
Rice Science ›› 2016, Vol. 23 ›› Issue (6): 334-338.DOI: 10.1016/j.rsci.2016.05.004
• Orginal Article • Previous Articles Next Articles
Quynh-Hoa Pham1, Xuan-An Tran1, Thi-Nha-Trang Nguyen1, Thi-Thuy-Anh Tran1, Hai-Yen Hoang1, Thi-Hong-Van Nguyen1, Thi-Hanh Tang2, Thi-Phuc Do1
Received:2016-02-25
Accepted:2016-05-06
Online:2016-12-12
Published:2016-08-10
Quynh-Hoa Pham, Xuan-An Tran, Thi-Nha-Trang Nguyen, Thi-Thuy-Anh Tran, Hai-Yen Hoang, Thi-Hong-Van Nguyen, Thi-Hanh Tang, Thi-Phuc Do. Investigation of Polymorphisms in Coding Region of OsHKT1 in Relation to Salinity in Rice[J]. Rice Science, 2016, 23(6): 334-338.
Add to citation manager EndNote|Ris|BibTeX
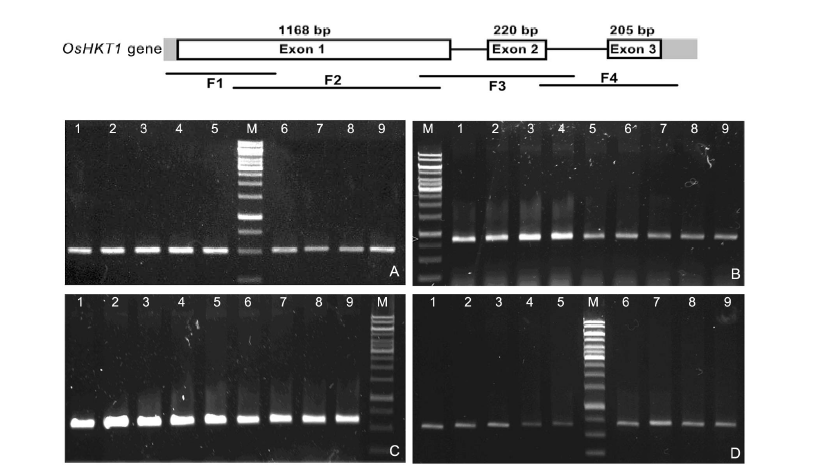
Fig. 1. PCR products using four pairs of primers. A, PCR product of 499 bp using primer pair 1 (F1); B, PCR product of 828 bp using primer pair 2 (F2); C, PCR product of 603 bp using primer pair 3 (F3); D, PCR product of 668 bp using primer pair 4 (F4). M, 1 kb ladder marker; Lanes 1 to 9, Nipponbare, Nep Non Tre, Chiem Cu, Re Nuoc, Hom Rau, Nep Oc, Ngoi, Dau An Do and Nep Deo Dang, respectively.
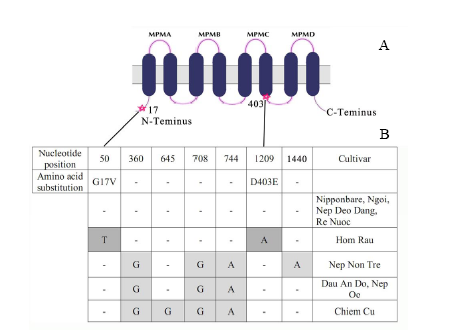
Fig. 2. Polymorphism in OsHKT1 sequence. A, Schematic representation of polymorphism locations in the protein. The position of changes in amino acid are indicated with a star; MPM, transmembrane-pore loop-transmembrane domain. B, Nucleotide sequence of OsHKT1 was compared among the cultivars with the Nipponbare sequence. Non-synonymous substitutions are indicated in dark grey, and the encoded amino acids are listed above. Synonymous substitutions are indicated in light grey. G17V means that glycine is replaced by valine; D403E means that aspartic acid is replaced by glutamic acid.
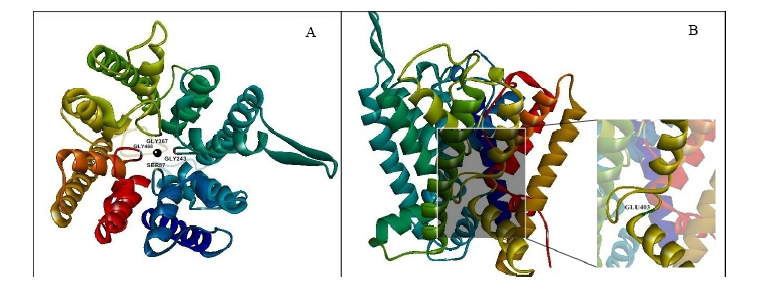
Fig. 3. 3D ribbon model of OsHKT1 protein visualized by Discovery studio 4.5 visualizer. A, 3D ribbon model of Ni-OsHKT1 protein visualized from the top that shows the pore. Four residues in Ser-Gly-Gly-Gly ion selection motif are labeled and Na+ ion is indicated as a black round circle at the center of the pore. B, Visualization of Ho-OsHKT1 transporter from the side that shows the zoom in D403E variance position (green) (G17V substitution belongs to the signal peptide so they are not modeled in 3D structure of OsHKT1 transporter).
| Position of nucleotide variance | Former codon | Usage frequency of former codon (‰) | Substituted codon | Usage frequency of substituted codon (‰) | Amino acid | Cultivar |
| 360 | CUA | 7.7 | CUG | 21.0 | Leu | Nep Non Tre, Dau An Do, Nep Oc, Chiem Cu |
| 645 | UCA | 12.4 | UCG | 12.3 | Leu | Chiem Cu |
| 708 | ACA | 11.6 | ACG | 11.4 | Thr | Nep Non Tre, Dau An Do, Nep Oc, Chiem Cu |
| 744 | ACG | 11.4 | ACA | 11.6 | Thr | Nep Non Tre, Dau An Do, Nep Oc, Chiem Cu |
| 1440 | AAG | 32.3 | CAA | 13.5 | Lys | Nep Non Tre |
| Values in bold meant that the substituted nucleotide caused an increase in codon usage frequence. | ||||||
Table 1 Changes in codon usage bias by nucleotide substitutions in nine rice cultivars
| Position of nucleotide variance | Former codon | Usage frequency of former codon (‰) | Substituted codon | Usage frequency of substituted codon (‰) | Amino acid | Cultivar |
| 360 | CUA | 7.7 | CUG | 21.0 | Leu | Nep Non Tre, Dau An Do, Nep Oc, Chiem Cu |
| 645 | UCA | 12.4 | UCG | 12.3 | Leu | Chiem Cu |
| 708 | ACA | 11.6 | ACG | 11.4 | Thr | Nep Non Tre, Dau An Do, Nep Oc, Chiem Cu |
| 744 | ACG | 11.4 | ACA | 11.6 | Thr | Nep Non Tre, Dau An Do, Nep Oc, Chiem Cu |
| 1440 | AAG | 32.3 | CAA | 13.5 | Lys | Nep Non Tre |
| Values in bold meant that the substituted nucleotide caused an increase in codon usage frequence. | ||||||
| 1 | Baxter I, Brazelton J N, Yu D, Huang Y S, Lahner B, Yakubova E, Li Y, Bergelson J, Borevitz J O, Nordborg M, Vitek O, Salt D E.2010. A coastal cline in sodium accumulation inArabidopsis thaliana is driven by natural variation of the sodium transporter AtHKT1;1. PLoS Genet, 6: e1001193. |
| 2 | Berthomieu P, Conejero G, Nublat A, Brackenbury W J, Lambert C, Savio C, Uozumi N, Oiki S, Yamada K, Cellier F, Gosti F, Simonneau T, Essah P A, Tester M, Very A A, Sentenac H, Casse F.2003. Functional analysis ofAtHKT1 in Arabidopsis shows that Na+ recirculation by the phloem is crucial for salt tolerance. EMBO J, 22(9): 2004-2014. |
| 3 | Brady K U, Kruckeberg A R, Bradshaw Jr H D.2005. Evolutionary ecology of plant adaptation to serpentine soils.Annu Rev Ecol Evol Syst, 36: 243-266. |
| 4 | Chinnusamy V, Jagendorf A, Zhu J K.2005. Understanding and improving salt tolerance in plants.Crop Sci, 45: 437-448. |
| 5 | Corpet F.1988. Multiple sequence alignment with hierarchical clustering.Nucl Acids Res, 16(22): 10881-10890. |
| 6 | Do T P, Nguyen T T T.2014. Polymorphism analysis ofOsHKT1 gene in rice(Oryza sativa). VNU J Sci: Natl Sci Technol, 30: 253-259. (in Vietnamese with English abstract) |
| 7 | Garciadeblás B, Senn M E, Banuelos M A, Rodriguez-Navarro A.2003. Sodium transport and HKT transporters: The rice model.Plant J, 34: 788-801. |
| 8 | Golldack D, Su H, Quigley F, Kamasani U R, Munoz-Garay C, Balderas E, Popova O V, Bennett J, Bohnert H J, Pantoja O.2002. Characterization of a HKT-type transporter in rice as a general alkali cation transporter.Plant J, 31(4): 529-542. |
| 9 | Hall T A.1999. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT.Nucl Acids Symp Ser, 41: 95-98. |
| 10 | Horie T, Yoshida K, Nakayama H, Yamada K, Oiki S, Shinmyo A.2001. Two types of HKT transporters with different properties of Na+ and K+ transport inOryza sativa. Plant J, 27(2): 129-138. |
| 11 | Horie T, Schroeder J I.2004. Sodium transporters in plants: Diverse genes and physiological functions.Plant Physiol, 136: 2457-2462. |
| 12 | Jabnoune M, Espeout S, Mieulet D, Fizames C, Verdeil J L, Conéjéro G, Rodríguez-Navarro A, Sentenac H, Guiderdoni E, Abdelly C, Very A A.2009. Diversity in expression patterns and functional properties in the rice HKT transporter family.Plant Physiol, 150(4): 1955-1971. |
| 13 | Kelley L A, Mezulis S, Yates C M, Wass M N, Sternberg M J E.2015. The Phyre2 web portal for protein modeling, prediction and analysis.Nat Protoc, 10: 845-858. |
| 14 | Maser P, Hosoo Y, Goshima S, Horie T, Eckelman B, Yamada K, Yoshida K, Bakker E R, Shinmyo A, Oiki S, Schroeder J I, Uozumi N.2002. Glycine residues in potassium channel-like selectivity filters determine potassium selectivity in four-loop-per-subunit HKT transporters from plants.Proc Natl Acad Sci USA, 99(9): 6428-6433. |
| 15 | Munns R.2002. Comparative physiology of salt and water stress.Plant Cell Environ, 25(2): 239-250. |
| 16 | Munns R, James R A, Laeuchli A.2006. Approaches to increasing the salt tolerance of wheat and other cereals.J Exp Bot, 57(5): 1025-1043. |
| 17 | Oomen R J F J, Benito B, Sentenac H, Rodrıguez-Navarro A, Talon M, Very A A, Domingo C.2012. HKT2;2/1, a K+-permeable transporter identified in a salt-tolerant rice cultivar through surveys of natural genetic polymorphism.Plant J, 71(5): 750-762. |
| 18 | Rains D W, Epstein E.1965. Transport of sodium in plant tissue.Science, 148: 1611. |
| 19 | Ren Z H, Gao J P, Li L G, Cai X L, Huang W, Chao D Y, Zhu M Z, Wang Z Y, Luan S, Lin H X.2005. A rice quantitative trait locus for salt tolerance encodes a sodium transporter.Nat Genet, 37: 1141-1146. |
| 20 | Rus A M, Bressan R A, Hasegawa P M.2005. Unraveling salt tolerance in crops.Nat Genet, 37: 1029-1030. |
| 21 | Schroeder J I, Ward J M, Gassmann W.1994. Perspectives on the physiology and structure of inward rectifying K+ channels in higher plants: Biophysical implications for K+ uptake.Ann Rev Biophys Biomol Struct, 23: 441-471. |
| 22 | Tarczynski M C, Jensen R G, Bohnert H J.1993. Stress protection of transgenic tobacco by production of the osmolyte mannitol.Science, 259: 508-510. |
| 23 | Tester M, Davenport R.2003. Na+ tolerance and Na+ transport in higher plants.Ann Bot, 91(5): 503-527. |
| 24 | Tran X A, Nguyen T N T, Do T P.2015. Evaluation of salt tolerance of some Vietnamese rice varieties.VNU J Sci: Natl Sci Technol, 31: 1-7. (in Vietnamese with English abstract) |
| 25 | Uozumi N, Kim E J, Rubio F, Yamaguchi T, Muto S, Tsuboi A, Bakker E P, Nakamura T, Schroeder J I.2000. TheArabidopsis HKT1 gene homolog mediates inward Na+ currents in Xenopus laevis oocytes and Na+ uptake in Saccharomyces cerevisiae. Plant Physiol, 122(4): 1249-1260. |
| 26 | Volkmar K M, Hu Y, Steppuhn H.1999. Physiological responses of plants to salinity: A review.Can J Plant Sci, 78: 19-27. |
| 27 | Yu C H, Dang Y K, Zhou Z P, Wu C, Zhao F Z, Sachs M S, Liu Y.2015. Codon usage influences the local rate of translation elongation to regulate co-translational protein folding.Mol Cell, 59(5): 744-754. |
| 28 | (Managing Editor: Wang Caihong) |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [13] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [14] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| [15] | Zhang Guomei, Li Han, Liu Shanshan, Zhou Xuming, Lu Mingyang, Tang Liang, Sun Lihua. Water Extract of Rice False Smut Balls Activates Nrf2/HO-1 and Apoptosis Pathways, Causing Liver Injury [J]. Rice Science, 2023, 30(5): 473-485. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||