
Rice Science ›› 2026, Vol. 33 ›› Issue (2): 186-202.DOI: 10.1016/j.rsci.2026.01.001
收稿日期:2025-06-26
接受日期:2025-12-12
出版日期:2026-03-28
发布日期:2026-04-01
. [J]. Rice Science, 2026, 33(2): 186-202.
| SUB/AG | Gene | Location (bp) | Note | Reference |
|---|---|---|---|---|
| AG | OsmiR393a | Chr. 01: 2 309 149-2 309 263 | MicroRNA393a | Guo et al, |
| AG | OsMYBS1 | Chr. 01: 18 755 268-18 756 900 | MYB family transcription factor | Hong et al, |
| AG | OsCBL10 | Chr. 01: 29 568 547-29 571 484 | CBL protein 10 | Ye et al, |
| AG | OsHXK6 | Chr. 01: 31 009 006-31 014 001 | Hexokinase gene | Hsu and Tung, |
| AG | OsCNL1 | Chr. 03: 1 706 051-1 708 336 | Peroxisomal cinnamate:CoA ligase 1 | Wang et al, |
| AG | OsEBP89 | Chr. 03: 4 343 725-4 345 716 | EREBP transcription factor | Zhang et al, |
| AG | Osa-miR167b | Chr. 03: 30 546 928-30 547 090 | MicroRNA167b | Lee et al, |
| AG | OsSnRK1A, OsSnRK1αA, OSK1 | Chr. 05: 26 346 997-26 351 611 | Sucrose non-fermenting-1 related protein kinase 1 | Lu et al, |
| AG | OsFLZ18 | Chr. 06: 1 361 364-1 362 509 | FLZ protein | Zhang et al, |
| AG | OsGH3.8, OsGH3-8, OsMGH3 | Chr. 07: 24 149 649-24 152 079 | Indole-3-acetic acid-amido (IAA) synthetase, IAA inactivation‐related gene | Jain et al, |
| AG | OsCLSY1 | Chr. 07: 29 465 173-29 475 499 | CLASSY1 | Castano-Duque et al, |
| AG | RAmy3D, AmyII-4, OsAmy3D | Chr. 08: 23 340 676-23 343 533 | Alpha-amylase isozyme 3D | Kumar et al, |
| AG | OsTPP7 | Chr. 09: 12 251 875-12 254 087 | Trehalose-6-phosphate phosphatase | Kretzschmar et al, |
| AG | OsbZIP72 | Chr. 09: 17 189 976-17 194 013 | bZIP transcription factor | Wang et al, |
| AG | OsCNL2 | Chr. 09: 22 082 637-22 084 897 | Peroxisomal cinnamate:CoA ligase 2 | Wang et al, |
| AG | OsMYBS2 | Chr. 10: 22 174 581-22 176 688 | MYB family transcription factor | Chen et al, |
| AG | OsCIPK15 | Chr. 11: 630 580-633 695 | CBL-interacting protein kinase 15 | Lee et al, |
| AG | ADH1 | Chr. 11: 5 712 641-5 716 288 | Alcohol dehydrogenase gene | Saika et al, |
| AG | OsUGT75A | Chr. 11: 14 848 226-14 849 740 | UDP-glucosyltransferase | He et al, |
| AG | OsGF14h | Chr. 11: 23 553 560-23 558 464 | 14-3-3/GF14 protein | Sun et al, |
| SUB | OsACS5 | Chr. 01: 5 002 118-5 005 727 | Aminocyclopropane-1-carboxylate synthase | Zhou et al, |
| SUB | SLRL1 | Chr. 01: 26 044 166-26 045 843 | GRAS-domain protein, SLR1 like-1 | Fukao and Bailey-Serres, |
| SUB | sd1, OsGA20ox2 | Chr. 01: 38 382 382-38 385 504 | DGWG dwarf, semidwarf-1, and gibberellin 20-oxidase gene | Kuroha et al, |
| SUB | CDPK5 | Chr. 02: 28 086 381-28 092 152 | Group I calcium-dependent protein kinase | Yamauchi et al, |
| SUB | OsABA8ox1, CYP707A5 | Chr. 02: 28 995 717-28 998 581 | ABA 8′-hydroxylas | Yang and Choi, Saika et al, |
| SUB | OsMAPK3 | Chr. 03: 9 847 700-9 850 473 | Mitogen-activated protein | Singh and Sinha, |
| SUB | OsEIL1, OsEIL1a | Chr. 03: 11 771 261-11 773 192 | Ethylene-insensitive | Kurokawa et al, |
| SUB | OsERF66 | Chr. 03: 12 713 329-12 714 584 | Group VII ethylene response factor | Lin et al, |
| SUB | OsERS1 | Chr. 03: 28 169 807-28 174 250 | Ethylene responsive sensor 1 | Yu et al, |
| SUB | OsETR2 | Chr. 04: 4 738 375-4 742 348 | Ethylene responsive sensor 2 | Yu et al, |
| SUB | OsCDPK13, OsCPK13, OsCDPK7 | Chr. 04: 29 531 223-29 536 492 | Calcium-dependent protein kinase | Yamauchi et al, |
| SUB | OsHIGD2 | Chr. 07: 28 484 045-28 485 201 | Oryza sativa HIGD | Hwang and Choi, |
| SUB | OsERF67 | Chr. 07: 28 540 697-28 541 796 | Group VII ethylene response factor | Lin et al, |
| SUB | Sub1 | Chr. 09: 6 387 891-6 389 789 | Submergence 1 | Xu et al, |
| SUB | OsGGT | Chr. 10: 21 798 920-21 802 986 | Submergence-induced gene, homologous to glycogenin glucosyltransferase | Qi et al, |
| SUB | OsHSD1, DRP7, LGF1 | Chr. 11: 17 785 138-17 792 672 | Hydroxysteroid dehydrogenase, leaf gas film 1, and dripping wet leaf 7 | Kurokawa et al, |
| SUB | OsrbohH, Osrboh9 | Chr. 12: 21 648 123-21 653 570 | NADPH oxidase, respiratory burst oxidase homolog | Yamauchi et al, |
| SUB | SNORKEL1 | Chr. 12: 24 966 354-26 108 004 | Ethylene response factor | Hattori et al, |
Table 1. Details of physiological and molecularly regulated genes involved in anaerobic germination (AG) and submergence (SUB) stresses.
| SUB/AG | Gene | Location (bp) | Note | Reference |
|---|---|---|---|---|
| AG | OsmiR393a | Chr. 01: 2 309 149-2 309 263 | MicroRNA393a | Guo et al, |
| AG | OsMYBS1 | Chr. 01: 18 755 268-18 756 900 | MYB family transcription factor | Hong et al, |
| AG | OsCBL10 | Chr. 01: 29 568 547-29 571 484 | CBL protein 10 | Ye et al, |
| AG | OsHXK6 | Chr. 01: 31 009 006-31 014 001 | Hexokinase gene | Hsu and Tung, |
| AG | OsCNL1 | Chr. 03: 1 706 051-1 708 336 | Peroxisomal cinnamate:CoA ligase 1 | Wang et al, |
| AG | OsEBP89 | Chr. 03: 4 343 725-4 345 716 | EREBP transcription factor | Zhang et al, |
| AG | Osa-miR167b | Chr. 03: 30 546 928-30 547 090 | MicroRNA167b | Lee et al, |
| AG | OsSnRK1A, OsSnRK1αA, OSK1 | Chr. 05: 26 346 997-26 351 611 | Sucrose non-fermenting-1 related protein kinase 1 | Lu et al, |
| AG | OsFLZ18 | Chr. 06: 1 361 364-1 362 509 | FLZ protein | Zhang et al, |
| AG | OsGH3.8, OsGH3-8, OsMGH3 | Chr. 07: 24 149 649-24 152 079 | Indole-3-acetic acid-amido (IAA) synthetase, IAA inactivation‐related gene | Jain et al, |
| AG | OsCLSY1 | Chr. 07: 29 465 173-29 475 499 | CLASSY1 | Castano-Duque et al, |
| AG | RAmy3D, AmyII-4, OsAmy3D | Chr. 08: 23 340 676-23 343 533 | Alpha-amylase isozyme 3D | Kumar et al, |
| AG | OsTPP7 | Chr. 09: 12 251 875-12 254 087 | Trehalose-6-phosphate phosphatase | Kretzschmar et al, |
| AG | OsbZIP72 | Chr. 09: 17 189 976-17 194 013 | bZIP transcription factor | Wang et al, |
| AG | OsCNL2 | Chr. 09: 22 082 637-22 084 897 | Peroxisomal cinnamate:CoA ligase 2 | Wang et al, |
| AG | OsMYBS2 | Chr. 10: 22 174 581-22 176 688 | MYB family transcription factor | Chen et al, |
| AG | OsCIPK15 | Chr. 11: 630 580-633 695 | CBL-interacting protein kinase 15 | Lee et al, |
| AG | ADH1 | Chr. 11: 5 712 641-5 716 288 | Alcohol dehydrogenase gene | Saika et al, |
| AG | OsUGT75A | Chr. 11: 14 848 226-14 849 740 | UDP-glucosyltransferase | He et al, |
| AG | OsGF14h | Chr. 11: 23 553 560-23 558 464 | 14-3-3/GF14 protein | Sun et al, |
| SUB | OsACS5 | Chr. 01: 5 002 118-5 005 727 | Aminocyclopropane-1-carboxylate synthase | Zhou et al, |
| SUB | SLRL1 | Chr. 01: 26 044 166-26 045 843 | GRAS-domain protein, SLR1 like-1 | Fukao and Bailey-Serres, |
| SUB | sd1, OsGA20ox2 | Chr. 01: 38 382 382-38 385 504 | DGWG dwarf, semidwarf-1, and gibberellin 20-oxidase gene | Kuroha et al, |
| SUB | CDPK5 | Chr. 02: 28 086 381-28 092 152 | Group I calcium-dependent protein kinase | Yamauchi et al, |
| SUB | OsABA8ox1, CYP707A5 | Chr. 02: 28 995 717-28 998 581 | ABA 8′-hydroxylas | Yang and Choi, Saika et al, |
| SUB | OsMAPK3 | Chr. 03: 9 847 700-9 850 473 | Mitogen-activated protein | Singh and Sinha, |
| SUB | OsEIL1, OsEIL1a | Chr. 03: 11 771 261-11 773 192 | Ethylene-insensitive | Kurokawa et al, |
| SUB | OsERF66 | Chr. 03: 12 713 329-12 714 584 | Group VII ethylene response factor | Lin et al, |
| SUB | OsERS1 | Chr. 03: 28 169 807-28 174 250 | Ethylene responsive sensor 1 | Yu et al, |
| SUB | OsETR2 | Chr. 04: 4 738 375-4 742 348 | Ethylene responsive sensor 2 | Yu et al, |
| SUB | OsCDPK13, OsCPK13, OsCDPK7 | Chr. 04: 29 531 223-29 536 492 | Calcium-dependent protein kinase | Yamauchi et al, |
| SUB | OsHIGD2 | Chr. 07: 28 484 045-28 485 201 | Oryza sativa HIGD | Hwang and Choi, |
| SUB | OsERF67 | Chr. 07: 28 540 697-28 541 796 | Group VII ethylene response factor | Lin et al, |
| SUB | Sub1 | Chr. 09: 6 387 891-6 389 789 | Submergence 1 | Xu et al, |
| SUB | OsGGT | Chr. 10: 21 798 920-21 802 986 | Submergence-induced gene, homologous to glycogenin glucosyltransferase | Qi et al, |
| SUB | OsHSD1, DRP7, LGF1 | Chr. 11: 17 785 138-17 792 672 | Hydroxysteroid dehydrogenase, leaf gas film 1, and dripping wet leaf 7 | Kurokawa et al, |
| SUB | OsrbohH, Osrboh9 | Chr. 12: 21 648 123-21 653 570 | NADPH oxidase, respiratory burst oxidase homolog | Yamauchi et al, |
| SUB | SNORKEL1 | Chr. 12: 24 966 354-26 108 004 | Ethylene response factor | Hattori et al, |
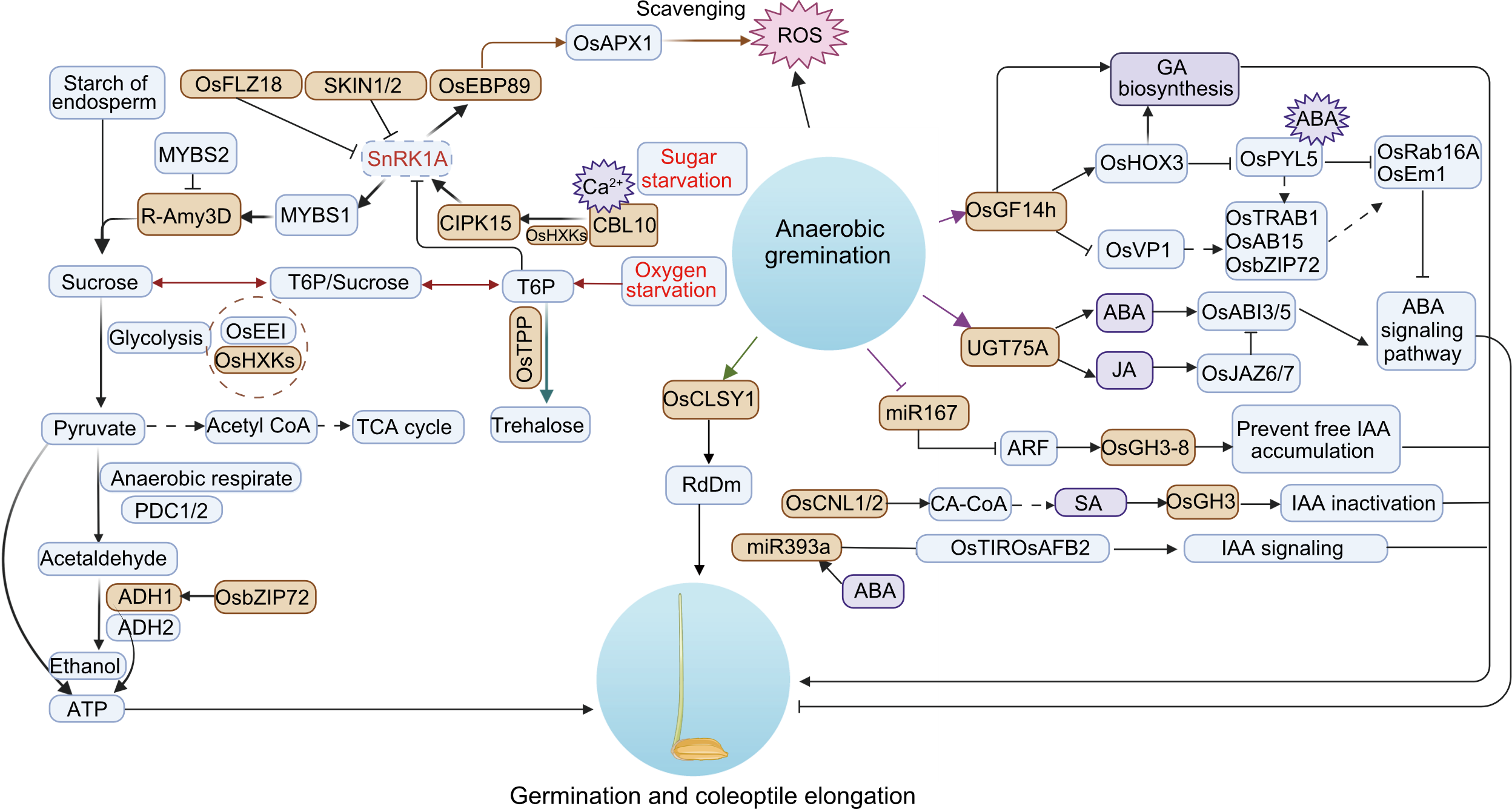
Fig. 1. Regulatory mechanisms during the germination stage under anaerobic stress. ABA, Abscisic acid; ATP, Adenosine triphosphate; CA-CoA, Cinnamoyl-CoA; GA, Gibberellin; IAA, Indole-3-acetic acid; JA, Jasmonic acid; ROS, Reactive oxygen species; SA, Salicylic acid; T6P, Trehalose 6-phosphate; TCA, Tricarboxylic acid cycle. The solid arrows, dashed arrows, and arrowheads represent step-by-step biochemical reactions, simplified biochemical reactions, and invalid biochemical reaction, respectively. Different colored arrows represent different biochemical pathways.
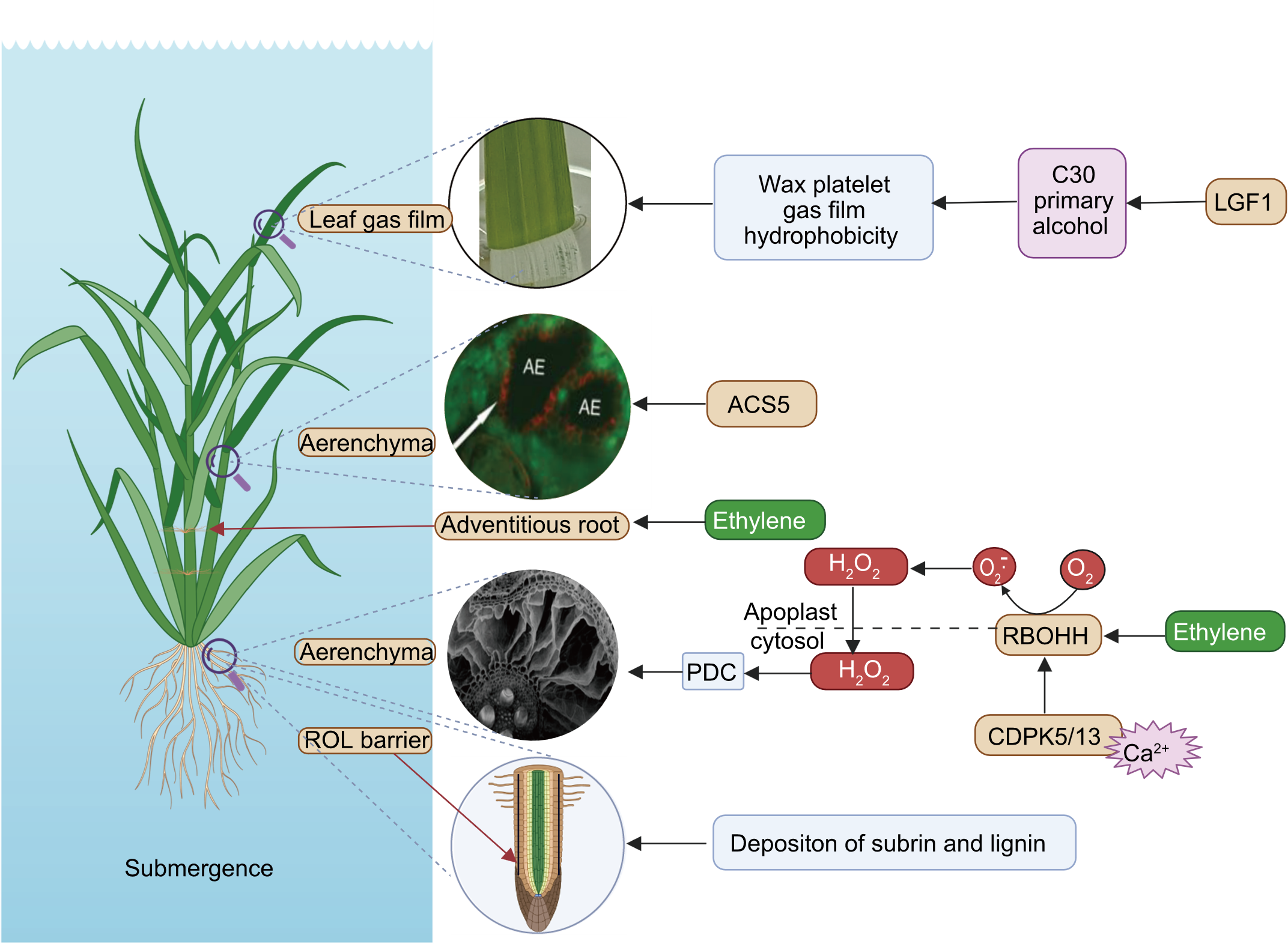
Fig. 3. Morphological traits associated with submergence adaptation in rice. LGF1, Leaf gas film 1; ROL, Radial oxygen loss; PDC, Pyruvate decarboxylase; CDPK, Calcium-dependent protein kinase; ACS5, 1-Aminocyclopropane-1-carboxylate synthase 5; RBOHH, Respiratory burst oxidase homolog; AE, Aerenchyma.
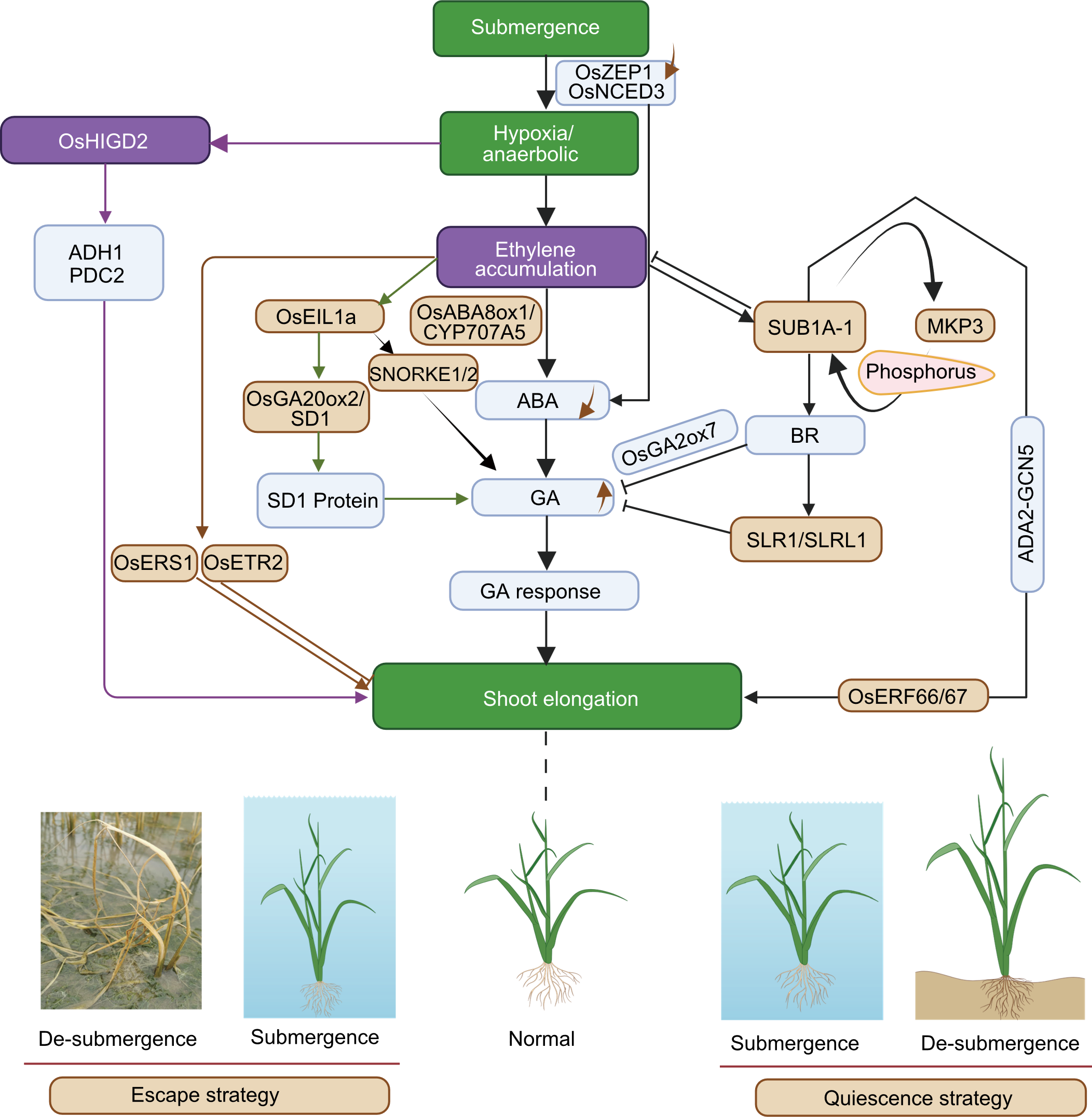
Fig. 4. Genetic and molecular regulatory mechanisms underlying two survival strategies of rice under submergence. ABA, Abscisic acid; BR, Brassinosteroids; GA, Gibberellin. The solid arrows and arrowheads represent step-by-step biochemical reactions and invalid biochemical reaction, respectively. Different colored arrows represent different biochemical pathways.
| [1] | Alpuerto J B, Fukuda M, Li S, et al. 2022. The submergence tolerance regulator SUB1A differentially coordinates molecular adaptation to submergence in mature and growing leaves of rice (Oryza sativa L.). Plant J, 110(1): 71-87. |
| [2] | Angaji S A, Septiningsih E M, Mackill D J, et al. 2010. QTLs associated with tolerance of flooding during germination in rice (Oryza sativa L.). Euphytica, 172(2): 159-168. |
| [3] | Ashraf M A, Ahmad M S A, Ashraf M, et al. 2011. Alleviation of waterlogging stress in upland cotton (Gossypium hirsutum L.) by exogenous application of potassium in soil and as a foliar spray. Crop Pasture Sci, 62(1): 25-38. |
| [4] | Bailey-Serres J, Fukao T, Ronald P, et al. 2010. Submergence tolerant rice: SUB1’s journey from landrace to modern cultivar. Rice, 3(2/3): 138-147. |
| [5] | Bailey-Serres J, Fukao T, Gibbs D J, et al. 2012. Making sense of low oxygen sensing. Trends Plant Sci, 17(3): 129-138. |
| [6] | Bairagi S, Mishra A K, Durand-Morat A. 2020. Climate risk management strategies and food security: Evidence from Cambodian rice farmers. Food Policy, 95: 101935. |
| [7] | Bhaduri D, Chakraborty K, Nayak A K, et al. 2020. Alteration in plant spacing improves submergence tolerance in Sub1 and non-Sub1 rice (cv. IR64) by better light interception and effective carbohydrate utilisation under stress. Funct Plant Biol, 47(10): 891-903. |
| [8] | bin Rahman A N M R, Zhang J H. 2023. Trends in rice research: 2030 and beyond. Food Energy Secur, 12(2): e390. |
| [9] | Castano-Duque L, Ghosal S, Quilloy F A, et al. 2021. An epigenetic pathway in rice connects genetic variation to anaerobic germination and seedling establishment. Plant Physiol, 186(2): 1042-1059. |
| [10] | Catling D. 1992. Rice in Deep Water. London, UK: Macmillan Press. |
| [11] | Chakraborty M S, Jena P, Bhaduri D. 2022. Tolerance Mechanism of Rice in Submergence and Stagnant Flooding Stress. India: ICAR-National Rice Research Institute. |
| [12] | Chatterjee Y, Tomar S, Mishra M, et al. 2025. OsLdh7 overexpression in rice confers submergence tolerance by regulating key metabolic pathways: Anaerobic glycolysis, ethanolic fermentation and amino acid metabolism. Plant Cell Environ, 48(4): 2804-2820. |
| [13] | Chen Y S, Ho T D, Liu L H, et al. 2019. Sugar starvation-regulated MYBS2 and 14-3-3 protein interactions enhance plant growth, stress tolerance, and grain weight in rice. Proc Natl Acad Sci USA, 116: 21925-21935. |
| [14] | Cho J I, Ryoo N, Eom J S, et al. 2009. Role of the rice hexokinases OsHXK5 and OsHXK 6 as glucose sensors. Plant Physiol, 149(2): 745-759. |
| [15] | Colmer T D. 2003. Long-distance transport of gases in plants: A perspective on internal aeration and radial oxygen loss from roots. Plant Cell Environ, 26(1): 17-36. |
| [16] | Colmer T D, Pedersen O. 2008a. Underwater photosynthesis and respiration in leaves of submerged wetland plants: Gas films improve CO2 and O2 exchange. New Phytol, 177(4): 918-926. |
| [17] | Colmer T D, Pedersen O. 2008b. Oxygen dynamics in submerged rice (Oryza sativa). New Phytol, 178(2): 326-334. |
| [18] | Darko Asante M, Ipinyomi S O, Abe A, et al. 2021. Genetic variability for and tolerance to anaerobic germination in rice (Oryza sativa L.). J Crop Improv, 35(6): 832-847. |
| [19] | Das D P, Panda D, Nagaraju M, et al. 2004. Antioxidant enzymes and aldehyde releasing capacity of rice cultivars (Oryza sativa L.) as determinants of anaerobic seedling establishment capacity. Bulg J Plant Physiol, 30(1/2): 34-44. |
| [20] | Dhankher O P, Foyer C H. 2018. Climate resilient crops for improving global food security and safety. Plant Cell Environ, 41(5): 877-884. |
| [21] | Diagne A, Amovin-Assagba E, Futakuchi K, et al. 2013. Estimation of cultivated area, number of farming households and yield for major rice-growing environments in Africa. In: Wopereis M CS, Johnson DE, AhmadiN, et al. Realizing Africa’s Rice Promise. Wallingfird UK: CABI: 35-45. |
| [22] | Djurhuus D L E, Song Z W, Andersen A G, et al. 2025. The relationship between anaerobic germination capacity and submergence tolerance in rice seedlings. Rice, 18(1): 45. |
| [23] | Doley D, Barua M, Sarma D, et al. 2018. Screening and enhancement of anaerobic germination of rice genotypes by pre-sowing seed treatments. Curr Sci, 115(6): 1185-1190. |
| [24] | dos Santos R S, Farias D D, Pegoraro C, et al. 2017. Evolutionary analysis of the SUB1 locus across the Oryza genomes. Rice, 10(1): 4. |
| [25] | Ella E S, Kawano N, Yamauchi Y, et al. 2003. Blocking ethylene perception enhances flooding tolerance in rice seedlings. Funct Plant Biol, 30(7): 813-819. |
| [26] | Ella E S, Dionisio-Sese M L, Ismail A M. 2011. Seed pre-treatment in rice reduces damage, enhances carbohydrate mobilization and improves emergence and seedling establishment under flooded conditions. AoB Plants, 2011: plr007. |
| [27] | Emerick K, Ronald P C. 2019. Sub1 rice: Engineering rice for climate change. Cold Spring Harb Perspect Biol, 11(12): a034637. |
| [28] | Fu J, Jian Y W, Wang X H, et al. 2023. Extreme rainfall reduces one-twelfth of China’s rice yield over the last two decades. Nat Food, 4(5): 416-426. |
| [29] | Fukagawa N K, Ziska L H. 2019. Rice: Importance for global nutrition. J Nutr Sci Vitaminol, 65: S2-S3. |
| [30] | Fukao T, Xu K N, Ronald P C, et al. 2006. A variable cluster of ethylene response factor-like genes regulates metabolic and developmental acclimation responses to submergence in rice. Plant Cell, 18(8): 2021-2034. |
| [31] | Fukao T, Bailey-Serres J. 2008a. Submergence tolerance conferred by Sub1A is mediated by SLR1 and SLRL1 restriction of gibberellin responses in rice. Proc Natl Acad Sci USA, 105(43): 16814-16819. |
| [32] | Fukao T, Bailey-Serres J. 2008b. Ethylene: A key regulator of submergence responses in rice. Plant Sci, 175(1/2): 43-51. |
| [33] | Fukao T, Yeung E, Bailey-Serres J. 2011. The submergence tolerance regulator SUB1A mediates crosstalk between submergence and drought tolerance in rice. Plant Cell, 23(1): 412-427. |
| [34] | Ghosal S, Quilloy F A, Casal C, et al. 2020. Trait-based mapping to identify the genetic factors underlying anaerobic germination of rice: Phenotyping, G×E, and QTL mapping. BMC Genet, 21(1): 6. |
| [35] | Guo F, Han N, Xie Y K, et al. 2016. The miR393a/target module regulates seed germination and seedling establishment under submergence in rice (Oryza sativa L.). Plant Cell Environ, 39(10): 2288-2302. |
| [36] | Hattori Y, Nagai K, Furukawa S, et al. 2009. The ethylene response factors SNORKEL1 and SNORKEL2 allow rice to adapt to deep water. Nature, 460: 1026-1030. |
| [37] | He X, Hayes D, Zhang W D. 2020. The Impact of Flooding on China’s Agricultural Production and Food Security in 2020. Iowa State University: Center for Agricultural and Rural Development, Agricultural Policy Review: 1-11. |
| [38] | He Y Q, Sun S, Zhao J, et al. 2023. UDP-glucosyltransferase OsUGT75A promotes submergence tolerance during rice seed germination. Nat Commun, 14(1): 2296. |
| [39] | Hong Y F, Ho T D, Wu C F, et al. 2012. Convergent starvation signals and hormone crosstalk in regulating nutrient mobilization upon germination in cereals. Plant Cell, 24(7): 2857-2873. |
| [40] | Hsu S K, Tung C W. 2015. Genetic mapping of anaerobic germination-associated QTLs controlling coleoptile elongation in rice. Rice, 8(1): 38. |
| [41] | Hussain S, Khan F, Hussain H A, et al. 2016a. Physiological and biochemical mechanisms of seed priming-induced chilling tolerance in rice cultivars. Front Plant Sci, 7: 116. |
| [42] | Hussain S, Yin H Q, Peng S B, et al. 2016b. Comparative transcriptional profiling of primed and non-primed rice seedlings under submergence stress. Front Plant Sci, 7: 1125. |
| [43] | Hussain W, Anumalla M, Ismail A M, et al. 2024. Revisiting FR13A for submergence tolerance: Beyond the SUB1A gene. J Exp Bot, 75(18): 5477-5483. |
| [44] | Hwang S T, Choi D. 2016. A novel rice protein family of OsHIGDs may be involved in early signalling of hypoxia-promoted stem growth in deepwater rice. Plant Cell Rep, 35(10): 2021-2031. |
| [45] | Ismail A M, Ella E S, Vergara G V, et al. 2009. Mechanisms associated with tolerance to flooding during germination and early seedling growth in rice (Oryza sativa). Ann Bot, 103(2): 197-209. |
| [46] | Ismail A M, Johnson D E, Ella E S, et al. 2012. Adaptation to flooding during emergence and seedling growth in rice and weeds, and implications for crop establishment. Aob Plants, 2012: pls019. |
| [47] | Ismail A M, Singh U S, Singh S, et al. 2013. The contribution of submergence-tolerant (Sub1) rice varieties to food security in flood-prone rainfed lowland areas in Asia. Field Crops Res, 152: 83-93. |
| [48] | Ismail A M. 2018. Submergence tolerance in rice: Resolving a pervasive quandary. New Phytol, 218(4): 1298-1300. |
| [49] | Jain M, Kaur N, Tyagi A K, et al. 2006. The auxin-responsive GH3 gene family in rice (Oryza sativa). Funct Integr Genomics, 6(1): 36-46. |
| [50] | Jiang L, Hong M Y, Wang C M, et al. 2004. Quantitative trait loci and epistatic analysis of seed anoxia germinability in rice (Oryza sativa). Rice Sci, 11(3): 238-244. |
| [51] | John D, Shylaraj K S. 2020. Mechanism of anoxic tolerance in backcross lines developed through Jyothi × Swarna-Sub 1 under submergence stress. Indian J Genet Plant Breed, 80(4): 375-383. |
| [52] | Khalil M I, Hassan M M, Samanta S C, et al. 2024. Unraveling the genetic enigma of rice submergence tolerance: Shedding light on the role of ethylene response factor-encoding gene SUB1A-1. Plant Physiol Biochem, 206: 108224. |
| [53] | Khanna A, Anumalla M, Ramos J, et al. 2024. Genetic gains in IRRI’s rice salinity breeding and elite panel development as a future breeding resource. Theor Appl Genet, 137(2): 37. |
| [54] | Kotula L, Ranathunge K, Schreiber L, et al. 2009. Functional and chemical comparison of apoplastic barriers to radial oxygen loss in roots of rice (Oryza sativa L.) grown in aerated or deoxygenated solution. J Exp Bot, 60(7): 2155-2167. |
| [55] | Kretzschmar T, Pelayo M A F, Trijatmiko K R, et al. 2015. A trehalose-6-phosphate phosphatase enhances anaerobic germination tolerance in rice. Nat Plants, 1: 15124. |
| [56] | Kumar D, Das P K, Sarmah B K. 2020. Identification of novel hypoxia-responsive factors in deep-water rice conferring tolerance to flood during germination. Biol Plant, 64: 244-252. |
| [57] | Kuroha T, Ashikari M. 2020. Molecular mechanisms and future improvement of submergence tolerance in rice. Mol Breed, 40(4): 41. |
| [58] | Kuroha T, Nagai K, Gamuyao R, et al. 2018. Ethylene-gibberellin signaling underlies adaptation of rice to periodic flooding. Science, 361: 181-186. |
| [59] | Kurokawa Y, Nagai K, Huan P D, et al. 2018. Rice leaf hydrophobicity and gas films are conferred by a wax synthesis gene (LGF1) and contribute to flood tolerance. New Phytol, 218(4): 1558-1569. |
| [60] | Lee K W, Chen P W, Lu C A, et al. 2009. Coordinated responses to oxygen and sugar deficiency allow rice seedlings to tolerate flooding. Sci Signal, 2(91): ra61. |
| [61] | Lee K W, Chen P W, Yu S M. 2014. Metabolic adaptation to sugar/O2 deficiency for anaerobic germination and seedling growth in rice. Plant Cell Environ, 37(10): 2234-2244. |
| [62] | Lee K W, Chen J J W, Wu C S, et al. 2023. Auxin plays a role in the adaptation of rice to anaerobic germination and seedling establishment. Plant Cell Environ, 46(4): 1157-1175. |
| [63] | Li D D, Liu K, Zhao C C, et al. 2023. GWAS combined with WGCNA of transcriptome and metabolome to excavate key candidate genes for rice anaerobic germination. Rice, 16(1): 49. |
| [64] | Li D D, Liu K, Su L, et al. 2025. Genome-wide association study reveals that enolase gene OsEE1 regulates coleoptile elongation in rice under anaerobic conditions. Crop J, 13(1): 215-226. |
| [65] | Li G T, An L N, Yang W N, et al. 2025. Integrated biotechnological and AI innovations for crop improvement. Nature, 643: 925-937. |
| [66] | Lim M N, Lee S E, Yim H K, et al. 2013. Differential anoxic expression of sugar-regulated genes reveals diverse interactions between sugar and anaerobic signaling systems in rice. Mol Cells, 36(2): 169-176. |
| [67] | Lin C C, Chao Y T, Chen W C, et al. 2019. Regulatory cascade involving transcriptional and N-end rule pathways in rice under submergence. Proc Natl Acad Sci USA, 116(8): 3300-3309. |
| [68] | Liu D, Ma M J, Yu F J, et al. 2025. Identification of submergence tolerance QTLs/genes during seed germination in 432 rice varieties by GWAS. BMC Plant Biol, 25(1): 1172. |
| [69] | Lu C A, Ho T H D, Ho S L, et al. 2002. Three novel MYB proteins with one DNA binding repeat mediate sugar and hormone regulation of α-amylase gene expression. Plant Cell, 14(8): 1963-1980. |
| [70] | Lu C A, Lin C C, Lee K W, et al. 2007. The SnRK1A protein kinase plays a key role in sugar signaling during germination and seedling growth of rice. Plant Cell, 19(8): 2484-2499. |
| [71] | Lu X L, Yang L, Shen L, et al. 2025. Genome-wide association study uncovers a novel gene responsible for rice seedling submergence tolerance. Plant Biotechnol J, 23(9): 4092-4108. |
| [72] | Ma Y M, Zhao J L, Fu H, et al. 2021. Genome-wide identification, expression and functional analysis reveal the involvement of FCS-like zinc finger gene family in submergence response in rice. Rice, 14(1): 76. |
| [73] | Magneschi L, Perata P. 2009. Rice germination and seedling growth in the absence of oxygen. Ann Bot, 103(2): 181-196. |
| [74] | Matsukura C, Kawai M, Toyofuku K, et al. 2000. Transverse vein differentiation associated with gas space formation: Fate of the middle cell layer in leaf sheath development of rice. Ann Bot, 85(1): 19-27. |
| [75] | Meresa H, Tischbein B, Mekonnen T. 2022. Climate change impact on extreme precipitation and peak flood magnitude and frequency: Observations from CMIP6 and hydrological models. Nat Hazards, 111(3): 2649-2679. |
| [76] | Miro B, Longkumer T, Entila F D, et al. 2017. Rice seed germination underwater: Morpho-physiological responses and the bases of differential expression of alcoholic fermentation enzymes. Front Plant Sci, 8: 1857. |
| [77] | Mishra A K, Bairagi S, Velasco M L, et al. 2018. Impact of access to capital and abiotic stress on production efficiency: Evidence from rice farming in Cambodia. Land Use Policy, 79: 215-222. |
| [78] | Mommer L, Pons T L, Wolters-Arts M, et al. 2005. Submergence-induced morphological, anatomical, and biochemical responses in a terrestrial species affect gas diffusion resistance and photosynthetic performance. Plant Physiol, 139(1): 497-508. |
| [79] | Mondal S, Khan M I R, Entila F, et al. 2020. Responses of AG1 and AG2 QTL introgression lines and seed pre-treatment on growth and physiological processes during anaerobic germination of rice under flooding. Sci Rep, 10(1): 10214. |
| [80] | Muhammad I I, Kong S L, Akmar Abdullah S N, et al. 2019. RNA-seq and ChIP-seq as complementary approaches for comprehension of plant transcriptional regulatory mechanism. Int J Mol Sci, 21(1): 167. |
| [81] | Mwakyusa L, Heredia M C, Kilasi N L, et al. 2023. Screening of potential donors for anaerobic stress tolerance during germination in rice. Front Plant Sci, 14: 1261101. |
| [82] | Myles S, Peiffer J, Brown P J, et al. 2009. Association mapping: Critical considerations shift from genotyping to experimental design. Plant Cell, 21(8): 2194-2202. |
| [83] | Nakamura M, Noguchi K. 2020. Tolerant mechanisms to O2 deficiency under submergence conditions in plants. J Plant Res, 133(3): 343-371. |
| [84] | Narsai R, Howell K A, Carroll A, et al. 2009. Defining core metabolic and transcriptomic responses to oxygen availability in rice embryos and young seedlings. Plant Physiol, 151(1): 306-322. |
| [85] | Narsai R, Edwards J M, Roberts T H, et al. 2015. Mechanisms of growth and patterns of gene expression in oxygen-deprived rice coleoptiles. Plant J, 82(1): 25-40. |
| [86] | Nghi K N, Tagliani A, Mariotti L, et al. 2021. Auxin is required for the long coleoptile trait in japonica rice under submergence. New Phytol, 229(1): 85-93. |
| [87] | Niroula R K, Pucciariello C, Ho V T, et al. 2012. SUB1A-dependent and -independent mechanisms are involved in the flooding tolerance of wild rice species. Plant J, 72(2): 282-293. |
| [88] | Okishio T, Sasayama D, Hirano T, et al. 2014. Growth promotion and inhibition of the Amazonian wild rice species Oryza grandiglumis to survive flooding. Planta, 240(3): 459-469. |
| [89] | Oladosu Y, Rafii M Y, Arolu F, et al. 2020. Submergence tolerance in rice: Review of mechanism, breeding and, future prospects. Sustainability, 12(4): 1632. |
| [90] | Pan J W, Sharif R, Xu X W, et al. 2021. Mechanisms of waterlogging tolerance in plants: Research progress and prospects. Front Plant Sci, 11: 627331. |
| [91] | Panda D, Sarkar R K. 2013. Characterization of leaf gas exchange and anti-oxidant defense of rice (Oryza sativa L.) cultivars differing in submergence tolerance owing to complete submergence and consequent re-aeration. Agric Res, 2(4): 301-308. |
| [92] | Panda D, Sarkar R K. 2014. Mechanism associated with nonstructural carbohydrate accumulation in submergence tolerant rice (Oryza sativa L.) cultivars. J Plant Interact, 9(1): 62-68. |
| [93] | Panda D, Barik J. 2021. Flooding tolerance in rice: Focus on mechanisms and approaches. Rice Sci, 28(1): 43-57. |
| [94] | Pandey G K, Jisha V, Dampanaboina L, et al. 2015. Overexpression of an AP2/ERF type transcription factor OsEREBP1 confers biotic and abiotic stress tolerance in rice. PLoS One, 10(6): e0127831. |
| [95] | Pedersen O, Rich S M, Colmer T D. 2009. Surviving floods: Leaf gas films improve O2 and CO2 exchange, root aeration, and growth of completely submerged rice. Plant J, 58(1): 147-156. |
| [96] | Pucciariello C. 2020. Molecular mechanisms supporting rice germination and coleoptile elongation under low oxygen. Plants, 9(8): 1037. |
| [97] | Qi X H, Li Q Q, Ma X T, et al. 2019. Waterlogging-induced adventitious root formation in cucumber is regulated by ethylene and auxin through reactive oxygen species signalling. Plant Cell Environ, 42(5): 1458-1470. |
| [98] | Qi Y H, Kawano N, Yamauchi Y, et al. 2005. Identification and cloning of a submergence-induced gene OsGGT (glycogenin glucosyltransferase) from rice (Oryza sativa L.) by suppression subtractive hybridization. Planta, 221(3): 437-445. |
| [99] | Rauf M, Choi Y M, Lee S, et al. 2019. Evaluation of anaerobic germinability in various rice subpopulations: Identifying genotypes suitable for direct-seeded rice cultivation. Euphytica, 215(2): 19. |
| [100] | Reddy C K V, Ranjith P, Panda S, et al. 2024. Exploring rice genotypes for anaerobic germination: Towards sustainable direct seeding. Plant Sci Today, 11(3): 79-87. |
| [101] | Reynoso M A, Kajala K, Bajic M, et al. 2019. Evolutionary flexibility in flooding response circuitry in angiosperms. Science, 365: 1291-1295. |
| [102] | Saika H, Matsumura H, Takano T, et al. 2006. A point mutation of Adh1 gene is involved in the repression of coleoptile elongation under submergence in rice. Breed Sci, 56(1): 69-74. |
| [103] | Saika H, Okamoto M, Miyoshi K, et al. 2007. Ethylene promotes submergence-induced expression of OsABA8ox1, a gene that encodes ABA 8′-hydroxylase in rice. Plant Cell Physiol, 48(2): 287-298. |
| [104] | Sanchez D, Ben Sadoun S, Mary-Huard T, et al. 2023. Improving the use of plant genetic resources to sustain breeding programs’ efficiency. Proc Natl Acad Sci USA, 120(14): e2205780119. |
| [105] | Schmitz A J, Folsom J J, Jikamaru Y, et al. 2013. SUB1A-mediated submergence tolerance response in rice involves differential regulation of the brassinosteroid pathway. New Phytol, 198(4): 1060-1070. |
| [106] | Seck P A, Diagne A, Mohanty S, et al. 2012. Crops that feed the world 7: Rice. Food Secur, 4(1): 7-24. |
| [107] | Setter T L, Waters I, Sharma S K, et al. 2009. Review of wheat improvement for waterlogging tolerance in Australia and India: The importance of anaerobiosis and element toxicities associated with different soils. Ann Bot, 103(2): 221-235. |
| [108] | Sevanthi A M, Sinha S K, Sureshkumar V, et al. 2021. Integration of dual stress transcriptomes and major QTLs from a pair of genotypes contrasting for drought and chronic nitrogen starvation identifies key stress responsive genes in rice. Rice, 14(1): 49. |
| [109] | Shanmugam A, Manivelan K, Deepika K, et al. 2023. Unraveling the genetic potential of native rice (Oryza sativa L.) landraces for tolerance to early-stage submergence. Front Plant Sci, 14: 1083177. |
| [110] | Shingaki-Wells R, Millar A H, Whelan J, et al. 2014. What happens to plant mitochondria under low oxygen? An omics review of the responses to low oxygen and reoxygenation. Plant Cell Environ, 37(10): 2260-2277. |
| [111] | Singh A, Septiningsih E M, Balyan H S, et al. 2017. Genetics, physiological mechanisms and breeding of flood-tolerant rice (Oryza sativa L.). Plant Cell Physiol, 58(2): 185-197. |
| [112] | Singh D, Sinha S, Satyendra, et al. 2026. Enhancing submergence tolerance in rice: Metabolic and physiological insights from Sub1 introgressed lines. J Plant Biochem Biotechnol, https://doi.org/10.1007/s13562-025-00996-3. |
| [113] | Singh N, Dang T T M, Vergara G V, et al. 2010. Molecular marker survey and expression analyses of the rice submergence-tolerance gene SUB1A. Theor Appl Genet, 121(8): 1441-1453. |
| [114] | Singh P, Sinha A K. 2016. A positive feedback loop governed by SUB1A1 interaction with MITOGEN-ACTIVATED PROTEIN KINASE3 imparts submergence tolerance in rice. Plant Cell, 28(5): 1127-1143. |
| [115] | Singh S, MacKill D J, Ismail A M. 2014. Physiological basis of tolerance to complete submergence in rice involves genetic factors in addition to the SUB1 gene. AoB Plants, 6: plu060. |
| [116] | Stallworth S, Shrestha S, Schumaker B, et al. 2021. Screening diverse weedy rice (Oryza sativa ssp.) mini germplasm for tolerance to heat and complete submergence stress during seedling stage. Front Agron, 3: 642335. |
| [117] | Steffens B, Geske T, Sauter M. 2011. Aerenchyma formation in the rice stem and its promotion by H2O2. New Phytol, 190(2): 369-378. |
| [118] | Su L, Yang J, Li D D, et al. 2021. Dynamic genome-wide association analysis and identification of candidate genes involved in anaerobic germination tolerance in rice. Rice, 14(1): 1. |
| [119] | Subbaiah C C, Sachs M M. 2003. Molecular and cellular adaptations of maize to flooding stress. Ann Bot, 91(2): 119-127. |
| [120] | Sun J, Zhang G C, Cui Z B, et al. 2022. Regain flood adaptation in rice through a 14-3-3 protein OsGF14h. Nat Commun, 13(1): 5664. |
| [121] | Sun Z G, Wang B X, Zhou Z L, et al. 2021. Screening of germplasm resources and QTL mapping for germinability under submerged condition in rice (Oryza sativa L.). Acta Agron Sin, 47(1): 61-70. (in Chinese with English abstract) |
| [122] | Sun Z G, Liu Y, Li J F, et al. 2023. Identification and evaluation method for germinability under submerged condition in rice and germplasm screening. China Rice, 29(4): 53-58. (in Chinese with English abstract) |
| [123] | Takahashi H, Greenway H, Matsumura H, et al. 2014. Rice alcohol dehydrogenase 1 promotes survival and has a major impact on carbohydrate metabolism in the embryo and endosperm when seeds are germinated in partially oxygenated water. Ann Bot, 113(5): 851-859. |
| [124] | Tan B C, Joseph L M, Deng W T, et al. 2003. Molecular characterization of the Arabidopsis 9-cis epoxycarotenoid dioxygenase gene family. Plant J, 35(1): 44-56. |
| [125] | Thapa R, Tabien R E, Thomson M J, et al. 2024. Genetic factors underlying anaerobic germination in rice: Genome-wide association study and transcriptomic analysis. Plant Genome, 17(1): e20261. |
| [126] | Toojinda T, Siangliw M, Tragoonrung S, et al. 2003. Molecular genetics of submergence tolerance in rice: QTL analysis of key traits. Ann Bot, 91 (2): 243-253. |
| [127] | Verma V K, Sandhu N. 2024. Understanding anaerobic germination in direct-seeded rice: A genomic mapping approach. BMC Plant Biol, 24(1): 1194. |
| [128] | Vinitha A, Vijayalakshmi D, Raveendran M, et al. 2023. Quantifying enzyme activities under anaerobic germination in traditional rice landraces to identify donors for direct seeded rice cultivation. AATCC Review, 11(3): 482-488. |
| [129] | Voesenek L A C J, Colmer T D, Pierik R, et al. 2006. How plants cope with complete submergence. New Phytol, 170(2): 213-226. |
| [130] | Wang S, Liu W N, He Y, et al. 2021. bZIP72 promotes submerged rice seed germination and coleoptile elongation by activating ADH1. Plant Physiol Biochem, 169: 112-118. |
| [131] | Wang Y K, Jin G C, Song S Y, et al. 2024. A peroxisomal cinnamate: CoA ligase-dependent phytohormone metabolic cascade in submerged rice germination. Dev Cell, 59(11): 1363-1378. |
| [132] | Winkel A, Colmer T D, Pedersen O. 2011. Leaf gas films of Spartina anglica enhance rhizome and root oxygen during tidal submergence. Plant Cell Environ, 34(12): 2083-2092. |
| [133] | Würschum T. 2012. Mapping QTL for agronomic traits in breeding populations. Theor Appl Genet, 125(2): 201-210. |
| [134] | Xu K N, MacKill D J. 1996. A major locus for submergence tolerance mapped on rice chromosome 9. Mol Breed, 2(3): 219-224. |
| [135] | Xu K N, Xu X, Fukao T, et al. 2006. Sub1A is an ethylene-response-factor-like gene that confers submergence tolerance to rice. Nature, 442: 705-708. |
| [136] | Yamauchi T, Yoshioka M, Fukazawa A, et al. 2017. An NADPH oxidase RBOH functions in rice roots during lysigenous aerenchyma formation under oxygen-deficient conditions. Plant Cell, 29(4): 775-790. |
| [137] | Yamauchi T, Colmer T D, Pedersen O, et al. 2018. Regulation of root traits for internal aeration and tolerance to soil waterlogging-flooding stress. Plant Physiol, 176(2): 1118-1130. |
| [138] | Yang J, Wei J, Xu J F, et al. 2022. Mapping QTLs for anaerobic tolerance at germination and bud stages using new high density genetic map of rice. Front Plant Sci, 13: 985080. |
| [139] | Yang S H, Choi D. 2006. Characterization of genes encoding ABA 8′-hydroxylase in ethylene-induced stem growth of deepwater rice (Oryza sativa L.). Biochem Biophys Res Commun, 350(3): 685-690. |
| [140] | Ye N H, Wang F Z, Shi L, et al. 2018. Natural variation in the promoter of rice calcineurin B-like protein 10 (OsCBL10) affects flooding tolerance during seed germination among rice subspecies. Plant J, 94(4): 612-625. |
| [141] | Yeung E, Bailey-Serres J, Sasidharan R. 2019. After the deluge: Plant revival post-flooding. Trends Plant Sci, 24(5): 443-454. |
| [142] | Yin M, Zheng Z Z, Zhang Y, et al. 2024. Identification of key genes and pathways for anaerobic germination tolerance in rice using weighted gene co-expression network analysis (WGCNA) in association with quantitative trait locus (QTL) mapping. Rice, 17(1): 37. |
| [143] | Yu J M, Buckler E S. 2006. Genetic association mapping and genome organization of maize. Curr Opin Biotechnol, 17(2): 155-160. |
| [144] | Yu M D, Yau C P, Yip W K. 2017. Differentially localized rice ethylene receptors OsERS1 and OsETR2 and their potential role during submergence. Plant Signal Behav, 12(8): e1356532. |
| [145] | Yuan L B, Chen M X, Wang L N, et al. 2023. Multi-stress resilience in plants recovering from submergence. Plant Biotechnol J, 21(3): 466-481. |
| [146] | Yuan S, Linquist B A, Wilson L T, et al. 2021. Sustainable intensification for a larger global rice bowl. Nat Commun, 12(1): 7163. |
| [147] | Zhang G C, Liu Z M, Liu Y H, et al. 2021. iTRAQ-based proteomics investigation of critical response proteins in embryo and coleoptile during rice anaerobic germination. Rice Sci, 28(4): 391-401. |
| [148] | Zhang M C, Lu Q, Wu W, et al. 2017. Association mapping reveals novel genetic loci contributing to flooding tolerance during germination in indica rice. Front Plant Sci, 8: 678. |
| [149] | Zhang Y, Li J, Chen S J, et al. 2020. An APETALA2/ethylene responsive factor, OsEBP89 knockout enhances adaptation to direct-seeding on wet land and tolerance to drought stress in rice. Mol Genet Genomics, 295(4): 941-956. |
| [150] | Zhou Z Y, de Almeida Engler J, Rouan D, et al. 2002. Tissue localization of a submergence-induced 1-aminocyclopropane-1-carboxylic acid synthase in rice. Plant Physiol, 129(1): 72-84. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||