
Rice Science ›› 2019, Vol. 26 ›› Issue (2): 118-124.DOI: 10.1016/j.rsci.2018.04.007
收稿日期:2018-02-26
接受日期:2018-04-27
出版日期:2019-03-04
发布日期:2018-12-18
. [J]. Rice Science, 2019, 26(2): 118-124.
| Primer | Sequence (5′-3′) | Purpose |
| T1-F | GGCACACGTCCCAGCTCGCCACCT | Vector construction |
| T1-R | AAACAGGTGGCGAGCTGGGACGTG | Vector construction |
| T2-F | GCCGTCCGTCGTCGTGTACGCCAC | Vector construction |
| T2-R | AAACGTGGCGTACACGACGACGGA | Vector construction |
| Wx-F | TAGCCACCCAAGAAACT | Amplification of the fragments across sgRNAs |
| Wx-R | CTACCTCAGCCACAACG | Amplification of the fragments across sgRNAs |
| hyg283-F | TCCGGAAGTGCTTGACATT | Vector deletion (HPTII) |
| hyg283-R | GTCGTCCATCACAGTTTGC | Vector deletion (HPTII) |
| Cas9-F | TTCGACCAGTCCAAGAACGG | Vector deletion (Cas9) ( |
| Cas9-R | CTTGACCTTGGTGAGCTCGT | Vector deletion (Cas9) ( |
Table 1 Primers used in this study
| Primer | Sequence (5′-3′) | Purpose |
| T1-F | GGCACACGTCCCAGCTCGCCACCT | Vector construction |
| T1-R | AAACAGGTGGCGAGCTGGGACGTG | Vector construction |
| T2-F | GCCGTCCGTCGTCGTGTACGCCAC | Vector construction |
| T2-R | AAACGTGGCGTACACGACGACGGA | Vector construction |
| Wx-F | TAGCCACCCAAGAAACT | Amplification of the fragments across sgRNAs |
| Wx-R | CTACCTCAGCCACAACG | Amplification of the fragments across sgRNAs |
| hyg283-F | TCCGGAAGTGCTTGACATT | Vector deletion (HPTII) |
| hyg283-R | GTCGTCCATCACAGTTTGC | Vector deletion (HPTII) |
| Cas9-F | TTCGACCAGTCCAAGAACGG | Vector deletion (Cas9) ( |
| Cas9-R | CTTGACCTTGGTGAGCTCGT | Vector deletion (Cas9) ( |
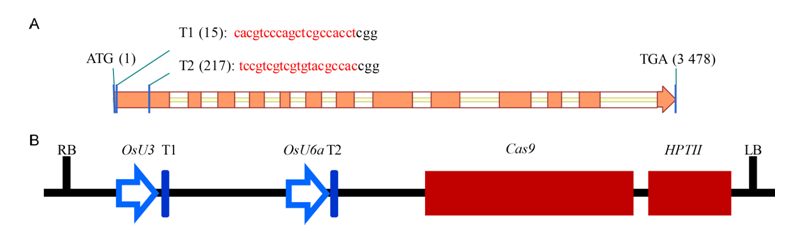
Fig. 1 Schematic diagrams of the Wx and CRISPR/Cas9 vector.A, Overview of the Wx gene structure and target sites for sgRNAs (T1 and T2). Exons and introns in Wx gene are depicted by orange rectangles and white rectangles, respectively. The target sequences of sgRNAs are shown at the top of the gene structure in red. B, The structure of the Wx gene editing system. T1 and T2 are driven by U3 and U6a promoters, respectively. RB, Right border; LB, Left border; Cas9, CRISPR associated protein 9 gene; HPTII, Hygromycin-resistance gene.
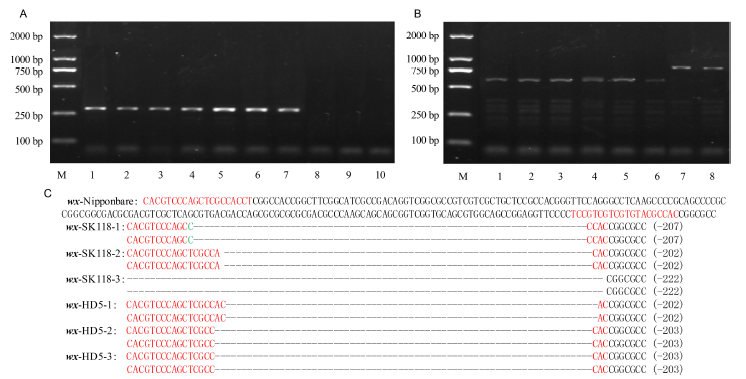
Fig. 2 CRISPR/Cas9-induced mutations in the Wx gene.A, PCR analysis of HPTII in T0 lines (part of the picture). M, DNA marker; 1, wx-SK118-1; 2, wx-SK118-2; 3, wx-SK118-3; 4, wx-HD5-1; 5, wx-HD5-2; 6, wx-HD5-3; 7, Positive control of pYLCRISPR/Cas9-MH-Wx; 8, Negative control of SK118; 9, Negative control of HD5; 10, Negative control of ddH2O. B, Polymerase chain reaction assays of Wx in T0 lines. M, DNA marker; 1, wx-SK118-1; 2, wx-SK118-2; 3, wx-SK118-3; 4, wx-HD5-1; 5, wx-HD5-2; 6, wx-HD5-3; 7, Control of SK118; 8, Control of HD5. C, Sequencing results of T0 mutant lines. sgRNA is shown in red letter; Insertion sequence is shown in green letter; Deletion sequence is shown by dashed line.
| Line | No. of examined plants | No. of transgene-free plants |
| wx-SK118-1 | 10 | 3 |
| wx-SK118-2 | 23 | 5 |
| wx-SK118-3 | 85 | 10 |
| wx-HD5-1 | 53 | 6 |
| wx-HD5-2 | 135 | 17 |
| wx-HD5-3 | 47 | 13 |
Table 2 Segregation of transgene-free rice in T1 generation.
| Line | No. of examined plants | No. of transgene-free plants |
| wx-SK118-1 | 10 | 3 |
| wx-SK118-2 | 23 | 5 |
| wx-SK118-3 | 85 | 10 |
| wx-HD5-1 | 53 | 6 |
| wx-HD5-2 | 135 | 17 |
| wx-HD5-3 | 47 | 13 |
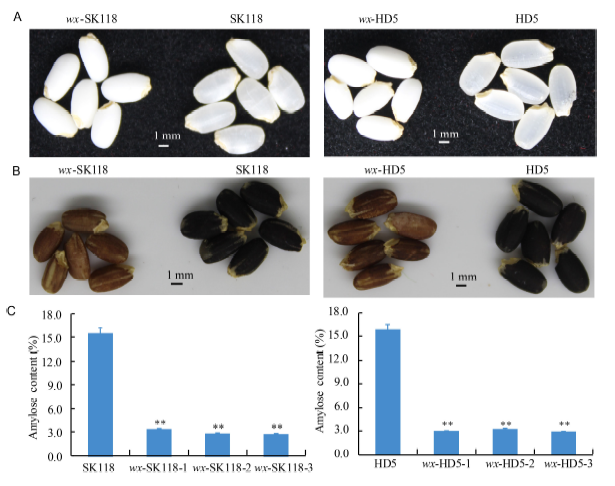
Fig. 3 Phenotype of the Wx mutants.A, Intact polished rice. B, Grains stained with iodine solution. C, Amylose content of Wx mutants. Data represent Mean ± SD (n = 3). **, Significant at the 0.01 level by the t-test.
| [1] | Baysal C, Bortesi L, Zhu C, Farre G, Schillberg S, Christou P.2016. CRISPR/Cas9 activity in the rice OsBEIIb gene does not induce off-target effects in the closely related paralog OsBEIIa. Mol Breeding, 36: 108. |
| [2] | Cai J W, Man J M, Huang J, Liu Q Q, Wei W X, Wei C X.2015. Relationship between structure and functional properties of normal rice starches with different amylose contents.Carbohyd Polym, 125: 35-44. |
| [3] | Chen M H, Bergman C, Pinson S, Fjellstrom R.2008. Waxy gene haplotypes: Associations with apparent amylose content and the effect by the environment in an international rice germplasm collection.J Cereal Sci, 47(3): 536-545. |
| [4] | Cho S W, Kim S, Kim J M, Kim JS.2013. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease.Nat Biotechnol, 31: 230-232. |
| [5] | Fasahat P, Rahman S, Ratnam W.2014. Genetic controls on starch amylose content in wheat and rice grains.J Genet, 93(1): 279-292. |
| [6] | Ishizaki T.2016. CRISPR/Cas9 in rice can induce new mutations in later generations, leading to chimerism and unpredicted segregation of the targeted mutation.Mol Breeding, 36(12): 165. |
| [7] | Juliano B O, Perez C M, Blakeney A B, Castillo D T, Kongseree N, Laignelet B, Lapis E T, Murty V V S, Paule C M, Webb B D.1981. International cooperative testing on the amylose content of milled rice.Starch-Stärke, 33(5): 157-162. |
| [8] | Kasem S, Waters D L E, Rice N F, Shapter F M, Henry R J.2011. The endosperm morphology of rice and its wild relatives as observed by scanning electron microscopy.Rice, 4(1): 12-20. |
| [9] | Kawagoe Y, Kubo A, Satoh H, Takaiwa F, Nakamura Y.2005. Roles of isoamylase and ADP-glucose pyrophosphorylase in starch granule synthesis in rice endosperm.Plant J, 42: 164-174. |
| [10] | Kiswara G, Lee J H, Hur Y J, Cho J H, Lee J Y, Kim S Y, Sohn Y B, Song Y C, Nam M H, Yun B W.2014. Genetic analysis and molecular mapping of low amylose genedu12(t) in rice, 127(1): 51-57. |
| [11] | Li Q L, Zhang D B, Chen M J, Liang W Q, Wei J J, Qi Y P, Yuan Z.2016. Development of japonica photo-sensitive genic male sterile rice lines by editing carbon starved anther using CRISPR/Cas9. J Genet Genom, 43(6): 415-419. |
| [12] | Li X F, Zhou W J, Ren Y K, Tian X J, Lv T X, Wang Z Y, Fang J, Chu C C, Yang J, Bu Q Y.2017. High-efficiency breeding of early-maturing rice cultivars via CRISPR/Cas9-mediated genome editing.J Genet Genom, 44(3): 175-178. |
| [13] | Lu G, Moriyama E N.2004. Vector NTI, a balanced all-in-one sequence analysis suite.Brief Bioinform, 5(4): 378-388. |
| [14] | Ma X L, Zhang Q Y, Zhu Q L, Liu W, Chen Y, Qiu R, Wang B, Yang Z F, Li H Y, Lin Y R, Xie Y Y, Shen R X, Chen S F, Wang Z, Chen Y L, Guo J X, Chen L T, Zhao X C, Dong Z C, Liu Y G.2015. A robust CRISPR/Cas9 system for convenient, high- efficiency multiplex genome editing in monocot and dicot plants.Mol Plant, 8(8): 1274-1284. |
| [15] | Mikami M, Toki S, Endo M.2015. Comparison of CRISPR/Cas9 expression constructs for efficient targeted mutagenesis in rice.Plant Mol Biol, 88(6): 561-572. |
| [16] | Murray M G, Thompson W F.1980. Rapid isolation of high molecular weight plant DNA.Nucleic Acids Res, 8(19): 4321-4325. |
| [17] | Nieves-cordones M, Mohamed S, Tanoi K, Kobayashi N I, Takagi K, Vernet A, Guiderdoni E, Périn C, Sentenac H, Véry A A.2017. Production of low-Cs(+) rice plants by inactivation of the K(+) transporter OsHAK1 with the CRISPR-Cas system.Plant J, 92(1): 43-56. |
| [18] | Peng B, Sun Y F, Pang R H, Li H L, Song X H, Yuan H Y, Zhang S H, Zhou Q Y, Li Q R, Li D, Song S Z.2016. Study on chalkiness character and endosperm structure of rice grain in different japonica varieties. Acta Agric Zhejiang, 28(11): 1803-1811. (in Chinese with English abstract) |
| [19] | Shahid S, Begum R, Razzaque S, Jesmin, Seraj Z I.2016. Variability in amylose content of Bangladeshi rice cultivars due to unique SNPs in Waxy allele. J Cereal Sci, 71: 1-9. |
| [20] | Shen L, Hua Y F, Fu Y P, Jian L, Liu Q, Jiao X Z, Xin G W, Wang J J, Wang X C, Yan C J, Wang K J.2017. Rapid generation of genetic diversity by multiplex CRISPR/Cas9 genome editing in rice.Sci China Life Sci, 60(5): 506-515. |
| [21] | Sun Y W, Jiao G A, Liu Z P, Zhang X, Li J Y, Guo X P, Du W M, Du J L, Francis F, Zhao Y D, Xia L Q.2017. Generation of high- amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes.Front Plant Sci, 8: 298. |
| [22] | Tang L, Mao B G, Li Y K, Lv Q M, Zhang L P, Chen C Y, He H J, Wang W P, Zeng X F, Shao Y, Pan Y L, Hu Y Y, Peng Y, Fu X Q, Li H Q, Xia S T, Zhao B R.2017. Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep, 7(1): 14438. |
| [23] | Toru T, Naoko F.2017. Thermal and rheological characteristics of mutant rice starches with widespread variation of amylose content and amylopectin structure.Food Hydrocolloid, 62: 83-93. |
| [24] | Wamnugu P, Ndjiondjop M N, Furtado A, Henry R.2017. Sequencing of bulks of segregants allows dissection of genetic control of amylose content in rice.Plant Biotechnol J, 16: 100-110. |
| [25] | Wang B K, Zhang H, Hong R K, Zhang J W, Yang R, Luo Q, Zeng Q C.2018. Wx gene editing via CRISPR/Cas9 system in rice.Chin J Rice Sci, 32(1): 35-42. (in Chinese with English abstract) |
| [26] | Wang F J, Wang C L, Liu P Q, Lei C L, Hao W, Gao Y, Liu Y G, Zhao K J.2016. Enhanced rice blast resistance by CRISPR/ Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS One, 11(4): e0154027. |
| [27] | Wang F Q, Fan F J, Li W Q, Zhu J Y, Wang J, Zhong W G, Yang J.2016. Knock out efficiency analysis of Pi21 gene using CRISPR/Cas9 in rice. Chin J Rice Sci, 30(5): 469-478. (in Chinese with English abstract) |
| [28] | Wang K, Hasjim J, Wu A C, Li E, Henry R J, Gilbert R G.2015. Roles of GBSSI and SSIIa in determining amylose fine structure. Carbohyd Polym, 127: 264-274. |
| [29] | Wang M G, Mao Y F, Lu Y M, Tao X P, Zhu J K.2017a. Multiplex gene editing in rice using the CRISPR-Cpf1 system.Mol Plant, 10(7): 1011-1013. |
| [30] | Wang Q X, Zhao H, Jiang J P, Xu J Y, Xie W B, Fu X K, Liu C, He Y Q, Wang G W.2017b. Genetic architecture of natural variation in rice nonphotochemical quenching capacity revealed by genome-wide association study.Front Plant Sci, 8: 1773. |
| [31] | Wang Y, Geng L Z, Yuan M L, Wei J, Jin C, Li M, Yu K, Zhang Y, Jin H B, Wang E, Chai Z J, Fu X D, Li X G.2017c. Deletion of a target gene in indica rice via CRISPR/Cas9. Plant Cell Rep, 36(8): 1333-1343. |
| [32] | Wu Y P, Pu C H, Lin H Y, Huang H Y, Huang Y C, Hong C Y, Chang M C, Lin Y R.2015. Three novel alleles of floury endosperm2 (FLO2) confer dull grains with low amylose content in rice. Plant Sci, 233: 44-52. |
| [33] | Xie Y Y, Niu B X, Long Y M, Li G S, Tang J T, Zhang Y L, Ren D, Liu Y G, Chen L T.2017. Suppression or knockout of SaF/SaM overcomes the Sa-mediated hybrid male sterility in rice. J Integr Plant Biol, 59(9): 669-679. |
| [34] | Xu R F, Yang Y C, Qin R Y, Li H, Qiu C H, Li L, Wei P C, Yang J B.2016. Rapid improvement of grain weight via highly efficient CRISPR/Cas9-mediated multiplex genome editing in rice.J Genet Genomics, 43(8): 529-532. |
| [35] | Yang X H, Nong B X, Xia X Z, Zhang Z Q, Zeng Y, Liu K Q, Deng G F, Li D T.2016. Rapid identification of a new gene influencing low amylose content in rice landraces (Oryza sativa L.) using genome-wide association study with specific-locus amplified fragment sequencing. Genome, 60(6): 465-472. |
| [36] | Yin X J, Biswal A K, Dionora J, Perdigon K M, Balahadia C P, Mazumdar S, Chater C, Lin H C, Coe R A, Kretzschmar T, Guick J E, Quick P W, Bandyopadhyay A.2017. CRISPR-Cas9 and CRISPR-Cpf1 mediated targeting of a stomatal developmental gene EPFL9 in rice. Plant Cell Rep, 36(5): 745-757. |
| [37] | Zhang H, Xu H, Feng M J, Zhu Y.2018. Suppression of OsMADS7 in rice endosperm stabilizes amylose content under high temperature stress. Plant Biotechnol J, 16(1): 18-26. |
| [38] | Zhang J S, Zhang H, Botella J R, Zhu J K.2018. Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Integr Plant Biol, 60(5): 369-375. |
| [39] | Zheng X L, Yang S X, Zhang D W, Zhong Z H, Tang X, Deng K J, Zhou J P, Qi Y P, Zhang Y.2016. Effective screen of CRISPR/Cas9-induced mutants in rice by single-strand conformation polymorphism.Plant Cell Rep, 35(7): 1545-1554. |
| [40] | Zhou H, He M, Li J, Chen L, Huang Z F, Zheng S Y, Zhu L Y, Ni E, Jiang D G, Zhao B R, Zhuang C X.2016. Development of commercial thermo-sensitive genic male sterile rice accelerates hybrid rice breeding using the CRISPR/Cas9-mediatedTMS5 editing system. Sci Rep, 6: 37395. |
| [1] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages[J]. Rice Science, 2023, 30(6): 7-. |
| [2] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage[J]. Rice Science, 2023, 30(6): 12-. |
| [3] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis[J]. Rice Science, 2023, 30(6): 8-. |
| [4] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines[J]. Rice Science, 2023, 30(6): 6-. |
| [5] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice[J]. Rice Science, 2023, 30(6): 4-. |
| [6] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications[J]. Rice Science, 2023, 30(6): 9-. |
| [7] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage[J]. Rice Science, 2023, 30(6): 14-. |
| [8] | LIU Tingting, ZOU Jinpeng, YANG Xi, WANG Kejian, RAO Yuchun, WANG Chun. Development and Application of Prime Editing in Plants[J]. Rice Science, 2023, 30(6): 3-. |
| [9] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance[J]. Rice Science, 2023, 30(6): 10-. |
| [10] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review[J]. Rice Science, 2023, 30(6): 5-. |
| [11] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats[J]. Rice Science, 2023, 30(6): 11-. |
| [12] | . [J]. Rice Science, 2021, 28(3): 217-232. |
| [13] | . [J]. Rice Science, 2019, 26(2): 77-87. |
| [14] | . [J]. Rice Science, 2019, 26(2): 109-117. |
| [15] | . [J]. Rice Science, 2019, 26(2): 88-97. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||