
Rice Science ›› 2022, Vol. 29 ›› Issue (6): 559-568.DOI: 10.1016/j.rsci.2022.05.003
• Research Paper • Previous Articles Next Articles
Lü Jun#, Liu Jinhui#, Chen Lin, Sun Jiawei, Su Qin, Li Shihui, Yang Jianhua, Zhang Wenqing( )
)
Received:2022-02-24
Accepted:2022-05-11
Online:2022-11-28
Published:2022-09-09
Contact:
Zhang Wenqing
About author:#These authors contributed equally to this work
Lü Jun, Liu Jinhui, Chen Lin, Sun Jiawei, Su Qin, Li Shihui, Yang Jianhua, Zhang Wenqing. Screening of Brown Planthopper Resistant miRNAs in Rice and Their Roles in Regulation of Brown Planthopper Fecundity[J]. Rice Science, 2022, 29(6): 559-568.
Add to citation manager EndNote|Ris|BibTeX
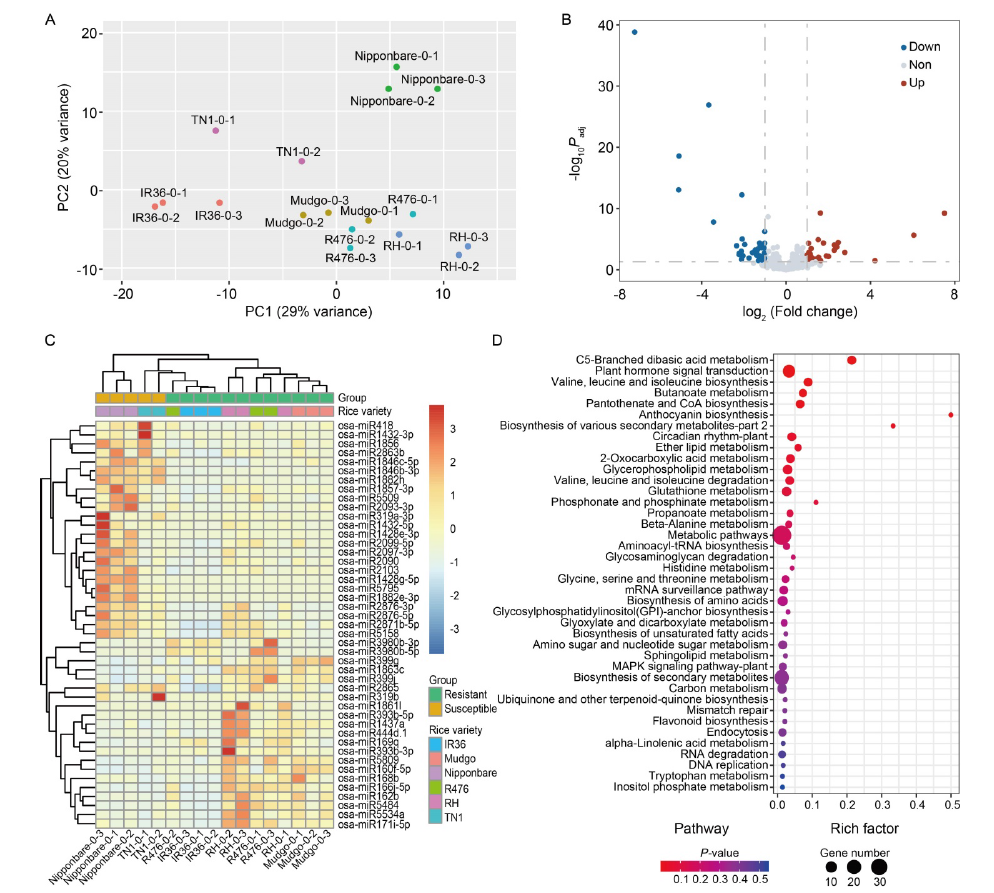
Fig. 1. miRNAs differentially expressed in brown planthopper (BPH)-susceptible and BPH-resistant rice varieties before BPH-feeding. A, Principal component analysis (PCA) of miRNAs in rice samples before BPH-feeding. B, Volcano plot of differential miRNA expression in BPH-resistant and BPH-susceptible rice varieties. ‘down’ and ‘up’ mean miRNA expression was down-regulated and up-regulated in BPH-resistant rice varieties compared with BPH-susceptible rice varieties, respectively, and ‘non’ means the expression of miRNA was not significantly different between the BPH-resistant and BPH-susceptible rice varieties. C, Heatmap of differential miRNA expression profile. D, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment of target genes of differential miRNAs. RH, Rathu Heenati.
| Rice variety | After BPH-feeding at 8 h / Before BPH-feeding (0 h) | |||
|---|---|---|---|---|
| TN1 | miR5792 | miR5489 | miR160f-5p | miR3980b-5p |
| miR531c | miR1861l | |||
| miR5501 | miR5801a | miR5795 | miR1882e-3p | |
| Nipponbare | miR160e-5p | miR160f-5p | miR393b-3p | miR160d-5p |
| miR394 | ||||
| IR36 | miR1850.3 | miR2863b | miR6249b | miR5337a |
| miR2876-5p | miR2876-3p | miR1846b-3p | miR2865 | |
| miR2092-3p | miR5795 | miR418 | ||
| miR3980b-5p | miR531c | miR399h | miR398b | |
| miR1850.1 | miR156l-5p | miR3980b-3p | miR1320-3p | |
| Mudgo | miR528-3p | miR168b | miR167h-3p | miR171e-5p |
| miR2102-3p | miR171d-5p | miR5809 | ||
| RH | miR1428e-3p | miR1428g-5p | miR319b | |
| miR5078 | miR398b | miR393b-3p | miR1430 | |
| miR166j-5p | miR5077 | miR164f | miR169g | |
| miR396f-3p | miR167j | miR1432-5p | miR169r-3p | |
| miR156j-3p | miR168a-5p | miR169i-5p.2 | miR166d-5p | |
| miR162b | miR156l-3p | miR399d | miR169r-5p | |
| miR810b.2 | miR396c-5p | miR444f | miR397a | |
| miR171h | miR393b-5p | miR7693-5p | miR397b | |
| miR444c.1 | miR810a | miR810b.1 | miR169e | |
| miR169c | miR164d | miR159b | miR169q | |
| Rice variety | After BPH-feeding at 32 h / Before BPH-feeding (0 h) | |||
| TN1 | miR3979-5p | miR398b | miR3980b-5p | miR531c |
| miR2876-3p | ||||
| IR36 | miR2863b | miR1320-3p | miR5805 | miR2092-3p |
| miR6256 | miR5806 | miR398b | miR5535 | |
| miR2922 | miR169q | miR171e-5p | miR3979-3p | |
| R476 | miR169e | miR5830 | miR6255 | miR171a |
| miR159d | miR396b-3p | miR5486 | miR2877 | |
| miR5490 | miR5081 | miR2106 | miR827 | |
| miR1882h | miR1882e-3p | miR156j-5p | miR438 | |
| miR169a | miR1320-3p | miR5822 | miR5527 | |
| miR390-3p | miR6253 | miR169p | miR164d | |
| miR5151 | miR417 | miR156l-5p | miR5494 | |
| miR5523 | miR160d-3p | miR5076 | miR5539b | |
| miR5073 | miR171d-5p | miR5075 | miR5829 | |
| miR1850.1 | miR166k-5p | miR2100-5p | miR166h-5p | |
| miR169r-3p | miR166e-5p | miR160f-3p | miR1861c | |
| miR395o | miR167a-3p | miR1850.2 | miR166b-5p | |
| miR2928 | miR810a | miR5790 | miR171e-5p | |
| miR2102-3p | ||||
| Mudgo | miR171a | miR5517 | miR1859 | miR2106 |
| miR5511 | miR160d-3p | miR166e-3p | miR1320-3p | |
| miR528-3p | miR396c-3p | miR2930 | miR2102-3p | |
| miR2926 | miR171e-5p | miR6246 | miR5073 | |
| miR171d-5p | miR408-5p | miR5153 | miR168b | |
| miR167h-3p | miR2924 | miR6249b | miR160f-5p | |
| miR1850.1 | miR167i-3p | miR1858b | miR160f-3p | |
| miR5809 | miR1437a | miR166l-5p | miR1846d-5p | |
| RH | miR529b | miR535-5p | ||
| miR396c-3p | miR166k-5p | miR171d-5p | miR530-3p | |
| miR166h-5p | miR160f-3p | miR172d-5p | miR810a | |
| miR5073 | miR1850.1 | miR166d-5p | miR6246 | |
| miR166b-5p | miR5485 | miR166e-5p | miR171e-5p | |
Table 1. Summary of rice miRNAs in response to brown planthopper (BPH)-feeding.
| Rice variety | After BPH-feeding at 8 h / Before BPH-feeding (0 h) | |||
|---|---|---|---|---|
| TN1 | miR5792 | miR5489 | miR160f-5p | miR3980b-5p |
| miR531c | miR1861l | |||
| miR5501 | miR5801a | miR5795 | miR1882e-3p | |
| Nipponbare | miR160e-5p | miR160f-5p | miR393b-3p | miR160d-5p |
| miR394 | ||||
| IR36 | miR1850.3 | miR2863b | miR6249b | miR5337a |
| miR2876-5p | miR2876-3p | miR1846b-3p | miR2865 | |
| miR2092-3p | miR5795 | miR418 | ||
| miR3980b-5p | miR531c | miR399h | miR398b | |
| miR1850.1 | miR156l-5p | miR3980b-3p | miR1320-3p | |
| Mudgo | miR528-3p | miR168b | miR167h-3p | miR171e-5p |
| miR2102-3p | miR171d-5p | miR5809 | ||
| RH | miR1428e-3p | miR1428g-5p | miR319b | |
| miR5078 | miR398b | miR393b-3p | miR1430 | |
| miR166j-5p | miR5077 | miR164f | miR169g | |
| miR396f-3p | miR167j | miR1432-5p | miR169r-3p | |
| miR156j-3p | miR168a-5p | miR169i-5p.2 | miR166d-5p | |
| miR162b | miR156l-3p | miR399d | miR169r-5p | |
| miR810b.2 | miR396c-5p | miR444f | miR397a | |
| miR171h | miR393b-5p | miR7693-5p | miR397b | |
| miR444c.1 | miR810a | miR810b.1 | miR169e | |
| miR169c | miR164d | miR159b | miR169q | |
| Rice variety | After BPH-feeding at 32 h / Before BPH-feeding (0 h) | |||
| TN1 | miR3979-5p | miR398b | miR3980b-5p | miR531c |
| miR2876-3p | ||||
| IR36 | miR2863b | miR1320-3p | miR5805 | miR2092-3p |
| miR6256 | miR5806 | miR398b | miR5535 | |
| miR2922 | miR169q | miR171e-5p | miR3979-3p | |
| R476 | miR169e | miR5830 | miR6255 | miR171a |
| miR159d | miR396b-3p | miR5486 | miR2877 | |
| miR5490 | miR5081 | miR2106 | miR827 | |
| miR1882h | miR1882e-3p | miR156j-5p | miR438 | |
| miR169a | miR1320-3p | miR5822 | miR5527 | |
| miR390-3p | miR6253 | miR169p | miR164d | |
| miR5151 | miR417 | miR156l-5p | miR5494 | |
| miR5523 | miR160d-3p | miR5076 | miR5539b | |
| miR5073 | miR171d-5p | miR5075 | miR5829 | |
| miR1850.1 | miR166k-5p | miR2100-5p | miR166h-5p | |
| miR169r-3p | miR166e-5p | miR160f-3p | miR1861c | |
| miR395o | miR167a-3p | miR1850.2 | miR166b-5p | |
| miR2928 | miR810a | miR5790 | miR171e-5p | |
| miR2102-3p | ||||
| Mudgo | miR171a | miR5517 | miR1859 | miR2106 |
| miR5511 | miR160d-3p | miR166e-3p | miR1320-3p | |
| miR528-3p | miR396c-3p | miR2930 | miR2102-3p | |
| miR2926 | miR171e-5p | miR6246 | miR5073 | |
| miR171d-5p | miR408-5p | miR5153 | miR168b | |
| miR167h-3p | miR2924 | miR6249b | miR160f-5p | |
| miR1850.1 | miR167i-3p | miR1858b | miR160f-3p | |
| miR5809 | miR1437a | miR166l-5p | miR1846d-5p | |
| RH | miR529b | miR535-5p | ||
| miR396c-3p | miR166k-5p | miR171d-5p | miR530-3p | |
| miR166h-5p | miR160f-3p | miR172d-5p | miR810a | |
| miR5073 | miR1850.1 | miR166d-5p | miR6246 | |
| miR166b-5p | miR5485 | miR166e-5p | miR171e-5p | |
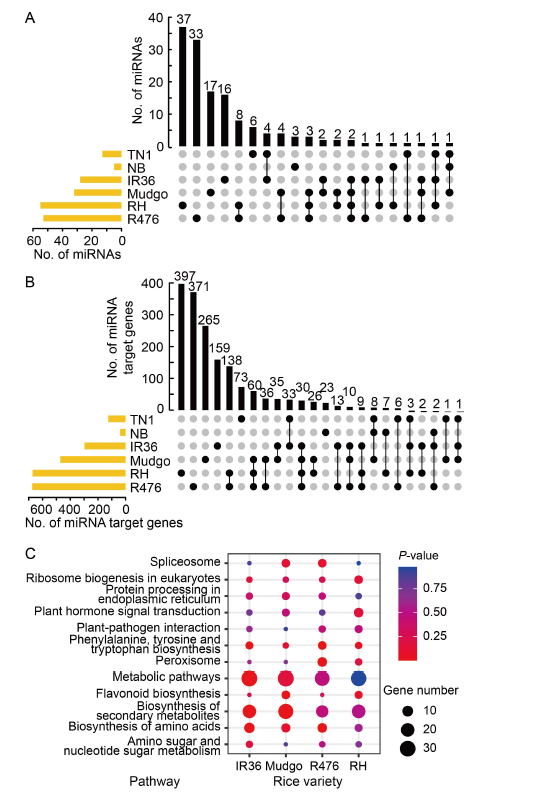
Fig. 2. Different miRNAs participate in rice-brown planthopper (BPH) interactions after BPH-feeding. A, UpSet diagram of miRNAs in rice response to BPH-feeding. ‘●’ indicates that the miRNAs appeared in only one rice variety, and ‘●-●’ indicates that the miRNAs appeared in two or more rice varieties. B, UpSet diagram of miRNA target genes in rice response to BPH- feeding. ‘●’ indicates that the miRNA target genes appeared in only one rice variety, and ‘●-●’ indicates that the miRNA target genes appeared in two or more rice varieties. C, Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment pathway is shared by four BPH-resistant rice varieties. NB, Nipponbare; RH, Rathu Heenati.
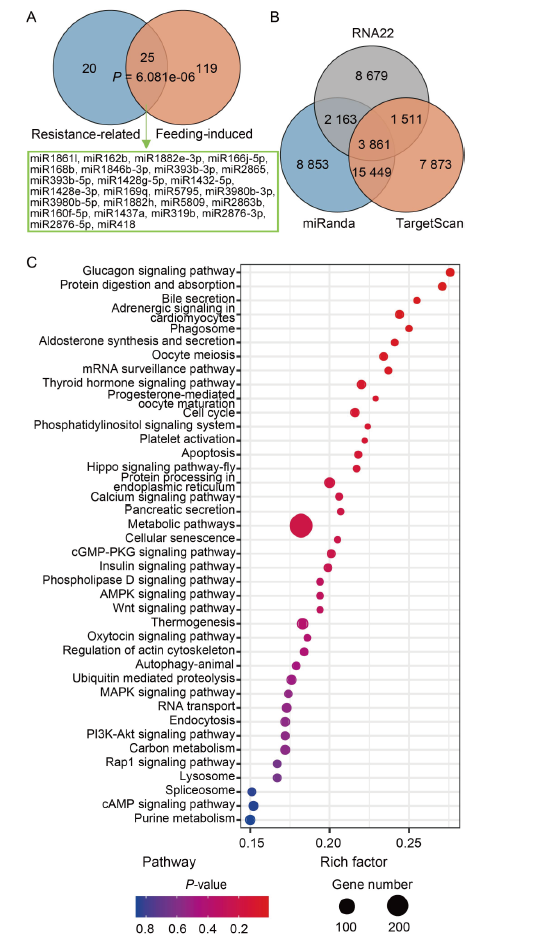
Fig. 3. Screening of potential cross-kingdom miRNAs in rice. A, Venn diagram of two sets of miRNAs. B, Venn diagram of target prediction of rice miRNAs using three bioinformatic algorithms (miRanda, TargetScan and RNA22) based on the total transcripts of Nilaparvata lugens. C, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment of target genes in N. lugens of miRNAs.
| miRNA name | No. of sequences in coding sequence region | No. of sequences in 3′-untranslated region | |||||
|---|---|---|---|---|---|---|---|
| miRanda | RNA22 | TargetScan | miRanda | RNA22 | TargetScan | ||
| miR1428g-5p | 2 | 1 | 2 | 2 | 1 | 3 | |
| miR1432-5p | 2 | 2 | 3 | 0 | 1 | 1 | |
| miR168b | 4 | 12 | 3 | 1 | 2 | 0 | |
| miR1882h | 4 | 4 | 2 | 2 | 0 | 0 | |
| miR2876-5p | 7 | 8 | 5 | 0 | 0 | 1 | |
| miR3980b-5p | 7 | 0 | 4 | 0 | 0 | 1 | |
| miR5795 | 8 | 11 | 8 | 0 | 1 | 0 | |
Table 2. Number of targets of rice miRNAs on fecundity of Nilaparvata lugens.
| miRNA name | No. of sequences in coding sequence region | No. of sequences in 3′-untranslated region | |||||
|---|---|---|---|---|---|---|---|
| miRanda | RNA22 | TargetScan | miRanda | RNA22 | TargetScan | ||
| miR1428g-5p | 2 | 1 | 2 | 2 | 1 | 3 | |
| miR1432-5p | 2 | 2 | 3 | 0 | 1 | 1 | |
| miR168b | 4 | 12 | 3 | 1 | 2 | 0 | |
| miR1882h | 4 | 4 | 2 | 2 | 0 | 0 | |
| miR2876-5p | 7 | 8 | 5 | 0 | 0 | 1 | |
| miR3980b-5p | 7 | 0 | 4 | 0 | 0 | 1 | |
| miR5795 | 8 | 11 | 8 | 0 | 1 | 0 | |
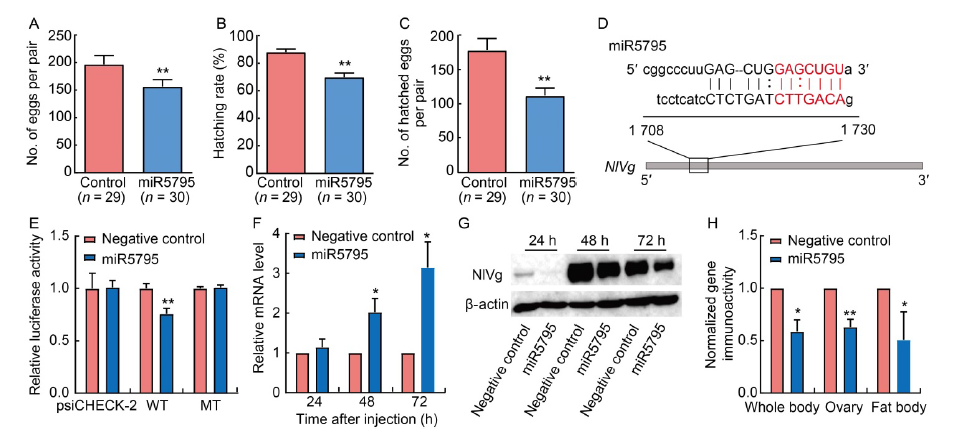
Fig. 4. miR5795 in rice may target vitellogenin gene in brown planthopper (BPH) and subsequently decrease its fecundity. A-C, The number of eggs per pair (A), hatching rate (B) and the number of hatched eggs per pair (C) of Nilaparvata lugens after injection of miR5795 mimics. D, Predicted binding sites between miR5795 and NlVg with energy of -87.65 kJ/mol. E, Dual-luciferase assay between miR5795 and NlVg. psiCHECK-2, Vector without insertion sites; WT, Wild type vector; MT, Mutation type vector. F, mRNA expression level of NlVg in whole bodies of BPH after injection of miR5795 mimics at 24, 48 and 72 h. G, Western blot for NlVg in whole bodies of BPH after injection of miR5795 mimics at 24, 48 and 72 h. β-actin was used as an internal reference. H, Protein expression levels of NlVg in whole bodies, ovaries and fat bodies of BPH after injection of miR5795 at 72 h. Data are Mean ± SE. n = 3 in A-C, F and H, and n = 9 in E. *, P < 0.05; **, P < 0.01.
| [1] | Agarwal V, Bell G W, Nam J W, Bartel D P. 2015. Predicting effective microRNA target sites in mammalian mRNAs. eLife, 4: e05005. |
| [2] | Akanksha S, Lakshmi V J, Singh A K, Deepthi Y, Chirutkar P M, Ramdeen, Balakrishnan D, Sarla N, Mangrauthia S K, Ram T. 2019. Genetics of novel brown planthopper Nilaparvata lugens (Stål) resistance genes in derived introgression lines from the interspecific cross O. sativa var. Swarna × O. nivara. J Genet, 98: 113. |
| [3] | Bao Y Y, Wang Y, Wu W J, Zhao D, Xue J, Zhang B Q, Shen Z C, Zhang C X. 2012. De novo intestine-specific transcriptome of the brown planthopper Nilaparvata lugens revealed potential functions in digestion, detoxification and immune response. Genomics, 99(4): 256-264. |
| [4] |
Bartel D P. 2004. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell, 116(2): 281-297.
PMID |
| [5] | Chen H, Stout M J, Qian Q, Chen F. 2012. Genetic, molecular and genomic basis of rice defense against insects. Crit Rev Plant Sci, 31(1): 74-91. |
| [6] | Chen J, Liang Z K, Liang Y K, Pang R, Zhang W Q. 2013. Conserved microRNAs miR-8-5p and miR-2a-3p modulate chitin biosynthesis in response to 20-hydroxyecdysone signaling in the brown planthopper, Nilaparvata lugens. Insect Biochem Mol Biol, 43(9): 839-848. |
| [7] |
Cheng X Y, Zhu L L, He G C. 2013. Towards understanding of molecular interactions between rice and the brown planthopper. Mol Plant, 6(3): 621-634.
PMID |
| [8] |
Chin A R, Fong M Y, Somlo G, Wu J, Swiderski P, Wu X W, Wang S E. 2016. Cross-kingdom inhibition of breast cancer growth by plant miR159. Cell Res, 26(2): 217-228.
PMID |
| [9] | Dai X B, Zhuang Z H, Zhao P X. 2018. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res, 46(W1): W49-W54. |
| [10] | Dai Z Y, Tan J, Zhou C, Yang X F, Yang F, Zhang S J, Sun S C, Miao X X, Shi Z Y. 2019. The OsmiR396-OsGRF8-OsF3H- flavonoid pathway mediates resistance to the brown planthopper in rice (Oryza sativa). Plant Biotechnol J, 17(8): 1657-1669. |
| [11] | Dong S Z, Ma Y, Hou Y, Yu X P, Ye G Y. 2011. Development of an ELISA for evaluating the reproductive status of female brown planthopper, Nilaparvata lugens, by measuring vitellogenin and vitellin levels. Entomol Exp Appl, 139(2): 103-110. |
| [12] | Du B, Zhang W L, Liu B F, Hu J, Wei Z, Shi Z Y, He R F, Zhu L L, Chen R Z, Han B, He G C. 2009. Identification and characterization of Bph14, a gene conferring resistance to brown planthopper in rice. Proc Natl Acad Sci USA, 106(52): 22163-22168. |
| [13] |
Dunoyer P, Schott G, Himber C, Meyer D, Takeda A, Carrington J C, Voinnet O. 2010. Small RNA duplexes function as mobile silencing signals between plant cells. Science, 328: 912-916.
PMID |
| [14] | Fujita D, Kohli A, Horgan F G. 2013. Rice resistance to planthoppers and leafhoppers. Crit Rev Plant Sci, 32(3): 162-191. |
| [15] |
Ge Y F, Han J Y, Zhou G X, Xu Y M, Ding Y, Shi M, Guo C K, Wu G. 2018. Silencing of miR156 confers enhanced resistance to brown planthopper in rice. Planta, 248(4): 813-826.
PMID |
| [16] | Guo J P, Xu C X, Wu D, Zhao Y, Qiu Y F, Wang X X, Ouyang Y D, Cai B D, Liu X, Jing S L, Shangguan X X, Wang H Y, Ma Y H, Hu L, Wu Y, Shi S J, Wang W L, Zhu L L, Xu X, Chen R Z, Feng Y Q, Du B, He G C. 2018. Bph6 encodes an exocyst- localized protein and confers broad resistance to planthoppers in rice. Nat Genet, 50(2): 297-306. |
| [17] | Jena K K, Kim S M. 2010. Current status of brown planthopper (BPH) resistance and genetics. Rice, 3(2/3): 161-171. |
| [18] | John B, Enright A J, Aravin A, Tuschl T, Sander C, Marks D S. 2004. Human microRNA targets. PLoS Biol, 2(11): e363. |
| [19] | Li C D, Wong A Y P, Wang S, Jia Q, Chuang W P, Bendena W G, Tobe S S, Yang S H, Chung G, Chan T F, Lam H M, Bede J C, Hui J H L. 2018. miRNA-mediated interactions in and between plants and insects. Int J Mol Sci, 19(10): 3239. |
| [20] |
Liu Y Q, Wu H, Chen H, Liu Y L, He J, Kang H Y, Sun Z G, Pan G, Wang Q, Hu J L, Zhou F, Zhou K N, Zheng X M, Ren Y L, Chen L M, Wang Y H, Zhao Z G, Lin Q B, Wu F Q, Zhang X, Guo X P, Cheng X N, Jiang L, Wu C Y, Wang H Y, Wan J M. 2015. A gene cluster encoding lectin receptor kinases confers broad-spectrum and durable insect resistance in rice. Nat Biotechnol, 33(3): 301-305.
PMID |
| [21] | Llave C, Xie Z X, Kasschau K D, Carrington J C. 2002. Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science, 297: 2053-2056. |
| [22] |
Loher P, Rigoutsos I. 2012. Interactive exploration of RNA22 microRNA target predictions. Bioinformatics, 28(24): 3322-3323.
PMID |
| [23] | Love M I, Huber W, Anders S. 2014. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol, 15(12): 550. |
| [24] |
Lucas-Barbosa D. 2016. Integrating studies on plant-pollinator and plant-herbivore interactions. Trends Plant Sci, 21(2): 125-133.
PMID |
| [25] | Lv Z Y, Wei Y, Wang D, Zhang C Y, Zen K, Li L M. 2014. Argonaute 2 in cell-secreted microvesicles guides the function of secreted miRNAs in recipient cells. PLoS One, 9(7): e103599. |
| [26] | Mi S J, Cai T, Hu Y G, Chen Y M, Hodges E, Ni F R, Wu L, Li S, Zhou H Y, Long C Z, Chen S, Hannon G J, Qi Y J. 2008. Sorting of small RNAs into Arabidopsis Argonaute complexes is directed by the 5′ terminal nucleotide. Cell, 133(1): 116-127. |
| [27] | Mitsumasu K, Azuma M, Niimi T, Yamashita O, Yaginuma T. 2008. Changes in the expression of soluble and integral-membrane trehalases in the midgut during metamorphosis in Bombyx mori. Zoolog Sci, 25(7): 693-698. |
| [28] |
Nanda S, Yuan S Y, Lai F X, Wang W X, Fu Q, Wan P J. 2020. Identification and analysis of miRNAs in IR56 rice in response to BPH infestations of different virulence levels. Sci Rep, 10(1): 19093.
PMID |
| [29] |
Rhoades M W, Reinhart B J, Lim L P, Burge C B, Bartel B, Bartel D P. 2002. Prediction of plant microRNA targets. Cell, 110(4): 513-520.
PMID |
| [30] |
Tan J Y, Wu Y, Guo J P, Li H M, Zhu L L, Chen R Z, He G C, Du B. 2020. A combined microRNA and transcriptome analyses illuminates the resistance response of rice against brown planthopper. BMC Genomics, 21(1): 144.
PMID |
| [31] | Wu S F, Zeng B, Zheng C, Mu X C, Zhang Y, Hu J, Zhang S, Gao C F, Shen J L. 2018. The evolution of insecticide resistance in the brown planthopper (Nilaparvata lugens Stål) of China in the period 2012-2016. Sci Rep, 8(1): 4586. |
| [32] | Wu Y, Lv W T, Hu L, Rao W W, Zeng Y, Zhu L L, He Y Q, He G C. 2017. Identification and analysis of brown planthopper- responsive microRNAs in resistant and susceptible rice plants. Sci Rep, 7(1): 8712. |
| [33] |
Yang L, Zhang W L. 2016. Genetic and biochemical mechanisms of rice resistance to planthopper. Plant Cell Rep, 35(8): 1559-1572.
PMID |
| [34] |
Zhang B H, Wang Q L, Pan X P. 2007. MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol, 210(2): 279-289.
PMID |
| [35] |
Zhang L, Hou D X, Chen X, Li D H, Zhu L Y, Zhang Y J, Li J, Bian Z, Liang X Y, Cai X, Yin Y, Wang C, Zhang T F, Zhu D H, Zhang D M, Xu J, Chen Q, Ba Y, Liu J, Wang Q, Chen J Q, Wang J, Wang M, Zhang Q P, Zhang J F, Zen K, Zhang C Y. 2012. Exogenous plant MIR168a specifically targets mammalian LDLRAP1: Evidence of cross-kingdom regulation by microRNA. Cell Res, 22(1): 107-126.
PMID |
| [36] |
Zhou G Y, Zhou Y, Chen X. 2017. New insight into inter-kingdom communication: Horizontal transfer of mobile small RNAs. Front Microbiol, 8: 768.
PMID |
| [37] |
Zhou Z, Li X H, Liu J X, Dong L, Chen Q, Liu J L, Kong H H, Zhang Q Y, Qi X, Hou D X, Zhang L, Zhang G Q, Liu Y C, Zhang Y J, Li J, Wang J, Chen X, Wang H, Zhang J F, Chen H L, Zen K, Zhang C Y. 2015. Honeysuckle-encoded atypical microRNA2911 directly targets influenza A viruses. Cell Res, 25(1): 39-49.
PMID |
| [38] | Zhu K G, Liu M H, Fu Z, Zhou Z, Kong Y, Liang H W, Lin Z G, Luo J, Zheng H Q, Wan P, Zhang J F, Zen K, Chen J, Hu F L, Zhang C Y, Ren J, Chen X. 2017. Plant microRNAs in larval food regulate honeybee caste development. PLoS Genet, 13(8): e1006946. |
| [1] | Wang Weixia, Zhu Tingheng, Wan Pinjun, Wei Qi, He Jiachun, Lai Fengxiang, Fu Qiang. Cloning and Functional Analysis of Calcineurin Subunits A and B in Development and Fecundity of Nilaparvata lugens (Stål) [J]. Rice Science, 2022, 29(2): 143-154. |
| [2] | Mamunur Rashid Md, Jahan Mahbuba, Shariful Islam Khandakar. Impact of Nitrogen, Phosphorus and Potassium on Brown Planthopper and Tolerance of Its Host Rice Plants [J]. Rice Science, 2016, 23(3): 119-131. |
| [3] | Singh Sarao Preetinder, Sanmallappa Bentur Jagadaish. Antixenosis and Tolerance of Rice Genotypes Against Brown Planthopper [J]. Rice Science, 2016, 23(2): 96-103. |
| [4] | Shu-hua Liu, Jian Tang, Ju Luo, Bao-jun Yang, Ai-ying Wang, Jin-cai Wu. Cloning and Characterization of karmoisin Homologue Gene (Nlka) in Two Brown Planthopper Strains with Different Eye Colors [J]. Rice Science, 2016, 23(2): 104-110. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||