
Rice Science ›› 2024, Vol. 31 ›› Issue (6): 634-637.DOI: 10.1016/j.rsci.2024.06.008
收稿日期:2024-02-19
接受日期:2024-06-07
出版日期:2024-11-28
发布日期:2024-12-10
. [J]. Rice Science, 2024, 31(6): 634-637.
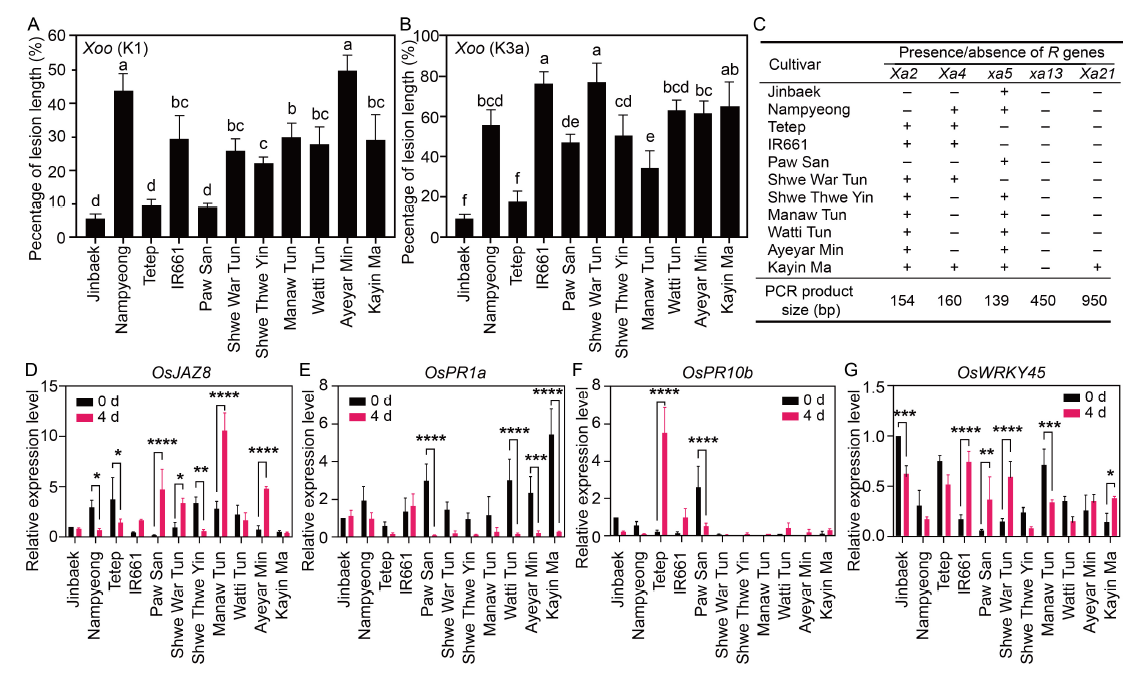
Fig. 1. Response of rice cultivars to Xanthomonas oryzae pv. oryzae (Xoo) infection. A-B, Percentage of lesion length of rice cultivars in response to infection by Xoo strains K1 (A) and K3a (B). C, PCR-based detection of variations in the expression of bacterial leaf blight resistance (R) genes (Xa2, Xa4, xa5, xa13, and Xa21) in 11 rice cultivars. ‘+’ and ‘-’ represent the presence and absence of the corresponding Xa gene, respectively. D-G, Transcript accumulation of resistance genes OsJAZ8 (D), OsPR1a (E), OsPR10b (F), and OsWRKY45 (G). Plants were inoculated with Xoo K1 and K3a strains by leaf clipping method. Leaves of the control plants were clipped by dipping the scissors in autoclaved distilled water. Leaf samples were collected for qRT-PCR after 4 d of inoculation. The rice actin gene (ACT1) was used as the internal control. Jinbaek and Nampyeong are bacterial leaf blight-resistant and -susceptible japonica rice lines, while Tetep and IR661 are bacterial leaf blight-resistant and -susceptible indica rice lines. Data in A and B are Mean ± SD (n = 27), and different lowercase letters indicate significant differences at the 0.05 level according to the Duncan’s multiple range test. Data in D-G are Mean ± SE (n = 3). *, **, ***, and **** indicate significant differences at the 0.05, 0.01, 0.001, and 0.0001 levels, respectively, followed by the Šídák’s multiple comparisons test.
| [1] | Ashiba R, Aiyanathan K E A, Kannan R, Pillai M A. 2020. Genotypic assessment of bacterial leaf blight resistance in indigenous rice (Oryza sativa L.) germplasm. Int J Curr Microbiol App Sci, 9(7): 210-227. |
| [2] | Bureau T E, Ronald P C, Wessler S R. 1996. A computer-based systematic survey reveals the predominance of small inverted- repeat elements in wild-type rice genes. Proc Natl Acad Sci USA, 93(16): 8524-8529. |
| [3] | Hajira S K, Sundaram R M, Laha G S, Yugander A, Balachandran S M, Viraktamath B C, Sujatha K, Balachiranjeevi C H, Pranathi K, Anila M, Bhaskar S, Abhilash V, Mahadevaswamy H K, Kousik M, Kumar T D, Harika G, Rekha G. 2016. A single-tube, functional marker-based multiplex PCR assay for simultaneous detection of major bacterial blight resistance genes Xa21, xa13 and xa5 in rice. Rice Sci, 23(3): 144-151. |
| [4] | Hou S, Tsuda K. 2022. Salicylic acid and jasmonic acid crosstalk in plant immunity. Essays Biochem, 66(5): 647-656. |
| [5] | Ji Z J, Yang S D, Zeng Y X, Liang Y, Yang C D, Qian Q. 2016. Pyramiding blast, bacterial blight and brown planthopper resistance genes in rice restorer lines. J Integr Agric, 15(7): 1432-1440. |
| [6] | Joshi R K, Nayak S. 2010. Gene pyramiding: A broad spectrum technique for developing durable stress resistance in crops. Biotechnol Mol Biol Rev, 5(3): 51-60. |
| [7] | Kazan K, Manners J M. 2012. JAZ repressors and the orchestration of phytohormone crosstalk. Trends Plant Sci, 17(1): 22-31. |
| [8] | Khush G S, Angeles E R. 1999. A new gene for resistance to race 6 of bacterial blight in rice, Oryza sativa L. Rice Genet Newsl, 16: 92-93. |
| [9] | Lee K S, Rasabandith S, Angeles E R, Khush G S. 2003. Inheritance of resistance to bacterial blight in 21 cultivars of rice. Phytopathology, 93(2): 147-152. |
| [10] | Li T, Zhan Z H, Lin Y N, Lin M J, Xie Q B, Chen Y H, He C Z, Tao J, Li C X. 2019. Biosynthesis of amino acids in Xanthomonas oryzae pv. oryzae is essential to its pathogenicity. Microorganisms, 7(12): 693. |
| [11] | Lwin C, Maung K, Murakami M, Hashimoto S. 2017. Scenarios of phosphorus flow from agriculture and domestic wastewater in Myanmar (2010-2100). Sustainability, 9(8): 1377. |
| [12] | Mishra D, Vishnupriya M R, Anil M G, Konda K, Raj Y, Sonti R V. 2013. Pathotype and genetic diversity amongst Indian isolates of Xanthomonas oryzae pv. oryzae. PLoS One, 8(11): e81996. |
| [13] | Noh T H, Song E S, Kim H I, Kang M H, Park Y J. 2016. Transcriptome-based identification of differently expressed genes from Xanthomonas oryzae pv. oryzae strains exhibiting different virulence in rice varieties. Int J Mol Sci, 17(2): 259. |
| [14] | Ogawa T, Lin L, Tabien R E, Khush G S. 1987. A new recessive gene for resistance to bacterial blight of rice. Rice Genet Newsl, 4(98): 100. |
| [15] | Oliva R, Ji C H, Atienza-Grande G, Huguet-Tapia J C, Perez- Quintero A, Li T, Eom J S, Li C H, Nguyen H, Liu B, Auguy F, Sciallano C, Luu V T, Dossa G S, Cunnac S, Schmidt S M, Slamet-Loedin I H, Vera Cruz C, Szurek B, Frommer W B, White F F, Yang B. 2019. Broad-spectrum resistance to bacterial blight in rice using genome editing. Nat Biotechnol, 37(11): 1344-1350. |
| [16] | Perumalsamy S, Bharani M, Sudha M, Nagarajan P, Arul L, Saraswathi R, Balasubramanian P, Ramalingam J. 2010. Functional marker- assisted selection for bacterial leaf blight resistance genes in rice (Oryza sativa L.). Plant Breed, 129(4): 400-406. |
| [17] | Petpisit V, Khush G S, Kauffman H E. 1977. Inheritance of resistance to bacterial blight in rice. Crop Sci, 17(4): cropsci1977.0011183X001700040018x. |
| [18] | Ronald P C, Albano B, Tabien R, Abenes L, Wu K S, McCouch S, Tanksley S D. 1992. Genetic and physical analysis of the rice bacterial blight disease resistance locus, Xa21. Mol Gen Genet, 236(1): 113-120. |
| [19] | Sanya D R A, Syed-Ab-Rahman S F, Jia A Q, Onésime D, Kim K M, Ahohuendo B C, Rohr J R. 2022. A review of approaches to control bacterial leaf blight in rice. World J Microbiol Biotechnol, 38(7): 113. |
| [20] | Suh J P, Noh T H, Kim K Y, Kim J J, Kim Y G, Jena K K. 2009. Expression levels of three bacterial blight resistance genes against K3a race of Korea by molecular and phenotype analysis in japonica rice (O. sativa L.). J Crop Sci Biotechnol, 12(3): 103-108. |
| [21] | Sun H, Zhu X L, Li C X, Ma Z M, Han X, Luo Y Y, Yang L, Yu J, Miao Y S. 2021. Xanthomonas effector XopR hijacks host actin cytoskeleton via complex coacervation. Nat Commun, 12(1): 4064. |
| [22] | Swamy P, Panchbhai A N, Dodiya P, Naik V, Panchbhai S D, Zehr U B, Azhakanandam K, Char B R. 2006. Evaluation of bacterial blight resistance in rice lines carrying multiple resistance genes and Xa21 transgenic lines. Curr Sci, 90(6): 818-824. |
| [23] | Tu J, Ona I, Zhang Q, Mew T W, Khush G S, Datta S K. 1998. Transgenic rice variety ‘IR72’ with Xa21 is resistant to bacterial blight. Theor Appl Genet, 97: 31-36. |
| [24] | Vieira P S, Bonfim I M, Araujo E A, Melo R R, Lima A R, Fessel M R, Paixão D A A, Persinoti G F, Rocco S A, Lima T B, Pirolla R A S, Morais M A B, Correa J B L, Zanphorlin L M, Diogo J A, Lima E A, Grandis A, Buckeridge M S, Gozzo F C, Benedetti C E, Polikarpov I, Giuseppe P O, Murakami M T. 2021. Xyloglucan processing machinery in Xanthomonas pathogens and its role in the transcriptional activation of virulence factors. Nat Commun, 12(1): 4049. |
| [25] | Vikal Y, Chawla H, Rajiv S, Lore J S, Singh K. 2014. Mapping of bacterial blight resistance gene xa8in rice (Oryza sativa L.). Indian J Genet Plant Breed, 74: 589-595. |
| [26] | Wang B, Wu G C, Zhang Y Q, Qian G L, Liu F Q. 2018. Dissecting the virulence-related functionality and cellular transcription mechanism of a conserved hypothetical protein in Xanthomonas oryzae pv. oryzae. Mol Plant Pathol, 19(8): 1859-1872. |
| [27] | Wang G L, Wu C, Zeng L, He C, Baraoidan M, de Assis Goes da Silva F, Williams C E,, Ronald P C, Leung H. 2004. Isolation and characterization of rice mutants compromised in Xa21-mediated resistance to X. oryzae pv. oryzae. Theor Appl Genet, 108(3): 379-384. |
| [28] | Zheng C K, Wang C L, Yu Y J, Liang Y T, Zhao K J. 2009. Identification and molecular mapping of Xa32(t), a novel resistance gene for bacterial blight (Xanthomonas oryzae pv. oryzae) in rice. Acta Agron Sin, 35(7): 1173-1180. (in Chinese with English abstract) |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||