
Rice Science ›› 2023, Vol. 30 ›› Issue (6): 567-576.DOI: 10.1016/j.rsci.2023.07.001
收稿日期:2023-04-12
接受日期:2023-07-15
出版日期:2023-11-28
发布日期:2023-08-10
. [J]. Rice Science, 2023, 30(6): 567-576.
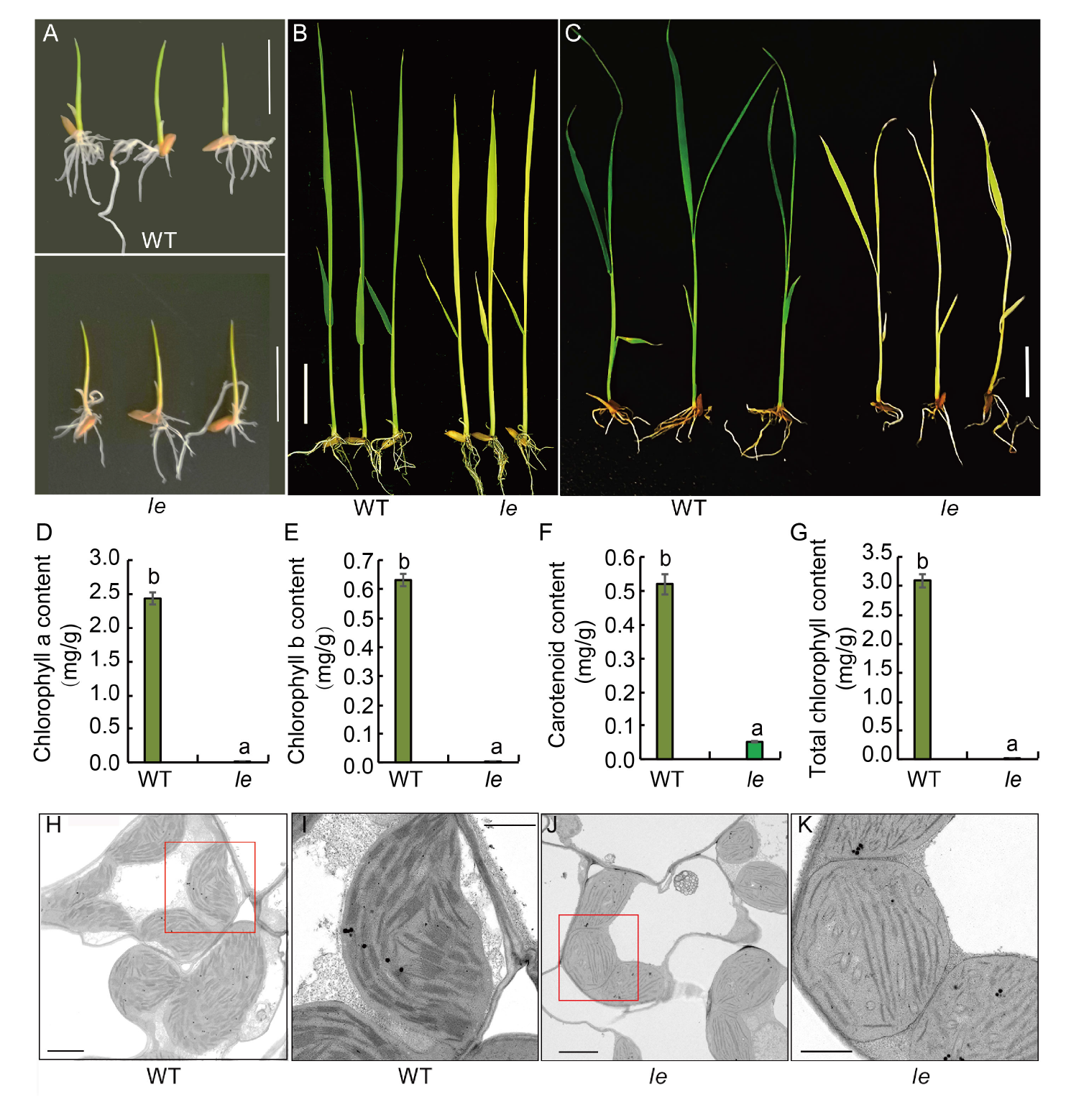
Fig. 1. Phenotype, chlorophyll content and ultrastructure of chloroplasts in wild type (WT) and le mutant. A?C, Phenotypes of WT and le mutant plants at 3 (A), 7 (B) and 14 (C) d after sowing. Scale bars, 2 cm. D?G, Contents of chlorophyll a (D), chlorophyll b (E), carotenoids (F) and total chlorophyll (G) in the mutant le and its WT. Data are Mean ± SD (n = 3). Different lowercase letters above the bars indicate statistical significances at the 0.05 probability level. H?K, Chloroplast ultrastructure of WT (H) and mutant le (J), and their enlarged parts highlighted by red boxes (I and K). Scale bars are 2 μm in H and J, and 1 μm in I and K.
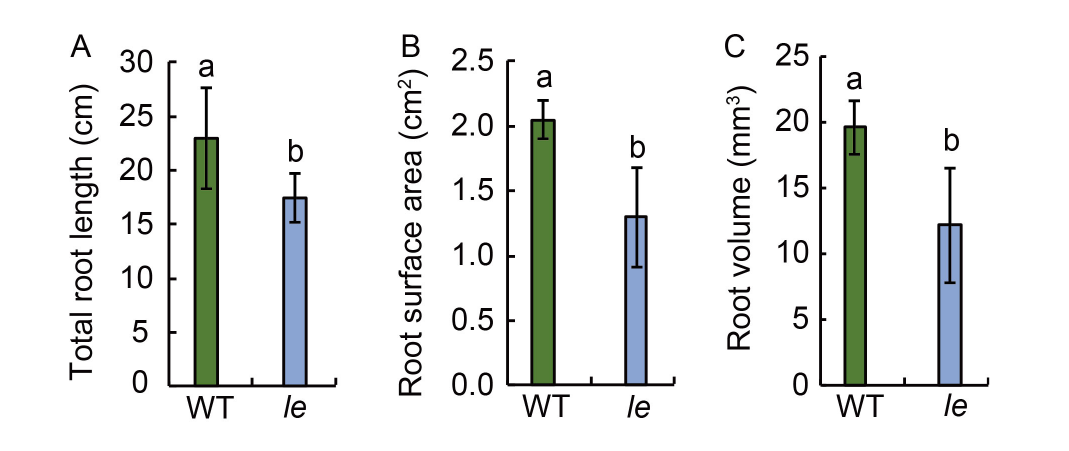
Fig. 2. Total root length (A), root surface area (B), and root volume (C) of seedlings of mutant le and its wild type (WT). Data are Mean ± SD (n = 3). Different lowercase letters above the bars indicate statistical significances at the 0.05 probability level.
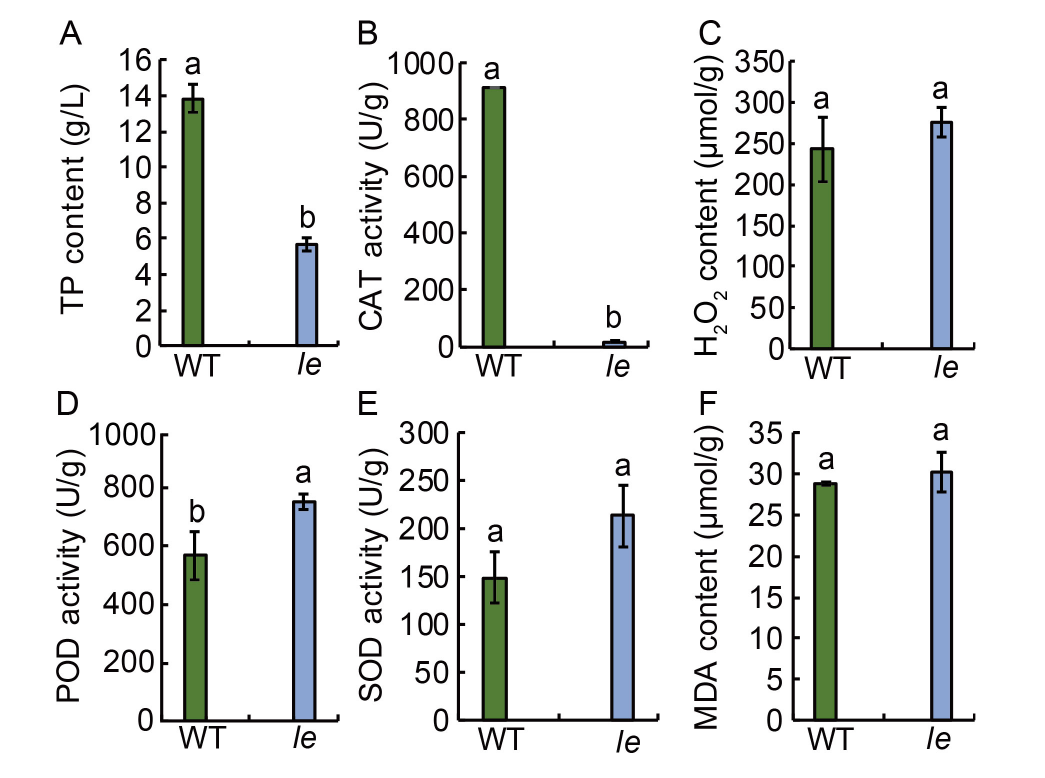
Fig. 3. Physiological and biochemical parameters of mutant le and its wild type (WT). A?F, Total intracellular protein (TP) content (A), catalase (CAT) activity (B), hydrogen peroxide (H2O2) content (C), peroxidase (POD) activity (D), superoxide dismutase (SOD) activity (E), and malondialdehyde (MDA) content (F) in le and WT. Data are Mean ± SD (n = 3). Different lowercase letters above the bars indicate statistical significances at the 0.05 probability level.
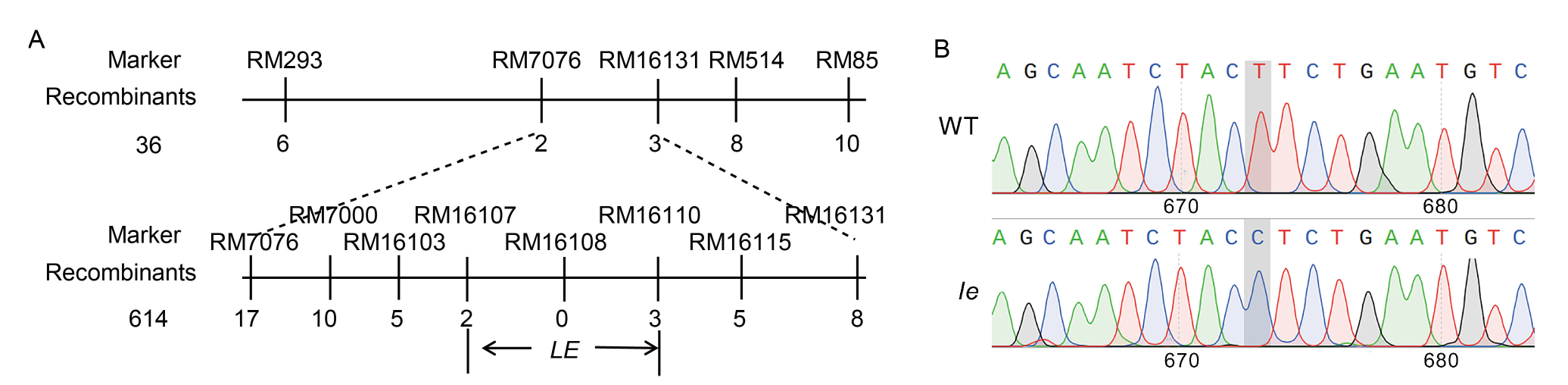
Fig. 4. Fine mapping of LE gene conferring lethal etiolated mutation (A) and differences in gene sequence of mutant le and its wild type (WT) at LOC_Os03g59640 (B).
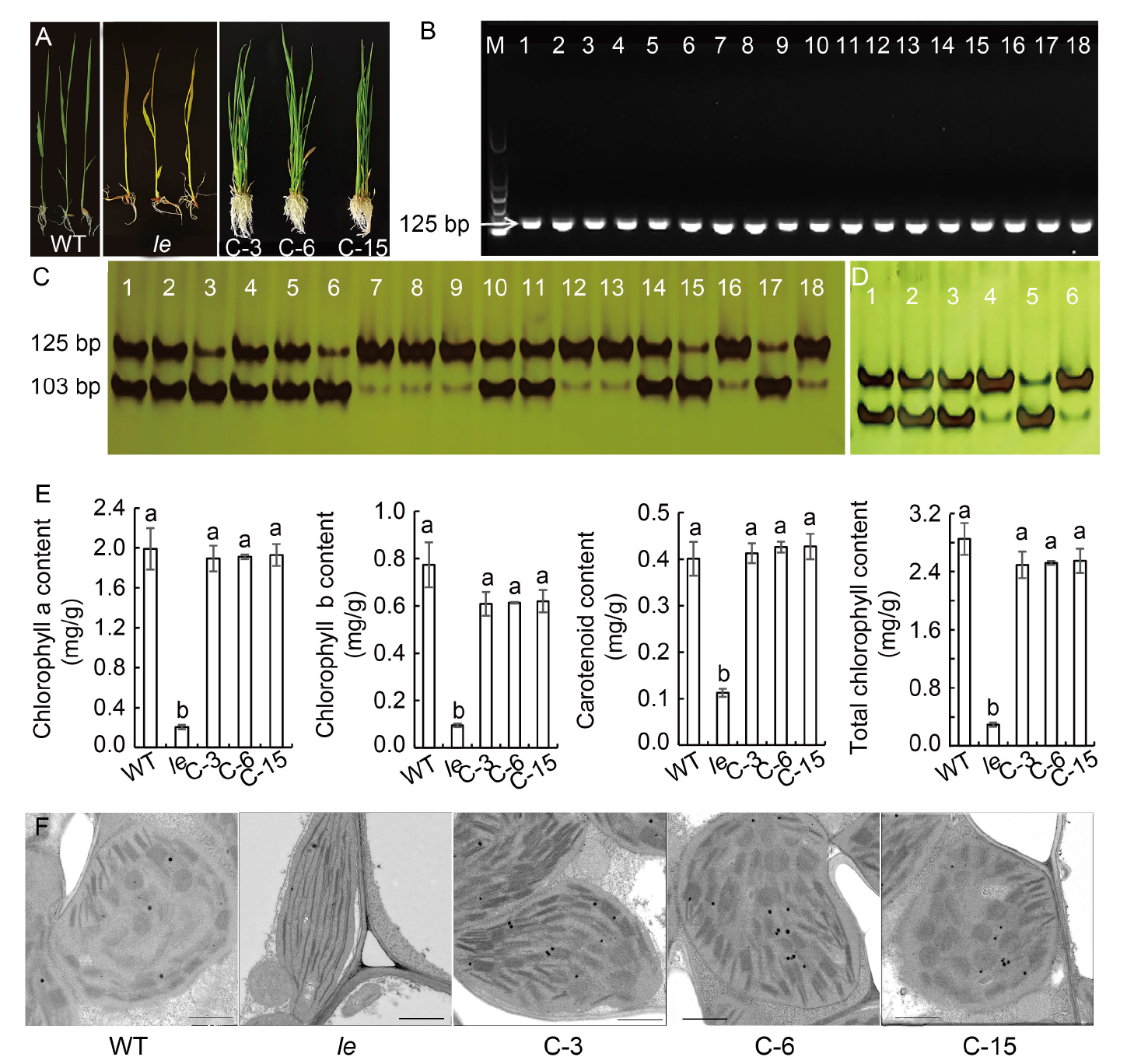
Fig. 5. Verification, phenotypes, chlo- rophyll contents, and ultrastructure of chloroplasts of complementary plants. A, Phenotypes of wild type (WT), le mutant and complementary plants (C-3, C-6, and C-15). B, PCR amplification results with primer D-1 using a 739-bp amplification product as template. M, Marker; Lanes 1?15, Transformed plants (C1?C15); Lane 16, WT; Lane 17, le mutant; Lane 18, HM133 mutant. C, Digestion results of primer D-1-based amplification products by Kpn I. Lanes 1?15, Transformed plants (C1?C15); Lane 16, WT; Lane 17, le mutant; Lane 18, HM133 mutant. D, Digestion results of transformants by Kpn I. Lanes 1?6 represent C-3, C-6, C-15, WT, le, and HM133, respectively. E, Contents of chlorophyll a, chlorophyll b, carotenoids, and total chlorophyll. Data are Mean ± SD (n = 3). Different lowercase letters above the bars indicate statistical significances at the 0.05 probability level. F, Chloroplast ultrastructures of WT, le, C-3, C-6, and C-15. Scale bars, 1 μm.
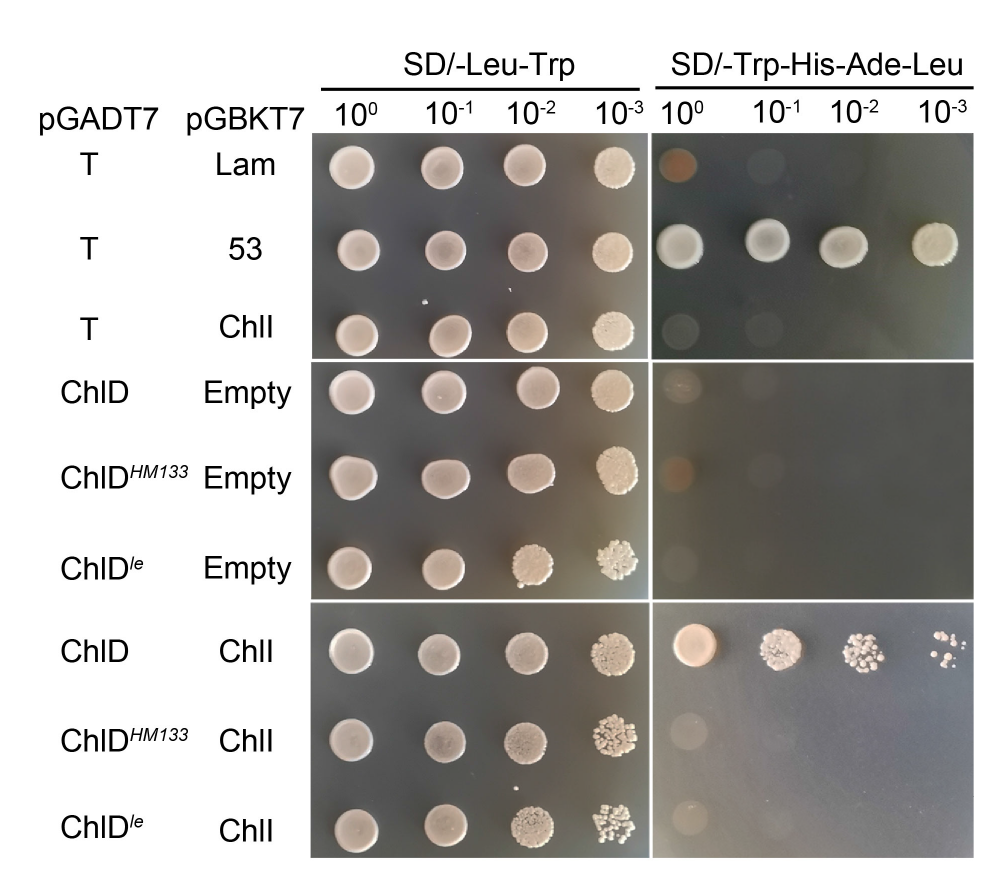
Fig. 6. Interaction of ChlDle and ChlI detected by yeast two-hybrid assay. The interaction of pGBKT7Lam and pGADT7T is used as negative control, and the interaction of pGBKT753 and pGADT7T is used as positive controls.
| [1] | Adams N B P, Vasilev C, Brindley A A, Hunter C N. 2016. Nanomechanical and thermophoretic analyses of the nucleotide- dependent interactions between the AAA+ subunits of Mg chelatase. J Am Chem Soc, 138(20): 6591-6597. |
| [2] | Brindley A A, Adams N B P, Hunter C N, Reid J D. 2015. Five glutamic acid residues in the C-terminal domain of the ChlD subunit play a major role in conferring Mg2+ cooperativity upon magnesium chelatase. Biochemistry, 54(44): 6659-6662. |
| [3] | Chen J H, Chen S T, He N Y, Wang Q L, Zhao Y, Gao W, Guo F Q. 2020. Nuclear-encoded synthesis of the D1 subunit of photosystem II increases photosynthetic efficiency and crop yield. Nat Plants, 6(5): 570-580. |
| [4] | Deng X J, Zhang H Q, Wang Y, He F, Liu J L, Xiao X, Shu Z F, Li W, Wang G H, Wang G L. 2014. Mapped clone and functional analysis of leaf-color gene Ygl7 in a rice hybrid (Oryza sativa L. ssp. indica). PLoS One, 9(6): e99564. |
| [5] | Farmer D A, Brindley A A, Hitchcock A, Jackson P J, Johnson B, Dickman M J, Hunter C N, Reid J D, Adams N B P. 2019. The ChlD subunit links the motor and porphyrin binding subunits of magnesium chelatase. Biochem J, 476(13): 1875-1887. |
| [6] | Gibson L C D, Jensen P E, Hunter C N. 1999. Magnesium chelatase from Rhodobacter sphaeroides: Initial characterization of the enzyme using purified subunits and evidence for a BchI-BchD complex. Biochem J, 337(2): 243. |
| [7] | He J Y, Wang Y Y, Ren Y F, Zhou G Q, Yang L J. 2009. Effect of cadmium on root morphology and physiological characteristics of rice seedlings. Ecol Environ Sci, 18(5): 1863-1868. (in Chinese with English abstract) |
| [8] | Ishijima S, Uchibori A, Takagi H, Maki R, Ohnishi M. 2003. Light- induced increase in free Mg2+ concentration in spinach chloroplasts: Measurement of free Mg2+ by using a fluorescent probe and necessity of stromal alkalinization. Arch Biochem Biophys, 412(1): 126-132. |
| [9] | Jensen P E, Gibson L C D, Henningsen K W, Hunter C N. 1996. Expression of the chlI, chlD, and chlH genes from the Cyanobacterium synechocystis PCC6803 in Escherichia coli and demonstration that the three cognate proteins are required for magnesium-protoporphyrin chelatase activity. J Biol Chem, 271(28): 16662-16667. |
| [10] | Jensen P E, Gibson L C, Hunter C N. 1999. ATPase activity associated with the magnesium-protoporphyrin IX chelatase enzyme of Synechocystis PCC6803: Evidence for ATP hydrolysis during Mg2+ insertion, and the MgATP-dependent interaction of the ChlI and ChlD subunits. Biochem J, 339: 127-134. |
| [11] | Kang S J, Fang Y X, Zou G X, Ruan B P, Zhao J A, Dong G J, Yan M X, Gao Z Y, Zhu L. 2015. White-green leaf gene encoding protochlorophyllide oxidoreductase B is involved in chlorophyll synthesis of rice. Crop Sci, 55(1): 284-293. |
| [12] | Lake V, Olsson U, Willows R D, Hansson M. 2004. ATPase activity of magnesium chelatase subunit I is required to maintain subunit D in vivo. Eur J Biochem, 271(11): 2182-2188. |
| [13] | Lee S, Kim J H, Yoo E S, Lee C H, Hirochika H, An G. 2005. Differential regulation of chlorophyll a oxygenase genes in rice. Plant Mol Biol, 57(6): 805-818. |
| [14] | Lundqvist J, Elmlund H, Wulff R P, Berglund L, Elmlund D, Emanuelsson C, Hebert H, Willows R D, Hansson M, Lindahl M, Al-Karadaghi S. 2010. ATP-induced conformational dynamics in the AAA+ motor unit of magnesium chelatase. Structure, 18(3): 354-365. |
| [15] | Luo S, Luo T, Peng P, Li Y P, Li X G. 2015. Activity regulation and functions of Mg chelatase. Plant Physiol J, 51(6): 806-812. (in Chinese with English abstract) |
| [16] | Luo S, Luo T, Liu Y N, Li Z W, Fan S Y, Wu C J. 2018. N-terminus plus linker domain of Mg-chelatase D subunit is essential for Mg-chelatase activity in Oryza sativa. Biochem Biophys Res Commun, 497(2): 749-755. |
| [17] | Luo T, Fan T T, Liu Y N, Rothbart M, Yu J, Zhou S X, Grimm B, Luo M Z. 2012. Thioredoxin redox regulates ATPase activity of magnesium chelatase CHLI subunit and modulates redox- mediated signaling in tetrapyrrole biosynthesis and homeostasis of reactive oxygen species in pea plants. Plant Physiol, 159(1): 118-130. |
| [18] | Luo T, Luo S, Araújo W L, Schlicke H, Rothbart M, Yu J, Fan T T, Fernie A R, Grimm B, Luo M Z. 2013. Virus-induced gene silencing of pea CHLI and CHLD affects tetrapyrrole biosynthesis, chloroplast development and the primary metabolic network. Plant Physiol Biochem, 65: 17-26. |
| [19] | Lv X G, Shi Y F, Xu X A, Wei Y L, Wang H M, Zhang X B, Wu J L. 2015. Oryza sativa chloroplast signal recognition particle 43 (OscpSRP43) is required for chloroplast development and photosynthesis. PLoS One, 10(11): e0143249. |
| [20] | Masuda T. 2008. Recent overview of the Mg branch of the tetrapyrrole biosynthesis leading to chlorophylls. Photosynth Res, 96(2): 121-143. |
| [21] | Papenbrock J, Mock H P, Kruse E, Grimm B. 1999. Expression studies in tetrapyrrole biosynthesis: Inverse maxima of magnesium chelatase and ferrochelatase activity during cyclic photoperiods. Planta, 208(2): 264-273. |
| [22] | Reid J D, Hunter C N. 2002. Current understanding of the function of magnesium chelatase. Biochem Soc Trans, 30(4): 643-645. |
| [23] | Ruan B P, Gao Z Y, Zhao J, Zhang B, Zhang A P, Hong K, Yang S L, Jiang H Z, Liu C L, Chen G, Peng Y L, Dong G J, Guo L B, Xu Z J, Qian Q. 2017. The rice YGL gene encoding an Mg2+- chelatase ChlD subunit is affected by temperature for chlorophyll biosynthesis. J Plant Biol, 60(4): 314-321. |
| [24] | Sakuraba Y, Rahman M L, Cho S H, Kim Y S, Koh H J, Yoo S C, Paek N C. 2013. The rice faded green leaf locus encodes protochlorophyllide oxidoreductase B and is essential for chlorophyll synthesis under high light conditions. Plant J, 74(1): 122-133. |
| [25] | Sawicki A, Zhou S X, Kwiatkowski K, Luo M Z, Willows R D. 2017. 1-N-histidine phosphorylation of ChlD by the AAA+ ChlI2 stimulates magnesium chelatase activity in chlorophyll synthesis. Biochem J, 474(12): 2095-2105. |
| [26] | Shang H H. 2022. Gene isolation and functional analysis of a spotted-leaf mutant spl26in rice. Wuhan, China: Huazhong Agricultural University. (in Chinese with English abstract) |
| [27] | Shi Y F, He Y, Guo D, Lv X G, Huang Q N, Wu J L. 2016. Genetic analysis and gene mapping of a pale green leaf mutant HM133 in rice. Chin J Rice Sci, 30(6): 603-610. (in Chinese with English abstract) |
| [28] | Sun X Q, Wang B, Xiao Y H, Wan C M, Deng X J, Wang P R. 2011. Genetic analysis and fine mapping of gene ygl98 for yellow-green leaf of rice. Acta Agron Sin, 37(6): 991-997. (in Chinese with English abstract) |
| [29] | Walker C J, Willows R D. 1997. Mechanism and regulation of Mg-chelatase. Biochem J, 327(Pt 2): 321-333. |
| [30] | Wang P R, Gao J X, Wan C M, Zhang F T, Xu Z J, Huang X Q, Sun X Q, Deng X J. 2010. Divinyl chlorophyll(ide) a can be converted to monovinyl chlorophyll(ide) a by a divinyl reductase in rice. Plant Physiol, 153(3): 994-1003. |
| [31] | Willows R D, Gibson L C D, Kanangara C G, Hunter C N, von Wettstein D. 1996. Three separate proteins constitute the magnesium chelatase of Rhodobacter sphaeroides. Eur J Biochem, 235(1/2): 438-443. |
| [32] | Wu Z M, Zhang X, He B, Diao L P, Sheng S L, Wang J L, Guo X P, Su N, Wang L F, Jiang L, Wang C M, Zhai H Q, Wan J M. 2007. A chlorophyll-deficient rice mutant with impaired chlorophyllide esterification in chlorophyll biosynthesis. Plant Physiol, 145(1): 29-40. |
| [33] | Zhang H, Liu L L, Cai M H, Zhu S S, Zhao J Y, Zheng T H, Xu X Y, Zeng Z Q, Niu J, Jiang L, Chen S H, Wan J M. 2015. A point mutation of magnesium chelatase OsCHLI gene dampens the interaction between CHLI and CHLD subunits in rice. Plant Mol Biol Rep, 33(6): 1975-1987. |
| [34] | Zhang H T, Li J J, Yoo J H, Yoo S C, Cho S H, Koh H J, Seo H S, Paek N C. 2006. Rice Chlorina-1 and Chlorina-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development. Plant Mol Biol, 62(3): 325-337. |
| [35] | Zhang T, Feng P, Li Y F, Yu P, Yu G L, Sang X C, Ling Y H, Zeng X Q, Li Y D, Huang J Y, Zhang T Q, Zhao F M, Wang N, Zhang C W, Yang Z L, Wu R H, He G H. 2018. VIRESCENT- ALBINO LEAF 1 regulates leaf colour development and cell division in rice. J Exp Bot, 69(20): 4791-4804. |
| [36] | Zhou S X, Sawicki A, Willows R D, Luo M Z. 2012. C-terminal residues of Oryza sativa GUN4 are required for the activation of the ChlH subunit of magnesium chelatase in chlorophyll synthesis. FEBS Lett, 586(3): 205-210. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||