
Rice Science ›› 2026, Vol. 33 ›› Issue (2): 173-185.DOI: 10.1016/j.rsci.2025.11.003
收稿日期:2025-09-24
接受日期:2025-11-25
出版日期:2026-03-28
发布日期:2026-04-01
. [J]. Rice Science, 2026, 33(2): 173-185.
| Tissue/organ | Number of lysine acetylation proteins/sites | Percentage of proteins with multiple lysine acetylation sites (%) | Reference |
|---|---|---|---|
| Suspension cells | 44/60 | 25 | Nallamilli et al, |
| Embryo of seeds | 389/699 | ‒ | He et al, |
| 5-week-old seedlings | 716/1 337 | 39 | Xiong et al, |
| Developing anthers | 676/1 354 | 37 | Li et al, |
| Developing seeds | 692/1 003 | 28 | Meng et al, |
| 3-week-old leaves | 866/1 353 | 31 | Xue et al, |
| 14-day-old leaves | 1 024/1 669 | ‒ | Zhou et al, |
| 14-day-old leaves | 1 952/4 868 | 47 | Xu et al, |
| Crown roots | 2 645/6 099 | 48 | Xu et al, |
Table 1. Lysine acetylomes in different tissues of rice.
| Tissue/organ | Number of lysine acetylation proteins/sites | Percentage of proteins with multiple lysine acetylation sites (%) | Reference |
|---|---|---|---|
| Suspension cells | 44/60 | 25 | Nallamilli et al, |
| Embryo of seeds | 389/699 | ‒ | He et al, |
| 5-week-old seedlings | 716/1 337 | 39 | Xiong et al, |
| Developing anthers | 676/1 354 | 37 | Li et al, |
| Developing seeds | 692/1 003 | 28 | Meng et al, |
| 3-week-old leaves | 866/1 353 | 31 | Xue et al, |
| 14-day-old leaves | 1 024/1 669 | ‒ | Zhou et al, |
| 14-day-old leaves | 1 952/4 868 | 47 | Xu et al, |
| Crown roots | 2 645/6 099 | 48 | Xu et al, |
| HDAC/HAT | Histone modification site | Biological function | Reference |
|---|---|---|---|
| OsHDA701 | H3K9ac | Regulation of immune responses | Chen et al, |
| OsHDA702 | H3K9, K14, K18ac, H4K5, K12, K16ac | Root growth | Chung et al, |
| OsHDA703 | H4K8, K12ac | Brassinosteroid signal transduction, growth, and heading date | Wang et al, |
| OsHDA704 | H4K8ac | Drought stress response | Guo Y L et al, |
| OsHDA705 | H4K8hib | Pathogen immune response | Xu et al, |
| OsHDA706 | H4K5, K8ac | Salt stress response | Liu et al, |
| OsHDA710 | H4K5, K16ac | Response to salt stress | Ullah et al, |
| OsSRT701 | H3K9ac | Seed development | Zhang et al, |
| OsSRT702 | H4K5, K8ac | Regulate immunity | Chen X Y et al, |
| OsHDT702 | H4K5, K16ac | Regulate plant innate immunity | Ding et al, |
| OsGCN5 | H3K18ac, K27ac | ‒ | Xue et al, |
| OsHAG704 | H4K5ac, K16ac | Cell division | Xu et al, |
Table 2. Types of histone modifications regulated by rice histone deacetylases (HDACs)/histone acetyltransferases (HATs).
| HDAC/HAT | Histone modification site | Biological function | Reference |
|---|---|---|---|
| OsHDA701 | H3K9ac | Regulation of immune responses | Chen et al, |
| OsHDA702 | H3K9, K14, K18ac, H4K5, K12, K16ac | Root growth | Chung et al, |
| OsHDA703 | H4K8, K12ac | Brassinosteroid signal transduction, growth, and heading date | Wang et al, |
| OsHDA704 | H4K8ac | Drought stress response | Guo Y L et al, |
| OsHDA705 | H4K8hib | Pathogen immune response | Xu et al, |
| OsHDA706 | H4K5, K8ac | Salt stress response | Liu et al, |
| OsHDA710 | H4K5, K16ac | Response to salt stress | Ullah et al, |
| OsSRT701 | H3K9ac | Seed development | Zhang et al, |
| OsSRT702 | H4K5, K8ac | Regulate immunity | Chen X Y et al, |
| OsHDT702 | H4K5, K16ac | Regulate plant innate immunity | Ding et al, |
| OsGCN5 | H3K18ac, K27ac | ‒ | Xue et al, |
| OsHAG704 | H4K5ac, K16ac | Cell division | Xu et al, |
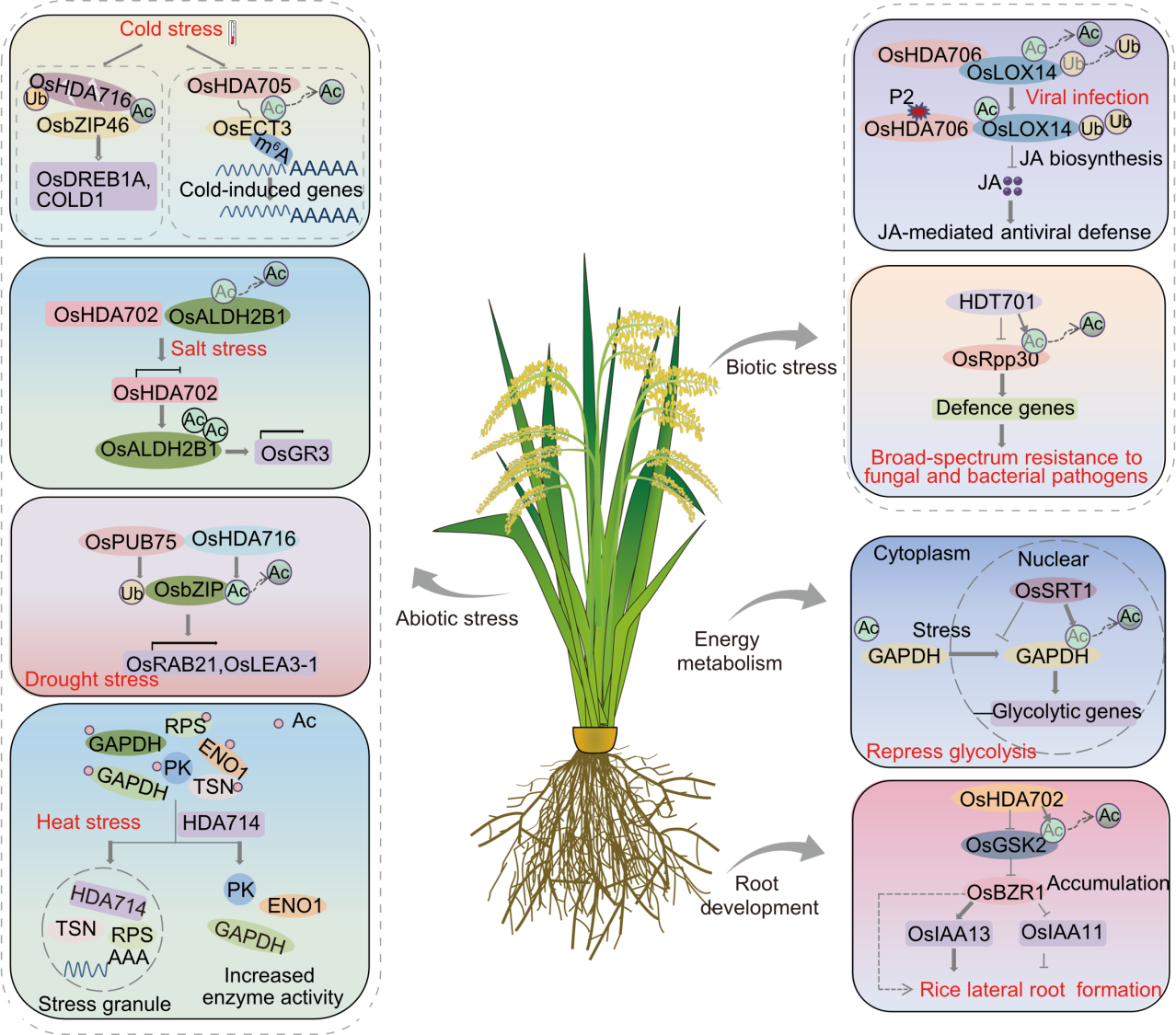
Fig. 1. Overview of role of rice histone deacetylases involved in environmental stress response and growth through non-histone acetylation. Ac, Acetylation; Ub, Ubiquitination; JA, Jasmonic acid; PK, Pyruvate kinase; ENO1, Enolase1.
| Modifying enzyme type | Enzyme | Non-histone substrate | Function of acetylation | Mechanism of action | Reference |
|---|---|---|---|---|---|
| Deacetylase | OsHDAC1/ OsHDA702 | OsGSK2 | Kinase activity | Control lateral root formation in rice | Hou et al, |
| OsALDH2B1 | Transcriptional activity, protein stability | Responses to salt stress | Wu et al, | ||
| OsHDA705 | OsECT3 | m6A binding activity | Regulate rice cold tolerance | Ma et al, | |
| OsHDA706 | OsLOX14 | Protein stability | Broad-spectrum antiviral immunity | Yang et al, | |
| OsHDA714 | Ribosomal protein | Protein abundance or stability | ‒ | Xu et al, | |
| OsHDA714 | Glycolytic enzyme | Enzyme activity | Confer plant tolerance to heat stress | Chen Z T et al, | |
| OsHDA716 | OsbZIP46 | Transcriptional activity, protein stability | Response to drought stress; repress rice chilling tolerance | b | |
| OsHDT701 | OsRpp30 | ‒ | Broad-spectrum resistance to fungal and bacterial pathogens | Li et al, | |
| OsSRT1 | GAPDH | Transcriptional activity | Repress glycolysis | Zhang et al, | |
| Acetyltransferase | GCN5 | ADA2 | Protein Stability | Metabolic control | Yu et al, |
| N-terminal acetyltransferase | NatA | GLO1/GLO5 | Protein degradation | Promote rice thermoresponsive growth | Li et al, |
| NatA | OsHYPK | Protein level | Coordinate plant development and abiotic stress responses | Gong et al, |
Table 3. Lysine acetylation modification of non-histone proteins in rice.
| Modifying enzyme type | Enzyme | Non-histone substrate | Function of acetylation | Mechanism of action | Reference |
|---|---|---|---|---|---|
| Deacetylase | OsHDAC1/ OsHDA702 | OsGSK2 | Kinase activity | Control lateral root formation in rice | Hou et al, |
| OsALDH2B1 | Transcriptional activity, protein stability | Responses to salt stress | Wu et al, | ||
| OsHDA705 | OsECT3 | m6A binding activity | Regulate rice cold tolerance | Ma et al, | |
| OsHDA706 | OsLOX14 | Protein stability | Broad-spectrum antiviral immunity | Yang et al, | |
| OsHDA714 | Ribosomal protein | Protein abundance or stability | ‒ | Xu et al, | |
| OsHDA714 | Glycolytic enzyme | Enzyme activity | Confer plant tolerance to heat stress | Chen Z T et al, | |
| OsHDA716 | OsbZIP46 | Transcriptional activity, protein stability | Response to drought stress; repress rice chilling tolerance | b | |
| OsHDT701 | OsRpp30 | ‒ | Broad-spectrum resistance to fungal and bacterial pathogens | Li et al, | |
| OsSRT1 | GAPDH | Transcriptional activity | Repress glycolysis | Zhang et al, | |
| Acetyltransferase | GCN5 | ADA2 | Protein Stability | Metabolic control | Yu et al, |
| N-terminal acetyltransferase | NatA | GLO1/GLO5 | Protein degradation | Promote rice thermoresponsive growth | Li et al, |
| NatA | OsHYPK | Protein level | Coordinate plant development and abiotic stress responses | Gong et al, |
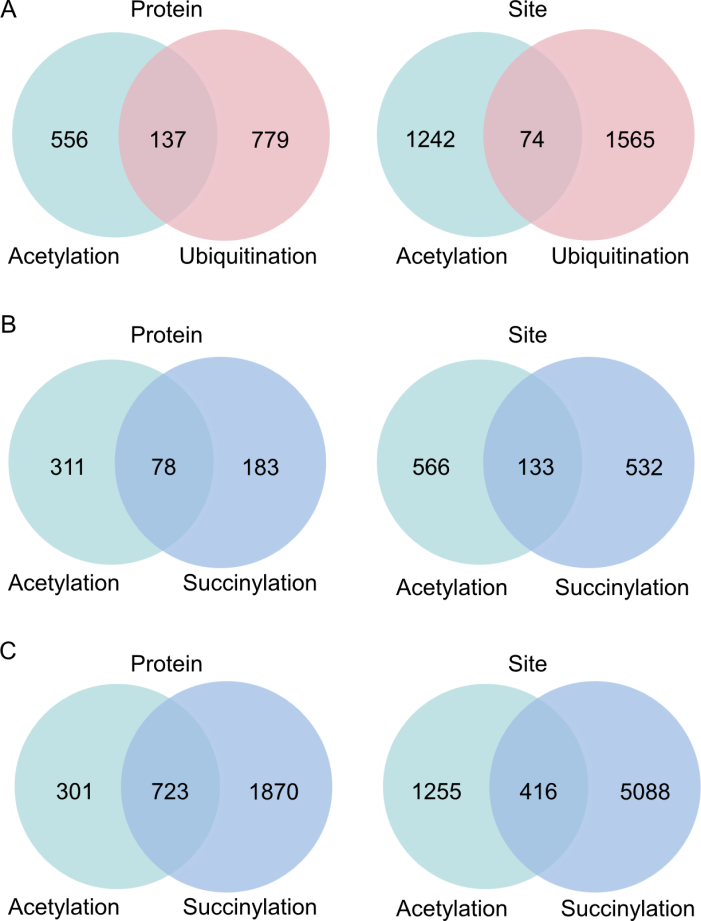
Fig. 2. Comparing sites and proteins between acetylation and other modification types. A, Comparison of acetylated and ubiquitinated groups in rice. B, Comparison of acetylated and succinylated groups in rice seeds. C, Comparison of acetylated and succinylated groups under oxidative stress in rice leaves.
| Classification of detection methods | Specific technology | Advantage | Limitation | Trend development |
|---|---|---|---|---|
| Antibody-dependent | Western blot, immunoprecipitation, immunofluorescence | Easy to operate; low cost | Dependence on antibody quality; low throughput; risk of false positives | Improving antibody specificity |
| Antibody-independent (Mass spectrometry-based) | TMT, iTRAQ, SILAC | High throughput; new site discovery; quantitative accuracy | Sample preparation is complex; require advanced data analysis | Single-cell resolution detection; plant system adaptation |
| Cutting-edge technology | Unlabeled single cell detection | Ultra-high sensitivity; spatial resolution | Immature technology; challenges in plant applications | Developing hypersensitive methods; establishing a spatiotemporal dynamic quantitative framework |
Table 4. Comparison of acetylation technologies.
| Classification of detection methods | Specific technology | Advantage | Limitation | Trend development |
|---|---|---|---|---|
| Antibody-dependent | Western blot, immunoprecipitation, immunofluorescence | Easy to operate; low cost | Dependence on antibody quality; low throughput; risk of false positives | Improving antibody specificity |
| Antibody-independent (Mass spectrometry-based) | TMT, iTRAQ, SILAC | High throughput; new site discovery; quantitative accuracy | Sample preparation is complex; require advanced data analysis | Single-cell resolution detection; plant system adaptation |
| Cutting-edge technology | Unlabeled single cell detection | Ultra-high sensitivity; spatial resolution | Immature technology; challenges in plant applications | Developing hypersensitive methods; establishing a spatiotemporal dynamic quantitative framework |
| [1] | Allfrey V G, Faulkner R, Mirsky A E. 1964. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA, 51(5): 786-794. |
| [2] | Balparda M, Elsässer M, Badia M B, et al. 2022. Acetylation of conserved lysines fine-tunes mitochondrial malate dehydrogenase activity in land plants. Plant J, 109(1): 92-111. |
| [3] | Cai J T, Culley M K, Zhao Y T, et al. 2018. The role of ubiquitination and deubiquitination in the regulation of cell junctions. Protein Cell, 9(9): 754-769. |
| [4] | Cai L J, Li B, Zhou Q T, et al. 2025. N-acetyltransferase 10 catalyzes RNA N4-acetylcytidine to regulate photosynthesis in rice. Cell Rep, 44(3): 115428. |
| [5] | Chen X Y, Duan Y H, Qiao F G, et al. 2022. A secreted fungal effector suppresses rice immunity through host histone hypoacetylation. New Phytol, 235(5): 1977-1994. |
| [6] | Chen X Y, Liu C, Wang H L, et al. 2024. Ustilaginoidea virens-secreted effector Uv1809 suppresses rice immunity by enhancing OsSRT2-mediated histone deacetylation. Plant Biotechnol J, 22(1): 148-164. |
| [7] | Chen Z T, Xu Q T, Wang J, et al. 2024. A histone deacetylase confers plant tolerance to heat stress by controlling protein lysine deacetylation and stress granule formation in rice. Cell Rep, 43(9): 114642. |
| [8] | Chung P J, Kim Y S, Jeong J S, et al. 2009. The histone deacetylase OsHDAC1 epigenetically regulates the OsNAC6 gene that controls seedling root growth in rice. Plant J, 59(5): 764-776. |
| [9] | Delker C, Stenzel I, Hause B, et al. 2006. Jasmonate biosynthesis in Arabidopsis thaliana: Enzymes, products, regulation. Plant Biol, 8(3): 297-306. |
| [10] | Ding B, del Rosario Bellizzi M, Ning Y S, et al. 2012. HDT701, a histone H4 deacetylase, negatively regulates plant innate immunity by modulating histone H4 acetylation of defense-related genes in rice. Plant Cell, 24(9): 3783-3794. |
| [11] | Feng C, Cai X W, Su Y N, et al. 2021. Arabidopsis RPD3-like histone deacetylases form multiple complexes involved in stress response. J Genet Genomics, 48(5): 369-383. |
| [12] | French M E, Koehler C F, Hunter T. 2021. Emerging functions of branched ubiquitin chains. Cell Discov, 7: 6. |
| [13] | Fukao T, Bailey-Serres J. 2004. Plant responses to hypoxia: Is survival a balancing act? Trends Plant Sci, 9(9): 449-456. |
| [14] | Gong X D, Huang Y Q, Liang Y, et al. 2022. OsHYPK-mediated protein N-terminal acetylation coordinates plant development and abiotic stress responses in rice. Mol Plant, 15(4): 740-754. |
| [15] | Guo M M, Wang S C, Liu H, et al. 2023. Histone deacetylase MdHDA6 is an antagonist in regulation of transcription factor MdTCP15 to promote cold tolerance in apple. Plant Biotechnol J, 21(11): 2254-2272. |
| [16] | Guo Y L, Tan Y Q, Qu M H, et al. 2023. OsWR2 recruits HDA704 to regulate the deacetylation of H4K8ac in the promoter of OsABI5 in response to drought stress. J Integr Plant Biol, 65(7): 1651-1669. |
| [17] | He D L, Wang Q, Li M, et al. 2016. Global proteome analyses of lysine acetylation and succinylation reveal the widespread involvement of both modification in metabolism in the embryo of germinating rice seed. J Proteome Res, 15(3): 879-890. |
| [18] | Hou J Q, Zheng X K, Ren R F, et al. 2022. The histone deacetylase 1/GSK3/SHAGGY-like kinase 2/BRASSINAZOLE-RESISTANT 1 module controls lateral root formation in rice. Plant Physiol, 189(2): 858-873. |
| [19] | Huang Y Q, Gong X D, Shi H, et al. 2025. OsHYPK/NatA-mediated N-terminal acetylation regulates the homeostasis of NLR immune protein to fine-tune rice immune responses and growth. Cell Rep, 44(5): 115719. |
| [20] | Jia M R, Meng X B, Song X G, et al. 2022. Chilling-induced phosphorylation of IPA1 by OsSAPK6 activates chilling tolerance responses in rice. Cell Discov, 8(1): 71. |
| [21] | Jones J D G, Dangl J L. 2006. The plant immune system. Nature, 444: 323-329. |
| [22] | Kovach M J, Sweeney M T, McCouch S R. 2007. New insights into the history of rice domestication. Trends Genet, 23(11): 578-587. |
| [23] | König A C, Hartl M, Boersema P J, et al. 2014. The mitochondrial lysine acetylome of Arabidopsis. Mitochondrion, 19: 252-260. |
| [24] | Li W, Xiong Y H, Lai L B, et al. 2021. The rice RNase P protein subunit Rpp30 confers broad-spectrum resistance to fungal and bacterial pathogens. Plant Biotechnol J, 19(10): 1988-1999. |
| [25] | Li X J, Ye J Y, Ma H, et al. 2018. Proteomic analysis of lysine acetylation provides strong evidence for involvement of acetylated proteins in plant meiosis and tapetum function. Plant J, 93(1): 142-154. |
| [26] | Li X T, Tang H S, Xu T, et al. 2024. N-terminal acetylation orchestrates glycolate-mediated ROS homeostasis to promote rice thermoresponsive growth. New Phytol, 243(5): 1742-1757. |
| [27] | Liang M T, Deng S L, Zhang Y, et al. 2025. Histone deacetylase 9 modulates the acetylation dynamics of phototropin 1 to fine-tune phototropic responses in plants. Plant Commun, 6(8): 101424. |
| [28] | Liang Z W, Huang Y S, Hao Y H, et al. 2025. The HISTONE ACETYLTRANSFERASE 1 interacts with CONSTANS to promote flowering in Arabidopsis. J Genet Genomics, 52(12): 1627-1637. |
| [29] | Lin A W, Man H Y. 2013. Ubiquitination of neurotransmitter receptors and postsynaptic scaffolding proteins. Neural Plast, 2013(1): 432057. |
| [30] | Liu K, Chen J J, Sun S, et al. 2023. Histone deacetylase OsHDA706 increases salt tolerance via H4K5/K8 deacetylation of OsPP2C49 in rice. J Integr Plant Biol, 65(6): 1394-1407. |
| [31] | Liu S C, Yu F C, Yang Z, et al. 2018. Establishment of dimethyl labeling-based quantitative acetylproteomics in Arabidopsis. Mol Cell Proteomics, 17(5): 1010-1027. |
| [32] | Liu Z, Zhang S M, Sun N, et al. 2015. Functional diversity of jasmonates in rice. Rice, 8: 5. |
| [33] | Ma N N, Song P Z, Liu Z Y, et al. 2025. Regulation of m6A RNA reader protein OsECT 3 activity by lysine acetylation in the cold stress response in rice. Nat Plants, 11(6): 1165-1180. |
| [34] | Meng X X, Lv Y D, Mujahid H, et al. 2018. Proteome-wide lysine acetylation identification in developing rice (Oryza sativa) seeds and protein co-modification by acetylation, succinylation, ubiquitination, and phosphorylation. BBA: Proteins Proteomics, 1866(3): 451-463. |
| [35] | Mun D G, Bhat F A, Joshi N, et al. 2024. Diversity of post-translational modifications and cell signaling revealed by single cell and single organelle mass spectrometry. Commun Biol, 7: 884. |
| [36] | Nallamilli B R R, Edelmann M J, Zhong X X, et al. 2014. Global analysis of lysine acetylation suggests the involvement of protein acetylation in diverse biological processes in rice (Oryza sativa). PLoS One, 9(2): e89283. |
| [37] | Narita T, Weinert B T, Choudhary C. 2019. Functions and mechanisms of non-histone protein acetylation. Nat Rev Mol Cell Biol, 20(3): 156-174. |
| [38] | Sang T, Zhang Z, Liu G T, et al. 2025. Navigating the landscape of plant proteomics. J Integr Plant Biol, 67(3): 740-761. |
| [39] | Satpathy S, Krug K, Beltran P M J, et al. 2021. A proteogenomic portrait of lung squamous cell carcinoma. Cell, 184: 4348-4371. |
| [40] | Sinha K, Kaur R, Singh N, et al. 2020. Mobilization of storage lipid reserve and expression analysis of lipase and lipoxygenase genes in rice (Oryza sativa var. Pusa Basmati 1) bran during germination. Phytochemistry, 180: 112538. |
| [41] | Soufi B, Soares N C, Ravikumar V, et al. 2012. Proteomics reveals evidence of cross-talk between protein modifications in bacteria: Focus on acetylation and phosphorylation. Curr Opin Microbiol, 15(3): 357-363. |
| [42] | Sun Y, Xie Z Z, Jin L, et al. 2024a. Histone deacetylase OsHDA716 represses rice chilling tolerance by deacetylating OsbZIP46 to reduce its transactivation function and protein stability. Plant Cell, 36(5): 1913-1936. |
| [43] | Sun Y, Gu X Y, Qu C F, et al. 2024b. OsPUB75-OsHDA716 mediates deactivation and degradation of OsbZIP46 to negatively regulate drought tolerance in rice. Plant Physiol, 197(1): kiae545. |
| [44] | Tang B Z, Liu C Y, Li Z Q, et al. 2021. Multilayer regulatory landscape during pattern-triggered immunity in rice. Plant Biotechnol J, 19(12): 2629-2645. |
| [45] | Tang N, Zhang H, Li X H, et al. 2012. Constitutive activation of transcription factor OsbZIP46 improves drought tolerance in rice. Plant Physiol, 158(4): 1755-1768. |
| [46] | Tong H N, Liu L C, Jin Y, et al. 2012. DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/ SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell, 24(6): 2562-2577. |
| [47] | Tuteja N. 2007. Abscisic acid and abiotic stress signaling. Plant Signal Behav, 2(3): 135-138. |
| [48] | Ullah F, Xu Q T, Zhao Y, et al. 2021. Histone deacetylase HDA710 controls salt tolerance by regulating ABA signaling in rice. J Integr Plant Biol, 63(3): 451-467. |
| [49] | Waadt R, Seller C A, Hsu P K, et al. 2022. Plant hormone regulation of abiotic stress responses. Nat Rev Mol Cell Biol, 23(10): 680-694. |
| [50] | Walley J W, Shen Z X, McReynolds M R, et al. 2018. Fungal-induced protein hyperacetylation in maize identified by acetylome profiling. Proc Natl Acad Sci USA, 115(1): 210-215. |
| [51] | Wang H C, Jiao X M, Kong X Y, et al. 2020. The histone deacetylase HDA703 interacts with OsBZR1 to regulate rice brassinosteroid signaling, growth and heading date through repression of Ghd7 expression. Plant J, 104(2): 447-459. |
| [52] | Wang Y F, Hou Y X, Qiu J H, et al. 2017. A quantitative acetylomic analysis of early seed development in rice (Oryza sativa L.). Int J Mol Sci, 18(7): 1376. |
| [53] | Wasternack C, Song S S. 2017. Jasmonates: Biosynthesis, metabolism, and signaling by proteins activating and repressing transcription. J Exp Bot, 68(6): 1303-1321. |
| [54] | Wu Y Q, Hou J Q, Xiao H Z, et al. 2025. OsHDAC1 deacetylates the aldehyde dehydrogenase OsALDH2B1, repressing OsGR3 and decreasing salt tolerance in rice. Plant Physiol, 198(1): kiaf149. |
| [55] | Xiong Y H, Peng X J, Cheng Z Y, et al. 2016. A comprehensive catalog of the lysine-acetylation targets in rice (Oryza sativa) based on proteomic analyses. J Proteomics, 138: 20-29. |
| [56] | Xu Q T, Liu Q, Chen Z T, et al. 2021. Histone deacetylases control lysine acetylation of ribosomal proteins in rice. Nucleic Acids Res, 49(8): 4613-4628. |
| [57] | Xu Q T, Wang Y J, Chen Z T, et al. 2023a. ROS-stimulated protein lysine acetylation is required for crown root development in rice. J Adv Res, 48: 33-46. |
| [58] | Xu Q T, Yue Y P, Liu B, et al. 2023b. ACL and HAT1 form a nuclear module to acetylate histone H4K5 and promote cell proliferation. Nat Commun, 14(1): 3265. |
| [59] | Xu Q T, Ma X, Wei X L, et al. 2025. Histone H4K8hib modification promotes gene expression and regulates rice immunity. Mol Plant, 18(1): 9-13. |
| [60] | Xue C, Liu S, Chen C, et al. 2018. Global proteome analysis links lysine acetylation to diverse functions in Oryza sativa. Proteomics, 18(1): 1700036. |
| [61] | Xue C, Zhao X R, Chen X, et al. 2024. Histone acetyltransferase GCN5 regulates rice growth and development and enhances salt tolerance. Rice Sci, 31(6): 688-699. |
| [62] | Yan Z, Shen Z, Gao Z F, et al. 2020. A comprehensive analysis of the lysine acetylome reveals diverse functions of acetylated proteins during de-etiolation in Zea mays. J Plant Physiol, 248: 153158. |
| [63] | Yang Z H, Du J, Tan X X, et al. 2024. Histone deacetylase OsHDA706 orchestrates rice broad-spectrum antiviral immunity and is impeded by a viral effector. Cell Rep, 43(3): 113838. |
| [64] | Yang Z R, Huang Y, Yang J L, et al. 2020. Jasmonate signaling enhances RNA silencing and antiviral defense in rice. Cell Host Microbe, 28(1): 89-103.e8. |
| [65] | Yu Y, Zhao F, Yue Y P, et al. 2024. Lysine acetylation of histone acetyltransferase adaptor protein ADA 2 is a mechanism of metabolic control of chromatin modification in plants. Nat Plants, 10(3): 439-452. |
| [66] | Zeng L R, Vega-Sánchez M E, Zhu T, et al. 2006. Ubiquitination-mediated protein degradation and modification: An emerging theme in plant-microbe interactions. Cell Res, 16(5): 413-426. |
| [67] | Zhang H, Lu Y, Zhao Y, et al. 2016. OsSRT1 is involved in rice seed development through regulation of starch metabolism gene expression. Plant Sci, 248: 28-36. |
| [68] | Zhang H, Zhao Y, Zhou D X. 2017. Rice NAD+-dependent histone deacetylase OsSRT1 represses glycolysis and regulates the moonlighting function of GAPDH as a transcriptional activator of glycolytic genes. Nucleic Acids Res, 45(21): 12241-12255. |
| [69] | Zhang M, Tan F Q, Fan Y J, et al. 2022. Acetylome reprograming participates in the establishment of fruit metabolism during polyploidization in citrus. Plant Physiol, 190(4): 2519-2538. |
| [70] | Zhang N, Hu J C, Liu Z S, et al. 2024. Sir2-mediated cytoplasmic deacetylation facilitates pathogenic fungi infection in host plants. New Phytol, 241(4): 1732-1746. |
| [71] | Zhang Z H, Tan M J, Xie Z Y, et al. 2011. Identification of lysine succinylation as a new post-translational modification. Nat Chem Biol, 7(1): 58-63. |
| [72] | Zhao Y, Hu Y F, Dai M Q, et al. 2009. The WUSCHEL-related homeobox gene WOX11 is required to activate shoot-borne crown root development in rice. Plant Cell, 21(3): 736-748. |
| [73] | Zhao Y, Cheng S F, Song Y L, et al. 2015. The interaction between rice ERF3 and WOX11 promotes crown root development by regulating gene expression involved in cytokinin signaling. Plant Cell, 27(9): 2469-2483. |
| [74] | Zhou H, Finkemeier I, Guan W X, et al. 2018. Oxidative stress-triggered interactions between the succinyl- and acetyl-proteomes of rice leaves. Plant Cell Environ, 41(5): 1139-1153. |
| [75] | Zhu J K. 2016. Abiotic stress signaling and responses in plants. Cell, 167(2): 313-324. |
| [76] | Zhu L Y, Cheng H, Peng G Q, et al. 2020. Ubiquitinome profiling reveals the landscape of ubiquitination regulation in rice young panicles. Genom Proteomics Bioinformatics, 18(3): 305-320. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||