
Rice Science ›› 2018, Vol. 25 ›› Issue (1): 57-60.DOI: 10.1016/j.rsci.2017.09.002
• • 上一篇
收稿日期:2017-07-18
接受日期:2017-09-25
出版日期:2018-01-28
发布日期:2017-11-16
. [J]. Rice Science, 2018, 25(1): 57-60.
| Isolate No. | Location a | Isolate No. | Location a |
|---|---|---|---|
| A5 | Danzhou, Hainan | B315 | Wuhua, Guangdong |
| A90 | Danzhou, Hainan | B316 | Wuhua, Guangdong |
| A92 | Danzhou, Hainan | B331 | Lianzhou, Guangdong |
| A107 | Ledong, Hainan | C40 | Minhou, Fujian |
| A132 | Haikou, Hainan | D164 | Hechi, Guangxi |
| A142 | Haikou, Hainan | D168 | Yangshuo, Guangxi |
| A154 | Wenchang, Hainan | D175 | Yangshuo, Guangxi |
| A182 | Tunchang, Hainan | D181 | Baise, Guangxi |
| B120 | Dongyuan, Guangdong | D186 | Baise, Guangxi |
| B237 | Maoming, Guangdong | E51 | Dali, Yunnan |
| B240 | Maoming, Guangdong | GD-2 | Lechang, Guangdong |
| B246 | Suixi, Guangdong | GD-11 | Ruyuan, Guangdong |
| B249 | Suixi, Guangdong | GD-61 | Guangzhou, Guangdong |
| B250 | Suixi, Guangdong | GD-118 | Doumen, Guangdong |
| B255 | Suixi, Guangdong | H1 | Xuancheng, Anhui |
| B261 | Conghua, Guangdong | H2 | Chaohu, Anhui |
| B265 | Conghua, Guangdong | H3 | Jingzhou, Hunan |
| B266 | Conghua, Guangdong | N1 | Hangzhou, Zhejiang |
| B270 | Xinxing, Guangdong | X-5 | Yangzhou, Jiangsu |
| B275 | Taishan, Guangdong | Y1 | Hangzhou, Zhejiang |
| B278 | Taishan, Guangdong | ZJG-15 | Zhangjiagang, Jiangsu |
| B297 | Lufeng, Guangdong |
Table 1 Details of Rhizoctonia solani AG-1 IA strains used in this study.
| Isolate No. | Location a | Isolate No. | Location a |
|---|---|---|---|
| A5 | Danzhou, Hainan | B315 | Wuhua, Guangdong |
| A90 | Danzhou, Hainan | B316 | Wuhua, Guangdong |
| A92 | Danzhou, Hainan | B331 | Lianzhou, Guangdong |
| A107 | Ledong, Hainan | C40 | Minhou, Fujian |
| A132 | Haikou, Hainan | D164 | Hechi, Guangxi |
| A142 | Haikou, Hainan | D168 | Yangshuo, Guangxi |
| A154 | Wenchang, Hainan | D175 | Yangshuo, Guangxi |
| A182 | Tunchang, Hainan | D181 | Baise, Guangxi |
| B120 | Dongyuan, Guangdong | D186 | Baise, Guangxi |
| B237 | Maoming, Guangdong | E51 | Dali, Yunnan |
| B240 | Maoming, Guangdong | GD-2 | Lechang, Guangdong |
| B246 | Suixi, Guangdong | GD-11 | Ruyuan, Guangdong |
| B249 | Suixi, Guangdong | GD-61 | Guangzhou, Guangdong |
| B250 | Suixi, Guangdong | GD-118 | Doumen, Guangdong |
| B255 | Suixi, Guangdong | H1 | Xuancheng, Anhui |
| B261 | Conghua, Guangdong | H2 | Chaohu, Anhui |
| B265 | Conghua, Guangdong | H3 | Jingzhou, Hunan |
| B266 | Conghua, Guangdong | N1 | Hangzhou, Zhejiang |
| B270 | Xinxing, Guangdong | X-5 | Yangzhou, Jiangsu |
| B275 | Taishan, Guangdong | Y1 | Hangzhou, Zhejiang |
| B278 | Taishan, Guangdong | ZJG-15 | Zhangjiagang, Jiangsu |
| B297 | Lufeng, Guangdong |
| Strain | Location a | dsRNA segment observed | ||
|---|---|---|---|---|
| Number | Size (kb) b | |||
| B275 | Taishan, Guangdong | 2 | 1.7, 2.0 | |
| A5 | Danzhou, Hainan | 3 | 1.7, 1.8, 9.0 | |
| A154 | Wenchang, Hainan | 2 | 1.8, 1.9 | |
| A182 | Tunchang, Hainan | 2 | 1.8, 1.9 | |
| B240 | Maoming, Guangdong | 2 | 1.8, 2.0 | |
| B261 | Conghua, Guangdong | 1 | 1.8 | |
| B266 | Conghua, Guangdong | 4 | 1.8, 2.0, 2.3, 9.0 | |
| B297 | Lufeng, Guangdong | 2 | 1.8, 1.9 | |
| B331 | Lianzhou, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| D168 | Yangshuo, Guangxi | 2 | 1.8, 2.0 | |
| D175 | Yangshuo, Guangxi | 2 | 1.8, 1.9 | |
| GD-11 | Ruyuan, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| H1 | Xuancheng, Anhui | 2 | 1.7, 1.8 | |
| H2 | Chaohu, Anhui | 2 | 1.8, 1.9 | |
| N1 | Hangzhou, Zhejiang | 4 | 1.8, 2.0, 2.3, 6.5 | |
| Y1 | Hangzhou, Zhejiang | 3 | 1.8, 2.0, 6.4 | |
Table 2 Characterization of dsRNA mycoviruses in Rhizoctonia solani AG-1 IA strains.
| Strain | Location a | dsRNA segment observed | ||
|---|---|---|---|---|
| Number | Size (kb) b | |||
| B275 | Taishan, Guangdong | 2 | 1.7, 2.0 | |
| A5 | Danzhou, Hainan | 3 | 1.7, 1.8, 9.0 | |
| A154 | Wenchang, Hainan | 2 | 1.8, 1.9 | |
| A182 | Tunchang, Hainan | 2 | 1.8, 1.9 | |
| B240 | Maoming, Guangdong | 2 | 1.8, 2.0 | |
| B261 | Conghua, Guangdong | 1 | 1.8 | |
| B266 | Conghua, Guangdong | 4 | 1.8, 2.0, 2.3, 9.0 | |
| B297 | Lufeng, Guangdong | 2 | 1.8, 1.9 | |
| B331 | Lianzhou, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| D168 | Yangshuo, Guangxi | 2 | 1.8, 2.0 | |
| D175 | Yangshuo, Guangxi | 2 | 1.8, 1.9 | |
| GD-11 | Ruyuan, Guangdong | 3 | 1.8, 2.0, 2.3 | |
| H1 | Xuancheng, Anhui | 2 | 1.7, 1.8 | |
| H2 | Chaohu, Anhui | 2 | 1.8, 1.9 | |
| N1 | Hangzhou, Zhejiang | 4 | 1.8, 2.0, 2.3, 6.5 | |
| Y1 | Hangzhou, Zhejiang | 3 | 1.8, 2.0, 6.4 | |
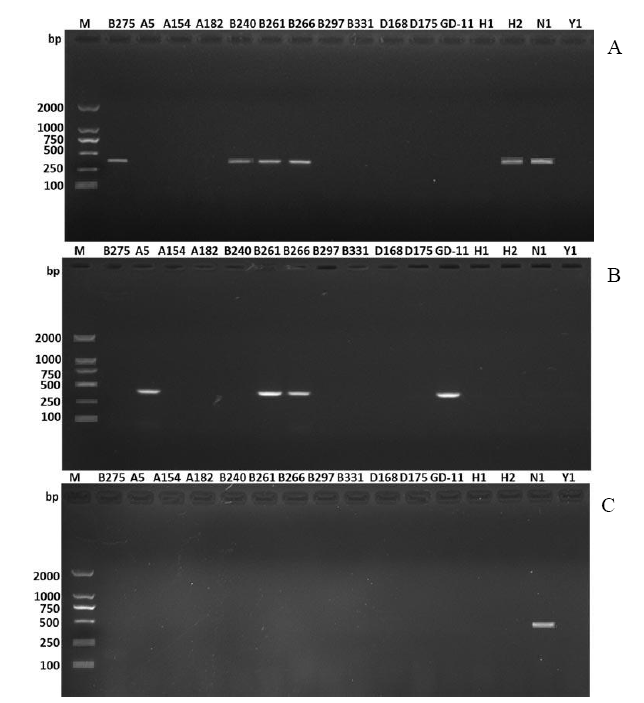
Fig. 2. RT-PCR amplification of three virus species (RsRV1, RsPV2 and RsRV-HN008) in 16 strains of Rhizoctonia solani. The species-specific RT-primers RsRV1F (5′-TGGCGTTGGCTG AGGCAAAGGT-3′) / RsRV1R (5′-GCGCATGATAGTGATGGATACG TGGATTAG-3′), RsPV2F (5′-GTTCAGCGACTACTACGACA-3′) / RsPV2R (5′-CCCCAGAGCATCGGTGTGACGCC-3′) and RsRV- HN008F (5′-GGTGATACGGCTGACAAA-3′) / RsRV-HN008R (5′-ATCG GTTGCCTAACTCCT-3′) designed based on the conserved sequences of RNA dependent RNA polymerase of the three viruses were used to amplify the corresponding conserved fragment of the viruses RsRV1 (A), RsPV2 (B) and RsRV-HN008 (C) in 16 strains of R. solani AG-1 IA, respectively.
| [1] | Bartholomäus A, Wibberg D, Winkler A, Pühler A, Schlüter A, Varrelmann M.2016. Deep sequencing analysis reveals the mycoviral diversity of the virome of an avirulent isolate of Rhizoctonia solani AG-2-2 IV. PLoS One, 11(11): e0165965. |
| [2] | Bharathan N, Saso H, Gudipati L, Bharathan S, Whited K, Anthony K.2005. Double-stranded RNA distribution and analysis among isolates of Rhizoctonia solani AG-2 to -13. Plant Pathol, 54(2): 196-203. |
| [3] | Castanho B, Butler E E.1978. Rhizoctonia decline: A degenerative disease of Rhizoctonia solani. Phytopathology, 68: 1505-1510. |
| [4] | Das S, Falloon R E, Stewart A, Pitman A R.2016. Novel mitoviruses in Rhizoctonia solani AG-3PT infecting potato. Fungal Biol, 120(3): 338-350. |
| [5] | Ghabrial S A.1998. Origin, adapation and evolutionary pathways of fungal viruses.Virus Genes, 16(1): 119-131. |
| [6] | Ghabrial S A, Suzuki N.2008. Fungal viruses. In: Mahy B W J, van Regenmortel M H V. Encyclopedia of Virology. 3rd edn. Oxford: Elsevier: 284-291. |
| [7] | Ghabrial S A, Suzuki N.2009. Viruses of plant pathogenic fungi.Annu Rev Phytopathol, 47: 353-384. |
| [8] | Lin Y H, Chiba S, Tani A, Kondo H, Sasaki A, Kanematsu S, Suzuki N.2012. A novel quadripartite dsRNA virus isolated from a phytopathogenic filamentous fungus,Rosellinia necatrix. Virology, 426(1): 42-50. |
| [9] | Liu L J, Xie J T, Cheng J S, Fu Y P, Li G Q, Yi X H, Jiang D H.2014. Fungal negative-stranded RNA virus that is related to bornaviruses and nyaviruses.Proc Natl Acad Sci USA, 111(33): 12205-12210. |
| [10] | Morris T J, Dodds J A.1979. Isolation and analysis of double stranded RNA from virus infected plant and fungal tissue.Phytopathology, 69(8): 854-858. |
| [11] | Nuss D L.2005. Hypovirulence: Mycoviruses at the fungal-plant interface.Nat Rev Microbiol, 3: 632-642. |
| [12] | Nuss D L.2010. Mycoviruses. In: Borkovich K A. Cellular and Molecular Biology of Filamentous Fungi. Washington, USA: American Society of Microbiology: 145-152. |
| [13] | Ogoshi A.1987. Ecology and pathogenicity of anastomosis and intraspecific groups of Rhizoctonia solani Kühn. Annu Rev Phytopathol, 25: 125-143. |
| [14] | Okada R, Kiyota E, Sabanadzovic S, Moriyama H, Fukuhara T, Saha P, Roossinck M J, Severin A, Valverde R A.2011. Bell pepper endornavirus: Molecular and biological properties, and occurrence in the genus Capsicum. J Gen Virol, 92: 2664-2673. |
| [15] | Pearson M N, Beever R E, Boine B, Arthur K.2009. Mycoviruses of filamentous fungi and their relevance to plant pathology.Mol Plant Pathol, 10(1): 115-128. |
| [16] | Vainio E J, Hyder R, Aday G, Hansen E, Piri T, Dogmus-Lehtijarvi T, Lehtijarvi A, Korhonen K, Hantula J.2012. Population structure of a novel putative mycovirus infecting the conifer root-rot fungus Heterobasidium annosum sensu lato. Virology, 422(2): 366-376. |
| [17] | Yu X, Li B, Fu Y P, Jiang D H, Ghabrial S A, Li G Q, Peng Y L, Xie J T, Cheng J S, Huang J, Yi X H.2010. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus.Proc Natl Acad Sci USA, 107(18): 8387-8392. |
| [18] | Yu X, Li B, Fu Y P, Xie J T, Cheng J S, Ghabrial S A, Li G Q, Yi X H, Jiang D H.2013. Extracellular transmission of a DNA mycovirus and its use as a nature fungicide.Proc Natl Acad Sci USA, 110(4): 1452-1457. |
| [19] | Zhang D T, Peng Z K, Cao Q Q, Yang M, Zheng L, Zhou E X.2012. Colonization of three antagonistic strains on rice plant and their biocontrol effects on rice sheath blight.J Northwest Agric Forest Univ, 40: 97-102. (in Chinese with English abstract) |
| [20] | Zheng L, Liu H Q, Zhang M L, Cao X, Zhou E X.2013. The complete genomic sequence of a novel mycovirus from Rhizoctonia solani AG-1 IA strain B275. Arch Virol, 158(7): 1609-1612. |
| [21] | Zheng L, Zhang M L, Chen Q G, Zhu M H, Zhou E X.2014. A novel mycovirus closely related to viruses in the genus Alphapartitivirus confers hypovirulence in the phytopathogenic fungus Rhizoctonia solani. Virology, 456/457(4): 220-226. |
| [22] | Zhong J, Chen C Y, Gao B D.2015. Genome sequence of a novel mycovirus of Rhizoctonia solani, a plant pathogenic fungus. Virus Genes, 51(1): 167-170. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||