
Rice Science ›› 2022, Vol. 29 ›› Issue (5): 402-406.DOI: 10.1016/j.rsci.2022.07.002
收稿日期:2021-12-18
接受日期:2022-02-15
出版日期:2022-09-28
发布日期:2022-07-04
. [J]. Rice Science, 2022, 29(5): 402-406.
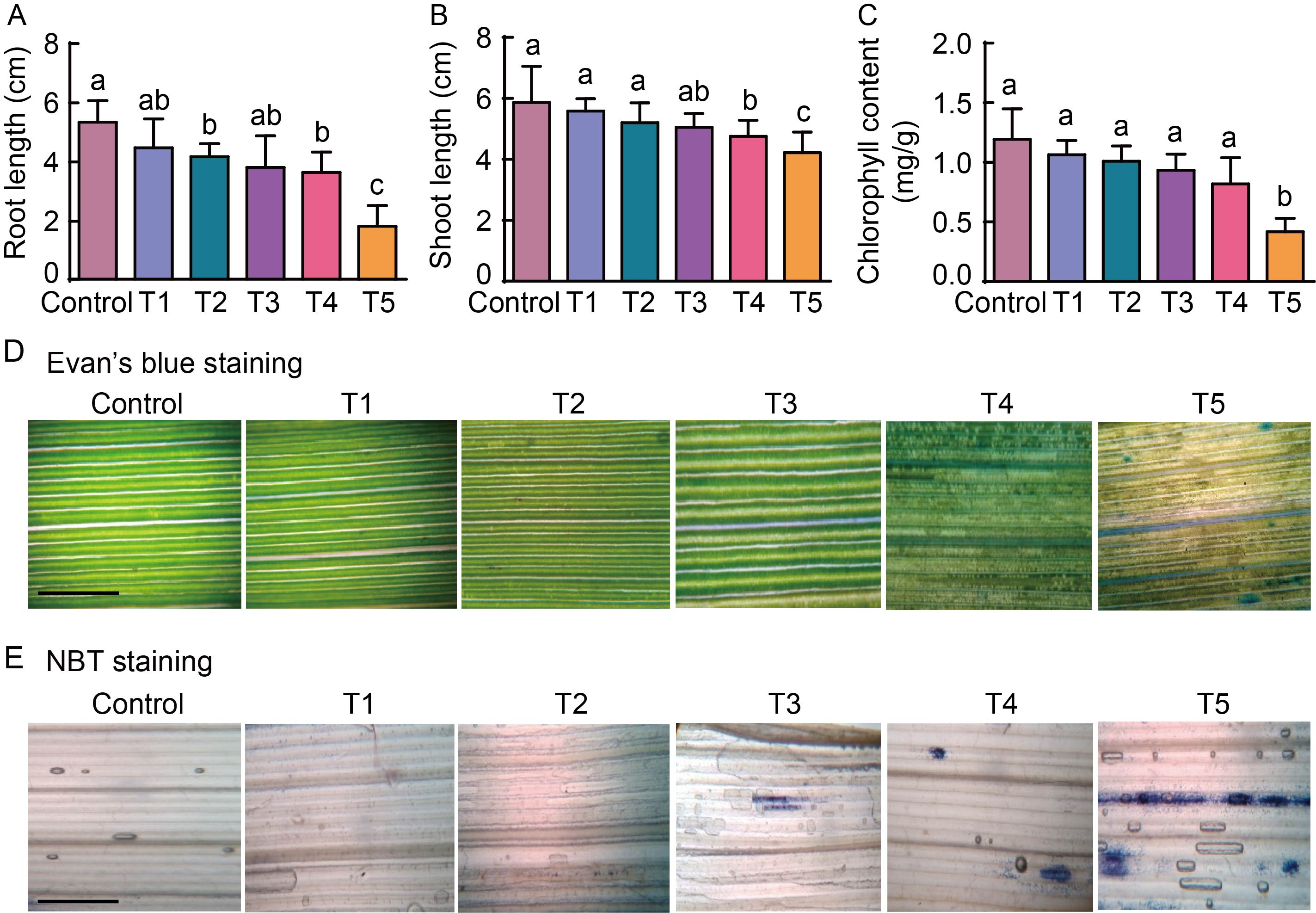
Fig. 1. Methyl viologen (MV) induced oxidative stress of rice seedlings under high light. A?C, Root (A) and shoot (B) lengths, and total chlorophyll content (C) of rice seedlings at different MV concen- trations. Ten seedlings with three replications were used for each con- centration. One way ANOVA was used to assess the statistical significance. Lowercase letters above the bars indicate significant differences at the 0.01 level. D and E, Membrane integrity measured by Evan’s blue staining (D) and levels of singlet O2·? by nitro blue tetrazolium chloride (NBT) staining (E). Scale bars, 25 μm. Seven-day-old rice seedlings were exposed to MV oxidative stress at different concentrations under high light conditions [600 µmol/(m2∙s)] for 16 h. T1 to T5 refer to treatments with MV concentrations of 2.5, 5.0, 10.0, 15.0 and 20.0 μmol/L, respectively.
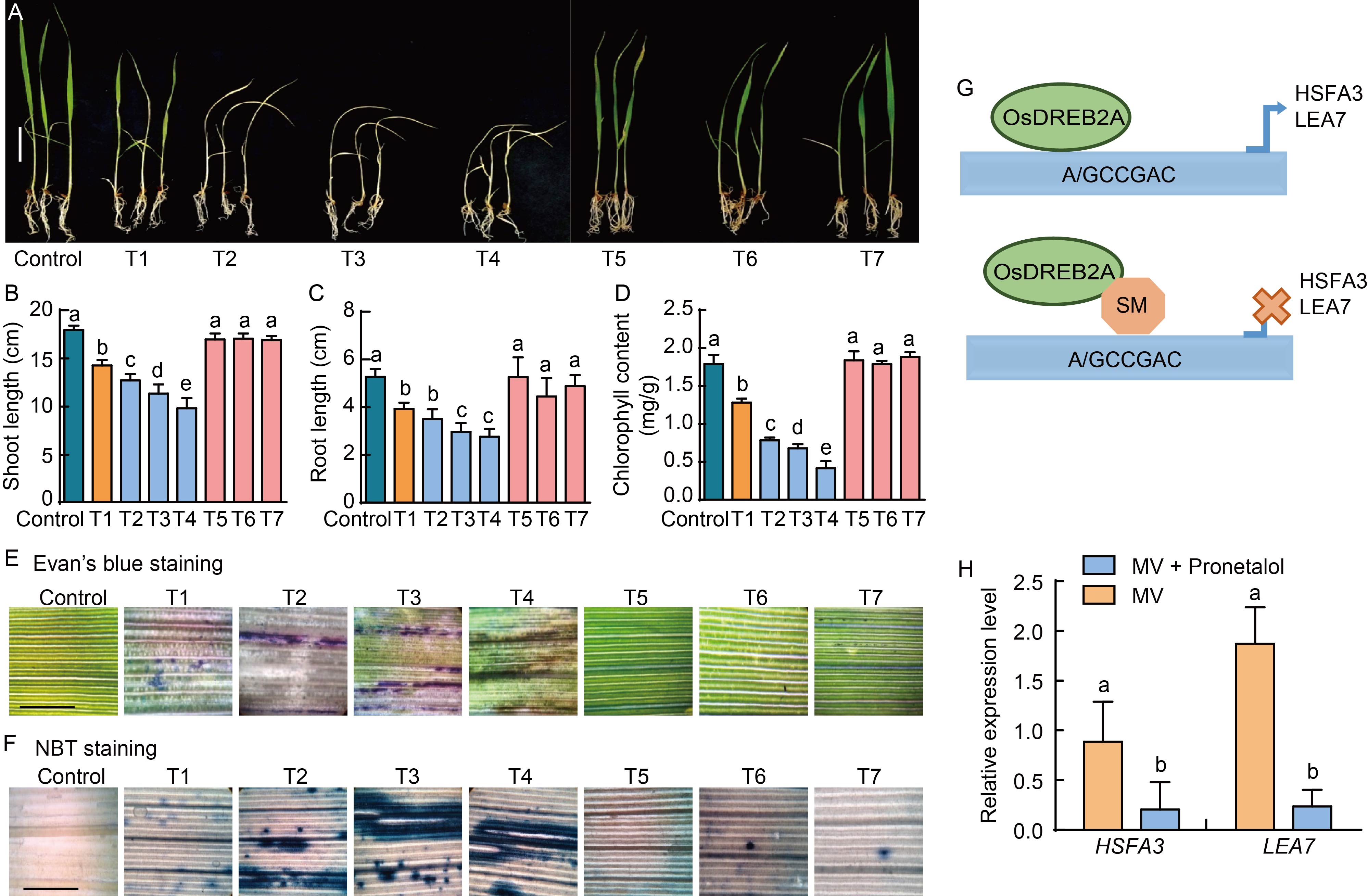
Fig. 2. Responses of rice seedlings treated with pronetalol (PRO) and methyl viologen (MV), and expression of OsDREB2A transcription factor target genes. A, Seedling growth at different PRO and MV concentrations. Scale bar, 1 cm. B?D, Shoot (B) and root (C) growth at different PRO and MV concentrations, and reduction in total chlorophyll content (D) over control. Ten seedlings with three replications were used for each concentration. E and F, Microscopic observation was taken for membrane integrity by Evan’s blue staining (E), and for levels of O2·? by nitro blue tetrazolium chloride (NBT) staining (F). Oxidative stress was imposed by MV at 20.0 µmol/L along with PRO (25.0, 50.0 and 75.0 µmol/L) for 10-day-old rice seedlings under high light conditions. Scale bars, 25 μm. G, Model depicting OsDREB2A binding to cis-elements of the target gene and small molecule (SM) interferences with cis-elements of target genes. H, Expression of OsDREB2A target genes HSFA3 and LEA7 by qRT-PCR. Minimum three replicates were used for each sample, and rice housekeeping gene ubiquitin was used for normalization. One-way ANOVA was used to assess the statistical significance. Different lowercase letters above the bars indicate significant differences at the 0.01 level. T1, 20.0 μmol/L MV; T2, 25.0 μmol/L PRO + 20.0 μmol/L MV; T3, 50.0 μmol/L PRO + 20.0 μmol/L MV; T4, 75.0 μmol/L PRO + 20.0 μmol/L MV; T5, 25.0 μmol/L PRO; T6, 50.0 μmol/L PRO; T7, 75.0 μmol/L PRO.
| [1] |
Babitha K C, Ramu S V, Pruthvi V, Mahesh P, Nataraja K N, Udayakumar M. 2013. Co-expression of AtbHLH17 and AtWRKY28 confers resistance to abiotic stress in Arabidopsis. Transgenic Res, 22(2): 327-341.
PMID |
| [2] |
Blackwell H E, Zhao Y D. 2003. Chemical genetic approaches to plant biology. Plant Physiol, 133(2): 448-455.
PMID |
| [3] | Botté C Y, Deligny M, Roccia A, Bonneau A L, Saïdani N, Hardré H, Aci S, Yamaryo-Botté Y, Jouhet J, Dubots E, Loizeau K, Bastien O, Bréhélin L, Joyard J, Cintrat J C, Falconet D, Block M A, Rousseau B, Lopez R, Maréchal E. 2011. Chemical inhibitors of monogalactosyldiacylglycerol synthases in Arabidopsis thaliana. Nat Chem Biol, 7(11): 834-842. |
| [4] |
Brooks A, Portis A R, Sharkey T D. 1988. Effects of irradiance and methyl viologen treatment on ATP, ADP, and activation of ribulose bisphosphate carboxylase in spinach leaves. Plant Physiol, 88(3): 850-853.
PMID |
| [5] |
Choi H, Kim J Y, Chang Y T, Nam H G. 2014. Forward chemical genetic screening. Methods Mol Biol, 1062: 393-404.
PMID |
| [6] | Cui M, Zhang W J, Zhang Q, Xu Z Q, Zhu Z G, Duan F P, Wu R. 2011. Induced over-expression of the transcription factor OsDREB2A improves drought tolerance in rice. Plant Physiol Biochem, 49(12): 1384-1391. |
| [7] |
Dejonghe W, Russinova E. 2017. Plant chemical genetics: From phenotype-based screens to synthetic biology. Plant Physiol, 174(1): 5-20.
PMID |
| [8] |
Halder V, Russinova E. 2019. Understanding the language of drugged plants. Nat Chem Biol, 15(11): 1025-1028.
PMID |
| [9] | Hasanuzzaman M, Borhannuddin Bhuyan M H M, Zulfiqar F, Raza A, Mohsin S M, Mahmud J A, Fujita M, Fotopoulos V. 2020. Reactive oxygen species and antioxidant defense in plants under abiotic stress: Revisiting the crucial role of a universal defense regulator. Antioxidants, 9(8): 681. |
| [10] |
Hicks G R, Raikhel N V. 2014. Plant chemical biology: Are we meeting the promise? Front Plant Sci, 5: 455.
PMID |
| [11] |
Huang W, Yang Y J, Zhang S B, Liu T. 2018. Cyclic electron flow around photosystem I promotes ATP synthesis possibly helping the rapid repair of photodamaged photosystem II at low light. Front Plant Sci, 9: 239.
PMID |
| [12] | Huo J Q, Zhao B, Zhang Z, Xing J H, Zhang J L, Dong J G, Fan Z J. 2018. Structure-based discovery and synthesis of potential transketolase inhibitors. Molecules, 23(9): 2116. |
| [13] | Javed T, Shabbir R, Ali A, Afzal I, Zaheer U, Gao S J. 2020. Transcription factors in plant stress responses: Challenges and potential for sugarcane improvement. Plants, 9(4): 491. |
| [14] | Khan B R, Faure L, Chapman K D, Blancaflor E B. 2017. A chemical genetic screen uncovers a small molecule enhancer of the N-acylethanolamine degrading enzyme, fatty acid amide hydrolase, in Arabidopsis. Sci Rep, 7: 41121. |
| [15] | Lascano M, Muñoz N, Robert G, Rodriguez M, Melchiorre M, Trippi V, Quero G. 2012. Paraquat: An oxidative stress inducer. In: Hasaneen M N. Herbicides: Properties, Synthesis and Control of Weeds. IntechOpen: 135-148. |
| [16] | Morimoto K, Mizoi J, Qin F, Kim J S, Sato H, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K. 2013. Stabilization of Arabidopsis DREB2A is required but not sufficient for the induction of target genes under conditions of stress. PLoS One, 8(12): e80457. |
| [17] |
Nakano T, Tanaka S, Ohtani M, Yamagami A, Takeno S, Hara N, Mori A, Nakano A, Hirose S, Himuro Y, Kobayashi M, Kushiro T, Demura T, Asami T, Osada H, Shinozaki K. 2018. FPX is a novel chemical inducer that promotes callus formation and shoot regeneration in plants. Plant Cell Physiol, 59(8): 1555-1567.
PMID |
| [18] | Nisarga K N, Vemanna R S, Kodekallu Chandrashekar B, Rao H, Vennapusa A R, Narasimaha A, Makarla U, Basavaiah M R. 2017. Aldo-ketoreductase 1 (AKR1) improves seed longevity in tobacco and rice by detoxifying reactive cytotoxic compounds generated during ageing. Rice, 10(1): 11. |
| [19] | Rehem B C, Bertolde F Z, de Almeida A A F. 2012. Regulation of gene expression in response to abiotic stress in plants. In: Bubulya P. Cell Metabolism: Cell Homeostasis and Stress Response. Rijeka, Croatia: InTech: 13-28. |
| [20] |
Rodríguez-Martínez J A, Peterson-Kaufman K J, Ansari A Z. 2010. Small-molecule regulators that mimic transcription factors. Biochim Biophys Acta, 1799: 768-774.
PMID |
| [21] |
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K. 2006. Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought- responsive gene expression. Plant Cell, 18(5): 1292-1309.
PMID |
| [22] | Song C, Chung W S, Lim C O. 2016. Overexpression of heat shock factor gene HsfA3 increases galactinol levels and oxidative stress tolerance in Arabidopsis. Mol Cells, 39(6): 477-483. |
| [23] | Sujeeth N, Mehterov N, Gupta S, Qureshi M K, Fischer A, Proost S, Omidbakhshfard M A, Obata T, Benina M, Staykov N, Balazadeh S, Walther D, Fernie A R, Mueller-Roeber B, Hille J, Gechev T S. 2020. A novel seed plants gene regulates oxidative stress tolerance in Arabidopsis thaliana. Cell Mol Life Sci, 77(4): 705-718. |
| [24] | Sujitha D, Jalendra Kumar H G, Thapliayal G, Vanitha P A, Uttarkar A Niranjan V, Rayalcheruvu U, Vemanna R S. 2022. Master regulators of oxidative stress associated genes improves stress adaptation in rice. Plant Mol Biol, (in press). |
| [25] |
Takaoka Y, Iwahashi M, Chini A, Saito H, Ishimaru Y, Egoshi S, Kato N, Tanaka M, Bashir K, Seki M, Solano R, Ueda M. 2018. A rationally designed JAZ subtype-selective agonist of jasmonate perception. Nat Commun, 9(1): 3654.
PMID |
| [26] | Vemanna R S, Swetha T N, Sheela S H, Babitha K C, Rohini S, Reddy M K, Tuteja N, Reddy C P, Prasad T G, Makarla U. 2016a. Simultaneous expression of regulatory genes associated with specific drought-adaptive traits improves drought adaptation in peanut. Plant Biotechnol J, 14(3): 1008-1020. |
| [27] | Vemanna R S, Paramanantham A, Ramegowda V, Mohan-Raju B, Makarla U, Senthil-Kumar M. 2016b. Transcriptome analysis of sunflower genotypes with contrasting oxidative stress tolerance reveals individual- and combined-biotic and abiotic stress tolerance mechanisms. PLoS One, 11(6): e0157522. |
| [28] | Vemanna R S, Babitha K C, Solanki J K, Reddy V A, Sarangi S K, Makarla U. 2017. Aldo-keto reductase-1 (AKR1) protect cellular enzymes from salt stress by detoxifying reactive cytotoxic compounds. Plant Physiol Biochem, 113: 177-186. |
| [29] | Vemanna R S, Vijayaraghavareedy P, Nisarga K N, Srivastava K R, Sheshshayee M S, Mysore K S, Makarla U. 2020. Carbonyl cytotoxicity affects plant cellular processes and detoxifying enzymes scavenge these compounds to improve stress tolerance. J Agric Food Chem, 68(23): 6237-6247. |
| [30] | Wei T L, Guo D L, Liu J H. 2021. Overexpression of PTRLEA7, a late embryogenesis abundant family gene from Poncirus trifoliata, confers enhanced drought tolerance by enhancing antioxidant capacity. Front Agr Sci Eng, 8(2): 236-246. |
| [31] | Wu Z, Liang J H, Wang C P, Zhao X, Zhong X H, Cao X, Li G Q, He J N, Yi M F. 2018. Overexpression of lily HsfA3s in Arabidopsis confers increased thermotolerance and salt sensitivity via alterations in proline catabolism. J Exp Bot, 69(8): 2005-2021. |
| [32] | Xiao B Z, Huang Y M, Tang N, Xiong L Z. 2007. Over-expression of a LEA gene in rice improves drought resistance under the field conditions. Theor Appl Genet, 115(1): 35-46. |
| [33] | Yoon Y, Seo D H, Shin H, Kim H J, Kim C M, Jang G. 2020. The role of stress-responsive transcription factors in modulating abiotic stress tolerance in plants. Agronomy, 10(6): 788. |
| [34] | Zhang X X, Tang Y J, Ma Q B, Yang C Y, Mu Y H, Suo H C, Luo L H, Nian H. 2013. OsDREB2A, a rice transcription factor, significantly affects salt tolerance in transgenic soybean. PLoS One, 8(12): e83011. |
| [35] | Ziadi A, Uchida N, Kato H, Hisamatsu R, Sato A, Hagihara S, Itami K, Torii K U. 2017. Discovery of synthetic small molecules that enhance the number of stomata: C-H functionalization chemistry for plant biology. Chem Commun, 53: 9632-9635. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||