
Rice Science ›› 2023, Vol. 30 ›› Issue (5): 379-404.DOI: 10.1016/j.rsci.2023.03.014
收稿日期:2022-07-05
接受日期:2023-03-08
出版日期:2023-09-28
发布日期:2023-08-14
. [J]. Rice Science, 2023, 30(5): 379-404.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| RGA1 | Os05g0333200 | Gα subunit | + | None | / | + | Proliferation | Oki et al, |
| GS3 | Os03g0407400 | Atypical Gγ subunit | − | + | − | − | Proliferation and elongation | Fan et al, |
| DEP1 | Os09g0441900 | Atypical Gγ subunit | + | / | / | / | / | Huang et al, |
| RGB1 | Os03g0669100 | Gβ subunit | + | / | / | / | Proliferation | Utsunomiya et al, |
| GGC2 | Os08g0456600 | Atypical Gγ subunit | + | / | / | / | / | Sun et al, |
| RGG1 | Os03g0635100 | Typical Gγ subunit | − | − | / | − | Proliferation | Sun et al, |
| RGG2 | Os02g0137900 | Typical Gγ subunit | − | − | − | − | Expansion | Sun et al, |
| OsMADS1/OsLG3b | Os03g0215400 | Transcription factor downstream of GS3 and DEP1 | + | / | + | + | Proliferation | Liu et al, |
Table 1. Main regulators in G protein signaling for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| RGA1 | Os05g0333200 | Gα subunit | + | None | / | + | Proliferation | Oki et al, |
| GS3 | Os03g0407400 | Atypical Gγ subunit | − | + | − | − | Proliferation and elongation | Fan et al, |
| DEP1 | Os09g0441900 | Atypical Gγ subunit | + | / | / | / | / | Huang et al, |
| RGB1 | Os03g0669100 | Gβ subunit | + | / | / | / | Proliferation | Utsunomiya et al, |
| GGC2 | Os08g0456600 | Atypical Gγ subunit | + | / | / | / | / | Sun et al, |
| RGG1 | Os03g0635100 | Typical Gγ subunit | − | − | / | − | Proliferation | Sun et al, |
| RGG2 | Os02g0137900 | Typical Gγ subunit | − | − | − | − | Expansion | Sun et al, |
| OsMADS1/OsLG3b | Os03g0215400 | Transcription factor downstream of GS3 and DEP1 | + | / | + | + | Proliferation | Liu et al, |
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| GW2 | Os02g0244100 | RING-type E3 ubiquitin ligase | − | − | − | − | Proliferation and expansion | Song et al, |
| HGW | Os06g0160400 | Ubiquitin-associated domain protein | None | + | + | / | Division and elongation | Li et al, |
| WTG1/ OsOTUB1 | Os08g0537800 | Deubiquitinating enzyme | + | − | − | − | Proliferation and expansion | Huang et al, et al, |
| OsUBP15/LG1 | Os02g0244300 | Deubiquitinating enzyme | + | + | + | + | Proliferation | Shi et al, |
| bZIP47 | Os06g0265400 | bZIP transcription factor | + | − | / | − | Proliferation | Hao et al, |
| OsUBC13 | Os01g0673600 | E2 ubiquitin-conjugating protein | / | + | / | + | / | Hao et al, |
| WG1/OsGRX8 | Os02g0512400 | Glutaredoxin protein/target of GW2 | − | + | / | + | Proliferation | Hao et al, |
Table 2. Main regulators in ubiquitination-related pathway for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| GW2 | Os02g0244100 | RING-type E3 ubiquitin ligase | − | − | − | − | Proliferation and expansion | Song et al, |
| HGW | Os06g0160400 | Ubiquitin-associated domain protein | None | + | + | / | Division and elongation | Li et al, |
| WTG1/ OsOTUB1 | Os08g0537800 | Deubiquitinating enzyme | + | − | − | − | Proliferation and expansion | Huang et al, et al, |
| OsUBP15/LG1 | Os02g0244300 | Deubiquitinating enzyme | + | + | + | + | Proliferation | Shi et al, |
| bZIP47 | Os06g0265400 | bZIP transcription factor | + | − | / | − | Proliferation | Hao et al, |
| OsUBC13 | Os01g0673600 | E2 ubiquitin-conjugating protein | / | + | / | + | / | Hao et al, |
| WG1/OsGRX8 | Os02g0512400 | Glutaredoxin protein/target of GW2 | − | + | / | + | Proliferation | Hao et al, |
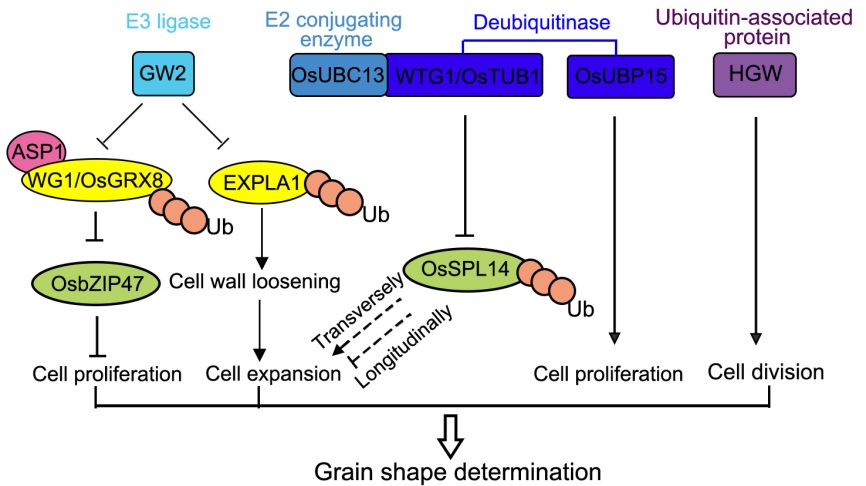
Fig. 1. Control of grain shape by ubiquitination-related pathway. GW2, a RING protein with E3 ubiquitin (Ub) ligase activity, interacts and catalyzes the ubiquitination of EXPLA1 and WG1/OsGRX8. EXPLA1 is a cell wall-loosening protein that increases cell expansion, while WG1/OsGRX8 acts as an adaptor protein to recruit the transcriptional co-repressor ASP1 to repress the transcription activity of OsbZIP47. OsbZIP47 limits grain growth by inhibiting cell proliferation. The deubiquitinase WTG1/OsTUB1 forms complex with E2 conjugating enzyme OsUBC13 to regulate grain width, maybe through promoting the ubiquitination-dependent proteasomal degradation of OsSPL14, and thus longitudinally increases cell length and transversely decreases cell width. Another deubiquitinase OsUBP15 positively regulates cell proliferation to determine grain shape. Moreover, a novel plant-specific ubiquitin-associated domain protein HGW (heading and grain weight) promotes latitudinal cell division to play a role in grain shape determination. Regular arrows indicate positive regulation, while block arrows indicate negative regulation.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| OsMKK4 | Os02g0787300 | MAPK kinase | + | + | / | + | Proliferation | Duan et al, |
| OsMAPK6 | Os06g0154500 | MAPK | + | + | / | + | Proliferation | Liu S Y et al, |
| OsWRKY53 | Os05g0343400 | Substrate of OsMAPK6 | + | + | / | / | Expansion | Tian et al, |
| GSN1/OsMKP1 | Os05g0115800 | MAPK phosphatase | − | − | / | − | Proliferation | Guo et al, |
| OsMKKK10 | Os04g0559800 | MAPK kinase kinase | + | + | / | + | Proliferation | Xu et al, |
Table 3. Main regulators in mitogen-activated protein kinase (MAPK) signaling pathway for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| OsMKK4 | Os02g0787300 | MAPK kinase | + | + | / | + | Proliferation | Duan et al, |
| OsMAPK6 | Os06g0154500 | MAPK | + | + | / | + | Proliferation | Liu S Y et al, |
| OsWRKY53 | Os05g0343400 | Substrate of OsMAPK6 | + | + | / | / | Expansion | Tian et al, |
| GSN1/OsMKP1 | Os05g0115800 | MAPK phosphatase | − | − | / | − | Proliferation | Guo et al, |
| OsMKKK10 | Os04g0559800 | MAPK kinase kinase | + | + | / | + | Proliferation | Xu et al, |
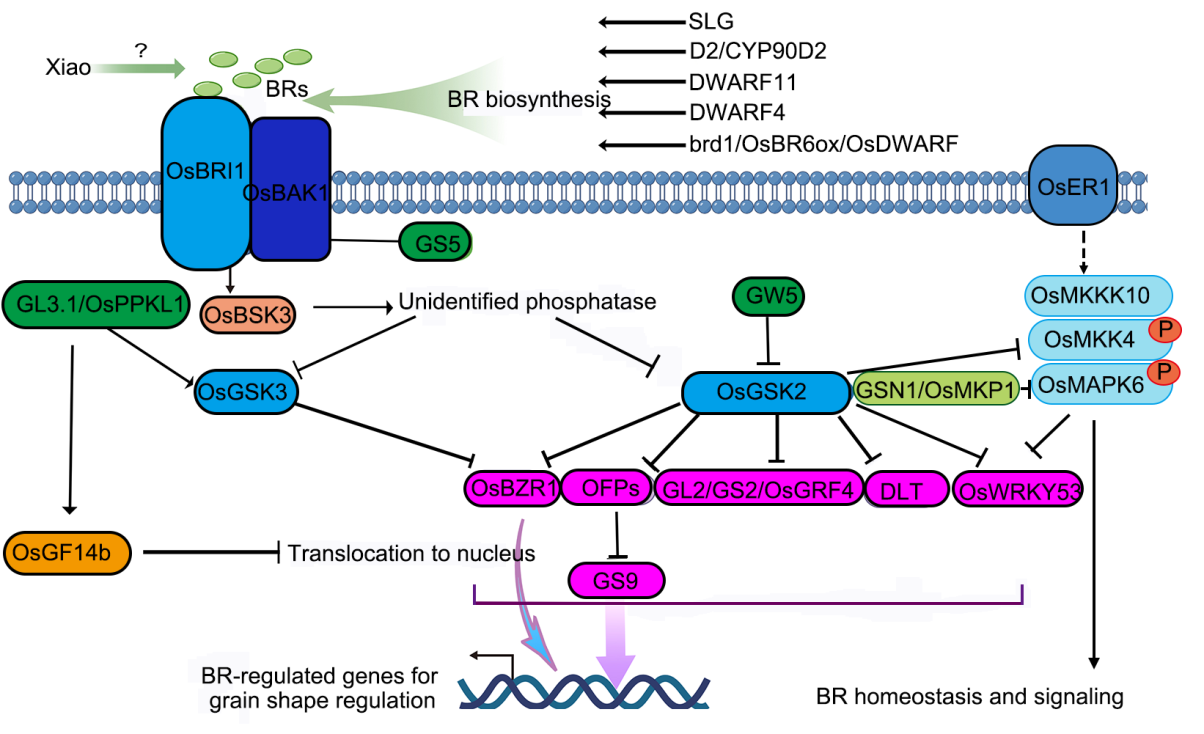
Fig. 2. Control of grain shape by brassinosteroid (BR) and mitogen-activated protein kinase (MAPK) signaling pathway. The rate limiting enzymes in BR synthesis, including SLG, D2/CYP90D2, DWARF11, DWARF4 and brd1/OsBR6ox/OsDWARF, are involved in grain shape determination through regulation of BR homeostasis. The BR signal is perceived by OsBRI1 and its co-receptor OsBAK1 and transduced to OsBSK3, which phosphorylates an unidentified phosphatase and further represses OsGSK2/OsSK22 and OsGSK3. OsGSK2 and OsGSK3 repress BR signaling by negatively regulating the transcription factors including OsBZR1, DLT, GL2/GS2/OsGRF4, OsWRKY53 and OFPs. OFPs repress the transcription activity of GS9. A protein phosphatase GL3.1/OsPPKL1 dephosphorylates but stabilizes OsGSK3, leading to accumulation of phosphorylated OsBZR1, which cannot activate BR-induced genes and thus suppressing BR signaling. GL3.1/OsPPKL1 also induces the phosphorylation of OsGF14b, which inhibits OsBZR1 to translocate to nucleus. OsGSK2 phosphorylates OsMKK4 to suppress OsMAPK6 activity, while the phosphorylation of OsWRKY53 by GSK2 lowers OsWRKY53 protein stability to regulate grain shape. GSN1/OsMKP1 inactivates OsMAPK6 via dephosphorylation and thus negatively regulates grain length and width formation. Regular arrows indicate positive regulation, while block arrows indicate negative regulation.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| GW8/OsSPL16 | Os08g0531600 | Target of miR156 | − | + | / | − | Proliferation and elongation | Wang et al, |
| GLW7/OsSPL13 | Os07g0505200 | Transcription factor, target of miR156 | + | None | + | + | Expansion | Si et al, |
| OsmiR396e | Os04g0674500 | miRNA396 | − | − | − | − | Elongation and expansion | Miao et al, |
| OsmiR396f | Os02g0829000 | miRNA396 | − | − | − | − | Elongation and expansion | Miao et al, |
| SPL4 | Os02g0829000 | Target of miR156 | − | − | + | − | Elongation and division | Hu et al, |
| OsmiR167a | miRNA167 | + | / | / | + | Elongation | Qiao et al, | |
| OsmiR529a | miRNA529 | + | − | / | − | / | Yan et al, | |
| OsmiR408 | miRNA408 | + | + | / | + | Elongation | Yang et al, | |
Table 4. Main regulators in microRNA process for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| GW8/OsSPL16 | Os08g0531600 | Target of miR156 | − | + | / | − | Proliferation and elongation | Wang et al, |
| GLW7/OsSPL13 | Os07g0505200 | Transcription factor, target of miR156 | + | None | + | + | Expansion | Si et al, |
| OsmiR396e | Os04g0674500 | miRNA396 | − | − | − | − | Elongation and expansion | Miao et al, |
| OsmiR396f | Os02g0829000 | miRNA396 | − | − | − | − | Elongation and expansion | Miao et al, |
| SPL4 | Os02g0829000 | Target of miR156 | − | − | + | − | Elongation and division | Hu et al, |
| OsmiR167a | miRNA167 | + | / | / | + | Elongation | Qiao et al, | |
| OsmiR529a | miRNA529 | + | − | / | − | / | Yan et al, | |
| OsmiR408 | miRNA408 | + | + | / | + | Elongation | Yang et al, | |
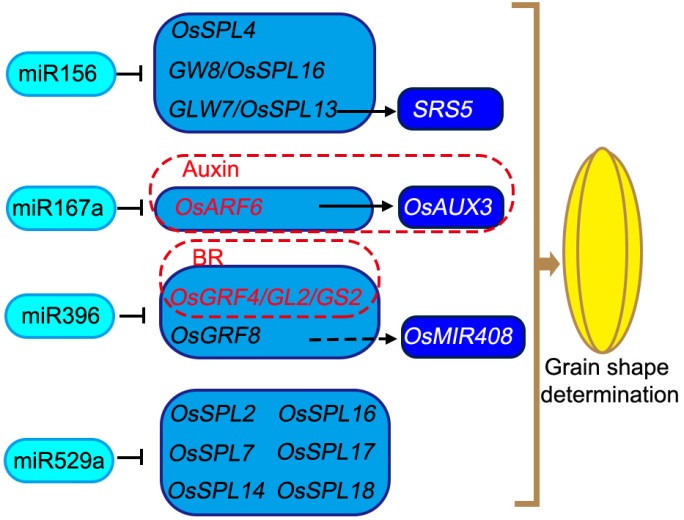
Fig. 3. MicroRNAs control rice grain shape by targeting key transcription factor-encoding genes. MicroRNA156 (miR156) targets and down-regulates the genes encoding transcription factors OsSPL4, GW8/OsSPL16 and GLW7/OsSPL13. GLW7/OsSPL13 promotes the gene expression of SRS5. miR167a targets and inhibits OsARF6, which encodes an upstream transcription factor promoting the expression of OsAUX3 in the auxin signaling pathway. miR396 directs OsGRF4/GL2/GS2 and OsGRF8 mRNA silencing. OsGRF4/GL2/GS2 is an important cross-connect node of brassinosteroid (BR) signaling and miRNA regulatory pathway to determine the grain shape. OsGRF8 directly binds to the OsmiR408 promoter and may promote the accumulation of OsmiR408. Moreover, miR529a is involved in the grain shape determination by directing other OsSPL family genes. Regular arrows indicate positive regulation, while block arrows indicate negative regulation.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| brd1/OsBR6ox/ OsDWARF | Os03g0602300 | BR-6-oxidase | + | + | + | + | / | Mori et al, |
| D2 | Os01g0197100 | Cytochrome P450 CYP90D2 | + | + | + | + | / | Hong et al, |
| D11 | Os04g0469800 | Cytochrome P450 CYP724B1 | + | + | + | + | / | Tanabe et al, |
| OsBRI1/D61 | Os01g0718300 | BR receptor | + | + | None | + | / | Morinaka et al, |
| GW5/GSE5 | Os05g0187500 | Calmodulin binding protein | − | − | − | − | Proliferation | Wan et al, |
| GS5 | Os05g0158500 | Putative serine carboxypeptidase | None | + | / | + | Division and expansion | Li et al, |
| GL3.1/OsPPKL1 | Os03g0646900 | Phosphatase kelch (PPKL) family-Ser/Thr phosphatase | + | + | + | + | Proliferation | Hu et al, |
| GL2/GS2/ OsGRF4/GLW2 | Os02g0701300 | Transcription factor | − | − | + | − | Division and expansion | Che et al, |
| SLG | Os08g0562500 | BAHD aceltransferase-like protein | + | - | / | + | Elongation | Feng et al, |
| GS9 | Os09g0448500 | Transcription factor | − | + | None | None | Proliferation | Zhao et al, |
Table 5. Main regulators in brassinosteroid (BR) biosynthesis and signaling pathways for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| brd1/OsBR6ox/ OsDWARF | Os03g0602300 | BR-6-oxidase | + | + | + | + | / | Mori et al, |
| D2 | Os01g0197100 | Cytochrome P450 CYP90D2 | + | + | + | + | / | Hong et al, |
| D11 | Os04g0469800 | Cytochrome P450 CYP724B1 | + | + | + | + | / | Tanabe et al, |
| OsBRI1/D61 | Os01g0718300 | BR receptor | + | + | None | + | / | Morinaka et al, |
| GW5/GSE5 | Os05g0187500 | Calmodulin binding protein | − | − | − | − | Proliferation | Wan et al, |
| GS5 | Os05g0158500 | Putative serine carboxypeptidase | None | + | / | + | Division and expansion | Li et al, |
| GL3.1/OsPPKL1 | Os03g0646900 | Phosphatase kelch (PPKL) family-Ser/Thr phosphatase | + | + | + | + | Proliferation | Hu et al, |
| GL2/GS2/ OsGRF4/GLW2 | Os02g0701300 | Transcription factor | − | − | + | − | Division and expansion | Che et al, |
| SLG | Os08g0562500 | BAHD aceltransferase-like protein | + | - | / | + | Elongation | Feng et al, |
| GS9 | Os09g0448500 | Transcription factor | − | + | None | None | Proliferation | Zhao et al, |
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| TGW6 | Os06g0623700 | Protein with IAA-glucose hydrolase activity | − | None | None | − | Proliferation and elongation | Ishimaru et al, |
| OsARF4 | Os01g0927600 | Transcription factor in auxin signaling | − | − | None | − | Expansion | Hu et al, |
| OsSK41/OsGSK5/GL3.3 | Os03g0841800 | GSK-like family member | − | − | / | − | Proliferation and elongation | Hu et al, et al, 2018 |
| qGL5/OsAUX3 | Os05g0447200 | Target genes in auxin signaling | − | None | / | − | Expansion | Qiao et al, |
Table 6. Main regulators in auxin biosynthesis and signaling pathway for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| TGW6 | Os06g0623700 | Protein with IAA-glucose hydrolase activity | − | None | None | − | Proliferation and elongation | Ishimaru et al, |
| OsARF4 | Os01g0927600 | Transcription factor in auxin signaling | − | − | None | − | Expansion | Hu et al, |
| OsSK41/OsGSK5/GL3.3 | Os03g0841800 | GSK-like family member | − | − | / | − | Proliferation and elongation | Hu et al, et al, 2018 |
| qGL5/OsAUX3 | Os05g0447200 | Target genes in auxin signaling | − | None | / | − | Expansion | Qiao et al, |
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| OsCKX4 | Os01g0940000 | Cytokinin oxidase/dehydrogenase | − | − | / | − | Division | Gao et al, |
| OsVQ13 | Os03g0100020 | JA responsive protein | + | + | / | / | / | Uji et al, |
| BG3/OsPUP4 | Os01g0680200 | Purine permease 4 | + | + | + | + | Division | Xiao et al, |
| OsPUP7 | Os05g0556800 | Purine permease 4 | + | + | / | + | Division | Xiao et al, |
| GW6 | Os06g0266800 | GA-regulated GAST family protein | + | + | / | + | Expansion | Shi et al, |
| OsPUP1 | Os03g0187800 | Purine permease 1 | None | − | / | - | Division | Xiao et al, |
| AGO2 | Os04g0615700 | ARGONAUTE protein | + | None | / | + | Division | Yin et al, |
Table 7. Other plant hormones regulation for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| OsCKX4 | Os01g0940000 | Cytokinin oxidase/dehydrogenase | − | − | / | − | Division | Gao et al, |
| OsVQ13 | Os03g0100020 | JA responsive protein | + | + | / | / | / | Uji et al, |
| BG3/OsPUP4 | Os01g0680200 | Purine permease 4 | + | + | + | + | Division | Xiao et al, |
| OsPUP7 | Os05g0556800 | Purine permease 4 | + | + | / | + | Division | Xiao et al, |
| GW6 | Os06g0266800 | GA-regulated GAST family protein | + | + | / | + | Expansion | Shi et al, |
| OsPUP1 | Os03g0187800 | Purine permease 1 | None | − | / | - | Division | Xiao et al, |
| AGO2 | Os04g0615700 | ARGONAUTE protein | + | None | / | + | Division | Yin et al, |
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| OsGIF1 | Os11g0615200 | Transcript co-activator | + | / | / | + | Proliferation | Duan et al, |
| OsGIF2 | Os12g0496900 | Transcript co-activator | + | / | / | + | Proliferation | Duan et al, |
| GW7/GL7/SLG7 | Os07g0603300 | TONNEAU1-recruiting motif protein | + | − | + | + | Proliferation | Wang S K et al, |
| OsFD2 | Os06g0720900 | bZIP transcription factor | − | − | − | − | Expansion and division | He et al, |
Table 8. Main regulators in other transcriptional regulatory pathway for rice grain shape.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| OsGIF1 | Os11g0615200 | Transcript co-activator | + | / | / | + | Proliferation | Duan et al, |
| OsGIF2 | Os12g0496900 | Transcript co-activator | + | / | / | + | Proliferation | Duan et al, |
| GW7/GL7/SLG7 | Os07g0603300 | TONNEAU1-recruiting motif protein | + | − | + | + | Proliferation | Wang S K et al, |
| OsFD2 | Os06g0720900 | bZIP transcription factor | − | − | − | − | Expansion and division | He et al, |
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| Cyclin-T1;3 | Os11g0157100 | Cell cycle regulator | + | / | / | / | Proliferation | Qi et al, |
| FUWA | Os02g0234200 | NHL domain-containing protein | + | − | + | − | Division | Chen et al, |
| GW6a/OsglHAT1 | Os06g0650300 | Histone H4 acetyltransferase | / | / | / | + | / | Song et al, |
| GF14f | Os03g0710800 | 14-3-3 protein | − | None | None | − | Expansion and division | Zhang et al, |
| MIS2/OsCR4 | Os03g0637800 | Receptor-like kinase | + | + | + | + | Expansion and division | Chun et al, |
| qTGW12a | Os12g0552600 | MATE transporter | / | + | / | + | / | Du et al, |
| OsDDM1b | Os03g0722400 | Subunit of chromatin remoeling complexes | + | + | / | / | Proliferation | Guo et al, |
| OsCEN2 | Os11g0152500 | CENTRORADIALIS family member | − | − | − | − | Expansion | He et al, |
| SMG3 | Os03g0410000 | MYB-like protein | − | − | / | − | Elongation and proliferation | Li et al, |
| qGSN5 | Os05g054860 | P-type PPR protein | + | + | / | + | Proliferation | Yuan et al, |
| SDR7-6 | Os07g0663900 | Short-chain alcohol dehydrogenase/reductase | None | + | + | None | / | Zheng et al, |
Table 9. Other regulators in rice grain shape determination.
| Gene | ID | Protein category | Regulatory role | Effect on cell | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Length | Width | Thickness | Weight | |||||
| Cyclin-T1;3 | Os11g0157100 | Cell cycle regulator | + | / | / | / | Proliferation | Qi et al, |
| FUWA | Os02g0234200 | NHL domain-containing protein | + | − | + | − | Division | Chen et al, |
| GW6a/OsglHAT1 | Os06g0650300 | Histone H4 acetyltransferase | / | / | / | + | / | Song et al, |
| GF14f | Os03g0710800 | 14-3-3 protein | − | None | None | − | Expansion and division | Zhang et al, |
| MIS2/OsCR4 | Os03g0637800 | Receptor-like kinase | + | + | + | + | Expansion and division | Chun et al, |
| qTGW12a | Os12g0552600 | MATE transporter | / | + | / | + | / | Du et al, |
| OsDDM1b | Os03g0722400 | Subunit of chromatin remoeling complexes | + | + | / | / | Proliferation | Guo et al, |
| OsCEN2 | Os11g0152500 | CENTRORADIALIS family member | − | − | − | − | Expansion | He et al, |
| SMG3 | Os03g0410000 | MYB-like protein | − | − | / | − | Elongation and proliferation | Li et al, |
| qGSN5 | Os05g054860 | P-type PPR protein | + | + | / | + | Proliferation | Yuan et al, |
| SDR7-6 | Os07g0663900 | Short-chain alcohol dehydrogenase/reductase | None | + | + | None | / | Zheng et al, |
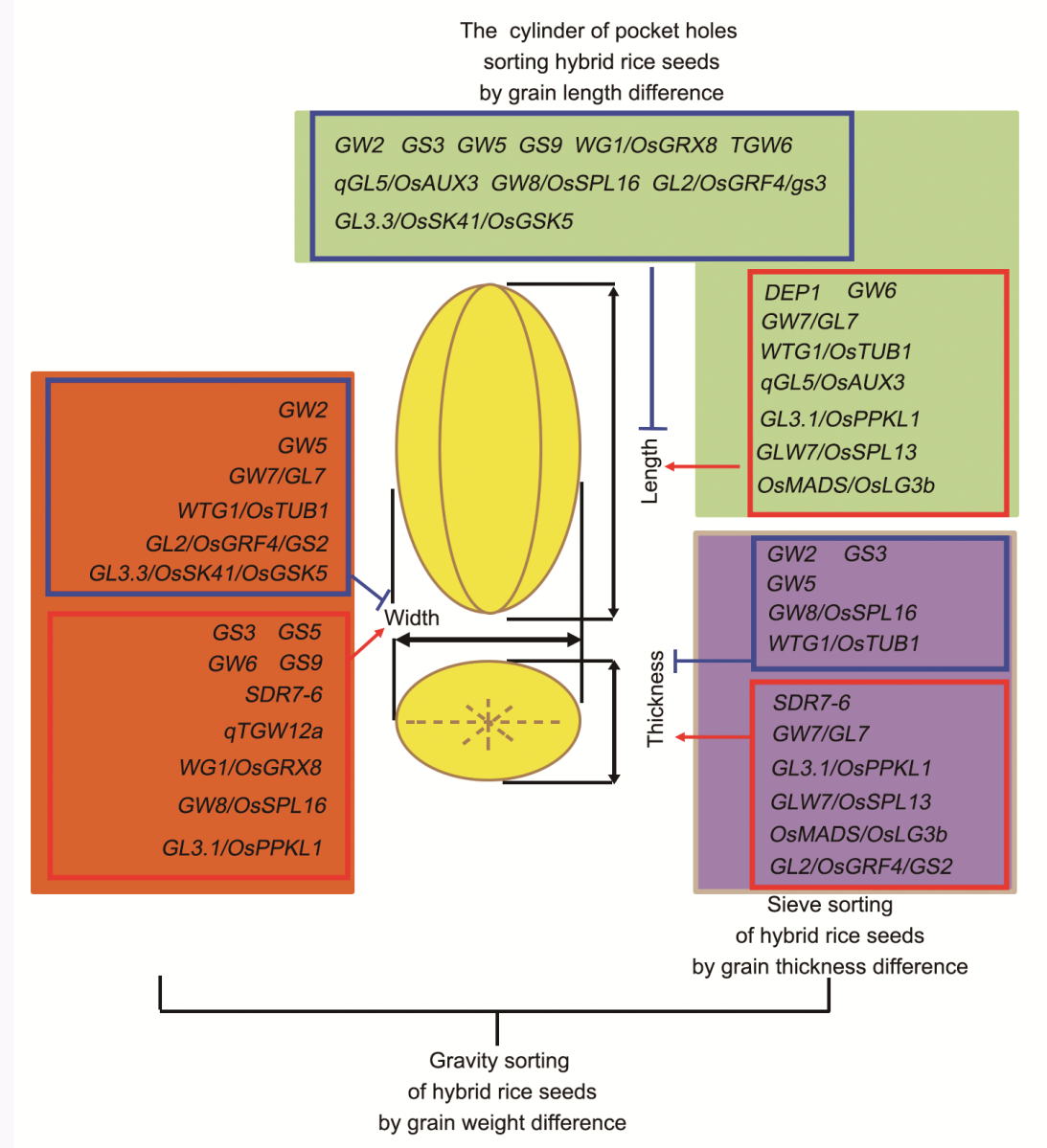
Fig. 4. Main genes regulating three-dimension of grain shape and their potential value in mechanized production of hybrid seeds. Genes with regulatory roles in the formation of grain length, grain width and grain thickness are highlighted in green, orange and purple, respectively. Red arrows indicate positive regulation, and blue block arrows indicate negative regulation. Red rectangles frame the genes with positive roles while blue rectangles frame genes with negative roles. The cylinder sorting, sieve sorting, gravity sorting systems can be used to separate hybrid rice seeds in dependence of grain length, grain thickness, and grain weight differences, respectively.
| [1] | Ali M L, McClung A M, Jia M H, Kimball, J A, McCouch S R, Eizenga G C. 2011. A rice diversity panel evaluated for genetic and agro-morphological diversity between subpopulations and its geographic distribution. Crop Sci, 51: 2021-2035. |
| [2] | Bai M Y, Zhang L Y, Gampala S S, Zhu S W, Song W Y, Chong K, Wang Z Y. 2007. Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc Natl Acad Sci USA, 104: 13839-13844. |
| [3] | Biswal A K, Wu T Y, Urano D, Pelissier R, Morel J B, Jones A M, Biswal A K,. 2022. Novel mutant alleles reveal a role of the extra- large G protein in rice grain filling, panicle architecture, plant growth, and disease resistance. Front Plant Sci, 12: 782960. |
| [4] | Butardo V M J, Sreenivasulu N, Juliano B O.2019. Improving rice grain quality: State-of-the-art and future prospects. Methods Mol Biol, 1892: 19-55. |
| [5] | Che R H, Tong H N, Shi B H, Liu Y Q, Fang S R, Liu D P, Xiao Y H, Hu B, Liu L C, Wang H R, Zhao M F, Chu C C. 2015. Control of grain size and rice yield by GL2-mediated brassinosteroid responses. Nat Plants, 2: 15195. |
| [6] | Chen J, Gao H, Zheng X M, Jin M N, Weng J F, Ma J, Ren Y L, Zhou K N, Wang Q, Wang J, Wang J L, Zhang X, Cheng Z J, Wu C Y, Wang H Y, Wan J M. 2015. An evolutionarily conserved gene, FUWA, plays a role in determining panicle architecture, grain shape and grain weight in rice. Plant J, 83(3): 427-438. |
| [7] | Chen J, Wang L H, Yuan M. 2021. Update on the roles of rice MAPK cascades. Int J Mol Sci, 22(4): 1679. |
| [8] | Chen K, Łyskowski A, Jaremko Ł, Jaremko M. 2021. Genetic and molecular factors determining grain weight in rice. Front Plant Sci, 12: 605799. |
| [9] | Choi B S, Kim Y J, Markkandan K, Koo Y J, Song J T, Seo H S. 2018. GW2 functions as an E3 ubiquitin ligase for rice expansin- like 1. Int J Mol Sci, 19(7): 1904. |
| [10] | Choi D, Kim J H, Kende H. 2004. Whole genome analysis of the OsGRF gene family encoding plant-specific putative transcription activators in rice (Oryza sativa L.). Plant Cell Physiol, 45(7): 897-904. |
| [11] |
Chun Y, Fang J J, Zafar S A, Shang J Y, Zhao J F, Yuan S J, Li X Y. 2020. MINI SEED 2 (MIS2) encodes a receptor-like kinase that controls grain size and shape in rice. Rice, 13(1): 7.
PMID |
| [12] | Dixit N, Dokku P, Amitha Mithra S V, Parida S K, Singh A K, Singh N K, Mohapatra T. 2013. Haplotype structure in grain weight gene GW2 and its association with grain characteristics in rice. Euphytica, 192(1): 55-61. |
| [13] |
Du Z X, Huang Z, Li J B, Bao J Z, Tu H, Zeng C H, Wu Z, Fu H H, Xu J, Zhou D H, Zhu C L, Fu J R, He H H. 2021. qTGW12a, a naturally varying QTL, regulates grain weight in rice. Theor Appl Genet, 134(9): 2767-2776.
PMID |
| [14] | Duan P G, Rao Y C, Zeng D L, Yang Y L, Xu R, Zhang B L, Dong G J, Qian Q, Li Y H. 2014. SMALL GRAIN 1, which encodes a mitogen-activated protein kinase kinase 4, influences grain size in rice. Plant J, 77(4): 547-557. |
| [15] | Duan P G, Ni S, Wang J M, Zhang B L, Xu R, Wang Y X, Chen H Q, Zhu X D, Li Y H. 2015. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat Plants, 2: 15203. |
| [16] | Duan P G, Xu J S, Zeng D L, Zhang B L, Geng M F, Zhang G Z, Huang K, Huang L J, Xu R, Ge S, Qian Q, Li Y H. 2017. Natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol Plant, 10(5): 685-694. |
| [17] | Duan P G, Li Y H. 2021. Size matters: G protein signaling is crucial for grain size control in rice. Mol Plant, 14(10): 1618-1620. |
| [18] |
Fan C C, Xing Y Z, Mao H L, Lu T T, Han B, Xu C G, Li X H, Zhang Q F. 2006. GS3, a major QTL for grain length and weight and minor QTL for grain width and thickness in rice, encodes a putative transmembrane protein. Theor Appl Genet, 112(6): 1164-1171.
PMID |
| [19] | Fan C C, Yu S B, Wang C R, Xing Y Z. 2009. A causal C-A mutation in the second exon of GS3 highly associated with rice grain length and validated as a functional marker. Theor Appl Genet, 118(3): 465-472. |
| [20] | Feng Z M, Wu C Y, Wang C M, Roh J, Zhang L, Chen J, Zhang S Z, Zhang H, Yang C Y, Hu J L, You X M, Liu X, Yang X M, Guo X P, Zhang X, Wu F Q, Terzaghi W, Kim S K, Jiang L, Wan J M. 2016. SLG controls grain size and leaf angle by modulating brassinosteroid homeostasis in rice. J Exp Bot, 67(14): 4241-4253. |
| [21] | Gao S P, Fang J, Xu F, Wang W, Sun X H, Chu J F, Cai B D, Feng Y Q, Chu C C. 2014. CYTOKININ OXIDASE/DEHYDROGENASE4 integrates cytokinin and auxin signaling to control rice crown root formation. Plant Physiol, 165(3): 1035-1046. |
| [22] | Gao X Y, Zhang J Q, Zhang X J, Zhou J, Jiang Z S, Huang P, Tang Z B, Bao Y M, Cheng J P, Tang H J, Zhang W H, Zhang H S, Huang J. 2019. Rice qGL3/OsPPKL1 functions with the GSK3/ SHAGGY-like kinase OsGSK3 to modulate brassinosteroid signaling. Plant Cell, 31(5): 1077-1093. |
| [23] | Gao X Y, Zhang J Q, Cai G, Du H Y, Li J B, Wang R Q, Wang Y J, Yin J, Zhang W C, Zhang H C, Huang J. 2022a. qGL3/OsPPKL1 induces phosphorylation of 14-3-3 protein OsGF14b to inhibit OsBZR1 function in brassinosteroid signaling. Plant Physiol, 188: 624-636. |
| [24] | Gao X Y, Zhang J Q, Li J B, Wang Y J, Zhang R, Du H Y, Yin J, Cai G, Wang R Q, Zhang B Y, Zhao Z, Zhang H C, Huang J, 2022b. The phosphoproteomic and interactomic landscape of qGL3/OsPPKL1-mediated brassinosteroid signaling in rice. Plant J, 109(5): 1048-1063. |
| [25] | Guo M L, Zhang W C, Mohammadi M A, He Z M, She Z Y, Yan M K, Shi C, Lin L W, Wang A Q, Liu J D, Tian D G, Zhao H M, Qin Y. 2022. OsDDM1b controls grain size by influencing cell cycling and regulating homeostasis and signaling of brassinosteroid in rice. Front Plant Sci, 13: 873993. |
| [26] | Guo T, Chen K, Dong N Q, Shi C L, Ye W W, Gao J P, Shan J X, Lin H X. 2018. GRAIN SIZE AND NUMBER1 negatively regulates the OsMKKK10-OsMKK4-OsMPK6 cascade to coordinate the trade-off between grain number per panicle and grain size in rice. Plant Cell, 30(4): 871-888. |
| [27] | Guo T, Lu Z Q, Shan J X, Ye W W, Dong N Q, Lin H X. 2020. ERECTA1 acts upstream of the OsMKKK10-OsMKK4-OsMPK6 cascade to control spikelet number by regulating cytokinin metabolism in rice. Plant Cell, 32(9): 2763-2779. |
| [28] |
Hao J Q, Wang D K, Wu Y B, Huang K, Duan P G, Li N, Xu R, Zeng D L, Dong G J, Zhang B L, Zhang L M, Inzé D, Qian Q, Li Y H. 2021. The GW2-WG1-OsbZIP47 pathway controls grain size and weight in rice. Mol Plant, 14(8): 1266-1280.
PMID |
| [29] |
Harberd N P. 2015. Shaping taste: The molecular discovery of rice genes improving grain size, shape and quality. J Genet Genomics, 42(11): 597-599.
PMID |
| [30] | Harlan J. 1992. Crops and Man. WI, USA: Am Soc Agronomy CSSA: 284. |
| [31] |
He Y, Li L Y, Shi W B, Tan J H, Luo X X, Zheng S Y, Chen W T, Li J, Zhuang C X, Jiang D G. 2022. Florigen repression complexes involving rice CENTRORADIALIS2 regulate grain size. Plant Physiol, 190(2): 1260-1274.
PMID |
| [32] |
Hirano K, Yoshida H, Aya K, Kawamura M, Hayashi M, Hobo T, Sato-Izawa K, Kitano H, Ueguchi-Tanaka M, Matsuoka M. 2017. Small organ size 1 and small organ size 2/dwarf and low-tillering form a complex to integrate auxin and brassinosteroid signaling in rice. Mol Plant, 10(4): 590-604.
PMID |
| [33] | Hong Z, Ueguchi-Tanaka M, Umemura K, Uozu S, Fujioka S, Takatsuto S, Yoshida S, Ashikari M, Kitano H, Matsuoka M. 2003. A rice brassinosteroid-deficient mutant, ebisu dwarf (d2), is caused by a loss of function of a new member of cytochrome P450. Plant Cell, 15(12): 2900-2910. |
| [34] | Hou J L, Xi X Y, Li S M, Li Z H, Li G, Fu Q, Li Y F, Yan Y T, Yang Q S, Yu H, Xia Q J, Liu B L, Li B, Li Y, Zhang G Y, Zhao S Q, Ni X M. 2019. Molecular markers and RAD sequencing assistant selection for improvement of grain size in two-line sterile rice. Mol Plant Breeding, 18(8): 2552-2558. (in Chinese with English abstract) |
| [35] | Hu J, Wang Y X, Fang Y X, Zeng L J, Xu J, Yu H P, Shi Z Y, Pan J J, Zhang D, Kang S J, Zhu L, Dong G J, Guo L B, Zeng D L, Zhang G H, Xie L H, Xiong G S, Li J Y, Qian Q. 2015. A rare allele of GS2 enhances grain size and grain yield in rice. Mol Plant, 8(10): 1455-1465. |
| [36] | Hu J H, Huang L Y, Chen G L, Liu H, Zhang Y S, Zhang R, Zhang S L, Liu J T, Hu Q Y, Hu F Y, Wang W, Ding Y. 2021. The elite alleles of OsSPL4 regulate grain size and increase grain yield in rice. Rice, 14(1): 90. |
| [37] | Hu Z J, He H H, Zhang S Y, Sun F, Xin X Y, Wang W X, Qian X, Yang J S, Luo X J. 2012. A Kelch motif-containing serine/ threonine protein phosphatase determines the large grain QTL trait in rice. J Integr Plant Biol, 54(12): 979-990. |
| [38] | Hu Z J, Lu S J, Wang M J, He H H, Sun L, Wang H R, Liu X H, Jiang L, Sun J L, Xin X Y, Kong W, Chu C C, Xue H W, Yang J S, Luo X J, Liu J X. 2018. A novel QTL qTGW3 encodes the GSK3/SHAGGY-like kinase OsGSK5/OsSK41 that interacts with OsARF4 to negatively regulate grain size and weight in rice. Mol Plant, 11(5): 736-749. |
| [39] | Huang K, Wang D K, Duan P G, Zhang B L, Xu R, Li N, Li Y H. 2017. Wide and thick grain 1, which encodes an otubain-like protease with deubiquitination activity, influences grain size and shape in rice. Plant J, 91(5): 849-860. |
| [40] | Huang X Z, Qian Q, Liu Z B, Sun H Y, He S Y, Luo D, Xia G M, Chu C C, Li J Y, Fu X D. 2009. Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet, 41(4): 494-497. |
| [41] |
Ishimaru K, Hirotsu N, Madoka Y, Murakami N, Hara N, Onodera H, Kashiwagi T, Ujiie K, Shimizu B I, Onishi A, Miyagawa H, Katoh E. 2013. Loss of function of the IAA-glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nat Genet, 45(6): 707-711.
PMID |
| [42] | Jiang Y H, Bao L, Jeong S Y, Kim S K, Xu C G, Li X H, Zhang Q F. 2012. XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice. Plant J, 70(3): 398-408. |
| [43] |
Kerr I D, Bennett M J. 2007. New insight into the biochemical mechanisms regulating auxin transport in plants. Biochem J, 401(3): 613-622.
PMID |
| [44] | Khush G S. 1997. Origin, dispersal, cultivation and variation of rice. Plant Mol Biol, 35(1/2): 25-34. |
| [45] |
Kim J H, Tsukaya H. 2015. Regulation of plant growth and development by the GROWTH-REGULATING FACTOR and GRF-INTERACTING FACTOR Duo. J Exp Bot, 66(20): 6093-6107.
PMID |
| [46] | Koh S, Lee S C, Kim M K, Koh J H, Lee S, An G, Choe S, Kim S R. 2007. T-DNA tagged knockout mutation of rice OsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses. Plant Mol Biol, 65: 453-466. |
| [47] | Lee C M, Park J, Kim B, Seo J, Lee G, Jang S, Koh H J. 2015. Influence of multi-gene allele combinations on grain size of rice and development of a regression equation model to predict grain parameters. Rice, 8(1): 33. |
| [48] | Lei J, Teng X, Wang Y F, Jiang X K, Zhao H H, Zheng X M, Ren Y L, Dong H, Wang Y L, Duan E C, Zhang Y Y, Zhang W W, Yang H, Chen X L, Chen R B, Zhang Y, Yu M Z, Xu S B, Bao X H, Zhang P C, Liu S J, Liu X, Tian Y L, Jiang L, Wang Y H, Wan J M. 2022. Plastidic pyruvate dehydrogenase complex E1 component subunit Alpha1 is involved in galactolipid biosynthesis required for amyloplast development in rice. Plant Biotechnol J, 20: 437-453. |
| [49] |
Leyser O. 2018. Auxin signaling. Plant Physiol, 176: 465-479.
PMID |
| [50] | Li D, Wang L, Wang M, Xu Y Y, Luo W, Liu Y J, Xu Z H, Li J, Chong K. 2009. Engineering OsBAK1 gene as a molecular tool to improve rice architecture for high yield. Plant Biotechnol J, 7(8): 791-806. |
| [51] | Li J, Chu H W, Zhang Y H, Mou T M, Wu C Y, Zhang Q F, Xu J. 2012. The rice HGW gene encodes a ubiquitin-associated (UBA) domain protein that regulates heading date and grain weight. PLoS One, 7(3): e34231. |
| [52] |
Li N, Li Y H. 2016. Signaling pathways of seed size control in plants. Curr Opin Plant Biol, 33: 23-32.
PMID |
| [53] |
Li N, Xu R, Li Y H. 2019. Molecular networks of seed size control in plants. Annu Rev Plant Biol, 70: 435-463.
PMID |
| [54] | Li Q F, Yu J W, Lu J, Fei H Y, Luo M, Cao B W, Huang L C, Zhang C Q, Liu Q Q. 2018. Seed-specific expression of OsDWF4, a rate-limiting gene involved in brassinosteroids biosynthesis, improves both grain yield and quality in rice. J Agric Food Chem, 66(15): 3759-3772. |
| [55] | Li R S, Li Z, Ye J, Yang Y Y, Ye J, Xu S L, Liu J R, Yuan X P, Wang Y P, Zhang M C, Yu H Y, Xu Q, Wang S, Yang Y L, Wang S, Wei X H, Feng Y. 2022. Identification of SMG3, a QTL coordinately controls grain size, grain number per panicle, and grain weight in rice. Front Plant Sci, 13: 880919. |
| [56] |
Li S C, Gao F Y, Xie K L, Zeng X H, Cao Y, Zeng J, He Z S, Ren Y, Li W B, Deng Q M, Wang S Q, Zheng A P, Zhu J, Liu H N, Wang L X, Li P. 2016. The OsmiR396c-OsGRF4-OsGIF1 regulatory module determines grain size and yield in rice. Plant Biotechnol J, 14(11): 2134-2146.
PMID |
| [57] |
Li X B, Tao Q D, Miao J, Yang Z F, Gu M H, Liang G H, Zhou Y. 2019. Evaluation of differential qPE9-1/DEP1 protein domains in rice grain length and weight variation. Rice, 12(1): 5.
PMID |
| [58] | Li Y B, Fan C C, Xing Y Z, Jiang Y H, Luo L J, Sun L, Shao D, Xu C J, Li X H, Xiao J H, He Y Q, Zhang Q F. 2011. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet, 43(12): 1266-1269. |
| [59] | Liu J F, Chen J, Zheng X M, Wu F Q, Lin Q B, Heng Y Q, Tian P, Cheng Z J, Yu X W, Zhou K N, Zhang X, Guo X P, Wang J L, Wang H Y, Wan J M. 2017. GW5 acts in the brassinosteroid signalling pathway to regulate grain width and weight in rice. Nat Plants, 3: 17043. |
| [60] | Liu L C, Tong H N, Xiao Y H, Che R H, Xu F, Hu B, Liang C Z, Chu J F, Li J Y, Chu C C. 2015. Activation of Big Grain1 significantly improves grain size by regulating auxin transport in rice. Proc Natl Acad Sci USA, 112: 11102-11107. |
| [61] |
Liu Q, Han R X, Wu K, Zhang J Q, Ye Y F, Wang S S, Chen J F, Pan Y J, Li Q, Xu X P, Zhou J W, Tao D Y, Wu Y J, Fu X D. 2018. G-protein βγ subunits determine grain size through interaction with MADS-domain transcription factors in rice. Nat Commun, 9(1): 852.
PMID |
| [62] | Liu S Y, Hua L, Dong S J, Chen H Q, Zhu X D, Jiang J E, Zhang F, Li Y H, Fang X H, Chen F. 2015. OsMAPK6, a mitogen- activated protein kinase, influences rice grain size and biomass production. Plant J, 84(4): 672-681. |
| [63] | Liu X Z, Zhang S W, Jiang Y L, Yan T W, Fang C W, Hou Q C, Wu S W, Xie K, An X L, Wan X Y. 2022. Use of CRISPR/ Cas9-Based gene editing to simultaneously mutate multiple homologous genes required for pollen development and male fertility in maize. Cells, 11(3): 439. |
| [64] | Ma G H, Yuan L P. 2015. Hybrid rice achievements, development and prospect in China. J Integr Agric, 2: 197-205. |
| [65] | Ma H G, Gao Y J, Wang Y G, Dai Y, Ma H X. 2022. Regulatory mechanisms of mitogen-activated protein kinase cascades in plants: More than sequential phosphorylation. Int J Mol Sci, 23(7): 3572. |
| [66] |
Mao H L, Sun S Y, Yao J L, Wang C R, Yu S B, Xu C G, Li X H, Zhang Q F. 2010. Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc Natl Acad Sci USA, 107: 19579-19584.
PMID |
| [67] | Miao C B, Wang D, He R Q, Liu S K, Zhu J K. 2020. Mutations in MIR396e and MIR396f increase grain size and modulate shoot architecture in rice. Plant Biotechnol J, 18(2): 491-501. |
| [68] |
Miao J, Yang Z F, Zhang D P, Wang Y Z, Xu M B, Zhou L H, Wang J, Wu S J, Yao Y L, Du X, Gu F F, Gong Z Y, Gu M H, Liang G H, Zhou Y. 2019. Mutation of RGG2, which encodes a type B heterotrimeric G protein γ subunit, increases grain size and yield production in rice. Plant Biotechnol J, 17(3): 650-664.
PMID |
| [69] | Mitchell J W, Mandava N, Worley J F, Plimmer J R, Smith M V. 1970. Brassins: A new family of plant hormones from rape pollen. Nature, 225: 1065-1066. |
| [70] |
Mori M, Nomura T, Ooka H, Ishizaka M, Yokota T, Sugimoto K, Okabe K, Kajiwara H, Satoh K, Yamamoto K, Hirochika H, Kikuchi S. 2002. Isolation and characterization of a rice dwarf mutant with a defect in brassinosteroid biosynthesis. Plant Physiol, 130(3): 1152-1161.
PMID |
| [71] |
Morinaka Y, Sakamoto T, Inukai Y, Agetsuma M, Kitano H, Ashikari M, Matsuoka M. 2006. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice. Plant Physiol, 141(3): 924-931.
PMID |
| [72] |
Nakamura A, Fujioka S, Sunohara H, Kamiya N, Hong Z, Inukai Y, Miura K, Takatsuto S, Yoshida S, Ueguchi-Tanaka M, Hasegawa Y, Kitano H, Matsuoka M. 2006. The role of OsBRI1 and its homologous genes, OsBRL1and OsBRL3, in rice. Plant Physiol, 140: 580-590.
PMID |
| [73] | Nayak A K, Anilkumar C, Behera S, Sah R P, Lavanya G R, Kumar A, Behera L, Tp M A. 2022. Genetic dissection of grain size traits through genome-wide association study vased on genic markers in rice. Rice Sci, 29(5): 462-472. |
| [74] | Ofoe R. 2021. Signal transduction by plant heterotrimeric G-protein. Plant Biol, 23(1): 3-10. |
| [75] | Oh M H, Honey S H, Tax F E. 2020. The control of cell expansion, cell division, and vascular development by brassinosteroids: A historical perspective. Int J Mol Sci, 21(5): 1743. |
| [76] |
Oki K, Fujisawa Y, Kato H, Iwasaki Y. 2005. Study of the constitutively active form of the alpha subunit of rice heterotrimeric G proteins. Plant Cell Physiol, 46(2): 381-386.
PMID |
| [77] |
Oki K, Kitagawa K, Fujisawa Y, Kato H, Iwasaki Y. 2009. Function of alpha subunit of heterotrimeric G protein in brassinosteroid response of rice plants. Plant Signal Behav, 4(2): 126-128.
PMID |
| [78] |
Pan W B, Wu Y R, Xie Q. 2019. Regulation of ubiquitination is central to the phosphate starvation response. Trends Plant Sci, 24: 755-769.
PMID |
| [79] | Qi P, Lin Y S, Song X J, Shen J B, Huang W, Shan J X, Zhu M Z, Jiang L W, Gao J P, Lin H X. 2012. The novel quantitative trait locus GL3.1 controls rice grain size and yield by regulating Cyclin-T1;3. Cell Res, 22(12): 1666-1680. |
| [80] |
Qiao J Y, Jiang H Z, Lin Y Q, Shang L G, Wang M, Li D M, Fu X D, Geisler M, Qi Y H, Gao Z Y, Qian Q. 2021. A novel miR167a-OsARF6-OsAUX3 module regulates grain length and weight in rice. Mol Plant, 14(10): 1683-1698.
PMID |
| [81] |
Reyes-Turcu F E, Ventii K H, Wilkinson K D. 2009. Regulation and cellular roles of ubiquitin-specific deubiquitinating enzymes. Annu Rev Biochem, 78: 363-397.
PMID |
| [82] | Rogers K, Chen X M. 2013. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell, 25(7): 2383-2399. |
| [83] | Sadras V O. 2007. Evolutionary aspects of the trade-off between seed size and number in crops. Field Crops Res, 100(2/3): 125-138. |
| [84] | Saichompoo U, Narumol P, Nakwilai P, Thongyos P, Nanta A, Tippunya P, Ruengphayak S, Itthisoponkul T, Bueraheng N, Cheabu S, Malumpong C. 2021. Breeding novel short grain rice for tropical region to combine important agronomical traits, biotic stress resistance and cooking quality in Koshihikari background. Rice Sci, 28(5): 479-792. |
| [85] |
Sakakibara H. 2006. Cytokinins: Activity, biosynthesis, and translocation. Annu Rev Plant Biol, 57: 431-449.
PMID |
| [86] |
Sakamoto T, Matsuoka M. 2008. Identifying and exploiting grain yield genes in rice. Curr Opin Plant Biol, 11(2): 209-214.
PMID |
| [87] | Shang J Y, He X J. 2022. Chromatin-remodeling complexes: Conserved and plant-specific subunits in Arbidopsis. J Integr Plant Biol, 64(2): 499-515. |
| [88] |
Shi C L, Ren Y L, Liu L L, Wang F, Zhang H, Tian P, Pan T, Wang Y F, Jing R N, Liu T Z, Wu F Q, Lin Q B, Lei C L, Zhang X, Zhu S S, Guo X P, Wang J L, Zhao Z C, Wang J, Zhai H Q, Cheng Z J, Wan J M. 2019. Ubiquitin specific protease 15 has an important role in regulating grain width and size in rice. Plant Physiol, 180(1): 381-391.
PMID |
| [89] | Shi C L, Dong N Q, Guo T, Ye W W, Shan J X, Lin H X. 2020. A quantitative trait locus GW6 controls rice grain size and yield through the gibberellin pathway. Plant J, 103(3): 1174-1188. |
| [90] |
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M. 2008. Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet, 40(8): 1023-1028.
PMID |
| [91] | Si L Z, Chen J Y, Huang X H, Gong H, Luo J H, Hou Q Q, Zhou T Y, Lu T T, Zhu J J, Shangguan Y Y, Chen E W, Gong C X, Zhao Q, Jing Y F, Zhao Y, Li Y, Cui L L, Fan D L, Lu Y Q, Weng Q J, Wang Y C, Zhan Q L, Liu K Y, Wei X H, An K, An G, Han B. 2016. OsSPL13 controls grain size in cultivated rice. Nat Genet, 48(4): 447-456. |
| [92] |
Smalle J, Vierstra R D. 2004. The ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol, 55: 555-590.
PMID |
| [93] | Song X J, Huang W, Shi M, Zhu M Z, Lin H X. 2007. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet, 39(5): 623-630. |
| [94] |
Song X J, Kuroha T, Ayano M, Furuta T, Nagai K, Komeda N, Segami S, Miura K, Ogawa D, Kamura T, Suzuki T, Higashiyama T, Yamasaki M, Mori H, Inukai Y, Wu J Z, Kitano H, Sakakibara H, Jacobsen S E, Ashikari M. 2015. Rare allele of a previously unidentified histone H4 acetyltransferase enhances grain weight, yield, and plant biomass in rice. Proc Natl Acad Sci USA, 112(1): 76-81.
PMID |
| [95] |
Spinner L, Gadeyne A, Belcram K, Goussot M, Moison M, Duroc Y, Eeckhout D, De Winne N, Schaefer E, Van De Slijke E, Persiau G, Witters E, Gevaert K, De Jaeger G, Bouchez D, van Damme D, Pastuglia M. 2013. A protein phosphatase 2A complex spatially controls plant cell division. Nat Commun, 4: 1863.
PMID |
| [96] |
Sun H Y, Qian Q, Wu K, Luo J J, Wang S S, Zhang C W, Ma Y F, Liu Q, Huang X Z, Yuan Q B, Han R X, Zhao M, Dong G J, Guo L B, Zhu X D, Gou Z H, Wang W, Wu Y J, Lin H X, Fu X D. 2014. Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat Genet, 46(6): 652-656.
PMID |
| [97] |
Sun S Y, Wang L, Mao H L, Shao L, Li X H, Xiao J H, Ouyang Y D, Zhang Q F. 2018. A G-protein pathway determines grain size in rice. Nat Commun, 9(1): 851.
PMID |
| [98] | Takano-Kai N, Jiang H, Kubo T, Sweeney M, Matsumoto T, Kanamori H, Padhukasahasram B, Bustamante C, Yoshimura A, Doi K, McCouch S. 2009. Evolutionary history of GS3, a gene conferring grain length in rice. Genetics, 182(4): 1323-1334. |
| [99] | Tan Y F, Xing Y Z, Li J X, Yu S B, Xu C G, Zhang Q F. 2000. Genetic bases of appearance quality of rice grains in Shanyou 63, an elite rice hybrid. Theor Appl Genet, 101(5): 823-829. |
| [100] | Tanabe S, Ashikari M, Fujioka S, Takatsuto S, Yoshida S, Yano M, Yoshimura A, Kitano H, Matsuoka M, Fujisawa Y, Kato H, Iwasaki Y. 2005. A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell, 17(3): 776-790. |
| [101] |
Tanaka A, Nakagawa H, Tomita C, Shimatani Z, Ohtake M, Nomura T, Jiang C J, Dubouzet J G, Kikuchi S, Sekimoto H, Yokota T, Asami T, Kamakura T, Mori M. 2009. BRASSINOSTEROID UPREGULATED1, encoding a helix-loop-helix protein, is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice. Plant Physiol, 151(2): 669-680.
PMID |
| [102] | Tang W B, Zhang G L, Deng H B. 2020. Technology exploration and practice of hybrid rice mechanized seed production. Chin J Rice Sci, 34(2): 95-103. (in Chinese with English abstract) |
| [103] | Tang W B, Chen X J, Zhang G L, Deng H B, Hu Y Y, Tian Y. 2022. Current situation and technical breakthrough of mechanized hybrid rice seed production. China Rice, 28(5): 20-27. (in Chinese with English abstract) |
| [104] |
Tao Y J, Miao J, Wang J, Li W Q, Xu Y, Wang F Q, Jiang Y J, Chen Z H, Fan F J, Xu M B, Zhou Y, Liang G H, Yang J. 2020. RGG1, involved in the cytokinin regulatory pathway, controls grain size in rice. Rice, 13(1): 76.
PMID |
| [105] |
Tian X J, Li X F, Zhou W J, Ren Y K, Wang Z Y, Liu Z Q, Tang J Q, Tong H N, Fang J, Bu Q Y. 2017. Transcription factor OsWRKY53 positively regulates brassinosteroid signaling and plant architecture. Plant Physiol, 175(3): 1337-1349.
PMID |
| [106] | Tian X J, He M L, Mei E Y, Zhang B W, Tang J Q, Xu M, Liu J L, Li X F, Wang Z Y, Tang W Q, Guan Q J, Bu Q Y. 2021. WRKY53 integrates classic brassinosteroid signaling and the mitogen-activated protein kinase pathway to regulate rice architecture and seed size. Plant Cell, 33(8): 2753-2775. |
| [107] |
Tong H N, Chu C C. 2018. Functional specificities of brassinosteroid and potential utilization for crop improvement. Trends Plant Sci, 23(11): 1016-1028.
PMID |
| [108] | Tong H N, Liu L C, Jin Y, Du L, Yin Y H, Qian Q, Zhu L H, Chu C C. 2012. DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell, 24(6): 2562-2577. |
| [109] | Uji Y, Kashihara K, Kiyama H, Mochizuki S, Akimitsu K, Gomi K. 2019. Jasmonic acid-induced VQ-motif-containing protein OsVQ13 influences the OsWRKY45 signaling pathway and grain size by associating with OsMPK6 in rice. Int J Mol Sci, 20(12): 2917. |
| [110] | Unnevehr L J, Duff B, Juliano B O. 1992. Consumer Demand for Rice Grain Quality. Manila, the Philippine: International Rice Research Institute. |
| [111] | Utsunomiya Y, Samejima C, Takayanagi Y, Izawa Y, Yoshida T, Sawada Y, Fujisawa Y, Kato H, Iwasaki Y. 2011. Suppression of the rice heterotrimeric G protein β-subunit gene, RGB1, causes dwarfism and browning of internodes and lamina joint regions. Plant J, 67(5): 907-916. |
| [112] |
Varshney V, Majee M. 2022. Emerging roles of the ubiquitin- proteasome pathway in enhancing crop yield by optimizing seed agronomic traits. Plant Cell Rep, 41(9): 1805-1826.
PMID |
| [113] | Verma A, Prakash G, Ranjan R, Tyagi A K, Agarwal P. 2021. Silencing of an ubiquitin ligase increases grain width and weight in indica rice. Front Genet, 11: 600378. |
| [114] | Wan X Y, Weng J F, Zhai H Q, Wang J K, Lei C L, Liu X L, Guo T, Jiang L, Su N, Wan J M. 2008. Quantitative trait loci (QTL) analysis for rice grain width and fine mapping of an identified QTL allele gw-5 in a recombination hotspot region on chromosome 5. Genetics, 179(4): 2239-2252. |
| [115] | Wang M, Qiao J Y, Yu C L, Chen H, Sun C D, Huang L Z, Li C Y, Geisler M, Qian Q, Jiang D A, Qi Y H. 2019. The auxin influx carrier, OsAUX3, regulates rice root development and responses to aluminium stress. Plant Cell Environ, 42(4): 1125-1138. |
| [116] | Wang S K, Wu K, Yuan Q B, Liu X Y, Liu Z B, Lin X Y, Zeng R Z, Zhu H T, Dong G J, Qian Q, Zhang G Q, Fu X D. 2012. Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet, 44(8): 950-954. |
| [117] | Wang S K, Li S, Liu Q, Wu K, Zhang J Q, Wang S S, Wang Y, Chen X B, Zhang Y, Gao C X, Wang F, Huang H X, Fu X D. 2015. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality. Nat Genet, 47(8): 949-954. |
| [118] |
Wang S S, Wu K, Qian Q, Liu Q, Li Q, Pan Y J, Ye Y F, Liu X Y, Wang J, Zhang J Q, Li S, Wu Y J, Fu X D. 2017. Non-canonical regulation of SPL transcription factors by a human OTUB1-like deubiquitinase defines a new plant type rice associated with higher grain yield. Cell Res, 27(9): 1142-1156.
PMID |
| [119] | Wang S X, Zong Y, Lin Q P, Zhang H W, Chai Z Z, Zhang D D, Chen K L, Qiu J L, Gao C X. 2020. Precise, predictable multi- nucleotide deletions in rice and wheat using APOBEC-Cas9. Nat Biotechnol, 38(12): 1460-1465. |
| [120] |
Wang Y D, Zhang T, Wang R C, Zhao Y D. 2018. Recent advances in auxin research in rice and their implications for crop improvement. J Exp Bot, 69(2): 255-263.
PMID |
| [121] | Wang Y X, Xiong G S, Hu J, Jiang L, Yu H, Xu J, Fang Y X, Zeng L J, Xu E B, Xu J, Ye W J, Meng X B, Liu R F, Chen H Q, Jing Y H, Wang Y H, Zhu X D, Li J Y, Qian Q. 2015. Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet, 47(8): 944-948. |
| [122] | Weng J F, Gu S H, Wan X Y, Gao H, Guo T, Su N, Lei C L, Zhang X, Cheng Z J, Guo X P, Wang J L, Jiang L, Zhai H Q, Wan J M. 2008. Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res, 18(12): 1199-1209. |
| [123] |
Workman J, Kingston R. 1998. Althernation of nucleosome structure as a mechanism of transcriptional regulation. Annu Rev Biochem, 67: 545-579.
PMID |
| [124] |
Wu J X, Qiu S J, Wang M L, Xu C J, Deng X W, Tang X Y. 2021. Construction of a weight-based seed sorting system for the third-generation hybrid rice. Rice, 14(1): 66.
PMID |
| [125] | Wu M L, Xie F P, Tang L, Yu Y H, Wang H M, Deng P, Liu Y. 2010. Design of separating cylinder of pocket hole separating machine on large-scale seeding production for hybrid rice. Transact Chin Society Agric Engin, 10(26): 1-4. (in English with Chinese abstract) |
| [126] | Xia D, Zhou H, Liu R J, Dan W H, Li P B, Wu B, Chen J X, Wang L Q, Gao G J, Zhang Q L, He Y Q. 2018. GL3.3, a novel QTL encoding a GSK3/SHAGGY-like kinase, epistatically interacts with GS3 to produce extra-long grains in rice. Mol Plant, 11(5): 754-756. |
| [127] |
Xiao Y H, Liu D P, Zhang G X, Tong H N, Chu C C. 2017. Brassinosteroids regulate OFP1, a DLT interacting protein, to modulate plant architecture and grain morphology in rice. Front Plant Sci, 8: 1698.
PMID |
| [128] | Xiao Y H, Liu D P, Zhang G X, Gao S P, Liu L C, Xu F, Che R H, Wang Y Q, Tong H N, Chu C C. 2019. Big Grain3, encoding a purine permease, regulates grain size via modulating cytokinin transport in rice. J Integr Plant Biol, 61(5): 581-597. |
| [129] | Xiao Y H, Zhang G X, Liu D P, Niu M, Tong H N, Chu C C. 2020a. GSK2 stabilizes OFP3 to suppress brassinosteroid responses in rice. Plant J, 102(6): 1187-1201. |
| [130] | Xiao Y H, Zhang J W, Yu G Y, Lu X D, Mei W T, Deng H B, Zhang G L, Chen G H, Chu C C, Tong H N, Tang W B. 2020b. Endoplasmic reticulum-localized PURINE PERMEASE1 regulates plant height and grain weight by modulating cytokinin distribution in rice. Front Plant Sci, 11: 618560. |
| [131] | Xie K B, Wu C Q, Xiong L Z. 2006. Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant Physiol, 142(1): 280-293. |
| [132] | Xu C J, Liu Y, Li Y B, Xu X D, Xu C G, Li X H, Xiao J H, Zhang Q F. 2015. Differential expression of GS5 regulates grain size in rice. J Exp Bot, 66(9): 2611-2623. |
| [133] | Xu E B, Wang Y X, Ni S, Chen H Q, Zhu X D. 2015. Application of recessive small grain gene in mechanical sorting of hybrid rice seeds. China Rice, 21(3): 8-11. (in Chinese) |
| [134] |
Xu R, Duan P G, Yu H Y, Zhou Z K, Zhang B L, Wang R C, Li J, Zhang G Z, Zhuang S S, Lyu J, Li N, Chai T Y, Tian Z X, Yao S G, Li Y H. 2018. Control of grain size and weight by the OsMKKK10-OsMKK4-OsMAPK6 signaling pathway in rice. Mol Plant, 11(6): 860-873.
PMID |
| [135] | Yan Y, Wei M X, Li Y, Tao H, Wu H Y, Chen Z F, Li C, Xu J H. 2021. MiR529a controls plant height, tiller number, panicle architecture and grain size by regulating SPL target genes in rice (Oryza sativa L.). Plant Sci, 302: 110728. |
| [136] | Yang C, Shen W J, He Y, Tian Z H, Li J X. 2016. OVATE family protein 8 positively mediates brassinosteroid signaling through interacting with the GSK3-like kinase in rice. PLoS Genet, 12(6): e1006118. |
| [137] |
Yang X F, Zhao X L, Dai Z Y, Ma F L, Miao X X, Shi Z Y. 2021. OsmiR396/growth regulating factor modulate rice grain size through direct regulation of embryo-specific miR408. Plant Physiol, 186(1): 519-533.
PMID |
| [138] | Yin W C, Xiao Y H, Niu M, Meng W J, Li L L, Zhang X X, Liu D P, Zhang G X, Qian Y W, Sun Z T, Huang R Y, Wang S P, Liu C M, Chu C C, Tong H N. 2020. ARGONAUTE2 enhances grain length and salt tolerance by activating BIG GRAIN3 to modulate cytokinin distribution in rice. Plant Cell, 32(7): 2292-2306. |
| [139] | Yu J, Miao J, Zhang Z, Xiong H, Zhu X, Sun X, Pan Y, Liang Y, Zhang Q, Abdul Rehman R M, Li J, Zhang H, Li Z. 2018. Alternative splicing of OsLG3b controls grain length and yield in japonica rice. Plant Biotechnol J, 16(9):1667-1678. |
| [140] |
Yuan H, Fan S J, Huang J, Zhan S J, Wang S F, Gao P, Chen W L, Tu B, Ma B T, Wang Y P, Qin P, Li S G. 2017. 08SG2/OsBAK1 regulates grain size and number, and functions differently in indica and japonica backgrounds in rice. Rice, 10(1): 25.
PMID |
| [141] | Yuan H, Gao P, Hu X L, Yuan M, Xu Z Y, Jin M Y, Song W C, Zhan S J, Zhu X B, Tu B, Li T, Wang Y P, Ma B T, Qin P, Chen W L, Li S G. 2022a. Fine mapping and candidate gene analysis of qGSN5, a novel quantitative trait locus coordinating grain size and grain number in rice. Theor Appl Genet, 135(1): 51-64. |
| [142] | Yuan H, Xu Z Y, Chen W L, Deng C Y, Liu Y, Yuan M, Gao P, Shi H, Tu B, Li T, Kang L Z, Ma B T, Wang Y P, Wang J, Chen X W, Li S G, Qin P. 2022b. OsBSK2, a putative brassinosteroid- signalling kinase, positively controls grain size in rice. J Exp Bot, 73(16): 5529-5542. |
| [143] |
Zhang B W, Wang X L, Zhao Z Y, Wang R J, Huang X H, Zhu Y L, Yuan L, Wang Y C, Xu X D, Burlingame A L, Gao Y J, Sun Y, Tang W Q. 2016. OsBRI1 activates BR signaling by preventing binding between the TPR and kinase domains of OsBSK3 via phosphorylation. Plant Physiol, 170(2): 1149-1161.
PMID |
| [144] | Zhang C, Xu Y Y, Guo S Y, Zhu J Y, Huan Q, Liu H H, Wang L, Luo G Z, Wang X J, Chong K. 2012. Dynamics of brassinosteroid response modulated by negative regulator LIC in rice. PLoS Genet, 8(4): e1002686. |
| [145] | Zhang D P, Zhang M Y, Liang J S. 2021. RGB1 regulates grain development and starch accumulation through its effect on OsYUC11-mediated auxin biosynthesis in rice endosperm cells. Front Plant Sci, 12: 585174. |
| [146] | Zhang H G, Wang R X, Xu Z P, Liu Q Q, Yan C J, Liang G H, Zhou Y, Tang S Z, Gu M H. 2021. Breeding of Yangnongjing 1030, a new medium maturity japonica rice variety with long grains. Jiangsu Agric Sci, 49(4): 62-65. (in Chinese with English abstract) |
| [147] | Zhang J Q, Gao X Y, Cai G, Wang Y J, Li J B, Du H Y, Wang R Q, Zhang H S, Huang J. 2021. An adenylate kinase OsAK3 involves brassinosteroid signaling and grain length in rice (Oryza sativa L.). Rice, 14(1): 105. |
| [148] |
Zhang L, Ma B, Bian Z, Li X Y, Zhang C Q, Liu J Y, Li Q, Liu Q Q, He Z H. 2020. Grain size selection using novel functional markers targeting 14 genes in rice. Rice, 13(1): 63.
PMID |
| [149] | Zhang X J, Wang J F, Huang J, Lan H X, Wang C L, Yin C F, Wu Y Y, Tang H J, Qian Q, Li J Y, Zhang H S. 2012. Rare allele of OsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc Natl Acad Sci USA, 109(52): 21534-21539. |
| [150] | Zhang Y D, Zhu Z, Zhao Q Y, Chen T, Yao S, Zhou L H, Zhao L, Zhao C F, Wang C L. 2016. Haplotypes of qGL3 and their roles in grain size regulation with GS3 alleles in rice. Genet Mol Res, 15(1): gmr.15017587. |
| [151] | Zhang Z X, Zhao H, Huang F L, Long J F, Song G, Lin W X. 2019. The 14-3-3 protein GF14f negatively affects grain filling of inferior spikelets of rice (Oryza sativa L.). Plant J, 99(2): 344-358. |
| [152] |
Zhang Z Y, Li J J, Tang Z S, Sun X M, Zhang H L, Yu J P, Yao G X, Li G L, Guo H F, Li J L, Wu H M, Huang H G, Xu Y W, Yin Z G, Qi Y H, Huang R F, Yang W C, Li Z C. 2018. Gnp4/LAX2, a RAWUL protein, interferes with the OsIAA3-OsARF25 interaction to regulate grain length via the auxin signaling pathway in rice. J Exp Bot, 69(20): 4723-4737.
PMID |
| [153] | Zhao D S, Li Q F, Zhang C Q, Zhang C, Yang Q Q, Pan L X, Ren X Y, Lu J, Gu M H, Liu Q Q. 2018. GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat Commun, 9(1): 1240. |
| [154] | Zhao J F, Wu C X, Yuan S J, Yin L, Sun W, Zhao Q L, Zhao B H, Li X Y. 2013. Kinase activity of OsBRI1 is essential for brassinosteroids to regulate rice growth and development. Plant Sci, 199/200: 113-120. |
| [155] |
Zhao Y D. 2010. Auxin biosynthesis and its role in plant development. Annu Rev Plant Biol, 61: 49-64.
PMID |
| [156] | Zheng Y M, Zhu Y S, Mao X H, Jiang M R, Wei Y D, Lian L, Xu H B, Chen L P, Xie H A, Lu G D, Zhang J F. 2022. SDR7-6, a short-chain alcohol dehydrogenase/reductase family protein, regulates light-dependent cell death and defence responses in rice. Mol Plant Pathol, 23(1): 78-91. |
| [157] | Zhou J Q, Zhang G L, Deng H B, Ming X Q, Lei B, Li F, Tang W B. 2022. Advantages of small grain male sterile lines in seed production for a new combination Zhuoliangyou 141 through the mixed-sowing manner. Acta Agron Sin, 48(2): 320-331. (in Chinese with English abstract) |
| [158] | Zhou Y, Zhu J Y, Li Z Y, Yi C D, Liu J, Zhang H G, Tang S Z, Gu M H, Liang G H. 2009. Deletion in a quantitative trait gene qPE9-1 associated with panicle erectness improves plant architecture during rice domestication. Genetics, 183(1): 315-324. |
| [159] | Zhou Y, Miao J, Gu H Y, Peng X R, Leburu M, Yuan F H, Gu H W, Gao Y, Tao Y J, Zhu J Y, Gong Z Y, Yi C D, Gu M H, Yang Z F, Liang G H. 2015. Natural variations in SLG7 regulate grain shape in rice. Genetics, 201(4): 1591-1599. |
| [160] | Zhou Y, Tao Y J, Zhu J Y, Miao J, Liu J, Liu Y H, Yi C D, Yang Z F, Gong Z Y, Liang G H. 2017. GNS4, a novel allele of DWARF11, regulates grain number and grain size in a high-yield rice variety. Rice, 10(1): 34. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||