
Rice Science ›› 2025, Vol. 32 ›› Issue (3): 277-282.DOI: 10.1016/j.rsci.2024.12.010
收稿日期:2024-10-20
接受日期:2024-12-25
出版日期:2025-05-28
发布日期:2025-06-16
. [J]. Rice Science, 2025, 32(3): 277-282.
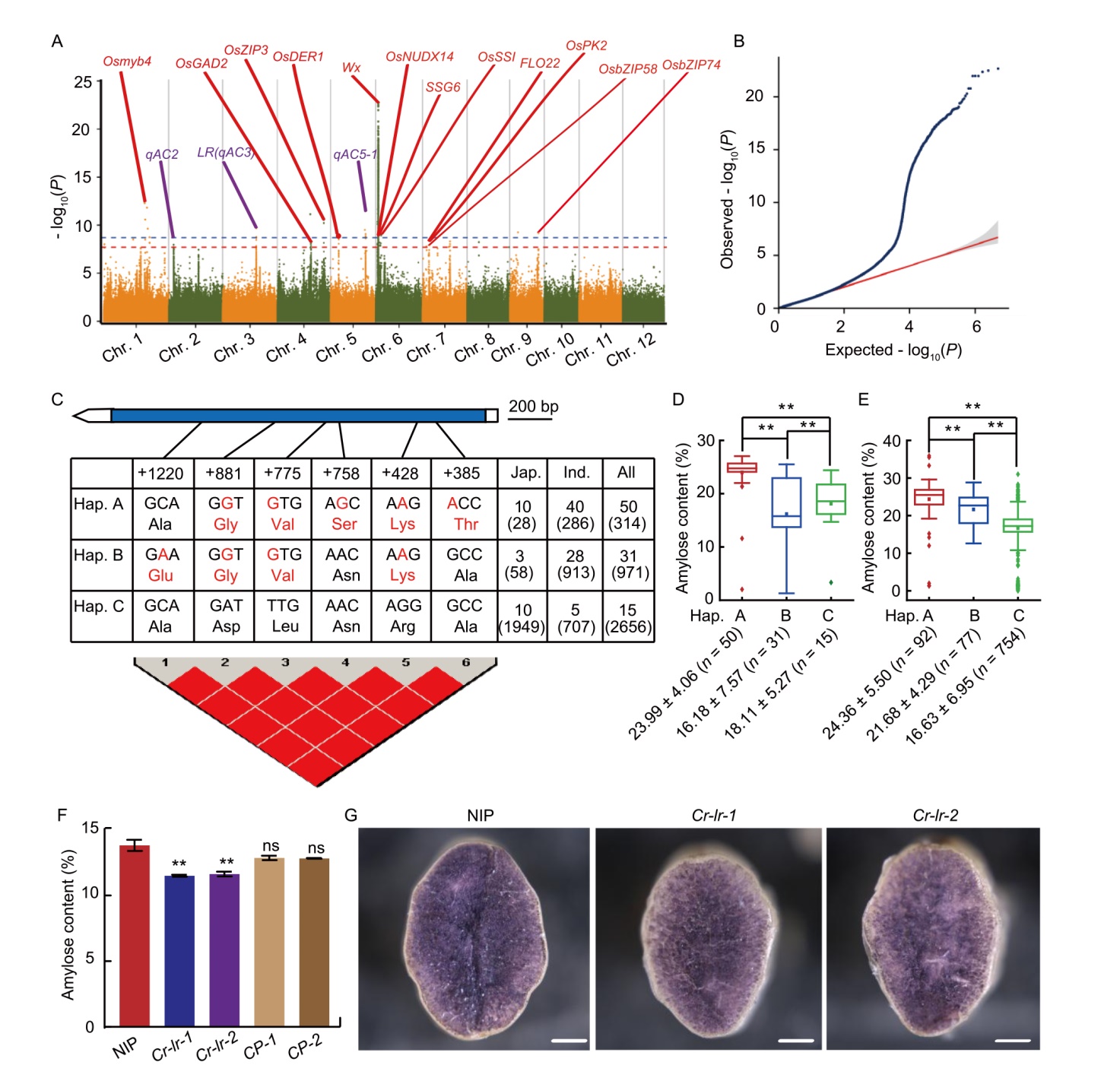
Fig. 1. Genome-wide association analysis of amylose content (AC) in Ting’s core collection and functional analysis of LR. A, Manhattan plots for AC. Genes in red represent reported genes, while purple represent candidate genes. Red and blue horizontal dashed lines indicate significance thresholds at P ≤ 1.93 × 10-8 and P ≤ 1.93 × 10-9, respectively. Chr., Chromosome. B, Quantile-quantile plot for AC using MLM (Q + K) model. Blue line represents the expected -log10(P), and red line represents the observed -log10(P). C, Haplotypes of LR. Blue frame represents the exon of LR. Red nucleotides represent natural variations, causing amino acid changes. Numbers represent counts of indica (Ind.), japonica (Jap.), and all rice in the Ting’s core collection and MBKBASE website (in parentheses), respectively. D and E, Comparison of ACs among different haplotypes in the Ting’s core germplasm (D) and MBKBASE website (E). Horizontal lines represent medians, nearest points represent means, and whiskers mark the range from the 5th to the 95th percentile of the total data. F, AC of wild type (Nipponbare, NIP), knockout (Cr-lr-1 and Cr-lr-2), and complementation (CP-1 and CP-2) mutants. G, Transverse sections stained with iodine in wild type (NIP) and knockout mutant (Cr-lr-1 and Cr-lr-2). Scale bars, 500 µm. In D-F, ** represents P < 0.01 by the Student’s t-test. ns, Not significant.
| [1] | Bai C, Wang G J, Feng X H, et al. 2024. OsMAPK6 phosphorylation and CLG1 ubiquitylation of GW6a non-additively enhance rice grain size through stabilization of the substrate. Nat Commun, 15(1): 4300. |
| [2] | Bella J, Hindle K L, McEwan P A, et al. 2008. The leucine-rich repeat structure. Cell Mol Life Sci, 65(15): 2307-2333. |
| [3] | Cai Y C, Li S F, Jiao G A, et al. 2018. OsPK2 encodes a plastidic pyruvate kinase involved in rice endosperm starch synthesis, compound granule formation and grain filling. Plant Biotechnol J, 16(11): 1878-1891. |
| [4] | Deng B W, Zhang Y N, Zhang F, et al. 2024. Genome-wide association study of cooked rice textural attributes and starch physicochemical properties in indica rice. Rice Sci, 31(3): 300-316. |
| [5] | Feng L, Gao Z R, Xiao G Q, et al. 2014. Leucine-rich repeat receptor-like kinase FON1 regulates drought stress and seed germination by activating the expression of ABA-responsive genes in rice. Plant Mol Biol Report, 32(6): 1158-1168. |
| [6] | Fu Y H, Hua Y H, Luo T T, et al. 2023. Generating waxy rice starch with target type of amylopectin fine structure and gelatinization temperature by waxy gene editing. Carbohydr Polym, 306: 120595. |
| [7] | Graham-Acquaah S, Mauromoustakos A, Cuevas R P, et al. 2020. Differences in physicochemical properties of commercial rice from urban markets in west Africa. J Food Sci Technol, 57(4): 1505-1516. |
| [8] | Holzwart E, Huerta A I, Glöckner N, et al. 2018. BRI1 controls vascular cell fate in the Arabidopsis root through RLP44 and phytosulfokine signaling. Proc Natl Acad Sci USA, 115(46): 11838-11843. |
| [9] | Ji D L, Xiao W H, Sun Z W, et al. 2023. Translocation and distribution of carbon-nitrogen in relation to rice yield and grain quality as affected by high temperature at early panicle initiation stage. Rice Sci, 30(6): 598-612. |
| [10] | Kang J F, Li J M, Gao S, et al. 2017. Overexpression of the leucine-rich receptor-like kinase gene LRK2 increases drought tolerance and tiller number in rice. Plant Biotechnol J, 15(9): 1175-1185. |
| [11] | Kawakatsu T, Takaiwa F. 2010. Differences in transcriptional regulatory mechanisms functioning for free lysine content and seed storage protein accumulation in rice grain. Plant Cell Physiol, 51(12): 1964-1974. |
| [12] | Lin F M, Li S, Wang K, et al. 2020. A leucine-rich repeat receptor-like kinase, OsSTLK, modulates salt tolerance in rice. Plant Sci, 296: 110465. |
| [13] | Lin L S, Li Z, Ning M Y, et al. 2024. A mutant allele of the Wx gene encoding granule-bound starch synthase I results in extremely low amylose content in rice. Plant Physiol, 196(4): 2296-2299. |
| [14] | Liu Y R, Zhang W, Wang Y H, et al. 2022. Nudix hydrolase 14 influences plant development and grain chalkiness in rice. Front Plant Sci, 13: 1054917. |
| [15] | Matsushima R, Maekawa M, Kusano M, et al. 2016. Amyloplast membrane protein SUBSTANDARD STARCH GRAIN6 controls starch grain size in rice endosperm. Plant Physiol, 170(3): 1445-1459. |
| [16] | Mikami I, Aikawa M, Hirano H Y, et al. 1999. Altered tissue-specific expression at the Wx gene of the opaque mutants in rice. Euphytica, 105(2): 91-97. |
| [17] | Mikami I, Uwatoko N, Ikeda Y, et al. 2008. Allelic diversification at the wx locus in landraces of Asian rice. Theor Appl Genet, 116(7): 979-989. |
| [18] | Nagar P, Sharma N, Jain M, et al. 2022. OsPSKR15, a phytosulfokine receptor from rice enhances abscisic acid response and drought stress tolerance. Physiol Plant, 174(1): e13569. |
| [19] | Nie S, Chen L, Zheng M H, et al. 2024. GWAS and transcriptomic analysis identify OsRING315 as a new candidate gene controlling amylose content and gel consistency in rice. Rice, 17(1): 38. |
| [20] | Qian D D, Chen G Q, Tian L H, et al. 2018. OsDER1 is an ER-associated protein degradation factor that responds to ER stress. Plant Physiol, 178(1): 402-412. |
| [21] | Sato H, Suzuki Y, Sakai M, et al. 2002. Molecular characterization of Wx-mq, a novel mutant gene for low-amylose content in endosperm of rice (Oryza sativa L.). Breed Sci, 52(2): 131-135. |
| [22] | Shi Y H, Zhang Y Y, Sun Y Y, et al. 2023. Natural variations of OsAUX5, a target gene of OsWRKY78, control the neutral essential amino acid content in rice grains. Mol Plant, 16(2): 322-336. |
| [23] | Suzuki A, Suzuki T, Tanabe F, et al. 1997. Cloning and expression of five myb-related genes from rice seed. Gene, 198(1/2): 393-398. |
| [24] | Tian S Q, Liang S, Qiao K, et al. 2019. Co-expression of multiple heavy metal transporters changes the translocation, accumulation, and potential oxidative stress of Cd and Zn in rice (Oryza sativa). J Hazard Mater, 380: 120853. |
| [25] | Tilman D, Balzer C, Hill J, et al. 2011. Global food demand and the sustainable intensification of agriculture. Proc Natl Acad Sci USA, 108(50): 20260-20264. |
| [26] | Tu B, Zhang T, Liu P, et al. 2024.The LCG1-OsBP5/OsEBP89-Wx module regulates the grain chalkiness and taste quality in rice Plant Biotechnol J: 1-15. |
| [27] | Wang J, Jiang Q H, Pleskot R, et al. 2023. TPLATE complex- dependent endocytosis attenuates CLAVATA1 signaling for shoot apical meristem maintenance. EMBO Rep, 24(9): e54709. |
| [28] | Xu Y, Lin Q P, Li X F, et al. 2021. Fine-tuning the amylose content of rice by precise base editing of the Wx gene. Plant Biotechnol J, 19(1): 11-13. |
| [29] | Yamakawa H, Hirose T, Kuroda M, et al. 2007. Comprehensive expression profiling of rice grain filling-related genes under high temperature using DNA microarray. Plant Physiol, 144(1): 258-277. |
| [30] | Yang H, Wang Y L, Tian Y L, et al. 2023. Rice FLOURY ENDOSPERM22, encoding a pentatricopeptide repeat protein, is involved in both mitochondrial RNA splicing and editing and is crucial for endosperm development. J Integr Plant Biol, 65(3): 755-771. |
| [31] | Yang J, Wang J, Fan F J, et al. 2013.Development of AS-PCR marker based on a key mutation confirmed by resequencing of Wxmp in Milky Princess and its application in japonica soft rice (Oryza sativa L.) breeding. Plant Breed, 132(6): 595-603. |
| [32] | Yao S, Zhang Y D, Liu Y Q, et al. 2020. Effects of soluble starch synthase genes on eating and cooking quality in semi waxy japonica rice with Wxmp. Food Prod Process Nutr, 2(1): 22. |
| [33] | Zhang C Q, Chen S J, Ren X Y, et al. 2017. Molecular structure and physicochemical properties of starches from rice with different amylose contents resulting from modification of OsGBSSI activity. J Agric Food Chem, 65(10): 2222-2232. |
| [34] | Zhang C Q, Zhu J H, Chen S J, et al. 2019. Wxlv, the ancestral allele of rice waxy gene. Mol Plant, 12(8): 1157-1166. |
| [35] | Zhang C Q, Yang Y, Chen S J, et al. 2021. A rare waxy allele coordinately improves rice eating and cooking quality and grain transparency. J Integr Plant Biol, 63(5): 889-901. |
| [36] | Zhang P, Li J Q, Li X L, et al. 2011. Population structure and genetic diversity in a rice core collection (Oryza sativa L.) investigated with SSR markers. PLoS One, 6(12): e27565. |
| [37] | Zhang P, Liu X D, Tong H H, et al. 2014. Association mapping for important agronomic traits in core collection of rice (Oryza sativa L.) with SSR markers. PLoS One, 9(10): e111508. |
| [38] | Zhang P, Zhong K Z, Zhong Z Z, et al. 2019a. Genome-wide association study of important agronomic traits within a core collection of rice (Oryza sativa L.). BMC Plant Biol, 19(1): 259. |
| [39] | Zhang P, Zhong K Z, Zhong Z Z, et al. 2019b. Mining candidate gene for rice aluminum tolerance through genome wide association study and transcriptomic analysis. BMC Plant Biol, 19(1): 490. |
| [40] | Zhou H, Xia D, Zhao D, et al. 2021. The origin of Wxla provides new insights into the improvement of grain quality in rice. J Integr Plant Biol, 63(5): 878-888. |
| No related articles found! |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||