
Rice Science ›› 2024, Vol. 31 ›› Issue (1): 87-102.DOI: 10.1016/j.rsci.2023.09.002
• Research Papers • Previous Articles Next Articles
Di Dongwei1, Ma Mingkun1,4, Zhang Xiaoyang1, Lu Yufang1, Herbert J. Kronzucker3, Shi Weiming1,2,4( )
)
Received:2023-06-01
Accepted:2023-08-11
Online:2024-01-28
Published:2024-02-06
Contact:
Shi Weiming (Di Dongwei, Ma Mingkun, Zhang Xiaoyang, Lu Yufang, Herbert J. Kronzucker, Shi Weiming. Potential Secretory Transporters and Biosynthetic Precursors of Biological Nitrification Inhibitor 1,9-Decanediol in Rice as Revealed by Transcriptome and Metabolome Analyses[J]. Rice Science, 2024, 31(1): 87-102.
Add to citation manager EndNote|Ris|BibTeX
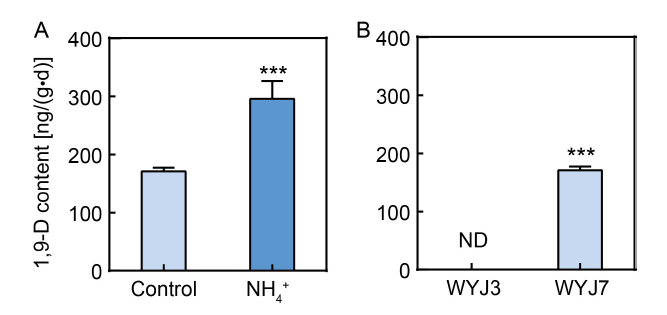
Fig. 1. Influence of NH4+ and variety on 1,9-decanediol (1,9-D) release in rice. A, 1,9-D release from Wuyunjing 7 (WYJ7) roots with or without 1 mmol/L NH4Cl (NH4+) treatment. B, 1,9-D release in Wuyunjing 3 (WYJ3) and WYJ7 grown in control media. Six-week-old seedlings were transferred to new solutions with 1 mmol/L CaCl2 (control) or 1 mmol/L NH4Cl (NH4+ treatment) and grown for another 24 h prior to exudate collection. ND indicates not detected. Data are Mean ± SD with four replications. ***, P < 0.001 (t-test).
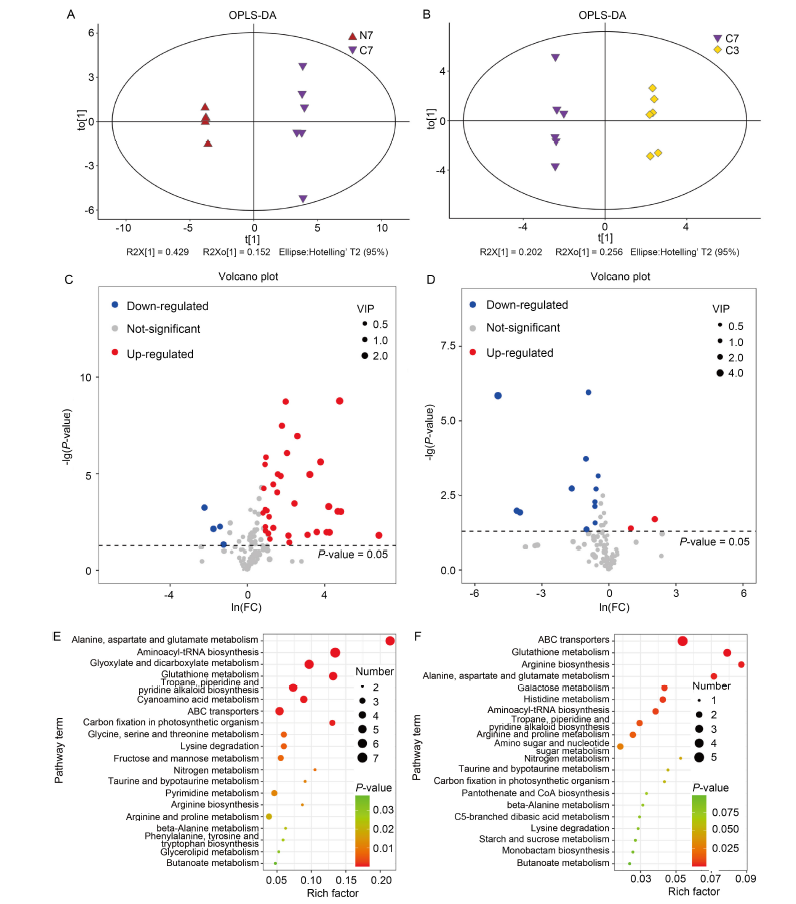
Fig. 2. Analysis of differentially synthesized metabolites (DSMs) based on gas chromatography-mass spectrometer (GC-MS). A and B, Orthogonal partial least squares-discriminant analysis (OPLS-DA) of NH4+-Wuyunjing 7 (N7)/CK-Wuyunjing 7 (C7) (A) and C7/CK- Wuyunjing 3 (C3) (B) groups. C and D, Volcano plots of DSMs in N7/C7 (C) and C7/C3 (D) groups. Each point in the volcano plot indicates an identified metabolite, and the red/blue/grey dots indicate up-/down-/unregulated metabolites. VIP, Variable importance of projection. E and F, Kyoto Encyclopedia of Genes and Genomes analysis of DSMs in N7/C7 (E) and C7/C3 (F) groups. The size and color of the dots indicate the number of DSMs and the significance (P < 0.05), respectively. ABC, ATP-binding cassette.
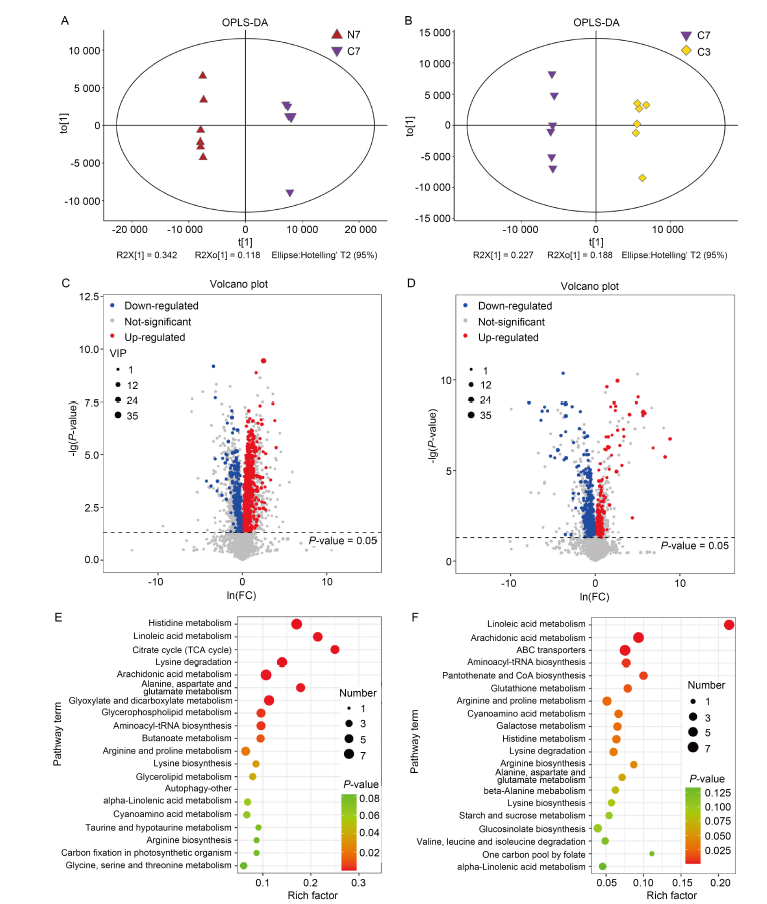
Fig. 3. Analysis of differentially synthesized metabolites (DSMs) based on liquid chromatography-mass spectrometer (LC-MS). A and B, Orthogonal partial least squares-discriminant analysis (OPLS-DA) of NH4+-Wuyunjing 7 (N7)/CK-Wuyunjing 7 (C7) (A) and C7/CK- Wuyunjing 3 (C3) (B) groups. C and D, Volcano plots of DSMs in N7/C7 (C) and C7/C3 (D) groups. Each point in the volcano plot indicates an identified metabolite, and the red/blue/grey dots indicate up-/down-/unregulated metabolites. VIP, Variable importance of projection. E and F, Kyoto Encyclopedia of Genes and Genomes analysis of DSMs in N7/C7 (E) and C7/C3 (F) groups. The size and color of the dots indicate the number of DSMs and the significance (P < 0.05), respectively. ABC, ATP-binding cassette.
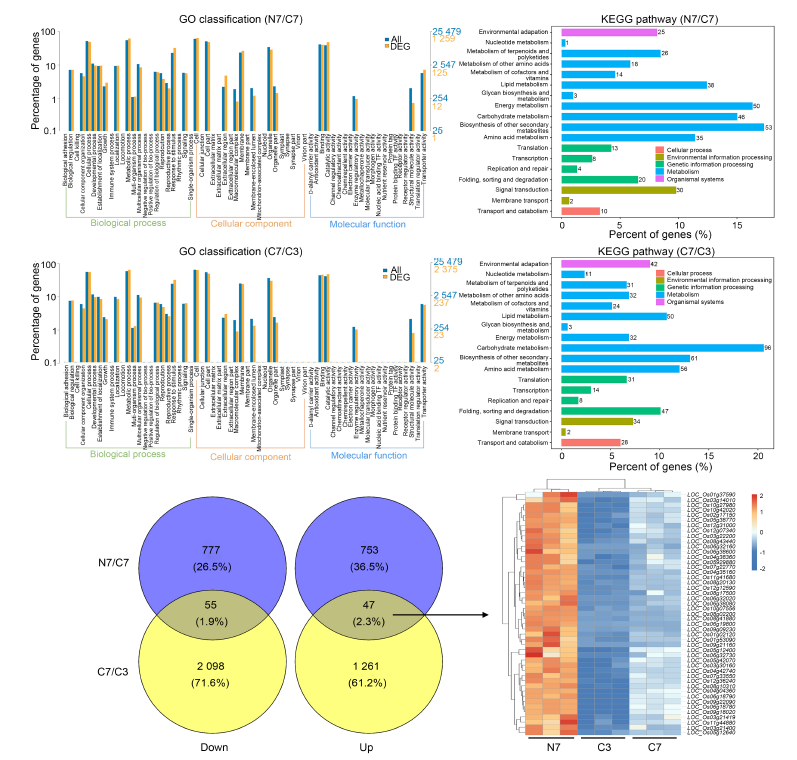
Fig. 4. Analysis of differentially expressed unigenes (DEGs) based on RNA-seq analysis. A and B, Gene Ontology (GO) classification of DEGs in NH4+-Wuyunjing 7 (N7)/CK-Wuyunjing 7 (C7) (A) and C7/CK-Wuyunjing 3 (C3) (B) groups. C and D, Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of DEGs in N7/C7 (C) and of C7/C3 (D) groups. E, Venn diagram showing co-regulated DEGs in N7/C7 and C7/C3 groups. F, Hierarchical clustering heatmap of co-upregulated DEGs in N7/C7 and C7/C3 groups.
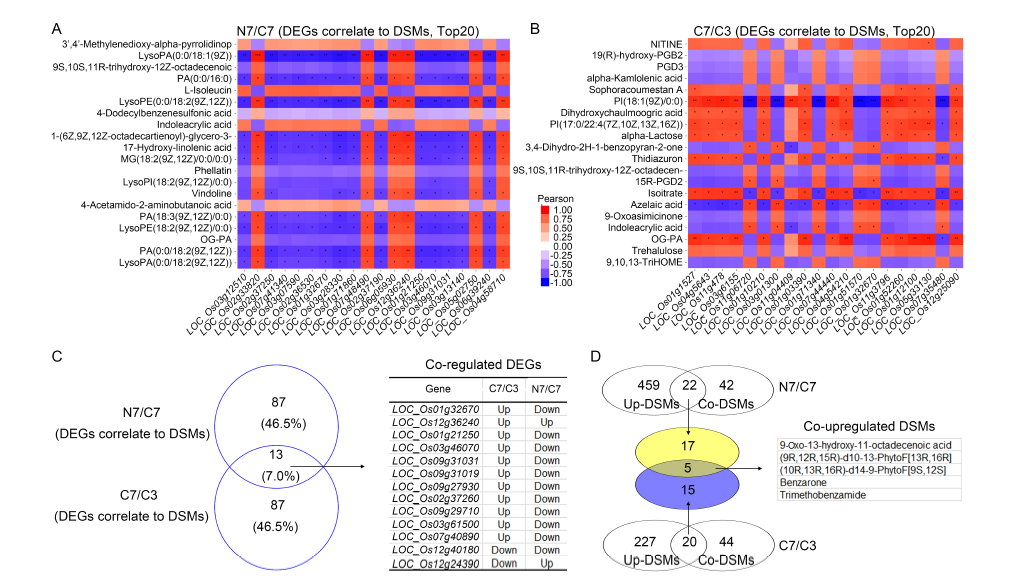
Fig. 5. Identification of co-regulated differentially expressed unigenes (DEGs) and differentially synthesized metabolites (DSMs) via integrated analysis of metabolome and transcriptome. A and B, Correlation analysis of top 20 DEGs and DSMs in NH4+-Wuyunjing 7 (N7)/CK-Wuyunjing 7 (C7) (A) and C7/CK-Wuyunjing 3 (C3) (B) groups. Red/blue represents a positive/negative correlation. *, **, and *** represent 0.05, 0.01, and 0.001 levels, respectively. C, Venn analysis of co-regulated DEGs of top 100 correlation between DEGs and DSMs in N7/C7 and C7/C3 groups. D, Venn analysis of co-upregulated DSMs in N7/C7 and C7/C3 groups.
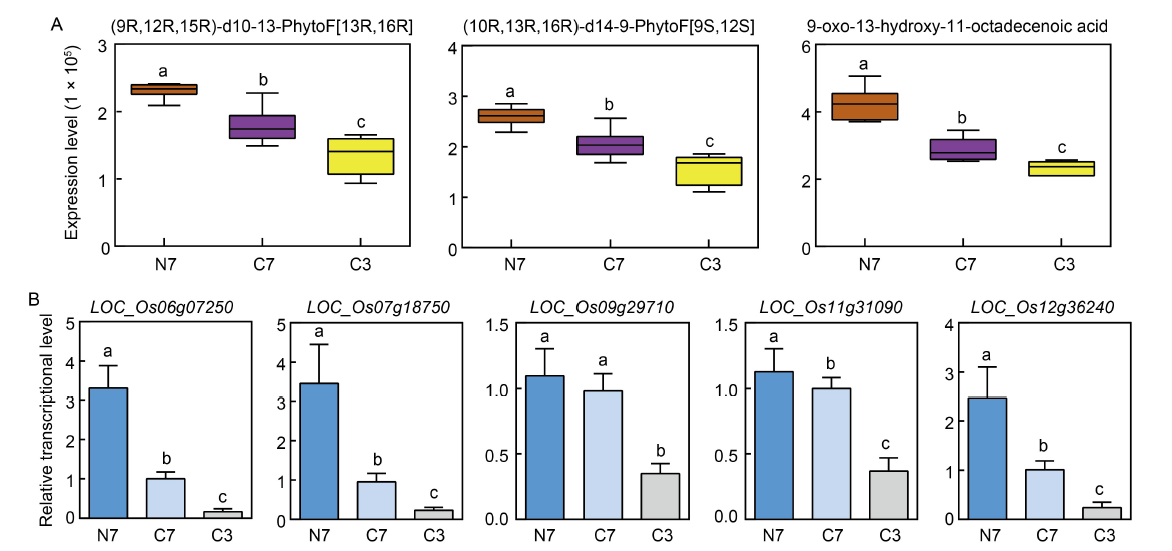
Fig. 6. Identification of possible genes involved in co-upregulated differentially expressed unigenes (DEGs) via qRT-PCR. A, Boxchart of three co-upregulated metabolites. B, qRT-PCR analysis of possible genes associated to DEGs. Six-week-old rice seedlings are transferred to fresh solutions with 1 mmol/L CaCl2 (control) or 1 mmol/L NH4Cl (NH4+ treatment) and grown for another 24 h prior to RNA extraction of roots. All data are normalized relative to OsUBI (LOC_Os03g13170). N7, C7, and C3 represent NH4+-Wuyunjing 7, CK-Wuyunjing 7, and CK-Wuyunjing 3, respectively. Values indicate Mean ± SD of three biological replicates. Error bars with different lowercase letters represent statistically significant differences (P < 0.05, Duncan’s test).
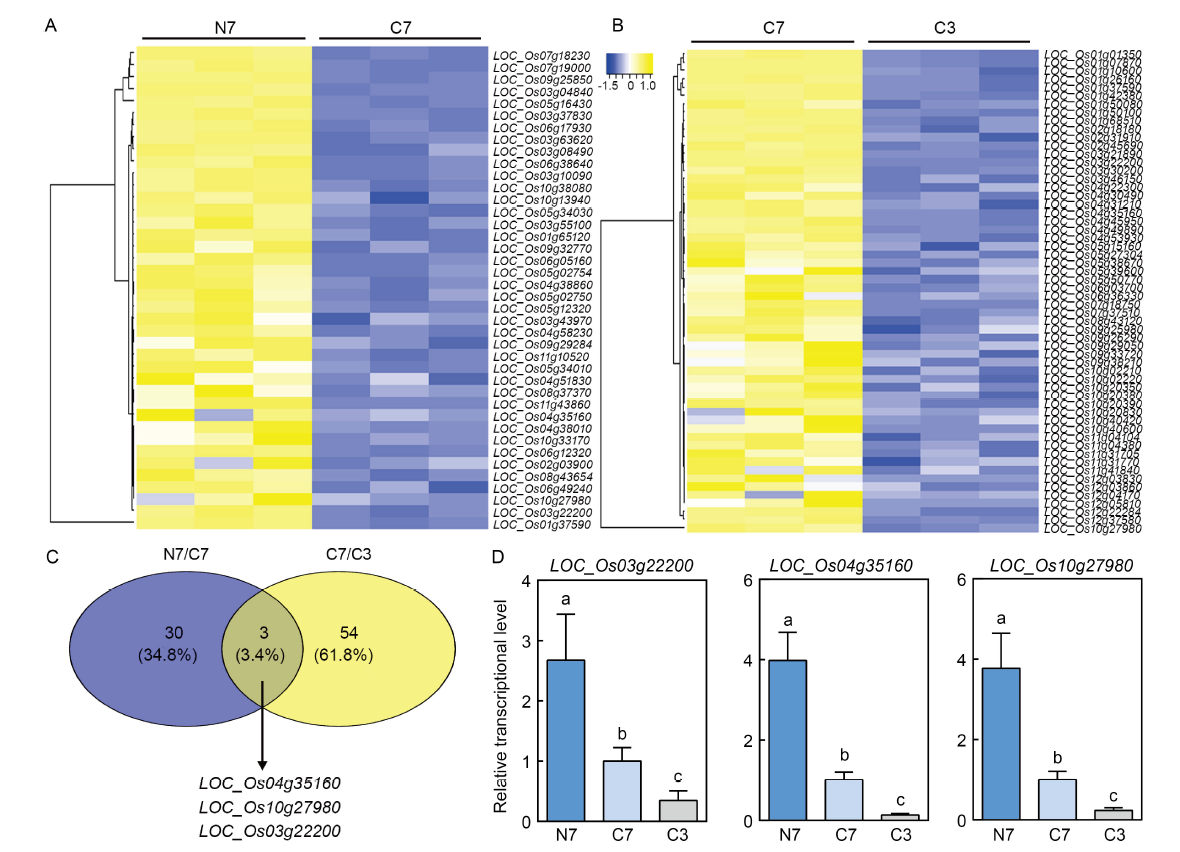
Fig. 7. Identification of possible 1,9-decanediol secretion-related genes. A and B, Hierarchical clustering heatmap of co-upregulated transporter-related differentially expressed unigenes (DEGs) in NH4+-Wuyunjing 7 (N7)/CK-Wuyunjing 7 (C7) (A) and C7/CK-Wuyunjing 3 (C3) (B) groups. C, Venn analysis of co-upregulated transporter-related genes in N7/C7 and C7/C3 groups. D, qRT-PCR analysis of co-upregulated genes. Six-week-old rice seedlings are transferred to fresh solutions with 1 mmol/L CaCl2 (control) or 1 mmol/L NH4Cl (NH4+ treatment) and grown for another 24 h prior to RNA extraction of roots. All data are normalized relative to OsUBI (LOC_Os03g13170). Values indicate Mean ± SD of three biological replicates. Error bars with different lowercase letters represent statistically significant differences (P < 0.05, Duncan’s test).
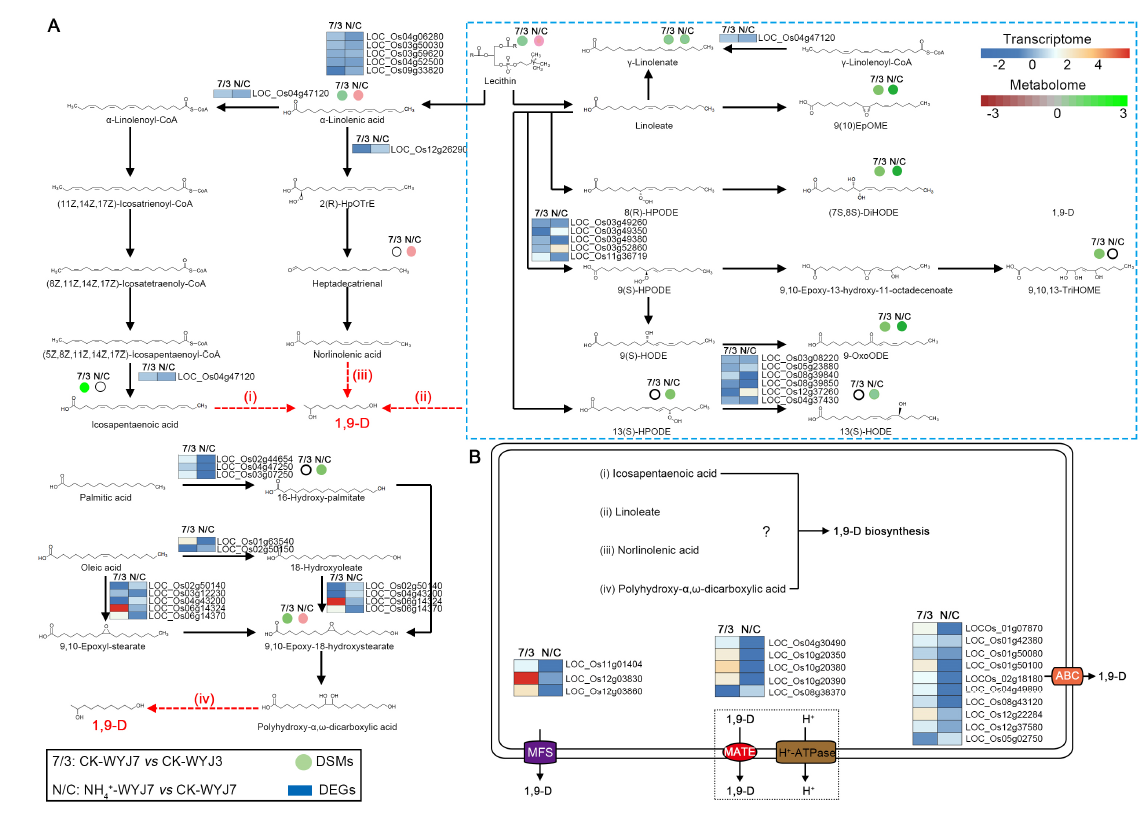
Fig. 8. Potential biosynthesis pathways and transporters responsible for 1,9-decanediol (1,9-D) synthesis and release in rice. A, Potential metabolic pathways involved in 1,9-D biosynthesis. B, Potential transporters responsible for 1,9-D release. ABC, ATP-binding cassette protein family; MATE, Multidrug and toxic compound extrusion family; MFS, Major facilitator superfamily. DSMs, Differentially synthesized metabolites; DEGs, Differentially expressed unigenes.
| [1] | Badri D V, Loyola-Vargas V M, Broeckling C D, De-la-Peña C, Jasinski M, Santelia D, Martinoia E, Sumner L W, Banta L M, Stermitz F, Vivanco J M. 2008. Altered profile of secondary metabolites in the root exudates of Arabidopsis ATP-binding cassette transporter mutants. Plant Physiol, 146(2): 762-771. |
| [2] | Buer C S, Muday G K, Djordjevic M A. 2007. Flavonoids are differentially taken up and transported long distances in Arabidopsis. Plant Physiol, 145(2): 478-490. |
| [3] | Chen Z C, Yamaji N, Motoyama R, Nagamura Y, Ma J F. 2012. Up-regulation of a magnesium transporter gene OsMGT1 is required for conferring aluminum tolerance in rice. Plant Physiol, 159(4): 1624-1633. |
| [4] | Cheng H, Kong W P, Tang T X, Ren K L, Zhang K L, Wei H X, Lin T. 2022. Identification of key gene networks controlling soluble sugar and organic acid metabolism during oriental melon fruit development by integrated analysis of metabolic and transcriptomic analyses. Front Plant Sci, 13: 830517. |
| [5] | Coskun D, Britto D T, Shi W M, Kronzucker H J. 2017a. How plant root exudates shape the nitrogen cycle. Trends Plant Sci, 22(8): 661-673. |
| [6] | Coskun D, Britto D T, Shi W M, Kronzucker H J. 2017b. Nitrogen transformations in modern agriculture and the role of biological nitrification inhibition. Nat Plants, 3: 17074. |
| [7] | Czarnota M A, Paul R N, Weston L A, Duke S O. 2003. Anatomy of sorgoleone-secreting root hairs of Sorghum species. Int J Plant Sci, 164(6): 861-866. |
| [8] | Di D W, Sun L, Zhang X N, Li G J, Kronzucker H J, Shi W M. 2018. Involvement of auxin in the regulation of ammonium tolerance in rice (Oryza sativa L.). Plant Soil, 432(1): 373-387. |
| [9] | Doshi R, McGrath A P, Piñeros M, Szewczyk P, Garza D M, Kochian L V, Chang G. 2017. Functional characterization and discovery of modulators of SbMATE, the agronomically important aluminium tolerance transporter from Sorghum bicolor. Sci Rep, 7: 17996. |
| [10] | Elrys A S, Desoky E S M, Abo El-Maati M F, Elnahal A S, Abdo A I, Raza S, Zhou J B. 2019. Can secondary metabolites extracted from Moringa seeds suppress ammonia oxidizers to increase nitrogen use efficiency and reduce nitrate contamination in potato tubers? Ecotoxicol Environ Saf, 185: 109689. |
| [11] | Elrys A S, Uwiragiye Y, Zhang Y H, Abdel-Fattah M K, Chen Z X, Zhang H M, Meng L, Wang J, Zhu T B, Cheng Y, Zhang J B, Cai Z C, Chang S X, Müller C. 2023. Expanding agroforestry can increase nitrate retention and mitigate the global impact of a leaky nitrogen cycle in croplands. Nat Food, 4(1): 109-121. |
| [12] | Fliegmann J, Schüler G, Boland W, Ebel J, Mithöfer A. 2003. The role of octadecanoids and functional mimics in soybean defense responses. Biol Chem, 384(3): 437-446. |
| [13] | Fu M Y, Yang X, Zheng J R, Wang L, Yang X Y, Tu Y, Ye J B, Zhang W W, Liao Y L, Cheng S Y, Xu F. 2021. Unraveling the regulatory mechanism of color diversity in Camellia japonica petals by integrative transcriptome and metabolome analysis. Front Plant Sci, 12: 685136. |
| [14] | Kaur-Bhambra J, Wardak D L R, Prosser J I, Gubry-Rangin C. 2022. Revisiting plant biological nitrification inhibition efficiency using multiple archaeal and bacterial ammonia-oxidising cultures. Biol Fertil Soils, 58(3): 241-249. |
| [15] | Kronzucker H J, Siddiqi M Y, Glass A D M. 1997. Conifer root discrimination against soil nitrate and the ecology of forest succession. Nature, 385: 59-61. |
| [16] | Kronzucker H J, Siddiqi M Y, Glass A D, Kirk G J. 1999. Nitrate- ammonium synergism in rice: A subcellular flux analysis. Plant Physiol, 119(3): 1041-1046. |
| [17] | Li T L, Wang Z G, Wang C X, Huang J Y, Feng Y F, Shen W S, Zhou M, Yang L Z. 2022. Ammonia volatilization mitigation in crop farming: A review of fertilizer amendment technologies and mechanisms. Chemosphere, 303: 134944. |
| [18] | Liu X Y, Sheng J, Curtiss R. 2011. Fatty acid production in genetically modified cyanobacteria. Proc Natl Acad Sci USA, 108(17): 6899-6904. |
| [19] | Lu X F, Vora H, Khosla C. 2008. Overproduction of free fatty acids in E. coli: Implications for biodiesel production. Metab Eng, 10(6): 333-339. |
| [20] | Lu Y F, Zhang X N, Jiang J F, Kronzucker H J, Shen W S, Shi W M. 2019. Effects of the biological nitrification inhibitor 1,9-decanediol on nitrification and ammonia oxidizers in three agricultural soils. Soil Biol Biochem, 129: 48-59. |
| [21] | Lu Y F, Zhang X N, Ma M K, Zu W J, Kronzucker H J, Shi W M. 2022. Syringic acid from rice as a biological nitrification and urease inhibitor and its synergism with 1,9-decanediol. Biol Fertil Soils, 58(3): 277-289. |
| [22] | Ma M K, Lu Y F, Di D W, Kronzucker H J, Dong G Q, Shi W M. 2023. The nitrification inhibitor 1,9-decanediol from rice roots promotes root growth in Arabidopsis through involvement of ABA and PIN2-mediated auxin signaling. J Plant Physiol, 280: 153891. |
| [23] | Min J, Shi W M. 2018. Nitrogen discharge pathways in vegetable production as non-point sources of pollution and measures to control it. Sci Total Environ, 613/614: 123-130. |
| [24] | Min J, Sun H J, Wang Y, Pan Y F, Kronzucker H J, Zhao D Q, Shi W M. 2021. Mechanical side-deep fertilization mitigates ammonia volatilization and nitrogen runoff and increases profitability in rice production independent of fertilizer type and split ratio. J Clean Prod, 316: 128370. |
| [25] | Nardi P, Laanbroek H J, Nicol G W, Renella G, Cardinale M, Pietramellara G, Weckwerth W, Trinchera A, Ghatak A, Nannipieri P. 2020. Biological nitrification inhibition in the rhizosphere: Determining interactions and impact on microbially mediated processes and potential applications. FEMS Microbiol Rev, 44(6): 874-908. |
| [26] | Nozoye T, Nagasaka S, Kobayashi T, Takahashi M, Sato Y, Sato Y, Uozumi N, Nakanishi H, Nishizawa N K. 2011. Phytosiderophore efflux transporters are crucial for iron acquisition in graminaceous plants. J Biol Chem, 286(7): 5446-5454. |
| [27] | O’Sullivan C A, Fillery I R P, Roper M M, Richards R A. 2016. Identification of several wheat landraces with biological nitrification inhibition capacity. Plant Soil, 404(1): 61-74. |
| [28] | Otaka J, Subbarao G V, Ono H, Yoshihashi T. 2022. Biological nitrification inhibition in maize: Isolation and identification of hydrophobic inhibitors from root exudates. Biol Fertil Soils, 58(3): 251-264. |
| [29] | Reiser S, Somerville C. 1997. Isolation of mutants of Acinetobacter calcoaceticus deficient in wax ester synthesis and complementation of one mutation with a gene encoding a fatty acyl coenzyme A reductase. J Bacteriol, 179(9): 2969-2975. |
| [30] | Ryan P R, Tyerman S D, Sasaki T, Furuichi T, Yamamoto Y, Zhang W H, Delhaize E. 2011. The identification of aluminium-resistance genes provides opportunities for enhancing crop production on acid soils. J Exp Bot, 62(1): 9-20. |
| [31] | Sivaguru M, Liu J P, Kochian L V. 2013. Targeted expression of SbMATE in the root distal transition zone is responsible for sorghum aluminum resistance. Plant J, 76(2): 297-307. |
| [32] | Souza E F C, Rosen C J, Venterea R T. 2021. Co-application of DMPSA and NBPT with urea mitigates both nitrous oxide emissions and nitrate leaching during irrigated potato production. Environ Pollut, 284: 117124. |
| [33] | Steen E J, Kang Y S, Bokinsky G, Hu Z H, Schirmer A, McClure A, Del Cardayre S B, Keasling J D. 2010. Microbial production of fatty-acid-derived fuels and chemicals from plant biomass. Nature, 463: 559-562. |
| [34] | Streubel J, Pesce C, Hutin M, Koebnik R, Boch J, Szurek B. 2013. Five phylogenetically close rice SWEET genes confer TAL effector-mediated susceptibility to Xanthomonas oryzae pv. oryzae. New Phytol, 200(3): 808-819. |
| [35] | Subbarao G V, Ito O, Sahrawat K L, Berry W L, Nakahara K, Ishikawa T, Watanabe T, Suenaga K, Rondon M, Rao I M. 2006. Scope and strategies for regulation of nitrification in agricultural systems: Challenges and opportunities. Crit Rev Plant Sci, 25(4): 303-335. |
| [36] | Subbarao G V, Tomohiro B, Masahiro K, Osamu I, Samejima H, Wang H Y, Pearse S J, Gopalakrishnan S, Nakahara K, Zakir Hossain A K M, Tsujimoto H, Berry W L. 2007. Can biological nitrification inhibition (BNI) genes from perennial Leymus racemosus (Triticeae) combat nitrification in wheat farming? Plant Soil, 299(1): 55-64. |
| [37] | Subbarao G V, Nakahara K, Ishikawa T, Yoshihashi T, Ito O, Ono H, Ohnishi-Kameyama M, Yoshida M, Kawano N, Berry W L. 2008. Free fatty acids from the pasture grass Brachiaria humidicola and one of their methyl esters as inhibitors of nitrification. Plant Soil, 313(1): 89-99. |
| [38] | Subbarao G V, Nakahara K, Hurtado M P, Ono H, Moreta D E, Salcedo A F, Yoshihashi A T, Ishikawa T, Ishitani M, Ohnishi- Kameyama M, Yoshida M, Rondon M, Rao I M, Lascano C E, Berry W L, Ito O. 2009. Evidence for biological nitrification inhibition in Brachiaria pastures. Proc Natl Acad Sci USA, 106: 17302-17307. |
| [39] | Subbarao G V, Nakahara K, Ishikawa T, Ono H, Yoshida M, Yoshihashi T, Zhu Y Y, Zakir H A K M, Deshpande S P, Hash C T, Sahrawat K L. 2013a. Biological nitrification inhibition (BNI) activity in sorghum and its characterization. Plant Soil, 366(1): 243-259. |
| [40] | Subbarao G V, Rao I M, Nakahara K, Sahrawat K L, Ando Y, Kawashima T. 2013b. Potential for biological nitrification inhibition to reduce nitrification and N2O emissions in pasture crop- livestock systems. Animal, 7: 322-332. |
| [41] | Subbarao G V, Sahrawat K L, Nakahara K, Rao I M, Ishitani M, Hash C T, Kishii M, Bonnett D G, Berry W L, Lata J C. 2013c. A paradigm shift towards low-nitrifying production systems: The role of biological nitrification inhibition (BNI). Ann Bot, 112(2): 297-316. |
| [42] | Subbarao G V, Yoshihashi T, Worthington M, Nakahara K, Ando Y, Sahrawat K L, Rao I M, Lata J C, Kishii M, Braun H J. 2015. Suppression of soil nitrification by plants. Plant Sci, 233: 155-164. |
| [43] | Subbarao G V, Kishii M, Bozal-Leorri A, Ortiz-Monasterio I, Gao X, Ibba M I, Karwat H, Gonzalez-Moro M B, Gonzalez-Murua C, Yoshihashi T, Tobita S, Kommerell V, Braun H J, Iwanaga M. 2021. Enlisting wild grass genes to combat nitrification in wheat farming: A nature-based solution. Proc Natl Acad Sci USA, 118(35): e2106595118. |
| [44] | Sun L, Lu Y F, Yu F W, Kronzucker H J, Shi W M. 2016. Biological nitrification inhibition by rice root exudates and its relationship with nitrogen-use efficiency. New Phytol, 212(3): 646-656. |
| [45] | Sun L, Di D W, Li G J, Kronzucker H J, Shi W M. 2017. Spatio- temporal dynamics in global rice gene expression (Oryza sativa L.) in response to high ammonium stress. J Plant Physiol, 212: 94-104. |
| [46] | Sun L, Di D W, Li G J, Li Y L, Kronzucker H J, Shi W M. 2020. Transcriptome analysis of rice (Oryza sativa L.) in response to ammonium resupply reveals the involvement of phytohormone signaling and the transcription factor OsJAZ9 in reprogramming of nitrogen uptake and metabolism. J Plant Physiol, 246/247: 153137. |
| [47] | Vioque J, Kolattukudy P E. 1997. Resolution and purification of an aldehyde-generating and an alcohol-generating fatty acyl-CoA reductase from pea leaves (Pisum sativumL.). Arch Biochem Biophys, 340(1): 64-72. |
| [48] | Wang M, Qiao J Y, Yu C L, Chen H, Sun C D, Huang L Z, Li C Y, Geisler M, Qian Q, Jiang D A, Qi Y H. 2019. The auxin influx carrier, OsAUX3, regulates rice root development and responses to aluminium stress. Plant Cell Environ, 42(4): 1125-1138. |
| [49] | Wang R, Ren C X, Dong S, Chen C, Xian B, Wu Q H, Wang J, Pei J, Chen J. 2021. Integrated metabolomics and transcriptome analysis of flavonoid biosynthesis in safflower (Carthamus tinctorius L.) with different colors. Front Plant Sci, 12: 712038. |
| [50] | Wang X, Bai J H, Wang C, Xie T, Wang W, Wang D W, Zhang G L. 2023. Two newly-identified biological nitrification inhibitors in Suaeda salsa: Synthetic pathways and influencing mechanisms. Chem Eng J, 454: 140172. |
| [51] | Weston L A, Ryan P R, Watt M. 2012. Mechanisms for cellular transport and release of allelochemicals from plant roots into the rhizosphere. J Exp Bot, 63(9): 3445-3454. |
| [52] | Woodward E E, Hladik M L, Kolpin D W. 2016. Nitrapyrin in streams: The first study documenting off-field transport of a nitrogen stabilizer compound. Environ Sci Technol Lett, 3(11): 387-392. |
| [53] | Ye R G, Wu Y R, Gao Z Y, Chen H, Jia L X, Li D M, Li X G, Qian Q, Qi Y H. 2021. Primary root and root hair development regulation by OsAUX4 and its participation in the phosphate starvation response. J Integr Plant Biol, 63(8): 1555-1567. |
| [54] | Yu C L, Sun C D, Shen C J, Wang S K, Liu F, Liu Y, Chen Y L, Li C Y, Qian Q, Aryal B, Geisler M, Jiang D A, Qi Y H. 2015. The auxin transporter, OsAUX1, is involved in primary root and root hair elongation and in Cd stress responses in rice (Oryza sativa L.). Plant J, 83(5): 818-830. |
| [55] | Zakir H A K M, Subbarao G V, Pearse S J, Gopalakrishnan S, Ito O, Ishikawa T, Kawano N, Nakahara K, Yoshihashi T, Ono H, Yoshida M. 2008. Detection, isolation and characterization of a root-exuded compound, methyl 3-(4-hydroxyphenyl) propionate, responsible for biological nitrification inhibition by sorghum (Sorghum bicolor). New Phytol, 180(2): 442-451. |
| [56] | Zhang M, Fan C H, Li Q L, Li B, Zhu Y Y, Xiong Z Q. 2015. A 2-yr field assessment of the effects of chemical and biological nitrification inhibitors on nitrous oxide emissions and nitrogen use efficiency in an intensively managed vegetable cropping system. Agric Ecosyst Environ, 201: 43-50. |
| [57] | Zhang M X, Zeng H Q, Afzal M R, Gao X, Li Y X, Subbarao G V, Zhu Y Y. 2022. BNI-release mechanisms in plant root systems: Current status of understanding. Biol Fertil Soils, 58(3): 225-233. |
| [58] | Zhang X N, Lu Y F, Yang T, Kronzucker H J, Shi W M. 2019. Factors influencing the release of the biological nitrification inhibitor 1,9-decanediol from rice (Oryza sativa L.) roots. Plant Soil, 436(1/2): 253-265. |
| [59] | Zhang Y T, Yang L W, Yang J J, Hu H L, Wei G Q, Cui J B, Xu J. 2022. Transcriptome and metabolome analyses reveal differences in terpenoid and flavonoid biosynthesis in Cryptomeria fortunei needles across different seasons. Front Plant Sci, 13: 862746. |
| [60] | Zhao H M, Ma T F, Wang X, Deng Y T, Ma H L, Zhang R S, Zhao J. 2015. OsAUX1 controls lateral root initiation in rice (Oryza sativa L.). Plant Cell Environ, 38(11): 2208-2222. |
| [61] | Zhao J, Dixon R A. 2009. MATE transporters facilitate vacuolar uptake of epicatechin 3ʹ-O-glucoside for proanthocyanidin biosynthesis in Medicago truncatula and Arabidopsis. Plant Cell, 21(8): 2323-2340. |
| [62] | Zheng Y N, Li L L, Liu Q, Yang J M, Wang X W, Liu W, Xu X, Liu H, Zhao G, Xian M. 2012. Optimization of fatty alcohol biosynthesis pathway for selectively enhanced production of C12/14 and C16/18 fatty alcohols in engineered Escherichia coli. Microb Cell Fact, 11: 65. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||