor R, Khanna A, Solanke A U, Krishnan S G, Singh A K, Sharma V, Sharma T R. 2017. Understanding host-pathogen interactions with expression profiling of NILs carrying rice-blast resistance Pi9 gene. Front Plant Sci, 8: 93. |
| [18] |
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna J A, Charpentier E. 2012. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science, 337: 816-821.
PMID |
| [19] | Kampinga H H, Andreasson C, Barducci A, Cheetham M E, Cyr D, Emanuelsson C, Genevaux P, Gestwicki J E, Goloubinoff P, Huerta-Cepas J, Kirstein J, Liberek K, Mayer M P, Nagata K, Nillegoda N B, Pulido P, Ramos C, de Los Rios P, Rospert S, Rosenzweig R, Sahi C, Taipale M, Tomiczek B, Ushioda R, Young J C, Zimmermann R, Zylicz A, Zylicz M, Craig E A, Marszalek J. 2019. Function, evolution, and structure of J-domain proteins. Cell Stress Chaperones, 24(1): 7-15. |
| [20] | Ko S H, Huang L M, Tarn W Y. 2019. The host heat shock protein MRJ/DNAJB6 modulates virus infection. Front Microbiol, 10: 2885. |
| [21] | Kusch S, Panstruga R. 2017. mlo-based resistance: An apparently universal “weapon” to defeat powdery mildew disease. Mol Plant Microbe Interact, 30(3): 179-189. |
| [22] |
Li W T, Chern M, Yin J J, Wang J, Chen X W. 2019. Recent advances in broad-spectrum resistance to the rice blast disease. Curr Opin Plant Biol, 50: 114-120.
PMID |
| [23] |
Luo Y, Fang B H, Wang W P, Yang Y, Rao L Q, Zhang C. 2019. Genome-wide analysis of the rice J-protein family: Identification, genomic organization, and expression profiles under multiple stresses. 3 Biotech, 9(10): 358.
PMID |
| [24] |
Miao J, Guo D S, Zhang J Z, Huang Q P, Qin G J, Zhang X, Wan J M, Gu H Y, Qu L J. 2013. Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res, 23(10): 1233-1236.
PMID |
| [25] |
Nakano T, Suzuki K, Fujimura T, Shinshi H. 2006. Genome-wide analysis of the ERF gene family in Arabidopsis and rice. Plant Physiol, 140(2): 411-432.
PMID |
| [26] |
Oliva R, Ji C H, Atienza-Grande G, Huguet-Tapia J C, Perez-Quintero A, Li T, Eom J S, Li C H, Nguyen H, Liu B, Auguy F, Sciallano C, Luu V T, Dossa G S, Cunnac S, Schmidt S M, Slamet-Loedin I H, Vera Cruz C, Szurek B, Frommer W B, White F F, Yang B. 2019. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotechnol, 37(11): 1344-1350.
PMID |
| [27] |
Parisi C, Tillie P, Rodríguez-Cerezo E. 2016. The global pipeline of GM crops out to 2020. Nat Biotechnol, 34(1): 31-36.
PMID |
| [28] | Park C J, Seo Y S. 2015. Heat shock proteins: A review of the molecular chaperones for plant immunity. Plant Pathol J, 31(4): 323-333. |
| [29] |
Phukan U J, Jeena G S, Tripathi V, Shukla R K. 2017. Regulation of Apetala2/ethylene response factors in plants. Front Plant Sci, 8: 150.
PMID |
| [30] |
Rashid M, He G Y, Yang G X, Hussain J, Yan X. 2012. AP2/ERF transcription factor in rice: Genome-wide canvas and syntenic relationships between monocots and eudicots. Evol Bioinform Online, 8: 321-355.
PMID |
| [31] | Ribot C, Hirsch J, Balzergue S, Tharreau D, Nottéghem J L, Lebrun M H, Morel J B. 2008. Susceptibility of rice to the blast fungus, Magnaporthe grisea. J Plant Physiol, 165(1): 114-124. |
| [32] |
Sallaud C, Lorieux M, Roumen E, Tharreau D, Berruyer R, Svestasrani P, Garsmeur O, Ghesquiere A, Notteghem J L. 2003. Identification of five new blast resistance genes in the highly blast-resistant rice variety IR64 using a QTL mapping strategy. Theor Appl Genet, 106(5): 794-803.
PMID |
| [33] | Sarkar N K, Thapar U, Kundnani P, Panwar P, Grover A. 2013. Functional relevance of J-protein family of rice (Oryza sativa). Cell Stress Chaperones, 18(3): 321-331. |
| [34] |
Srivastava R, Kumar R. 2018. The expanding roles of APETALA2/Ethylene Responsive Factors and their potential applications in crop improvement. Brief Funct Genomics, 18(4): 240-254.
PMID |
| [35] | Stam R, McDonald B A. 2018. When resistance gene pyramids are not durable: The role of pathogen diversity. Mol Plant Pathol, 19(3): 521-524. |
| [36] | Streubel J, Pesce C, Hutin M, Koebnik R, Boch J, Szurek B. 2013. Five phylogenetically close rice SWEET genes confer TAL effector-mediated susceptibility to Xanthomonas oryzae pv. oryzae. New Phytol, 200(3): 808-819. |
| [37] | Távora F T P K, Bevitori R, Mello R N, Cintra M M D F, Oliveira-Neto O B, Fontes W, Castro M S, Sousa M V, Franco O L, Mehta A. 2021. Shotgun proteomics coupled to transient- inducible gene silencing reveal rice susceptibility genes as new sources for blast disease resistance. J Proteomics, 241: 104223. |
| [38] |
van Schie C C N, Takken F L W. 2014. Susceptibility genes 101: How to be a good host. Annu Rev Phytopathol, 52: 551-581.
PMID |
| [39] | Verma A K, Tamadaddi C, Tak Y, Lal S S, Cole S J, Hines J K, Sahi C. 2019. The expanding world of plant J-domain proteins. Crit Rev Plant Sci, 38: 382-400. |
| [40] |
Win J, Chaparro-Garcia A, Belhaj K, Saunders D G O, Yoshida K, Dong S, Schornack S, Zipfel C, Robatzek S, Hogenhout S A, Kamoun S. 2012. Effector biology of plant-associated organisms: Concepts and perspectives. Cold Spring Harb Symp Quant Biol, 77: 235-247.
PMID |
| [41] | Wen C K. 2015. Ethylene in Plants. Berlin: Springer: 286. |
| [42] | Yang L T, Ding J Y, Zhang C M, Jia J W, Weng H B, Liu W X, Zhang D B. 2005. Estimating the copy number of transgenes in transformed rice by real-time quantitative PCR. Plant Cell Rep, 23(10/11): 759-763. |
| [43] | Zaidi S S E A, Mukhtar M S, Mansoor S. 2018. Genome editing: Targeting susceptibility genes for plant disease resistance. Trends Biotechnol, 36(9): 898-906. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||