
Rice Science ›› 2021, Vol. 28 ›› Issue (2): 114-118.DOI: 10.1016/j.rsci.2021.01.001
• Letter • Previous Articles Next Articles
Zhenhua Zhang, Yujun Zhu, Shilin Wang, Yeyang Fan, Jieyun Zhuang( )
)
Received:2020-03-17
Accepted:2020-08-05
Online:2021-03-28
Published:2021-03-28
Zhenhua Zhang, Yujun Zhu, Shilin Wang, Yeyang Fan, Jieyun Zhuang. Genetic Interaction of Hd1 with Ghd7, DTH8 and Hd2 Largely Determines Eco-Geographical Adaption of Rice Varieties in Southern China[J]. Rice Science, 2021, 28(2): 114-118.
Add to citation manager EndNote|Ris|BibTeX
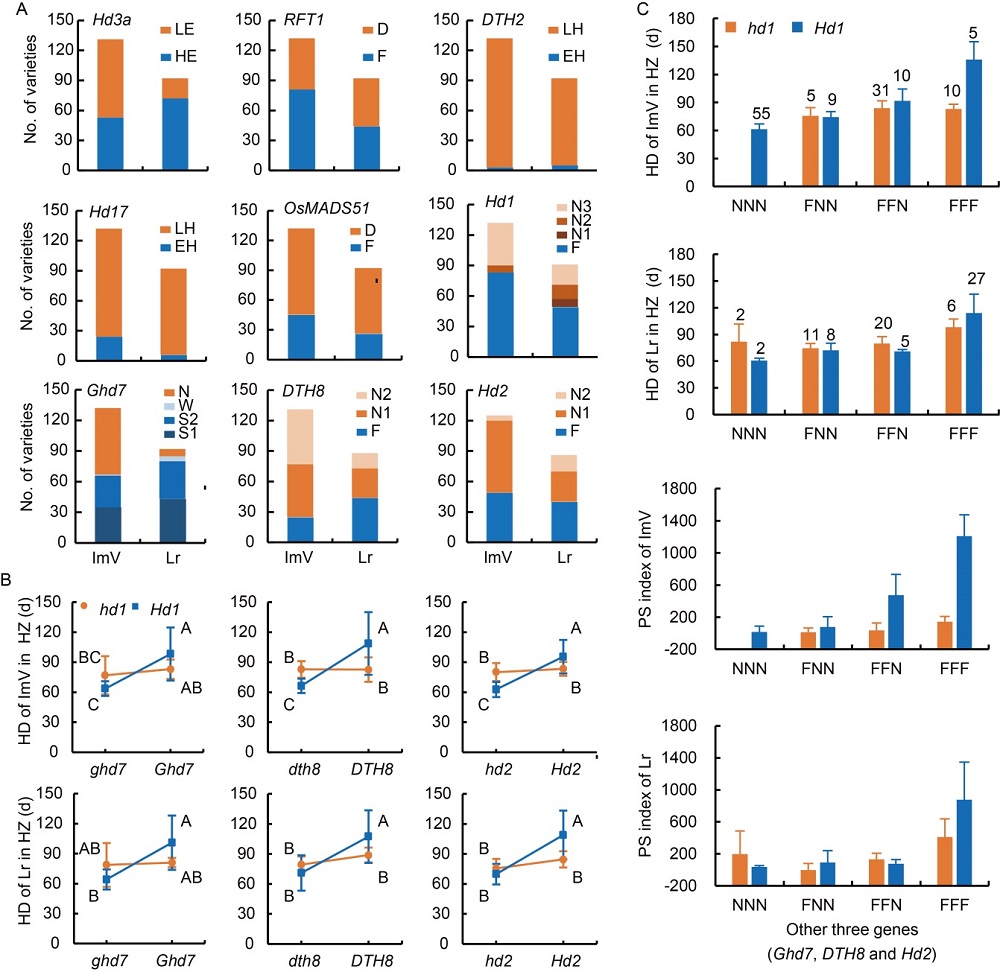
Fig. 1. Allelic variations of nine genes and genetic interactions among Hd1 and DTH8, Hd2 and Ghd7 in improved variety (ImV) and landrace (Lr).A, Allelic variation of nine genes. LE, Low expression; HE, High expression; D, Defective; F, Functional; LH, Late heading; EH, Early heading; N, Non-functional; W, Weak; S1, Strong 1; S2, Strong 2; N1, Non-functional-1; N2, Non-functional-2; N3, Non-functional-3.B, Heading date (HD) in Hangzhou (HZ) of genotypic groups in rice classified based on combinations of Hd1 and each of Ghd7, DTH8 and Hd2. Data are presented in Mean ± SD. Different capital letters indicate significant differences (P < 0.01) among mean values based on the Duncan’s multiple range test.C, Accumulation of the interaction of functional Ghd7, DTH8 and Hd2 with Hd1. PS index, Photoperiodic sensitivity index; NNN, FNN, FFN and FFF indicate the presence of functional genotypes at none, one, two and three of the Ghd7, DTH8 and Hd2 loci, respectively. Data are presented in Mean ± SD. Values above the bars indicate number of varieties.
| Gene | Trait | Improved variety | Landrace | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Group 1 | Group 2 | P | Effect | R2 | Group 1 | Group 2 | P | Effect | R2 | |||
| Hd3a | HD (HZ) | 77.4 ± 24.1 | 74.9 ± 13.3 | 0.4315 | 94.0 ± 24.3 | 74.9 ± 10.1 | 0.0016 | 19.1 | 11.0 | |||
| HD (LS) | 87.4 ± 11.9 | 93.7 ± 13.2 | 0.0049 | -6.3 | 6.1 | 90.4 ± 12.5 | 94.1 ± 12.6 | 0.2828 | ||||
| PS index | 214.3 ± 367.3 | 78.4 ± 192.4 | 0.0070 | 135.9 | 5.6 | 453.8 ± 495.3 | 63.5 ± 224.0 | 0.0029 | 390.3 | 10.2 | ||
| RFT1 | HD (HZ) | 77.0 ± 17.6 | 74.3 ± 19.6 | 0.4158 | 88.0 ± 21.3 | 91.9 ± 25.2 | 0.4337 | |||||
| HD (LS) | 93.5 ± 13.1 | 88.2 ± 12.4 | 0.0218 | 87.6 ± 14.3 | 94.1 ± 10.0 | 0.0166 | ||||||
| PS index | 118.7 ± 285.5 | 154.6 ± 278.4 | 0.4830 | 411.1 ± 526.0 | 354.3 ± 443.1 | 0.5907 | ||||||
| DTH2 | HD (HZ) | 91.9 ± 49.6 | 75.6 ± 17.3 | 0.1295 | 76.4 ± 7.9 | 90.9 ± 23.8 | 0.1796 | |||||
| HD (LS) | 77.7 ± 8.5 | 91.8 ± 13.0 | 0.0640 | 104.5 ± 15.0 | 90.4 ± 12.1 | 0.0272 | ||||||
| PS index | 535.6 ± 813.0 | 123.0 ± 259.0 | 0.0118 | -22.2 ± 65.7 | 400.2 ± 483.8 | 0.0863 | ||||||
| Hd17 | HD (HZ) | 75.6 ± 11.8 | 76.1 ± 19.5 | 0.9149 | 79.0 ± 8.6 | 90.9 ± 24.0 | 0.2298 | |||||
| HD (LS) | 95.7 ± 15.5 | 90.6 ± 12.4 | 0.0852 | 105.0 ± 13.0 | 90.2 ± 12.0 | 0.0094 | 14.8 | 7.9 | ||||
| PS index | 68.1 ± 133.3 | 146.3 ± 303.6 | 0.2294 | -41.2 ± 71.0 | 406.7 ± 483.3 | 0.0426 | ||||||
| OsMADS51 | HD (HZ) | 85.0 ± 18.0 | 71.6 ± 17.1 | 0.0001 | 13.4 | 11.1 | 96.3 ± 28.6 | 87.8 ± 20.9 | 0.1263 | |||
| HD (LS) | 100.5 ± 13.0 | 86.6 ± 10.2 | <0.0001 | 13.9 | 26.9 | 87.6 ± 14.2 | 92.3 ± 11.8 | 0.1260 | ||||
| PS index | 154.4 ± 322.9 | 121.3 ± 260.3 | 0.5286 | 498.3 ± 569.4 | 339.2 ± 443.5 | 0.1828 | ||||||
| Hd1 | HD (HZ) | 72.0 ± 21.4 | 82.9 ± 7.8 | 0.0009 | -10.9 | 8.2 | 97.9 ± 28.0 | 80.8 ± 10.9 | 0.0005 | 17.1 | 13.3 | |
| HD (LS) | 83.2 ± 6.1 | 105.7 ± 8.9 | <0.0001 | -22.5 | 69.5 | 85.1 ± 12.2 | 98.4 ± 8.6 | <0.0001 | -13.3 | 27.9 | ||
| PS index | 170.4 ± 341.7 | 67.8 ± 103.2 | 0.0452 | 593.2 ± 551.9 | 129.5 ± 177.6 | <0.0001 | 463.7 | 23.1 | ||||
| Ghd7 | HD (HZ) | 88.0 ± 17.1 | 64.1 ± 10.1 | <0.0001 | 23.9 | 42.3 | 91.8 ± 23.3 | 70.4 ± 15.1 | 0.0192 | |||
| HD (LS) | 100.9 ± 11.4 | 81.4 ± 4.7 | <0.0001 | 19.5 | 55.6 | 91.7 ± 12.7 | 85.0 ± 9.0 | 0.1765 | ||||
| PS index | 208.2 ± 353.9 | 54.5 ± 147.2 | 0.0017 | 153.7 | 7.5 | 403.7 ± 492.8 | 120.5 ± 184.1 | 0.1364 | ||||
| DTH8 | HD (HZ) | 94.1 ± 24.7 | 71.6 ± 13.5 | <0.0001 | 22.5 | 23.4 | 104.2 ± 25.8 | 76.8 ± 8.7 | <0.0001 | 27.4 | 34.8 | |
| HD (LS) | 94.6 ± 11.2 | 90.6 ± 13.4 | 0.1673 | 86.5 ± 14.1 | 95.5 ± 8.7 | 0.0007 | -9.0 | 13.5 | ||||
| PS index | 398.8 ± 467.5 | 67.9 ± 165.5 | <0.0001 | 330.9 | 21.5 | 690.9 ± 516.4 | 91.2 ± 126.1 | <0.0001 | 599.7 | 40.5 | ||
| Hd2 | HD (HZ) | 90.4 ± 18.1 | 64.7 ± 9.0 | <0.0001 | 25.7 | 46.7 | 98.6 ± 23.0 | 73.2 ± 9.1 | <0.0001 | 25.4 | 28.2 | |
| HD (LS) | 100.5 ± 12.1 | 84.5 ± 9.6 | <0.0001 | 16.0 | 35.4 | 90.3 ± 13.8 | 93.7 ± 9.9 | 0.2496 | ||||
| PS index | 252.5 ± 382.4 | 23.9 ± 79.4 | <0.0001 | 228.6 | 16.4 | 539.7 ± 501.5 | 54.3 ± 137.2 | <0.0001 | 485.4 | 24.3 | ||
Table S4. Phenotypic differences between two genotypic groups of indica rice at nine gene loci.
| Gene | Trait | Improved variety | Landrace | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Group 1 | Group 2 | P | Effect | R2 | Group 1 | Group 2 | P | Effect | R2 | |||
| Hd3a | HD (HZ) | 77.4 ± 24.1 | 74.9 ± 13.3 | 0.4315 | 94.0 ± 24.3 | 74.9 ± 10.1 | 0.0016 | 19.1 | 11.0 | |||
| HD (LS) | 87.4 ± 11.9 | 93.7 ± 13.2 | 0.0049 | -6.3 | 6.1 | 90.4 ± 12.5 | 94.1 ± 12.6 | 0.2828 | ||||
| PS index | 214.3 ± 367.3 | 78.4 ± 192.4 | 0.0070 | 135.9 | 5.6 | 453.8 ± 495.3 | 63.5 ± 224.0 | 0.0029 | 390.3 | 10.2 | ||
| RFT1 | HD (HZ) | 77.0 ± 17.6 | 74.3 ± 19.6 | 0.4158 | 88.0 ± 21.3 | 91.9 ± 25.2 | 0.4337 | |||||
| HD (LS) | 93.5 ± 13.1 | 88.2 ± 12.4 | 0.0218 | 87.6 ± 14.3 | 94.1 ± 10.0 | 0.0166 | ||||||
| PS index | 118.7 ± 285.5 | 154.6 ± 278.4 | 0.4830 | 411.1 ± 526.0 | 354.3 ± 443.1 | 0.5907 | ||||||
| DTH2 | HD (HZ) | 91.9 ± 49.6 | 75.6 ± 17.3 | 0.1295 | 76.4 ± 7.9 | 90.9 ± 23.8 | 0.1796 | |||||
| HD (LS) | 77.7 ± 8.5 | 91.8 ± 13.0 | 0.0640 | 104.5 ± 15.0 | 90.4 ± 12.1 | 0.0272 | ||||||
| PS index | 535.6 ± 813.0 | 123.0 ± 259.0 | 0.0118 | -22.2 ± 65.7 | 400.2 ± 483.8 | 0.0863 | ||||||
| Hd17 | HD (HZ) | 75.6 ± 11.8 | 76.1 ± 19.5 | 0.9149 | 79.0 ± 8.6 | 90.9 ± 24.0 | 0.2298 | |||||
| HD (LS) | 95.7 ± 15.5 | 90.6 ± 12.4 | 0.0852 | 105.0 ± 13.0 | 90.2 ± 12.0 | 0.0094 | 14.8 | 7.9 | ||||
| PS index | 68.1 ± 133.3 | 146.3 ± 303.6 | 0.2294 | -41.2 ± 71.0 | 406.7 ± 483.3 | 0.0426 | ||||||
| OsMADS51 | HD (HZ) | 85.0 ± 18.0 | 71.6 ± 17.1 | 0.0001 | 13.4 | 11.1 | 96.3 ± 28.6 | 87.8 ± 20.9 | 0.1263 | |||
| HD (LS) | 100.5 ± 13.0 | 86.6 ± 10.2 | <0.0001 | 13.9 | 26.9 | 87.6 ± 14.2 | 92.3 ± 11.8 | 0.1260 | ||||
| PS index | 154.4 ± 322.9 | 121.3 ± 260.3 | 0.5286 | 498.3 ± 569.4 | 339.2 ± 443.5 | 0.1828 | ||||||
| Hd1 | HD (HZ) | 72.0 ± 21.4 | 82.9 ± 7.8 | 0.0009 | -10.9 | 8.2 | 97.9 ± 28.0 | 80.8 ± 10.9 | 0.0005 | 17.1 | 13.3 | |
| HD (LS) | 83.2 ± 6.1 | 105.7 ± 8.9 | <0.0001 | -22.5 | 69.5 | 85.1 ± 12.2 | 98.4 ± 8.6 | <0.0001 | -13.3 | 27.9 | ||
| PS index | 170.4 ± 341.7 | 67.8 ± 103.2 | 0.0452 | 593.2 ± 551.9 | 129.5 ± 177.6 | <0.0001 | 463.7 | 23.1 | ||||
| Ghd7 | HD (HZ) | 88.0 ± 17.1 | 64.1 ± 10.1 | <0.0001 | 23.9 | 42.3 | 91.8 ± 23.3 | 70.4 ± 15.1 | 0.0192 | |||
| HD (LS) | 100.9 ± 11.4 | 81.4 ± 4.7 | <0.0001 | 19.5 | 55.6 | 91.7 ± 12.7 | 85.0 ± 9.0 | 0.1765 | ||||
| PS index | 208.2 ± 353.9 | 54.5 ± 147.2 | 0.0017 | 153.7 | 7.5 | 403.7 ± 492.8 | 120.5 ± 184.1 | 0.1364 | ||||
| DTH8 | HD (HZ) | 94.1 ± 24.7 | 71.6 ± 13.5 | <0.0001 | 22.5 | 23.4 | 104.2 ± 25.8 | 76.8 ± 8.7 | <0.0001 | 27.4 | 34.8 | |
| HD (LS) | 94.6 ± 11.2 | 90.6 ± 13.4 | 0.1673 | 86.5 ± 14.1 | 95.5 ± 8.7 | 0.0007 | -9.0 | 13.5 | ||||
| PS index | 398.8 ± 467.5 | 67.9 ± 165.5 | <0.0001 | 330.9 | 21.5 | 690.9 ± 516.4 | 91.2 ± 126.1 | <0.0001 | 599.7 | 40.5 | ||
| Hd2 | HD (HZ) | 90.4 ± 18.1 | 64.7 ± 9.0 | <0.0001 | 25.7 | 46.7 | 98.6 ± 23.0 | 73.2 ± 9.1 | <0.0001 | 25.4 | 28.2 | |
| HD (LS) | 100.5 ± 12.1 | 84.5 ± 9.6 | <0.0001 | 16.0 | 35.4 | 90.3 ± 13.8 | 93.7 ± 9.9 | 0.2496 | ||||
| PS index | 252.5 ± 382.4 | 23.9 ± 79.4 | <0.0001 | 228.6 | 16.4 | 539.7 ± 501.5 | 54.3 ± 137.2 | <0.0001 | 485.4 | 24.3 | ||
| Gene | Trait | Functional type | Improved variety | Landrace | |||
|---|---|---|---|---|---|---|---|
| Hd1 | HD (HZ) | Functional | 72.0 ± 21.4 | A | 97.9 ± 28.0 | A | |
| Non-functional-1 | - | - | 77.5 ± 6.0 | A | |||
| Non-functional-2 | 83.4 ± 7.9 | A | 80.7 ± 10.6 | A | |||
| Non-functional-3 | 79.7 ± 7.3 | A | 82.8 ± 13.4 | A | |||
| HD (LS) | Functional | 83.2 ± 6.1 | B | 85.1 ± 12.2 | B | ||
| Non-functional-1 | - | - | 102.7 ± 6.1 | A | |||
| Non-functional-2 | 106.4 ± 9.3 | A | 97.6 ± 8.7 | A | |||
| Non-functional-3 | 101.5 ± 4.7 | A | 97.1 ± 9.5 | A | |||
| PS index | Functional | 170.4 ± 341.7 | A | 593.2 ± 551.9 | A | ||
| Non-functional-1 | - | - | 8.3 ± 99.1 | B | |||
| Non-functional-2 | 67.4 ± 104.8 | A | 133.2 ± 150.7 | B | |||
| Non-functional-3 | 70.2 ± 100.5 | A | 189.5 ± 218.3 | AB | |||
| Ghd7 | HD (HZ) | Strong-1 | 92.5 ± 20.7 | A | 98.4 ± 27.4 | A | |
| Strong-2 | 83.2 ± 10.8 | A | 86.6 ± 16.8 | AB | |||
| Weak | - | - | 76.4 ± 7.9 | AB | |||
| Non-functional | 64.1 ± 10.1 | A | 70.4 ± 15.1 | B | |||
| HD (LS) | Strong-1 | 102.8 ± 13.1 | A | 89.3 ± 12.9 | AB | ||
| Strong-2 | 98.3 ± 8.6 | A | 92.7 ± 11.5 | AB | |||
| Weak | - | - | 104.5 ± 15.0 | A | |||
| Non-functional | 81.4 ± 4.7 | B | 85.0 ± 9.0 | B | |||
| PS index | Strong-1 | 242.6 ± 416.7 | A | 531.6 ± 548.1 | A | ||
| Strong-2 | 179.0 ± 275.4 | A | 315.9 ± 412.2 | A | |||
| Weak | - | - | -22.2 ± 65.7 | A | |||
| Non-functional | 54.5 ± 147.2 | A | 120.5 ± 184.1 | A | |||
| DTH8 | HD (HZ) | Functional | 94.1 ± 24.7 | A | 104.2 ± 25.8 | A | |
| Non-functional-1 | 69.9 ± 13.5 | B | 76.3 ± 8.1 | B | |||
| Non-functional-2 | 73.2 ± 24.7 | B | 77.6 ± 10.1 | B | |||
| HD (LS) | Functional | 94.6 ± 11.2 | A | 86.5 ± 14.1 | B | ||
| Non-functional-1 | 88.4 ± 11.5 | A | 93.3 ± 7.8 | AB | |||
| Non-functional-2 | 92.7 ± 11.2 | A | 100.0 ± 9.1 | A | |||
| PS index | Functional | 398.8 ± 467.5 | A | 690.9 ± 516.4 | A | ||
| Non-functional-1 | 71.0 ± 184.7 | B | 120.3 ± 119.5 | B | |||
| Non-functional-2 | 64.9 ± 146.3 | B | 30.7 ± 121.6 | B | |||
Table S5. Heading date (Hd) and photoperiodic sensitivity (PS) index of multiple allelic types of indica rice at DTH8, Ghd7 and Hd1.
| Gene | Trait | Functional type | Improved variety | Landrace | |||
|---|---|---|---|---|---|---|---|
| Hd1 | HD (HZ) | Functional | 72.0 ± 21.4 | A | 97.9 ± 28.0 | A | |
| Non-functional-1 | - | - | 77.5 ± 6.0 | A | |||
| Non-functional-2 | 83.4 ± 7.9 | A | 80.7 ± 10.6 | A | |||
| Non-functional-3 | 79.7 ± 7.3 | A | 82.8 ± 13.4 | A | |||
| HD (LS) | Functional | 83.2 ± 6.1 | B | 85.1 ± 12.2 | B | ||
| Non-functional-1 | - | - | 102.7 ± 6.1 | A | |||
| Non-functional-2 | 106.4 ± 9.3 | A | 97.6 ± 8.7 | A | |||
| Non-functional-3 | 101.5 ± 4.7 | A | 97.1 ± 9.5 | A | |||
| PS index | Functional | 170.4 ± 341.7 | A | 593.2 ± 551.9 | A | ||
| Non-functional-1 | - | - | 8.3 ± 99.1 | B | |||
| Non-functional-2 | 67.4 ± 104.8 | A | 133.2 ± 150.7 | B | |||
| Non-functional-3 | 70.2 ± 100.5 | A | 189.5 ± 218.3 | AB | |||
| Ghd7 | HD (HZ) | Strong-1 | 92.5 ± 20.7 | A | 98.4 ± 27.4 | A | |
| Strong-2 | 83.2 ± 10.8 | A | 86.6 ± 16.8 | AB | |||
| Weak | - | - | 76.4 ± 7.9 | AB | |||
| Non-functional | 64.1 ± 10.1 | A | 70.4 ± 15.1 | B | |||
| HD (LS) | Strong-1 | 102.8 ± 13.1 | A | 89.3 ± 12.9 | AB | ||
| Strong-2 | 98.3 ± 8.6 | A | 92.7 ± 11.5 | AB | |||
| Weak | - | - | 104.5 ± 15.0 | A | |||
| Non-functional | 81.4 ± 4.7 | B | 85.0 ± 9.0 | B | |||
| PS index | Strong-1 | 242.6 ± 416.7 | A | 531.6 ± 548.1 | A | ||
| Strong-2 | 179.0 ± 275.4 | A | 315.9 ± 412.2 | A | |||
| Weak | - | - | -22.2 ± 65.7 | A | |||
| Non-functional | 54.5 ± 147.2 | A | 120.5 ± 184.1 | A | |||
| DTH8 | HD (HZ) | Functional | 94.1 ± 24.7 | A | 104.2 ± 25.8 | A | |
| Non-functional-1 | 69.9 ± 13.5 | B | 76.3 ± 8.1 | B | |||
| Non-functional-2 | 73.2 ± 24.7 | B | 77.6 ± 10.1 | B | |||
| HD (LS) | Functional | 94.6 ± 11.2 | A | 86.5 ± 14.1 | B | ||
| Non-functional-1 | 88.4 ± 11.5 | A | 93.3 ± 7.8 | AB | |||
| Non-functional-2 | 92.7 ± 11.2 | A | 100.0 ± 9.1 | A | |||
| PS index | Functional | 398.8 ± 467.5 | A | 690.9 ± 516.4 | A | ||
| Non-functional-1 | 71.0 ± 184.7 | B | 120.3 ± 119.5 | B | |||
| Non-functional-2 | 64.9 ± 146.3 | B | 30.7 ± 121.6 | B | |||
| Collection | Cropping region | Cropping season | Number of varieties | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| hd1 group | Hd1 group | Total | ||||||||||
| NNN | FNN | FFN | FFF | NNN | FNN | FFN | FFF | |||||
| Improved variety | Central China | Early | 0 | 2 | 2 | 0 | 49 | 1 | 0 | 0 | 54 | |
| Central China | Middle | 0 | 0 | 9 | 3 | 0 | 1 | 0 | 0 | 13 | ||
| Central China | Late | 0 | 0 | 1 | 2 | 0 | 0 | 8 | 1 | 12 | ||
| South China | Early | 0 | 2 | 13 | 5 | 4 | 6 | 0 | 0 | 30 | ||
| South China | Middle | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 2 | ||
| South China | Late | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 4 | 6 | ||
| Southwest China | Early | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 2 | ||
| Southwest China | Middle | 0 | 1 | 4 | 0 | 0 | 1 | 0 | 0 | 6 | ||
| Total | 0 | 5 | 31 | 10 | 55 | 9 | 10 | 5 | 125 | |||
| Landrace | Central China | Early/Middle | 1 | 5 | 11 | 0 | 2 | 7 | 2 | 3 | 31 | |
| Central China | Late | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 9 | 11 | ||
| South China | Early/Middle | 0 | 2 | 2 | 0 | 0 | 0 | 0 | 0 | 4 | ||
| South China | Late | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 8 | 8 | ||
| Southwest China | Early/Middle | 0 | 4 | 6 | 1 | 0 | 1 | 3 | 0 | 15 | ||
| Southwest China | Late | 1 | 0 | 0 | 4 | 0 | 0 | 0 | 7 | 12 | ||
| Total | 2 | 11 | 20 | 6 | 2 | 8 | 5 | 27 | 81 | |||
Table 1. Ecotypic distribution of different combinations of Hd1, Ghd7, DTH8 and Hd2 functional types.
| Collection | Cropping region | Cropping season | Number of varieties | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| hd1 group | Hd1 group | Total | ||||||||||
| NNN | FNN | FFN | FFF | NNN | FNN | FFN | FFF | |||||
| Improved variety | Central China | Early | 0 | 2 | 2 | 0 | 49 | 1 | 0 | 0 | 54 | |
| Central China | Middle | 0 | 0 | 9 | 3 | 0 | 1 | 0 | 0 | 13 | ||
| Central China | Late | 0 | 0 | 1 | 2 | 0 | 0 | 8 | 1 | 12 | ||
| South China | Early | 0 | 2 | 13 | 5 | 4 | 6 | 0 | 0 | 30 | ||
| South China | Middle | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 2 | ||
| South China | Late | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 4 | 6 | ||
| Southwest China | Early | 0 | 0 | 0 | 0 | 2 | 0 | 0 | 0 | 2 | ||
| Southwest China | Middle | 0 | 1 | 4 | 0 | 0 | 1 | 0 | 0 | 6 | ||
| Total | 0 | 5 | 31 | 10 | 55 | 9 | 10 | 5 | 125 | |||
| Landrace | Central China | Early/Middle | 1 | 5 | 11 | 0 | 2 | 7 | 2 | 3 | 31 | |
| Central China | Late | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 9 | 11 | ||
| South China | Early/Middle | 0 | 2 | 2 | 0 | 0 | 0 | 0 | 0 | 4 | ||
| South China | Late | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 8 | 8 | ||
| Southwest China | Early/Middle | 0 | 4 | 6 | 1 | 0 | 1 | 3 | 0 | 15 | ||
| Southwest China | Late | 1 | 0 | 0 | 4 | 0 | 0 | 0 | 7 | 12 | ||
| Total | 2 | 11 | 20 | 6 | 2 | 8 | 5 | 27 | 81 | |||
| [1] | Du A P, Tian W, Wei M H, Yan W, He H, Zhou D, Huang X, Li S G, Ouyang X H.2017. The DTH8-Hd1 module mediates day-length- dependent regulation of rice flowering.Mol Plant, 10(7): 948-961. |
| [2] | Fujino K, Yamanouchi U, Yano M.2013. Roles of the Hd5 gene controlling heading date for adaptation to the northern limits of rice cultivation.Theor Appl Genet, 126(3): 611-618. |
| [3] | Gomez-Ariza J, Galbiati F, Goretti D, Brambilla V, Shrestha R, Pappolla A, Courtois B, Fornara F.2015. Loss of floral repressor function adapts rice to higher latitudes in Europe.J Exp Bot, 66(7): 2027-2039. |
| [4] | Goretti D, Martignago D, Landini M, Brambilla V, Gómez-Ariza J, Gnesutta N, Galbiati F, Collani S, Takagi H, Terauchi R, Mantovani R, Fornara F.2017. Transcriptional and post-transcriptional mechanisms limit heading date 1 (Hd1) function to adapt rice to high latitudes.PLoS Genet, 13(7): e1006530. |
| [5] | Li X F, Liu H Z, Wang M Q, Liu H L, Tian X J, Zhou W J, Lu T X, Wang Z Y, Chu C C, Fang J, Bu Q Y.2015. Combinations of Hd2 and Hd4 genes determine rice adaptability to Heilongjiang Province, northern limit of China.J Integr Plant Biol, 57(8): 698-707. |
| [6] | Li X F, Sun Y Q, Tian X J, Ren Y K, Tang J Q, Wang Z Y, Cheng Y Q, Bu Q Y.2018. Comprehensive identification of major flowering time genes and their combinations, which determined rice distribution in Northeast China.Plant Growth Regul, 84(3): 593-602. |
| [7] | Lin H X, Yamamoto T, Sasaki T, Yano M.2000. Characterization and detection of epistatic interactions of 3 QTLs, Hd1, Hd2, and Hd3, controlling heading date in rice using nearly isogenic lines.Theor Appl Genet, 101(7): 1021-1028. |
| [8] | Naranjo L, Talon M, Domingo C.2014. Diversity of floral regulatory genes of japonica rice cultivated at northern latitudes.BMC Genom, 15: 101. |
| [9] | Nemoto Y, Nonoue Y, Yano M, Izawa T.2016. Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7.Plant J, 86(3): 221-233. |
| [10] | Qin G, Ma Z F, Qing Y Y, Liu C, Zhang Y X, Huang D H.2016. Effect of changes of ecological condition of plant breeding base of Hainan on yield related traits of indica rice.Southwest China J Agric Sci, 29(1): 1-5. (in Chinese with English abstract) |
| [11] | Wang J L, Li J Q, Zhang H, Xu W, Meng Q.2001. The performance northern japonica rice in Hainan and field managements.J Shenyang Agric Univ, 32(2): 83-88. (in Chinese with English abstract) |
| [12] | Xu J F, Wei X J, Jiang L, Lu G W, Wang H J, Zhou Z L, Wan J M.2010. Genetic analysis of heading date of some early season indica rice cultivars and hybrid rice parent in China.Chin J Rice Sci, 24(3): 215-222. (in Chinese with English abstract) |
| [13] | Ye J, Niu X J, Yang Y L, Wang S, Xu Q, Yuan X P, Yu H Y, Wang Y P, Wang S, Feng Y, Wei X H.2018. Divergent Hd1, Ghd7, and DTH7 alleles control heading date and yield potential of japonica rice in northeast China.Front Plant Sci, 9: 35. |
| [14] | Zhang Z Y, Zhang B, Qi F X, Wu H, Li Z X, Xing Y Z.2019. Hd1 function conversion in regulating heading is dependent on gene combinations of Ghd7, Ghd8, and Ghd7.1 under long-day conditions in rice.Mol Breeding, 39: 92. |
| [15] | Zhao F Y, Liu C Z, Ma B, Hu J F, Tan K F, Yu K C, Wang C.2017. Main traits change of cold region japonica planted in Hainan Province.Heilongjiang Agric Sci, 5: 31-33. (in Chinese with English abstract) |
| [16] | Zhou J, Luo J M, Xu X R, Zhang Y L, Ma S, Lu H Y.2014. Strategic significance and development tend of Nanfan base construction.Trop Agric Engin, 38(3): 45-48. (in Chinese with English abstract) |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||