
Rice Science ›› 2021, Vol. 28 ›› Issue (2): 109-113.DOI: 10.1016/j.rsci.2020.09.010
• Letter • Next Articles
Fangping Li, Yanhao Gao, Bingqi Wu, Qingpei Cai, Pengling Zhan, Weifeng Yang, Wanxuan Shi, Xiaohua Li, Zifeng Yang, Quanya Tan, Xin Luan, Guiquan Zhang, Shaokui Wang( )
)
Received:2020-08-11
Accepted:2020-09-18
Online:2021-03-28
Published:2021-03-28
Fangping Li, Yanhao Gao, Bingqi Wu, Qingpei Cai, Pengling Zhan, Weifeng Yang, Wanxuan Shi, Xiaohua Li, Zifeng Yang, Quanya Tan, Xin Luan, Guiquan Zhang, Shaokui Wang. High-Quality de novo Genome Assembly of Huajingxian 74, a Receptor Parent of Single Segment Substitution Lines[J]. Rice Science, 2021, 28(2): 109-113.
Add to citation manager EndNote|Ris|BibTeX
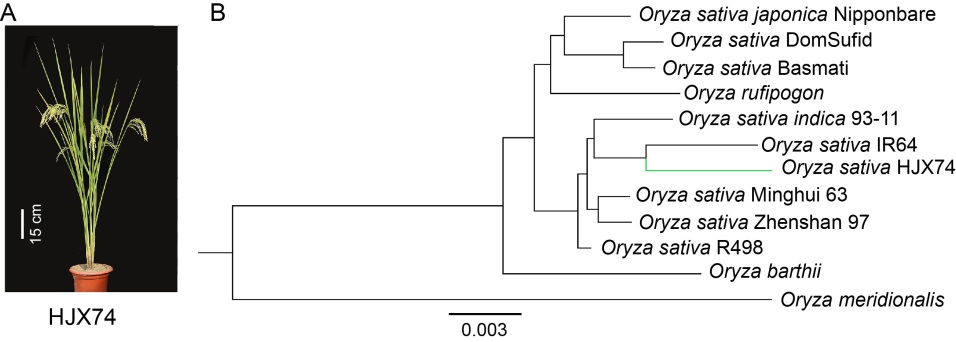
Fig. 1. Phenotype (A) and phylogeney (B) of HJX74 (Huajingxian 74).Phylogenetic tree constructed by the maximum- likelihood method using coding sequences of single copy lineal homologous genes (the genes were showed in Table S4). Totally 12 species or varieties were used for alignment, 9 of them are cultivated rice (Nipponbare, 93-11, R498, Zhenshan 97, IR64, Minghui 63, Basmati, DomSuid and HJX74) and the other 3 are wild rice (O. meridionalis, O. barthii and O. rufipogon).

Fig. 2. Characteristics of Huajingxian 74 (HJX74) genome and synteny examining, SNPs (single nucleotide polymorphisms) and InDels (Inserts/Deletes) mining, dN/dS comparison with Nipponbare (NPB) and R498. A, Distribution of SNPs and InDels between HJX74 and NPB (the data refer to Table S5).B, Distribution of SNPs and InDels between HJX74 and R498 (the data refer to Table S6).C, dN/dS distribution of different combinations. The vertical lines represent average values of dN/dS.D, Chromosomal synteny among HJX74 and two reference genomes of rice.E, Interactive dot plot between HJX74 and NPB.F, Interactive dot plot between HJX74 and R498. G, Characteristics of the HJX74 genome. Tracks from outside to inside are the 12 chromosomes of HJX74, GC content, long terminal repeat density, and simple sequence repeat density (the data refer to Table S11).
| Statistics without reference | HJX74 (contig level) | HJX74 | Nipponbare | R498 |
|---|---|---|---|---|
| Contigs | 155 | 108 | 63 | 14 |
| Contigs ≥ 1 000 bp | 155 | 108 | 63 | 14 |
| Contigs ≥ 5 000 bp | 141 | 94 | 62 | 14 |
| Contigs ≥ 10 000 bp | 135 | 88 | 51 | 14 |
| Contigs ≥ 25 000 bp | 126 | 80 | 20 | 14 |
| Contigs ≥ 50 000 bp | 109 | 69 | 18 | 14 |
| Largest contig | 26 609 203 | 44 230 994 | 43 270 923 | 44 361 539 |
| Total length | 399 000 942 | 398 866 991 | 375 049 285 | 390 983 850 |
| Total length ≥1 000 bp | 399 000 942 | 398 866 991 | 375 049 285 | 390 983 850 |
| Total length ≥5 000 bp | 398 955 832 | 398 821 873 | 375 045 049 | 390 983 850 |
| Total length ≥10 000 bp | 398 920 023 | 398 786 031 | 374 960 283 | 390 983 850 |
| Total length ≥25 000 bp | 398 753 016 | 398 632 314 | 374 514 817 | 390 983 850 |
| Total length ≥50 000 bp | 398 046 761 | 398 182 493 | 374 450 782 | 390 983 850 |
| N50 | 14 408 600 | 31 354 834 | 29 958 434 | 31 778 392 |
| N75 | 11 330 867 | 30 528 296 | 28 443 022 | 29 952 003 |
| L50 | 11 | 6 | 6 | 6 |
| L75 | 18 | 9 | 9 | 9 |
| GC (%) | 43.7 | 43.7 | 43.57 | 43.62 |
| # N’s (Mismatches) | 0 | 5 511 | 118 258 | 50 000 |
| # N’s per 100 kbp (Mismatches) | 0 | 1.38 | 31.53 | 12.79 |
Table S1. Comparison of contigs and scaffolds among Huajingxian 74 (HJX74) and two reference rice genomes (Nipponbare and R498).
| Statistics without reference | HJX74 (contig level) | HJX74 | Nipponbare | R498 |
|---|---|---|---|---|
| Contigs | 155 | 108 | 63 | 14 |
| Contigs ≥ 1 000 bp | 155 | 108 | 63 | 14 |
| Contigs ≥ 5 000 bp | 141 | 94 | 62 | 14 |
| Contigs ≥ 10 000 bp | 135 | 88 | 51 | 14 |
| Contigs ≥ 25 000 bp | 126 | 80 | 20 | 14 |
| Contigs ≥ 50 000 bp | 109 | 69 | 18 | 14 |
| Largest contig | 26 609 203 | 44 230 994 | 43 270 923 | 44 361 539 |
| Total length | 399 000 942 | 398 866 991 | 375 049 285 | 390 983 850 |
| Total length ≥1 000 bp | 399 000 942 | 398 866 991 | 375 049 285 | 390 983 850 |
| Total length ≥5 000 bp | 398 955 832 | 398 821 873 | 375 045 049 | 390 983 850 |
| Total length ≥10 000 bp | 398 920 023 | 398 786 031 | 374 960 283 | 390 983 850 |
| Total length ≥25 000 bp | 398 753 016 | 398 632 314 | 374 514 817 | 390 983 850 |
| Total length ≥50 000 bp | 398 046 761 | 398 182 493 | 374 450 782 | 390 983 850 |
| N50 | 14 408 600 | 31 354 834 | 29 958 434 | 31 778 392 |
| N75 | 11 330 867 | 30 528 296 | 28 443 022 | 29 952 003 |
| L50 | 11 | 6 | 6 | 6 |
| L75 | 18 | 9 | 9 | 9 |
| GC (%) | 43.7 | 43.7 | 43.57 | 43.62 |
| # N’s (Mismatches) | 0 | 5 511 | 118 258 | 50 000 |
| # N’s per 100 kbp (Mismatches) | 0 | 1.38 | 31.53 | 12.79 |
| Method (Database) | Data |
|---|---|
| BUSCO | C, 98.3%; S, 97.6%, D, 0.7%; F, 0.3%; M, 1.4%; n, 1440 |
| CEGMA | Complet: 230 (92.74%); Complete + Partial: 235 (94.76%) |
Table S2. Evaluation of Huangjingxian 74 genome assembly by Benchmarking Universal single-copy ortholog (BUSCO) and Core Eukaryotic Genes Mapping Approach (CEGMA).
| Method (Database) | Data |
|---|---|
| BUSCO | C, 98.3%; S, 97.6%, D, 0.7%; F, 0.3%; M, 1.4%; n, 1440 |
| CEGMA | Complet: 230 (92.74%); Complete + Partial: 235 (94.76%) |
| Species | Length | Intact | Total | Raw LAI | LAI |
|---|---|---|---|---|---|
| R498 (indica) | 390 983 850 | 0.0663 | 0.2828 | 23.44 | 23.94 |
| Nipponbare (japonica) | 375 049 285 | 0.0505 | 0.2452 | 20.58 | 22.59 |
| HJX74 (indica) | 398 866 991 | 0.0656 | 0.2887 | 22.73 | 23.42 |
Table S3. LTR-TRs assembly index (LAI) of R498, Nipponbare and Huajingxian 74 (HJX74).
| Species | Length | Intact | Total | Raw LAI | LAI |
|---|---|---|---|---|---|
| R498 (indica) | 390 983 850 | 0.0663 | 0.2828 | 23.44 | 23.94 |
| Nipponbare (japonica) | 375 049 285 | 0.0505 | 0.2452 | 20.58 | 22.59 |
| HJX74 (indica) | 398 866 991 | 0.0656 | 0.2887 | 22.73 | 23.42 |

Fig. S2. Cumulative sequence length and presence and absence variations (PAVs) distribution.A and B, Comparison of cumulative sequence length of HJX74 and two published reference rice genomes (NPB and R498); C and D, Comparison of PAVs length distribution in HJX74 and two published reference rice genomes (NPB and R498).
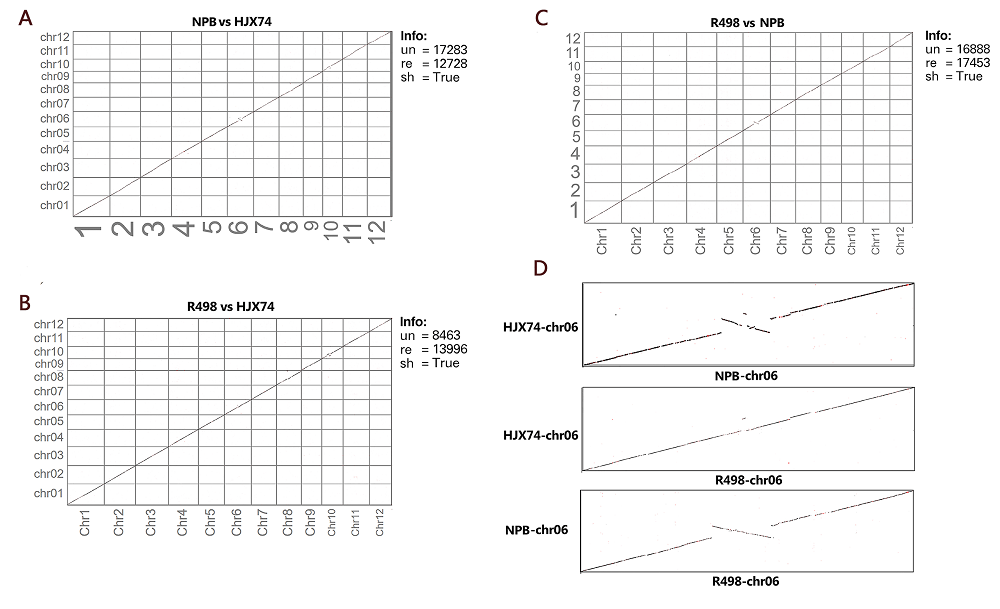
Fig. S3. Interactive dot plot of Huajingxian 74 (HJX74) and two reference rice genomes (R498 and Nipponbare, NPB).A to C, Paired comparison of HJX74 and two reference genomes on genome level; D, Comparison of HJX74 and two reference rice genomes on the region of chromosome 6(un, Unique alignment; re, Repetitive alignment; sh, Show repetitive alignment).
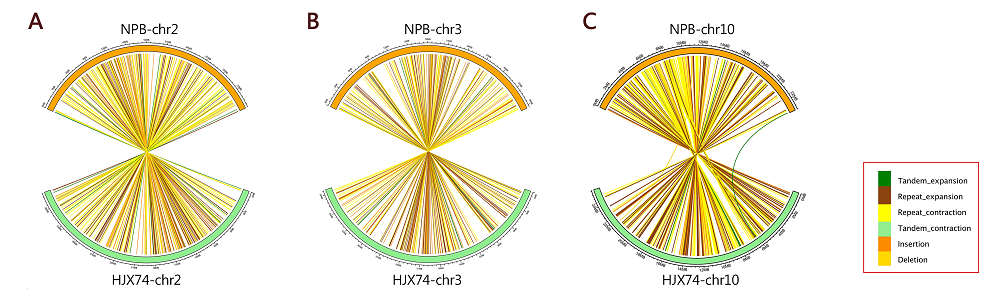
Fig. S4. Presence and absence variations (PAVs) types and distribution of some chromosomes with significantly different lengths between Huajingxian 74 (HJX74) and Nipponbare (NPB).A, Chromosome 2; B, Chromosome 3; C, Chromosome10.
| Species | Protein | Cluster | Singleton |
|---|---|---|---|
| Oryza barthii | 34 575 | 24 966 | 3 223 |
| Oryza meridionalis | 29 308 | 20 721 | 4 088 |
| Oryza rufipogon | 37 062 | 27 114 | 2 912 |
| Oryza sativa Nipponbare | 35 667 | 23 642 | 5 446 |
| Oryza sativa 93-11 | 37 878 | 25 644 | 3 690 |
| Oryza sativa Basmati | 41 259 | 29 417 | 4 210 |
| Oryza sativa DomSufid | 38 329 | 28 037 | 3 490 |
| Oryza sativa Minghui 63 | 83 258 | 40 508 | 13 514 |
| Oryza sativa R498 | 88 169 | 27 111 | 33 193 |
| Oryza sativa Zhenshan 97 | 84 957 | 40 646 | 14 163 |
| Oryza sativa Huajingxian 74 | 46 933 | 27 202 | 7 991 |
| Oryza sativa IR64 | 41 458 | 23 591 | 9 810 |
Table S7. Clustering of homologous genes of 12 species of rice.
| Species | Protein | Cluster | Singleton |
|---|---|---|---|
| Oryza barthii | 34 575 | 24 966 | 3 223 |
| Oryza meridionalis | 29 308 | 20 721 | 4 088 |
| Oryza rufipogon | 37 062 | 27 114 | 2 912 |
| Oryza sativa Nipponbare | 35 667 | 23 642 | 5 446 |
| Oryza sativa 93-11 | 37 878 | 25 644 | 3 690 |
| Oryza sativa Basmati | 41 259 | 29 417 | 4 210 |
| Oryza sativa DomSufid | 38 329 | 28 037 | 3 490 |
| Oryza sativa Minghui 63 | 83 258 | 40 508 | 13 514 |
| Oryza sativa R498 | 88 169 | 27 111 | 33 193 |
| Oryza sativa Zhenshan 97 | 84 957 | 40 646 | 14 163 |
| Oryza sativa Huajingxian 74 | 46 933 | 27 202 | 7 991 |
| Oryza sativa IR64 | 41 458 | 23 591 | 9 810 |
| Chromosome | Length of each chromosome (bp) | ||
|---|---|---|---|
| HJX74 | NPB | R498 | |
| 1 | 44 230 993 | 43 270 923 | 44 361 539 |
| 2 | 38 415 586 | 35 937 250 | 37 764 328 |
| 3 | 39 700 684 | 36 413 819 | 39 691 490 |
| 4 | 36 567 208 | 35 502 694 | 35 849 732 |
| 5 | 30 862 635 | 29 958 434 | 31 237 231 |
| 6 | 31 992 588 | 31 248 787 | 32 465 040 |
| 7 | 30 655 926 | 29 697 621 | 30 277 827 |
| 8 | 30 528 296 | 28 443 022 | 29 952 003 |
| 9 | 24 203 274 | 23 012 720 | 24 760 661 |
| 10 | 25 662 364 | 23 207 287 | 25 582 588 |
| 11 | 31 354 834 | 29 021 106 | 31 778 392 |
| 12 | 27 052 960 | 27 531 856 | 26 601 357 |
| Average difference | 1.390 Mb | 0.07543 Mb | |
Table S9. Chromosome lengths of Huajingxian 74 (HJX74), Nipponbare (NPB) and R498.
| Chromosome | Length of each chromosome (bp) | ||
|---|---|---|---|
| HJX74 | NPB | R498 | |
| 1 | 44 230 993 | 43 270 923 | 44 361 539 |
| 2 | 38 415 586 | 35 937 250 | 37 764 328 |
| 3 | 39 700 684 | 36 413 819 | 39 691 490 |
| 4 | 36 567 208 | 35 502 694 | 35 849 732 |
| 5 | 30 862 635 | 29 958 434 | 31 237 231 |
| 6 | 31 992 588 | 31 248 787 | 32 465 040 |
| 7 | 30 655 926 | 29 697 621 | 30 277 827 |
| 8 | 30 528 296 | 28 443 022 | 29 952 003 |
| 9 | 24 203 274 | 23 012 720 | 24 760 661 |
| 10 | 25 662 364 | 23 207 287 | 25 582 588 |
| 11 | 31 354 834 | 29 021 106 | 31 778 392 |
| 12 | 27 052 960 | 27 531 856 | 26 601 357 |
| Average difference | 1.390 Mb | 0.07543 Mb | |
| Repeat content | HJX74 | R498 | NPB | |||
|---|---|---|---|---|---|---|
| Length (bp) | Percentage of genome (%) | Length (bp) | Percentage of genome (%) | Length (bp) | Percentage of genome (%) | |
| Total | 110 814 118 | 27.78 | 109 473 086 | 28.00 | 90 414 977 | 24.12 |
| Gypsy | 84 080 899 | 21.08 | 82 619 222 | 21.13 | 64 570 822 | 17.22 |
| Copia | 13 457 780 | 3.37 | 12 661 044 | 3.24 | 13 948 467 | 3.72 |
| Unknown | 13 275 439 | 3.33 | 14 192 820 | 3.63 | 11 895 688 | 3.17 |
Table S10. Repeat content (LTR-RTs) in Huajingxian 74 (HJX74), R498 and Nipponbare (NPB).
| Repeat content | HJX74 | R498 | NPB | |||
|---|---|---|---|---|---|---|
| Length (bp) | Percentage of genome (%) | Length (bp) | Percentage of genome (%) | Length (bp) | Percentage of genome (%) | |
| Total | 110 814 118 | 27.78 | 109 473 086 | 28.00 | 90 414 977 | 24.12 |
| Gypsy | 84 080 899 | 21.08 | 82 619 222 | 21.13 | 64 570 822 | 17.22 |
| Copia | 13 457 780 | 3.37 | 12 661 044 | 3.24 | 13 948 467 | 3.72 |
| Unknown | 13 275 439 | 3.33 | 14 192 820 | 3.63 | 11 895 688 | 3.17 |
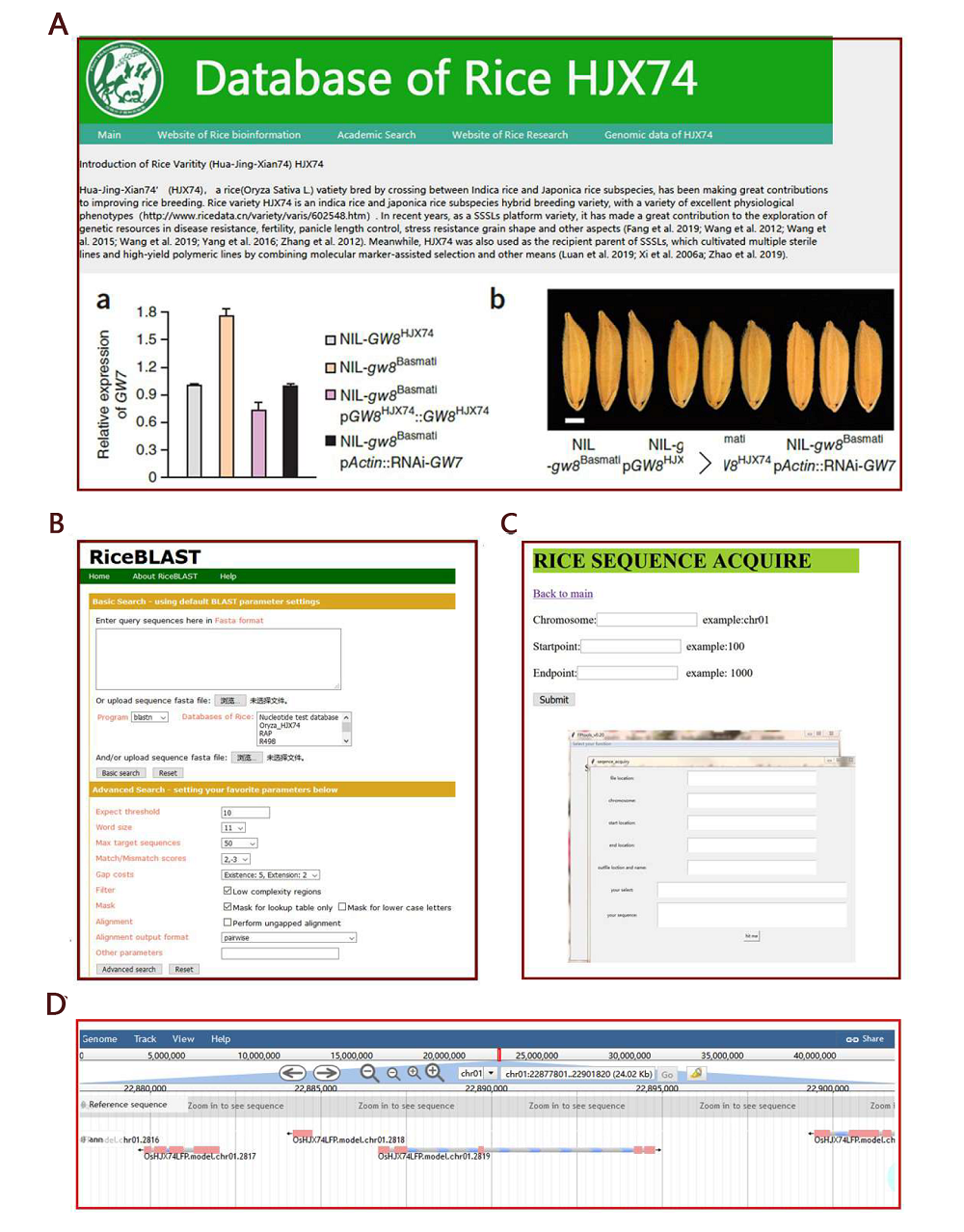
Fig. S5. Online accesible platfrom of HJX74 Genome data.A, Display of platform homepage; B, Platform sequence search (BLAST) function; C, Display of FPtools page and online sequence extraction page; D, Display of genes browsing page.
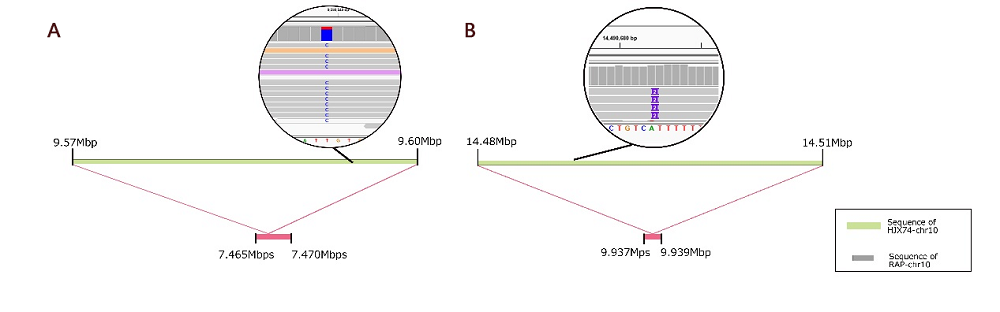
Fig. S6. Sequence difference in presence and absence variations (PAVs) locus.A, SNP detection in PAVs locus; B, Insertion sites detection in PAVs locus.
| [1] | Choi J Y, Lye Z N, Groen S C, Dai X G, Rughani P, Zaaijer S, Harrington E D, Juul S, Purugganan M D.2020. Nanopore sequencing-based genome assembly and evolutionary genomics of circum-basmati rice.Genome Biol, 21(1): 21. |
| [2] | Du H L, Yu Y, Ma Y F, Gao Q, Cao Y H, Chen Z, Ma B, Qi M, Li Y, Zhao X F, Wang J, Liu K F, Qin P, Yang X, Zhu L H, Li S G, Liang C Z.2017. Sequencing and de novo assembly of a near complete indica rice genome.Nat Commun, 8(1): 15324. |
| [3] | Fang C W, Li L, He R M, Wang D Q, Wang M, Hu Q, Ma Q R, Qin K Y, Feng X Y, Zhang G Q, Fu X L, Liu Z Q.2019. Identification of S23 causing both interspecific hybrid male sterility and environment-conditioned male sterility in rice.Rice, 12(1): 10. |
| [4] | Groen S C, Calic I, Joly-Lopez Z, Platts A E, Choi J Y, Natividad M, Dorph K, Mauck III W M, Bracken B, Cabral C L U, Kumar A, Torres R O, Satija R, Vergara G, Henry A, Franks S J, Purugganan M D.2020. The strength and pattern of natural selection on gene expression in rice.Nature, 578: 572-576. |
| [5] | Kawahara Y, de la Bastide M, Hamilton J P, Kanamori H, Mccombie W R, Ouyang S, Schwartz D C, Tanaka T, Wu J Z, Zhou S G, Childs K L, Davidson R M, Lin H N, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee S S, Kim J, Numa H, Itoh T, Buell C R, Matsumoto T.2013. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data.Rice, 6(1): 4. |
| [6] | Kim S, Park J, Yeom S I, Kim Y M, Seo E, Kim K T, Kim M S, Lee J M, Cheong K, Shin H S, Kim S B, Han K, Lee J, Park M, Lee H A, Lee H Y, Lee Y, Oh S, Lee J H, Choi E, Choi E, Lee S E, Jeon J, Kim H, Choi G, Song H, Lee J, Lee S C, Kwon J K, Lee H Y, Koo N, Hong Y, Kim R W, Kang W H, Huh J H, Kang B C, Yang T J, Lee Y H, Bennetzen J L, Choi D.2017. New reference genome sequences of hot pepper reveal the massive evolution of plant disease-resistance genes by retroduplication.Genome Biol, 18(1): 210-221. |
| [7] | Kryazhimskiy S, Plotkin J B.2008. The population genetics of dN/dS.PLoS Genet, 4(12): e1000304. |
| [8] | Li W, Li K, Zhang Q J, Zhu T, Zhang Y, Shi C, Liu Y L, Xia E H, Jiang J J, Shi C, Zhang L P, Huang H, Tong Y, Liu Y, Zhang D, Zhao Y, Jiang W K, Zhao Y J, Mao S Y, Jiao J Y, Xu P Z, Yang L L, Yin G Y, Gao L Z.2020. Improved hybrid de novo genome assembly and annotation of African wild rice, Oryza longistaminata, from Illumina and PacBio sequencing reads.Plant Genome, 13(1): e20001. |
| [9] | Luan X, Dai Z J, Yang W F, Tan Q Y, Lu Q, Guo J, Zhu H T, Liu G F, Wang S K, Zhang G Q.2019. Breeding by design of CMS lines on the platform of SSSL library in rice.Mol Breeding, 39(9): 126. |
| [10] | Luo Y C, Ma T C, Zhang A F, Ong K H, Luo Z X, Li Z F, Yang J B, Yin Z C.2017. Marker-assisted breeding of Chinese elite rice cultivar 9311 for disease resistance to rice blast and bacterial blight and tolerance to submergence.Mol Breeding, 37(8): 106. |
| [11] | Rhoads A, Au K F.2015. PacBio sequencing and its applications.Genom Proteom Bioinf, 13(5): 278-289. |
| [12] | Sakai H, Lee S S, Tanaka T, Numa H, Kim J, Kawahara Y, Wakimoto H, Yang C C, Iwamoto M, Abe T, Yamada Y, Muto A, Inokuchi H, Ikemura T, Matsumoto T, Sasaki T, Itoh T.2013. Rice Annotation Project Database (RAP-DB): An integrative and interactive database for rice genomics.Plant Cell Physiol, 54(2): e6. |
| [13] | Shao L, Xing F, Xu C H, Zhang Q H, Che J, Wang X M, Song J M, Li X H, Xiao J H, Chen L L, Ouyang Y D, Zhang Q F.2019. Patterns of genome-wide allele-specific expression in hybrid rice and the implications on the genetic basis of heterosis.Prol Natl Acad Sci USA, 116(12): 5653-5658. |
| [14] | Tanaka T, Nishijima R, Teramoto S, Kitomi Y, Hayashi T, Uga Y, Kawakatsu T.2020. De novo genome assembly of the indica rice variety IR64 using linked-read sequencing and nanopore sequencing. G3: Genes Genom Genet, 10(5): 1495-1501. |
| [15] | The Rice Chromosomes 11 and 12 Sequencing Consortia.2005. The sequence of rice chromosomes 11 and 12, rich in disease resistance genes and recent gene duplications.BMC Biol, 3(1): 20. |
| [16] | van Berkum N L, Lieberman-Aiden E, Williams L, Imakaev M, Gnirke A, Mirny L A, Dekker J, Lander E S.2010. Hi-C: A method to study the three-dimensional architecture of genomes.J Visualized Exp, 39: e1869. |
| [17] | Wang S K, Li S, Liu Q, Wu K, Zhang J Q, Wang S S, Wang Y, Chen X B, Zhang Y, Gao C X, Wang F, Huang H X, Fu X D.2015. The OsSPL16-GW7 regulatory module determines grain shape and simultaneously improves rice yield and grain quality.Nat Genet, 47(8): 949-954. |
| [18] | Wang X L, Liu G F, Wang Z Q, Chen S L, Xiao Y L, Yu C Y.2019. Identification and application of major quantitative trait loci for panicle length in rice (Oryza sativa) through single-segment substitution lines.Plant Breeding, 138(3): 299-308. |
| [19] | Wang Y, Coleman-Derr D, Chen G P, Gu Y Q.2015. OrthoVenn: A web server for genome wide comparison and annotation of orthologous clusters across multiple species.Nucl Acids Res, 43: W78-W84. |
| [20] | Wang Y P, Tang H B, Debarry J D, Tan X, Li J P, Wang X Y, Lee T H, Jin H Z, Marler B, Guo H, Kissinger J C, Paterson A H.2012. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity.Nucl Acids Res, 40(7): e49. |
| [21] | Xie X R, Du H L, Tang H W, Tang J N, Tan X Y, Liu W Z, Li T, Lin Z S, Liang C Z, Liu Y G.2020. A chromosome-level genome assembly of the wild rice Oryza rufipogon facilitates tracing the origins of Asian cultivated rice. Sci China: Life Sci, 5(1): 1-11. |
| [22] | Zhang G Q.2019. The platform of breeding by design based on the SSSL library in rice.Hereditas, 41(8): 754-760. |
| [23] | Zhang J W, Chen L L, Xing F, Kudrna D A, Yao W, Copetti D, Mu T, Li W M, Song J M, Xie W B, Lee S, Talag J, Shao L, An Y, Zhang C L, Ouyang Y D, Sun S, Jiao W B, Lv F, Du B G, Luo M Z, Maldonado C E, Goicechea J L, Xiong L Z, Wu C Y, Xing Y Z, Zhou D X, Yu S B, Zhao B, Wang G W, Yu Y, Luo Y J, Zhou Z W, Hurtado B E P, Danowitz A, Wing R A, Zhang Q F.2016. Extensive sequence divergence between the reference genomes of two elite indica rice varieties Zhenshan 97 and Minghui 63.Proc Natl Acad Sci USA, 113: E5163-E5171. |
| [24] | Zhang Q, Liang Z, Cui X A, Ji C M, Li Y, Zhang P X, Liu J R, Riaz A, Yao P, Liu M, Wang Y P, Lu T G, Yu H, Yang D L, Zheng H K, Gu X F.2018. N-6-methyladenine DNA methylation in japonica and indica rice genomes and its association with gene expression, plant development, and stress responses.Mol Plant, 11(12): 1492-1508. |
| [25] | Zhang T Z, Hu Y, Jiang W K, Fang L, Guan X Y, Chen J D, Zhang J B, Saski C A, Scheffler B E, Stelly D M, Hulse-Kemp A M, Wan Q, Liu B L, Liu C X, Wang S, Pan M Q, Wang Y K, Wang D W, Ye W X, Chang L J, Zhang W P, Song Q X, Kirkbride R C, Chen X Y, Dennis E, Llewellyn D J, Peterson D G, Thaxton P, Jones D C, Wang Q, Xu X Y, Zhang H, Wu H T, Zhou L, Mei G F, Chen S Q, Tian Y, Xiang D, Li X H, Ding J, Zuo Q Y, Tao L N, Liu Y C, Li J, Lin Y, Hui Y Y, Cao Z S, Cai C P, Zhu X F, Jiang Z, Zhou B L, Guo W Z, Li R Q, Chen Z J.2015. Sequencing of allotetraploid cotton (Gossypium hirsutum L. acc. TM-1) provides a resource for fiber improvement. Nat Biotechnol, 33(5): 531-537. |
| [26] | Zhao H W, Sun L L, Xiong T Y, Wang Z Q, Liao Y, Zou T, Zheng M M, Zhang Z, Pan X P, He N, Zhang G Q, Zhu H T, Liu Z Q, He P, Fu X L.2019. Genetic characterization of the chromosome single-segment substitution lines of O. glumaepatula and O. barthii and identification of QTLs for yield-related traits.Mol Breeding, 39(4): 51. |
| [27] | Zhou Y L, Xie Y H, Cai J L, Liu C B, Zhu H T, Jiang R, Zhong Y Y, Zhang G L, Tan B, Liu G F, Fu X L, Liu Z Q, Wang S K, Zhang G Q, Zeng R Z.2017. Substitution mapping of QTLs controlling seed dormancy using single segment substitution lines derived from multiple cultivated rice donors in seven cropping seasons.Theor Appl Genet, 130(6): 1191-1205. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||