
Rice Science ›› 2025, Vol. 32 ›› Issue (2): 137-142.DOI: 10.1016/j.rsci.2024.12.001
收稿日期:2024-07-12
接受日期:2024-10-10
出版日期:2025-03-28
发布日期:2025-04-14
. [J]. Rice Science, 2025, 32(2): 137-142.
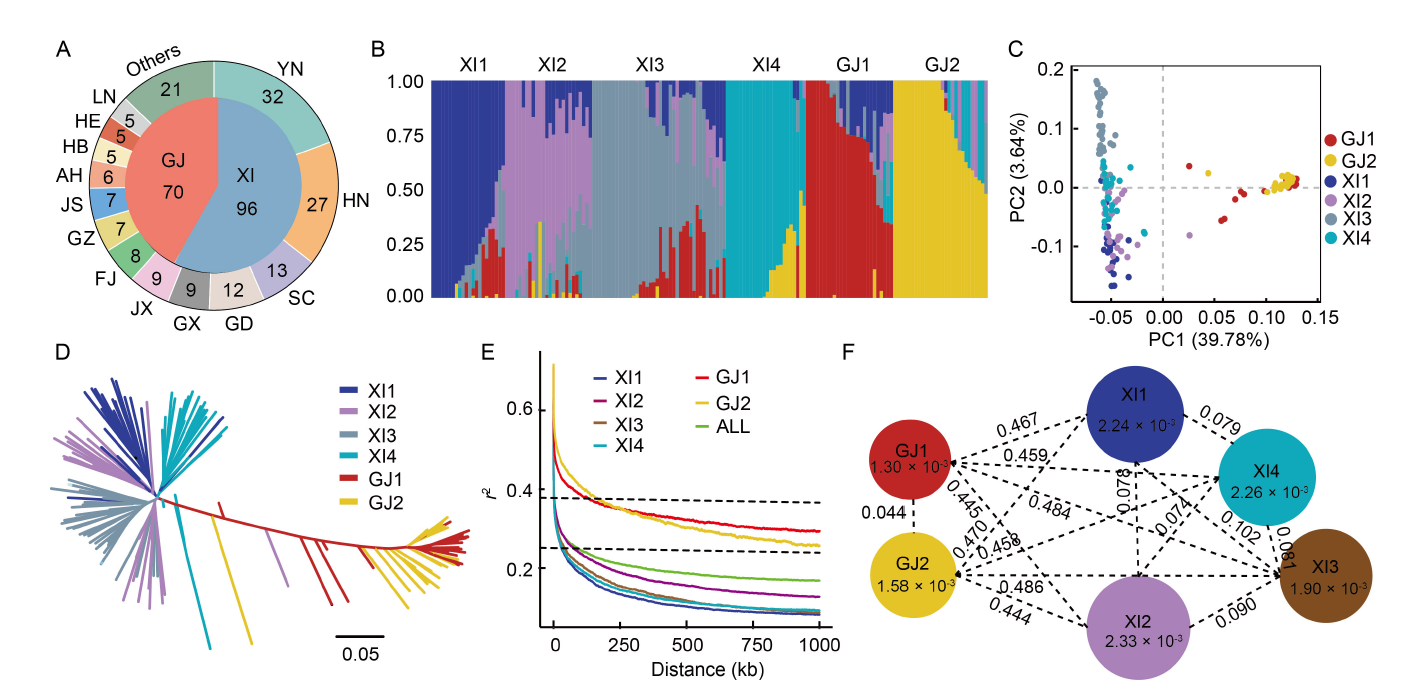
Fig. 1. Population structure and genetic diversity estimates for 166 mini-core Chinese rice germplasms. A, Geographic distribution of 166 rice mini-core germplasms in China. YN, Yunnan Province; HN, Hunan Province; SC, Sichuan Province; GD, Guangdong Province; GX, Guangxi Province; JX, Jiangxi Province; FJ, Fujian Province; GZ, Guizhou Province; JS, Jiangsu Province; AH, Anhui Province; HB, Hubei Province; HE, Hebei Province; LN, Liaoning Province. B, Estimated population structure for rice varieties (K = 6). C, Principal component analysis (PCA) plot of the first two principal components (PC1 and PC2) for rice varieties. D, Phylogenetic tree constructed based on inferred genetic distances. E, Genome-wide average linkage disequilibrium (LD) map for all germplasms (All) and the XI1, XI2, XI3, XI4, GJ1, and GJ2 subpopulations, respectively. The dotted line indicates that LD decrease to half of its maximum value, with the GJ subpopulation at 0.37 and the XI subpopulation at 0.25. F, Genetic diversity and subpopulation differentiation of 166 germplasms. The circle size and the number within each circle represent the genetic diversity (π) of each subpopulation; the number displayed on the horizontal line represents the calculated genetic differentiation coefficient (Fst) value between the two subpopulations. XI, Indica rice variety; GJ, Japonica rice variety.
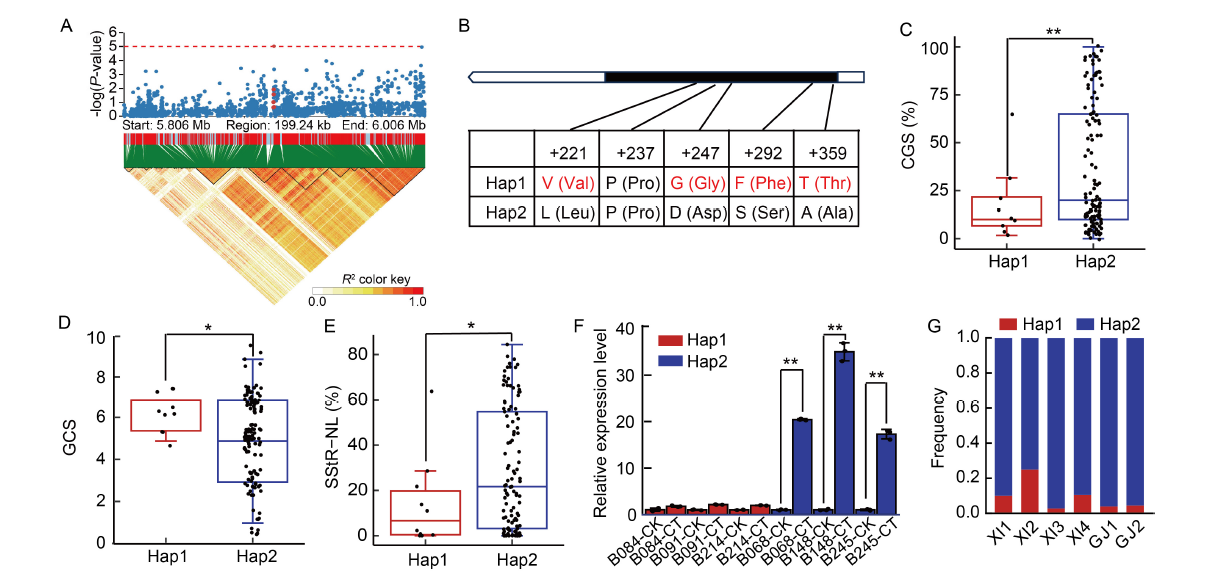
Fig. 2. Localization of candidate gene OsOPR6 on chromosome 6. A, Partial Manhattan plot (upper panel) and linkage disequilibrium heatmap (lower panel) surrounding the peak on chromosome 6. The red dot indicates the position of a missense variant within the OsOPR6 gene. B, Gene structure and polymorphisms identified in OsOPR6. C‒E, Boxplots depicting the germination rate at low temperature at the germination stage (CGS, C), the grade of cold tolerance at the seedling stage (GCS, D), and the proportion of seed-setting rate per panicle under natural low temperature conditions at the booting and flowering stage (SStR-NL, E) for different haplotypes of OsOPR6. * and ** indicate significant differences between haplotypes at P < 0.05 and P < 0.01, respectively, as determined by a t-test. F, Expression analysis of OsOPR6 by qRT-PCR. UBI was used as an internal control. Data represent Mean ± SD (n = 3). RNA was extracted from rice young leaves under control and cold stress conditions for 8 h. CK, Control; CT, Cold stress for 8 h. B084, Bawangbian 1; B091, Xiaohonggu; B214, Xuanenchangtanqingzhan; B068, Gaoyangdiandao dahongmang; B148, Lengshuinuo; B245, Shanjiugu. * and ** indicate significant differences between the control and stress conditions at P < 0.05 and P < 0.01, respectively, as determined by a t-test. G, Frequency distribution of haplotypes Hap1 and Hap2 across different subpopulations.
| [1] | Chen F, Dong G J, Wang F, et al. 2021. A β-ketoacyl carrier protein reductase confers heat tolerance via the regulation of fatty acid biosynthesis and stress signaling in rice. New Phytol, 232(2): 655-672. |
| [2] | Cui D, Zhou H, Ma X D, et al. 2022. Genomic insights on the contribution of introgressions from Xian/Indica to the genetic improvement of Geng/Japonica rice cultivars. Plant Commun, 3(3): 100325. |
| [3] | Fukuoka S, Saka N, Koga H, et al. 2009. Loss of function of a proline-containing protein confers durable disease resistance in rice. Science, 325: 998-1001. |
| [4] | Huang L J, Hua K, Xu R, et al. 2021. The LARGE2-APO1/APO2 regulatory module controls panicle size and grain number in rice. Plant Cell, 33(4): 1212-1228. |
| [5] | Ji C H, Ji Z Y, Liu B, et al. 2020. Xa1 allelic R genes activate rice blight resistance suppressed by interfering TAL effectors. Plant Commun, 1(4): 100087. |
| [6] | Jing H W, Yang X L, Zhang J, et al. 2015. Peptidyl-prolyl isomerization targets rice Aux/IAAs for proteasomal degradation during auxin signalling. Nat Commun, 6: 7395. |
| [7] | Lee S, Choi S C, An G. 2008. Rice SVP-group MADS-box proteins, OsMADS22 and OsMADS55, are negative regulators of brassinosteroid responses. Plant J, 54(1): 93-105. |
| [8] | Lee S K, Lee S M, Kim M H, et al. 2022. Genome-wide analysis of cyclic nucleotide-gated channel genes related to pollen development in rice. Plants, 11(22): 3145. |
| [9] | Li X M, Chao D Y, Wu Y, et al. 2015. Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat Genet, 47(7): 827-833. |
| [10] | Liu C T, Wu Y B, Wang X P. 2012. bZIP transcription factor OsbZIP52/RISBZ5: A potential negative regulator of cold and drought stress response in rice. Planta, 235(6): 1157-1169. |
| [11] | Liu X H, Lyu Y S, Yang W P, et al. 2020. A membrane-associated NAC transcription factor OsNTL3 is involved in thermotolerance in rice. Plant Biotechnol J, 18(5): 1317-1329. |
| [12] | Liu Y, Liu B, Zhu X Y, et al. 2013. Fine-mapping and molecular marker development for Pi56(t), a NBS-LRR gene conferring broad-spectrum resistance to Magnaporthe oryzae in rice. Theor Appl Genet, 126(4): 985-998. |
| [13] | Ma Y, Dai X Y, Xu Y Y, et al. 2015. COLD1 confers chilling tolerance in rice. Cell, 160(6): 1209-1221. |
| [14] | Mao D H, Xin Y Y, Tan Y J, et al. 2019. Natural variation in the HAN1 gene confers chilling tolerance in rice and allowed adaptation to a temperate climate. Proc Natl Acad Sci USA, 116(9): 3494-3501. |
| [15] | Okuyama Y, Kanzaki H, Abe A, et al. 2011. A multifaceted genomics approach allows the isolation of the rice Pia-blast resistance gene consisting of two adjacent NBS-LRR protein genes. Plant J, 66(3): 467-479. |
| [16] | Qin R, Zeng D D, Yang C C, et al. 2018. LTBSG1, a new allele of BRD2, regulates panicle and grain development in rice by brassinosteroid biosynthetic pathway. Genes, 9(6): 292. |
| [17] | Qu S H, Liu G F, Zhou B, et al. 2006. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine- rich repeat protein and is a member of a multigene family in rice. Genetics, 172(3): 1901-1914. |
| [18] | Richter A, Streubel J, Blücher C, et al. 2014. A TAL effector repeat architecture for frameshift binding. Nat Commun, 5: 3447. |
| [19] | Sasaki T, Burr B. 2000. International rice genome sequencing project: The effort to completely sequence the rice genome. Curr Opin Plant Biol, 3(2): 138-141. |
| [20] | Shao G N, Lu Z F, Xiong J S, et al. 2019. Tiller bud formation regulators MOC1 and MOC3 cooperatively promote tiller bud outgrowth by activating FON1 expression in rice. Mol Plant, 12(8): 1090-1102. |
| [21] | Shi C L, Dong N Q, Guo T, et al. 2020. A quantitative trait locus GW6 controls rice grain size and yield through the gibberellin pathway. Plant J, 103(3): 1174-1188. |
| [22] | Wang C L, Cui D, Tang C F, et al. 2022. Evaluation of rice landraces from different altitudes in Yunnan for cold tolerance at different growing stages. J Plant Genet Resour, 23(1): 83-91. (in Chinese with English abstract) |
| [23] | Wang Y K, Yuan G L, Yuan S H, et al. 2016. TaOPR2 encodes a 12-oxo-phytodienoic acid reductase involved in the biosynthesis of jasmonic acid in wheat (Triticum aestivum L.). Biochem Biophys Res Commun, 470(1): 233-238. |
| [24] | Wang Y W, Deng C, Ai P F, et al. 2021. ALM1, encoding a Fe- superoxide dismutase, is critical for rice chloroplast biogenesis and drought stress response. Crop J, 9(5): 1018-1029. |
| [25] | Wang Y X, Xiong G S, Hu J, et al. 2015. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet, 47(8): 944-948. |
| [26] | Xiong Y Z, Xie J, Zhang X B, et al. 2021. PLASTOCHRON1 regulates leaf inclination through brassinolide pathway in Oryza sativa. Crop Sci, 61(2): 1280-1288. |
| [27] | Yang L M, Liu H L, Zhao H W, et al. 2019. Mapping quantitative trait loci and meta-analysis for cold tolerance in rice at booting stage. Euphytica, 215(5): 89. |
| [28] | Yang L M, Wang J G, Han Z H, et al. 2021. Combining QTL-seq and linkage mapping to fine map a candidate gene in qCTS6 for cold tolerance at the seedling stage in rice. BMC Plant Biol, 21(1): 278. |
| [29] | Ye J H, Zhang M C, Yuan X P, et al. 2022. Genomic insight into genetic changes and shaping of major inbred rice cultivars in China. New Phytol, 236(6): 2311-2326. |
| [30] | Zhang J Y, Li X M, Lin H X, et al. 2019. Crop improvement through temperature resilience. Annu Rev Plant Biol, 70: 753-780. |
| [31] | Zhao J L, Zhang S H, Dong J F, et al. 2017. A novel functional gene associated with cold tolerance at the seedling stage in rice. Plant Biotechnol J, 15(9): 1141-1148. |
| [32] | Zong W B, Ren D, Huang M H, et al. 2021. Strong photoperiod sensitivity is controlled by cooperation and competition among Hd1, Ghd7 and DTH8 in rice heading. New Phytol, 229(3): 1635-1649. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||