
Rice Science ›› 2023, Vol. 30 ›› Issue (5): 437-448.DOI: 10.1016/j.rsci.2023.03.015
• Research Papers • Previous Articles Next Articles
Liu Qiao1, Qiu Linlin1, Hua Yangguang1, Li Jing2, Pang Bo1, Zhai Yufeng1, Wang Dekai1( )
)
Received:2023-01-09
Accepted:2023-03-30
Online:2023-09-28
Published:2023-08-14
Contact:
Wang Dekai (About author:First author contact:#These authors contributed equally to this work
Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice[J]. Rice Science, 2023, 30(5): 437-448.
Add to citation manager EndNote|Ris|BibTeX
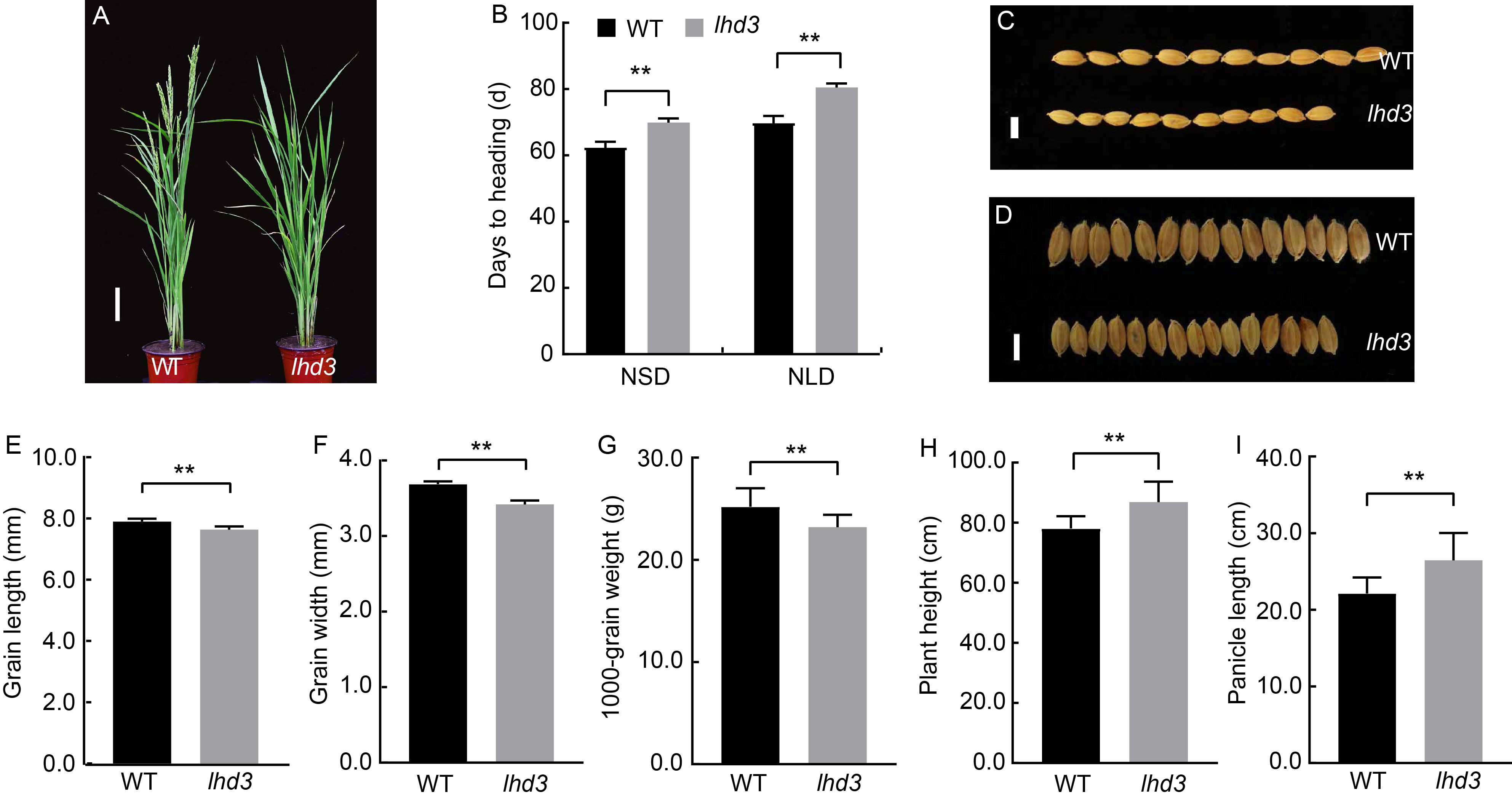
Fig. 1. Phenotypes and agronomic traits of lhd3. A, Phenotypes of wild type (WT) and mutant (lhd3) rice at the heading stage. Scale bar, 10 cm. B?I, Heading dates (B), grain lengths (C and E), grain widths (D and F), 1000-grain weights (G), plant heights (H) and panicle lengths (I) of WT and lhd3. Scale bars, 1 mm in C and D. NSD, Natural short-day; NLD, Natural long-day. Values are Mean ± SD (n = 10 in B, E, F, H and I; n = 3 in G). **, P < 0.01 compared with WT by the Student’s t-test.
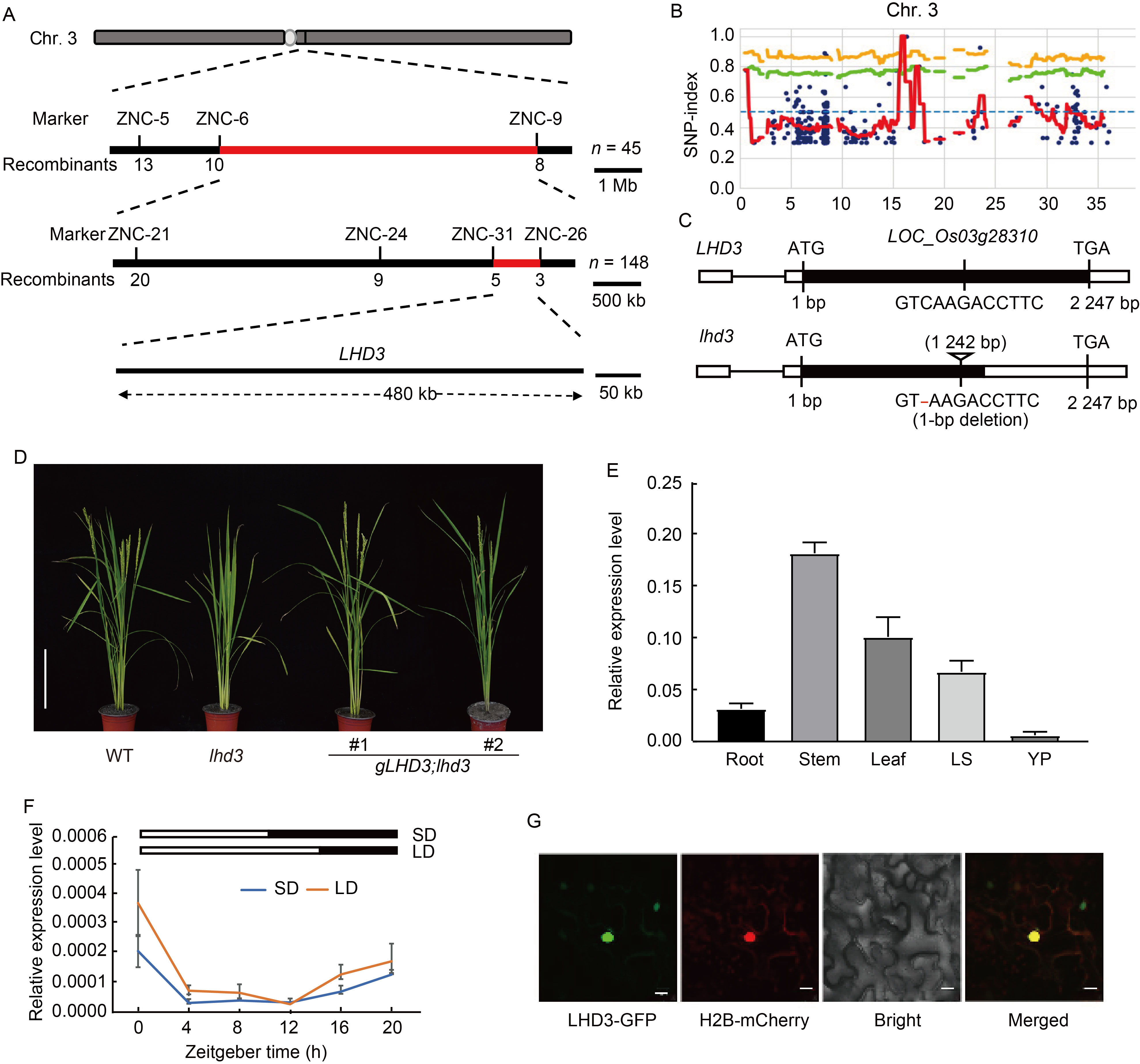
Fig. 2. Cloning and expression analysis of LHD3. A, Molecular mapping of LHD3. The numbers below the lines indicate the number of recombinants. B, Distribution of SNP-index between bulked pools lhd3 and Zhonghua 11 on rice chromosome 3 (Chr. 3). Blue dot, Variant; Orange line, p99; Green line, p95; Red line, SNP-index. C, Schematic diagrams of LHD3 and mutated gene. The 1-bp deletion at 1 242 bp of LHD3 leads to premature termination of the encoded amino acid of LOC_Os03g28310. D, Phenotypes of wild type (WT), mutant lhd3 and its complementary lines. #1 and #2 represent positive transgenic lines from the complementation test. Scale bar, 20 cm. E, Expression of LHD3 in different organs by qRT-PCR analysis. LS, Leaf sheath; YP, Young panicle. Samples were harvested at natural long-day condition. The rice Ubiquitin-5 gene was used as an internal control. Values are Mean ± SD of three independent experiments and three biological replicates. F, Diurnal expression of LHD3 under controlled short-day (SD) and long-day (LD) conditions. The rice Ubiquitin-5 gene was used as an internal control. Values are Mean ± SD of three independent experiments and two biological replicates. G, Subcellular localization of LHD3 in Nicotiana benthamiana mesophyll cells. H2B-mCherry is a nucleus marker. Scale bars, 20 μm.
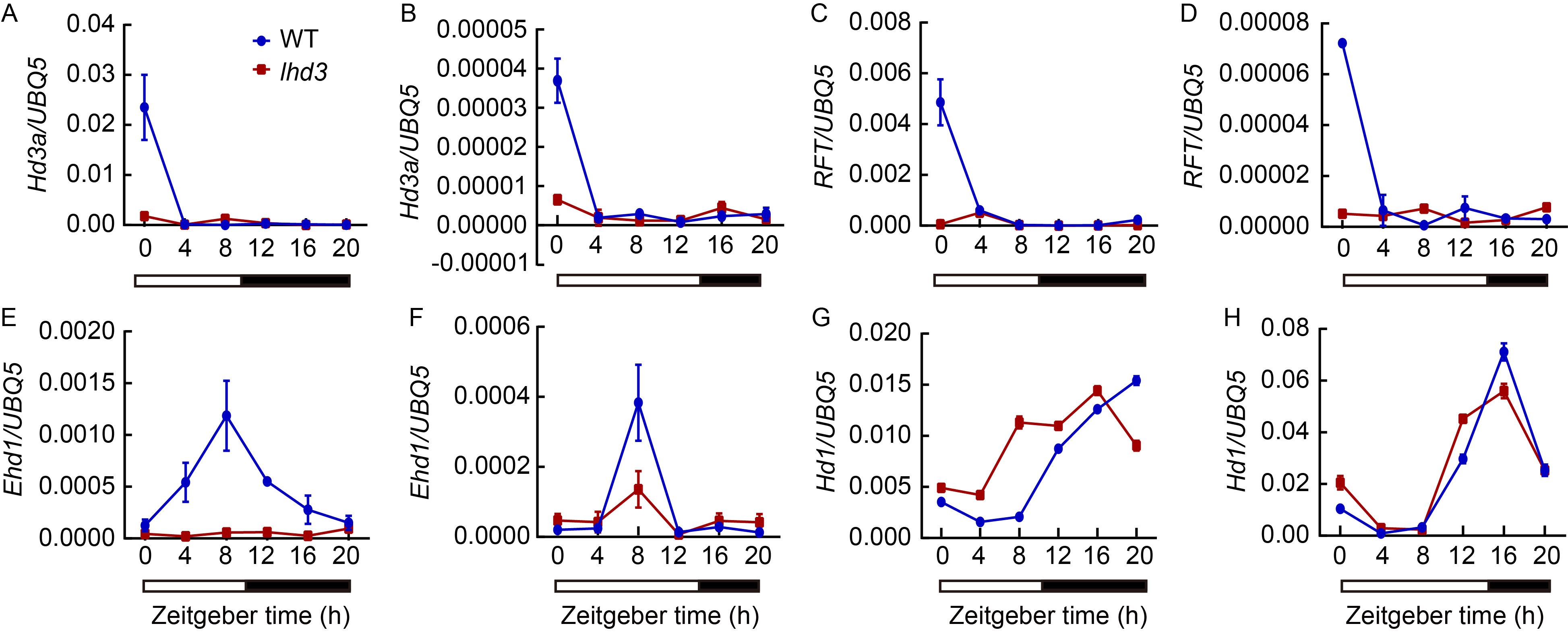
Fig. 3. Rhythmic expression pattern of flowering genes in wild type (WT) and mutant lhd3. A, C, E and G showed short-day conditions. B, D, F and H showed long-day conditions. The open and filled bars at the bottom represent the light and dark periods, respectively. The rice Ubiquitin-5 (UBQ5) gene was used as an internal control. Values are Mean ± SD of three independent experiments and three biological replicates.
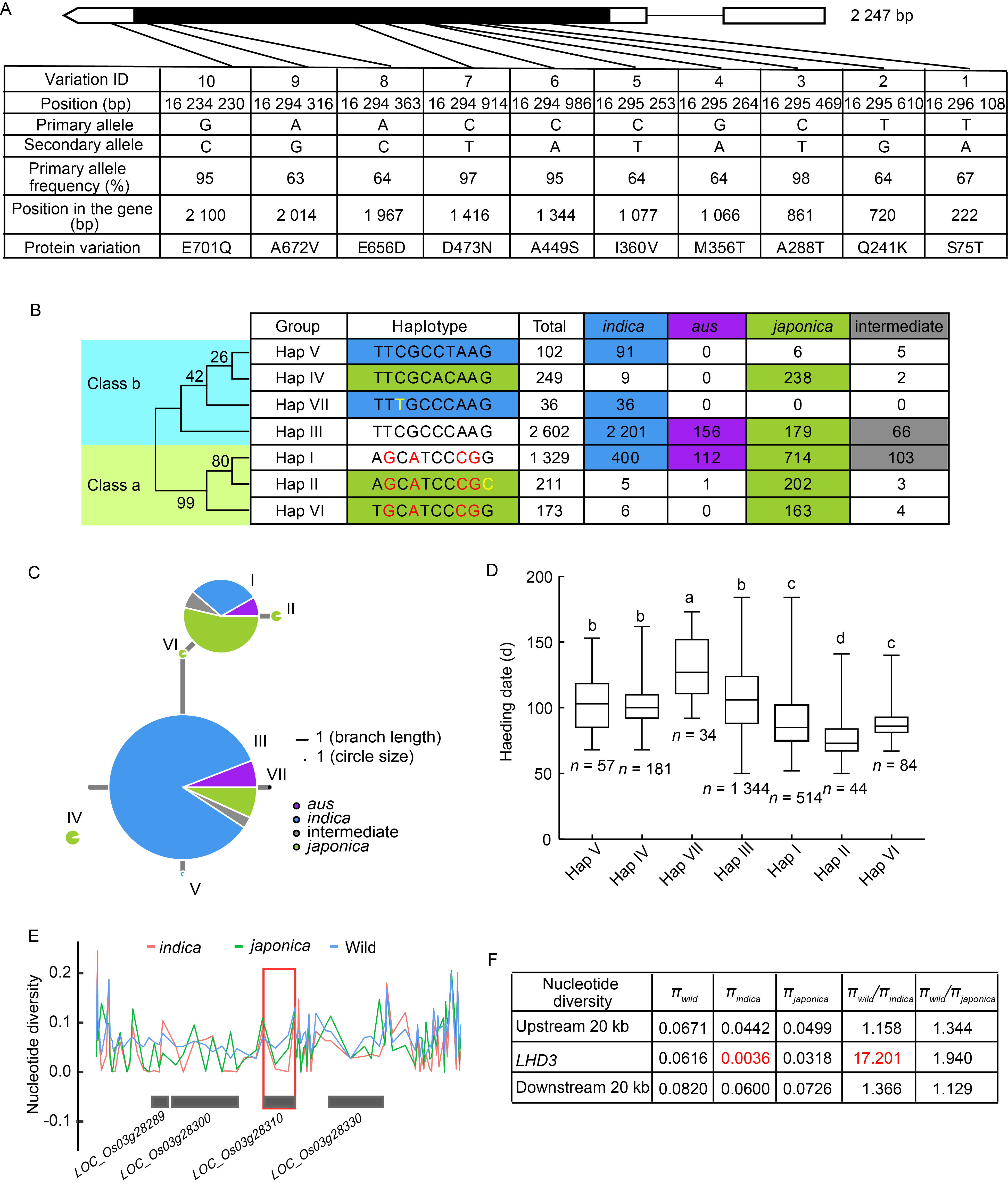
Fig. 4. Haplotype and nucleotide diversity analysis of LHD3. A, Natural variation in LHD3 genomic region revealed 7 haplotypes in 4 702 rice accessions (Oryza sativa ssp. indica, O. sativa ssp. japonica, O. sativa ssp. aus, and intermediate). B, Haplotype analysis of LHD3. The neighbor-joining phylogenetic tree was constructed using MEGA 11 software with 1 000 bootstrap replicates (left). C, Haplotype network analysis of LHD3 in 4 702 rice accessions. Allele frequencies are proportional to the size of circles. D, Box diagram of heading date in seven haplotypes. Different lowercase letters represent significant difference at 5% level according to the Duncan’s multiple range. E, Nucleotide diversity of LHD3 in wild, japonica and indica rice. Red box marked the position of LHD3. F, Nucleotide diversity, the ratios of πwild to πindica and πwild to πjaponica surrounding LHD3.
| [1] |
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R. 2012. Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol, 30(2): 174-178.
PMID |
| [2] |
Andrés F, Coupland G. 2012. The genetic basis of flowering responses to seasonal cues. Nat Rev Genet, 13(9): 627-639.
PMID |
| [3] |
Bian X F, Liu X, Zhao Z G, Jiang L, Gao H, Zhang Y H, Zheng M, Chen L M, Liu S J, Zhai H Q, Wan J M. 2011. Heading date gene, dth3 controlled late flowering in O. Glaberrima Steud. by down-regulating Ehd1. Plant Cell Rep, 30(12): 2243-2254.
PMID |
| [4] |
Bukau B, Weissman J, Horwich A. 2006. Molecular chaperones and protein quality control. Cell, 125(3): 443-451.
PMID |
| [5] | Chai J T, Zhu S S, Li C N, Wang C M, Cai M H, Zheng X M, Zhou L, Zhang H, Sheng P K, Wu M M, Jin X, Cheng Z J, Zhang X, Lei C L, Ren Y L, Lin Q B, Zhou S R, Guo X P, Wang J, Zhao Z C, Wan J M. 2021. OsRE1 interacts with OsRIP1 to regulate rice heading date by finely modulating Ehd1 expression. Plant Biotechnol J, 19(2): 300-310. |
| [6] | Chen K M, Holmström M, Raksajit W, Suorsa M, Piippo M, Aro E M. 2010. Small chloroplast-targeted DnaJ proteins are involved in optimization of photosynthetic reactions in Arabidopsis thaliana. BMC Plant Biol, 10: 43. |
| [7] | Cho L H, Yoon J, An G. 2017. The control of flowering time by environmental factors. Plant J, 90(4): 708-719. |
| [8] |
Christensen C A, Gorsich S W, Brown R H, Jones L G, Brown J, Shaw J M, Drews G N. 2002. Mitochondrial GFA2 is required for synergid cell death in Arabidopsis. Plant Cell, 14(9): 2215-2232.
PMID |
| [9] | Craig E A, Marszalek J. 2017. How do J-proteins get Hsp70 to do so many different things. Trends Biochem Sci, 42(5): 355-368. |
| [10] | Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A. 2004. Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes Dev, 18(8): 926-936. |
| [11] |
Fang N, Xu R, Huang L J, Zhang B L, Duan P G, Li N, Luo Y H, Li Y H. 2016. SMALL GRAIN 11 controls grain size, grain number and grain yield in rice. Rice, 9(1): 64.
PMID |
| [12] | Galbiati F, Chiozzotto R, Locatelli F, Spada A, Genga A, Fornara F. 2016. Hd3a, RFT1 and Ehd1 integrate photoperiodic and drought stress signals to delay the floral transition in rice. Plant Cell Environ, 39(9): 1982-1993. |
| [13] | Gao H, Zheng X M, Fei G L, Chen J, Jin M N, Ren Y L, Wu W X, Zhou K N, Sheng P K, Zhou F, Jiang L, Wang J, Zhang X, Guo X P, Wang J L, Cheng Z J, Wu C Y, Wang H Y, Wan J M. 2013. Ehd4 encodes a novel and Oryza-genus-specific regulator of photoperiodic flowering in rice. PLoS Genet, 9(2): e1003281. |
| [14] |
Harris C J, Scheibe M, Wongpalee S P, Liu W L, Cornett E M, Vaughan R M, Li X Q, Chen W, Xue Y, Zhong Z H, Yen L D, Barshop W D, Rayatpisheh S, Gallego-Bartolome J, Groth M, Wang Z H, Wohlschlegel J A, Du J M, Rothbart S B, Butter F, Jacobsen S E. 2018. A DNA methylation reader complex that enhances gene transcription. Science, 362: 1182-1186.
PMID |
| [15] | Hayama R, Yokoi S, Tamaki S, Yano M, Shimamoto K. 2003. Adaptation of photoperiodic control pathways produces short- day flowering in rice. Nature, 422: 719-722. |
| [16] |
Hiei Y, Ohta S, Komari T, Kumashiro T. 1994. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J, 6(2): 271-282.
PMID |
| [17] |
Hori K, Matsubara K, Yano M. 2016. Genetic control of flowering time in rice: Integration of Mendelian genetics and genomics. Theor Appl Genet, 129(12): 2241-2252.
PMID |
| [18] | Huang X H, Kurata N, Wei X H, Wang Z X, Wang A H, Zhao Q, Zhao Y, Liu K Y, Lu H Y, Li W J, Guo Y L, Lu Y Q, Zhou C C, Fan D L, Weng Q J, Zhu C R, Huang T, Zhang L, Wang Y C, Feng L, Furuumi H, Kubo T, Miyabayashi T, Yuan X P, Xu Q, Dong G J, Zhan Q L, Li C Y, Fujiyama A, Toyoda A, Lu T T, Feng Q, Qian Q, Li J Y, Han B. 2012. A map of rice genome variation reveals the origin of cultivated rice. Nature, 490: 497-501. |
| [19] |
Jung C, Müller A E. 2009. Flowering time control and applications in plant breeding. Trends Plant Sci, 14(10): 563-573.
PMID |
| [20] | Kampinga H H, Craig E A. 2010. The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat Rev Mol Cell Biol, 11(8): 579-592. |
| [21] | Kim S L, Lee S, Kim H J, Nam H G, An G. 2007. OsMADS51 is a short-day flowering promoter that functions upstream of Ehd1, OsMADS14, and Hd3a. Plant Physiol, 145(4): 1484-1494. |
| [22] | Koo B H, Yoo S C, Park J W, Kwon C T, Lee B D, An G, Zhang Z Y, Li J J, Li Z C, Paek N C. 2013. Natural variation in OsPRR37 regulates heading date and contributes to rice cultivation at a wide range of latitudes. Mol Plant, 6(6): 1877-1888. |
| [23] | Lee S, Kim J, Han J J, Han M J, An G. 2004. SOC1/ AGL20) ortholog in rice. Plant J, 38(5): 754-764. |
| [24] | Li X F, Tian X J, He M L, Liu X X, Li Z Y, Tang J Q, Mei E Y, Xu M, Liu Y X, Wang Z Y, Guan Q J, Meng W, Fang J, Zhang J, Bu Q Y. 2022. bZIP71 delays flowering by suppressing Ehd1 expression in rice. J Integr Plant Biol, 64(7): 1352-1363. |
| [25] | Lin B L, Wang J S, Liu H C, Chen R W, Meyer Y, Barakat A, Delseny M. 2001. Genomic analysis of the Hsp70 superfamily in Arabidopsis thaliana. Cell Stress Chaperones, 6(3): 201-208. |
| [26] |
Luo Y, Fang B H, Wang W P, Yang Y, Rao L Q, Zhang C. 2019. Genome-wide analysis of the rice J-protein family: Identification, genomic organization, and expression profiles under multiple stresses. 3 Biotech, 9(10): 358.
PMID |
| [27] |
Matsubara K, Yamanouchi U, Wang Z X, Minobe Y, Izawa T, Yano M. 2008. Ehd2, a rice ortholog of the maize INDETERMINATE1 gene, promotes flowering by up-regulating Ehd1. Plant Physiol, 148(3): 1425-1435.
PMID |
| [28] | Matsubara K, Yamanouchi U, Nonoue Y, Sugimoto K, Wang Z X, Minobe Y, Yano M. 2011. Ehd3, encoding a plant homeodomain finger-containing protein, is a critical promoter of rice flowering. Plant J, 66(4): 603-612. |
| [29] |
Matsubara K, Ogiso-Tanaka E, Hori K, Ebana K, Ando T, Yano M. 2012. Natural variation in Hd17, a homolog of Arabidopsis ELF3 that is involved in rice photoperiodic flowering. Plant Cell Physiol, 53(4): 709-716.
PMID |
| [30] | Nemoto Y, Nonoue Y, Yano M, Izawa T. 2016. Hd1, a CONSTANS ortholog in rice, functions as an Ehd1 repressor through interaction with monocot-specific CCT-domain protein Ghd7. Plant J, 86(3): 221-233. |
| [31] | Ohta M, Takaiwa F. 2020. OsERdj7 is an ER-resident J-protein involved in ER quality control in rice endosperm. J Plant Physiol, 245: 153109. |
| [32] | Park M Y, Kim S Y. 2014. The Arabidopsis J protein AtJ1 is essential for seedling growth, flowering time control and ABA response. Plant Cell Physiol, 55(12): 2152-2163. |
| [33] | Peng L T, Shi Z Y, Li L, Shen G Z, Zhang J L. 2007. Ectopic expression of OsLFL1in rice represses Ehd1 by binding on its promoter. Biochem Biophys Res Commun, 360(1): 251-256. |
| [34] | Rajan V B V, D’Silva P. 2009. Arabidopsis thaliana J-class heat shock proteins: Cellular stress sensors. Funct Integr Genomics, 9(4): 433-446. |
| [35] | Roberts J A, Evan D, Mcmanus M T, Rose J. 2018. Photoreceptors and light signaling pathways in plants. Annu Plant Rev, 10(21): 107-131. |
| [36] |
Rogers S O, Bendich A J. 1985. Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Mol Biol, 5(2): 69-76.
PMID |
| [37] |
Senguttuvel P, Sravanraju N, Jaldhani V, Divya B, Beulah P, Nagaraju P, Manasa Y, Prasad A S H, Brajendra P, Gireesh C, Anantha M S, Suneetha K, Sundaram R M, Madhav M S, Tuti M D, Subbarao L V, Neeraja C N, Bhadana V P, Rao P R, Voleti S R, Subrahmanyam D. 2021. Evaluation of genotype by environment interaction and adaptability in lowland irrigated rice hybrids for grain yield under high temperature. Sci Rep, 11(1): 15825.
PMID |
| [38] | Shen L S, Kang Y G G, Liu L, Yu H. 2011. The J-domain protein J3 mediates the integration of flowering signals in Arabidopsis. Plant Cell, 23(2): 499-514. |
| [39] | Shen T T, Wang L, Shang C H, Zhen Y C, Fang Y L, Wei L L, Zhou T, Bai J T, Li B. 2022. The Arabidopsis J-protein AtDjC5 facilitates thermotolerance likely by aiding in the ER stress response. Int J Mol Sci, 23(21): 13134. |
| [40] |
Sjögren L, Floris M, Barghetti A, Völlmy F, Linding R E, Brodersen P. 2018. Farnesylated heat shock protein 40 is a component of membrane-bound RISC in Arabidopsis. J Biol Chem, 293(43): 16608-16622.
PMID |
| [41] |
Song Y H, Shim J S, Kinmonth-Schultz H A, Imaizumi T. 2015. Photoperiodic flowering: Time measurement mechanisms in leaves. Annu Rev Plant Biol, 66: 441-464.
PMID |
| [42] |
Sun C H, Chen D, Fang J, Wang P R, Deng X J, Chu C C. 2014. Understanding the genetic and epigenetic architecture in complex network of rice flowering pathways. Protein Cell, 5(12): 889-898.
PMID |
| [43] | Tamadaddi C, Sagar V, Verma A K, Afsal F, Sahi C. 2021. Expansion of the evolutionarily conserved network of J-domain proteins in the Arabidopsis mitochondrial import complex. Plant Mol Biol, 105(4): 385-403. |
| [44] |
Tamadaddi C, Verma A K, Zambare V, Vairagkar A, Diwan D, Sahi C. 2022. J-like protein family of Arabidopsis thaliana: The enigmatic cousins of J-domain proteins. Plant Cell Rep, 41(6): 1343-1355.
PMID |
| [45] |
Tamaki S, Matsuo S, Wong H L, Yokoi S, Shimamoto K. 2007. Hd3a protein is a mobile flowering signal in rice. Science, 316: 1033-1036.
PMID |
| [46] |
Tsuji H, Taoka K I, Shimamoto K. 2011. Regulation of flowering in rice: Two florigen genes, a complex gene network, and natural variation. Curr Opin Plant Biol, 14(1): 45-52.
PMID |
| [47] | Wang F H, Tang Z B, Wang Y, Fu J, Yang W B, Wang S X, Wang Y T, Bai T, Huang Z B, Yin H Q, Wang Z F. 2022. Leaf Mutant 7encoding heat shock protein OsHSP40 regulates leaf size in rice. Int J Mol Sci, 23(8): 4446. |
| [48] | Wei F J, Tsai Y C, Wu H P, Huang L T, Chen Y C, Chen Y F, Wu C C, Tseng Y T, Hsing Y I C. 2016. Both Hd1 and Ehd1 are important for artificial selection of flowering time in cultivated rice. Plant Sci, 242: 187-194. |
| [49] |
Wu C Y, You C J, Li C S, Long T, Chen G X, Byrne M E, Zhang Q F. 2008. RID1, encoding a Cys2/His2-type zinc finger transcription factor, acts as a master switch from vegetative to floral development in rice. Proc Natl Acad Sci USA, 105(35): 12915-12920.
PMID |
| [50] |
Xing Y Z, Zhang Q F. 2010. Genetic and molecular bases of rice yield. Annu Rev Plant Biol, 61: 421-442.
PMID |
| [51] | Xue W Y, Xing Y Z, Weng X Y, Zhao Y, Tang W J, Wang L, Zhou H J, Yu S B, Xu C G, Li X H, Zhang Q F. 2008. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet, 40(6): 761-767. |
| [52] | Yan W H, Wang P, Chen H X, Zhou H J, Li Q P, Wang C R, Ding Z H, Zhang Y S, Yu S B, Xing Y Z, Zhang Q F. 2011. A major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height, and heading date in rice. Mol Plant, 4(2): 319-330. |
| [53] | Yan W H, Liu H Y, Zhou X C, Li Q P, Zhang J, Lu L, Liu T M, Liu H J, Zhang C J, Zhang Z Y, Shen G J, Yao W, Chen H X, Yu S B, Xie W B, Xing Y Z. 2013. Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res, 23(7): 969-971. |
| [54] | Yang K Z, Xia C, Liu X L, Dou X Y, Wang W, Chen L Q, Zhang X Q, Xie L F, He L Y, Ma X, Ye D. 2009. A mutation in THERMOSENSITIVE MALE STERILE 1, encoding a heat shock protein with DnaJ and PDI domains, leads to thermosensitive gametophytic male sterility in Arabidopsis. Plant J, 57(5): 870-882. |
| [55] | Yang Y Q, Qin Y X, Xie C G, Zhao F Y, Zhao J F, Liu D F, Chen S Y, Fuglsang A T, Palmgren M G, Schumaker K S, Deng X W, Guo Y. 2010. The Arabidopsis chaperone J3 regulates the plasma membrane H+-ATPase through interaction with the PKS5 kinase. Plant Cell, 22(4): 1313-1332. |
| [56] |
Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T. 2000. Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. Plant Cell, 12(12): 2473-2484.
PMID |
| [57] |
Zhang H, Zhu S S, Liu T Z, Wang C M, Cheng Z J, Zhang X, Chen L P, Sheng P K, Cai M H, Li C N, Wang J C, Zhang Z, Chai J T, Zhou L, Lei C L, Guo X P, Wang J L, Wang J, Jiang L, Wu C Y, Wan J M. 2019. DELAYED HEADING DATE1 interacts with OsHAP5C/D, delays flowering time and enhances yield in rice. Plant Biotechnol J, 17(2): 531-539.
PMID |
| [58] |
Zhao H, Li J C, Yang L, Qin G, Xia C J, Xu X B, Su Y M, Liu Y M, Ming L C, Chen L L, Xiong L Z, Xie W B. 2021. An inferred functional impact map of genetic variants in rice. Mol Plant, 14(9): 1584-1599.
PMID |
| [59] | Zhao J M, Huang X, Ouyang X H, Chen W L, Du A P, Zhu L, Wang S G, Deng X W, Li S G. 2012. OsELF3-1, an ortholog of Arabidopsis early flowering 3, regulates rice circadian rhythm and photoperiodic flowering. PLoS One, 7(8): e43705. |
| [60] |
Zhou S R, Zhu S S, Cui S, Hou H G, Wu H Q, Hao B Y, Cai L, Xu Z, Liu L L, Jiang L, Wang H Y, Wan J M. 2021. Transcriptional and post-transcriptional regulation of heading date in rice. New Phytol, 230(3): 943-956.
PMID |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||