
Rice Science ›› 2019, Vol. 26 ›› Issue (4): 229-238.DOI: 10.1016/j.rsci.2018.08.008
• Orginal Article • Previous Articles Next Articles
Cheabu Sulaiman1, Panichawong Nat1, Rattanametta Prisana1, Wasuri Boonthong2, Kasemsap Poonpipope3, Arikit Siwaret1,4, Vanavichit Apichart1,4, Malumpong Chanate1( )
)
Received:2018-03-31
Accepted:2018-08-22
Online:2019-07-28
Published:2019-04-04
Cheabu Sulaiman, Panichawong Nat, Rattanametta Prisana, Wasuri Boonthong, Kasemsap Poonpipope, Arikit Siwaret, Vanavichit Apichart, Malumpong Chanate. Screening for Spikelet Fertility and Validation of Heat Tolerance in a Large Rice Mutant Population[J]. Rice Science, 2019, 26(4): 229-238.
Add to citation manager EndNote|Ris|BibTeX
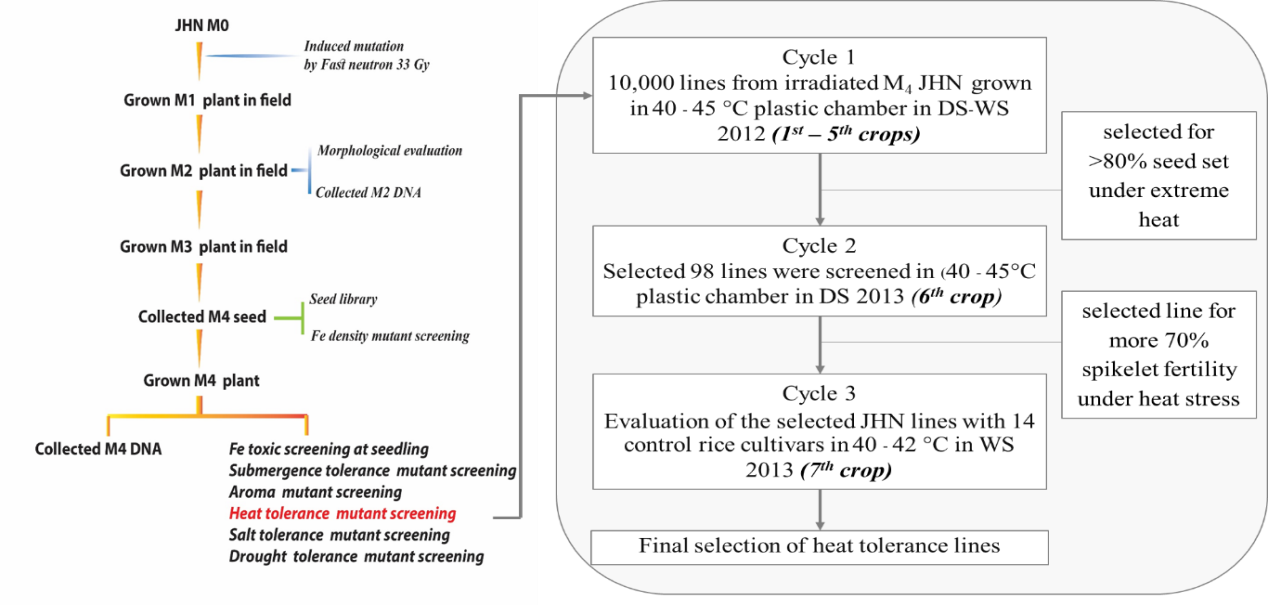
Supplemental Fig. 1 Schematic view of mutant screening for heat tolerance of a large Fast Neutron-treated population and the workflow of the present study, including screening and validation (Ruengphayak et al, 2015). In the first step, the heat tolerance as indicated by spikelet fertility was screened in 10,000 lines at high temperature. In the second step, lines were selected for >80% spikelet fertility (IRRI, 2013) under extreme heat. In the third part, the heat tolerance was validated by an evaluation of the selected JHN lines with rice cultivars.
| Experiment | Duration | Greenhouse | |||||
| Day time (6 h) | Night time | ||||||
| T (oC) | RH (%) | LI [µmol/(L·m2·s)] | T (oC) | RH (%) | |||
| Preliminary screening | April-December 2012 | 41.70 ± 1.90 | 56.52 ± 4.42 | 1 001.17 | 29.07 ± 2.28 | 82.18 ± 4.90 | |
| Repeat screening | April-May 2013 | 42.13 ± 1.15 | 61.51 ± 4.47 | 929.26 | 29.41 ± 1.10 | 86.34 ± 4.85 | |
| Validation screening | July-August 2013 | 41.44 ± 1.62 | 59.10 ± 3.44 | 997.34 | 28.35 ± 1.20 | 84.57 ± 5.86 | |
| Experiment | Duration | In field | |||||
| Day time (6 h) | Night time | ||||||
| T (oC) | RH (%) | LI [µmol/(L·m2·s)] | T (oC) | RH (%) | |||
| Preliminary screening | April-December 2012 | 35.42 ± 2.90 | 47.61 ± 7.76 | 1 667.13 | 28.68 ± 2.31 | 80.42 ± 6.49 | |
| Repeat screening | April-May 2013 | 35.24 ± 1.16 | 56.38 ± 5.81 | 1 468.89 | 27.97 ± 0.86 | 85.37 ± 7.01 | |
| Validation screening | July-August 2013 | 33.25 ± 1.26 | 55.37 ± 6.13 | 1 538.70 | 27.08 ± 0.95 | 83.45 ± 8.57 | |
Table 1 Temperature (T), relative humidity (RH) and light intensity (LI) for the screening experiments.
| Experiment | Duration | Greenhouse | |||||
| Day time (6 h) | Night time | ||||||
| T (oC) | RH (%) | LI [µmol/(L·m2·s)] | T (oC) | RH (%) | |||
| Preliminary screening | April-December 2012 | 41.70 ± 1.90 | 56.52 ± 4.42 | 1 001.17 | 29.07 ± 2.28 | 82.18 ± 4.90 | |
| Repeat screening | April-May 2013 | 42.13 ± 1.15 | 61.51 ± 4.47 | 929.26 | 29.41 ± 1.10 | 86.34 ± 4.85 | |
| Validation screening | July-August 2013 | 41.44 ± 1.62 | 59.10 ± 3.44 | 997.34 | 28.35 ± 1.20 | 84.57 ± 5.86 | |
| Experiment | Duration | In field | |||||
| Day time (6 h) | Night time | ||||||
| T (oC) | RH (%) | LI [µmol/(L·m2·s)] | T (oC) | RH (%) | |||
| Preliminary screening | April-December 2012 | 35.42 ± 2.90 | 47.61 ± 7.76 | 1 667.13 | 28.68 ± 2.31 | 80.42 ± 6.49 | |
| Repeat screening | April-May 2013 | 35.24 ± 1.16 | 56.38 ± 5.81 | 1 468.89 | 27.97 ± 0.86 | 85.37 ± 7.01 | |
| Validation screening | July-August 2013 | 33.25 ± 1.26 | 55.37 ± 6.13 | 1 538.70 | 27.08 ± 0.95 | 83.45 ± 8.57 | |
| Score | Spikelet fertility | Description |
| 1 | > 80% | Highly tolerant |
| 3 | 61%-80% | Tolerant |
| 5 | 41%-60% | Moderately tolerant |
| 7 | 11%-40% | Susceptible |
| 9 | < 11% | Highly susceptible |
Table 2 Heat tolerance scoring system in rice (IRRI, 2013).
| Score | Spikelet fertility | Description |
| 1 | > 80% | Highly tolerant |
| 3 | 61%-80% | Tolerant |
| 5 | 41%-60% | Moderately tolerant |
| 7 | 11%-40% | Susceptible |
| 9 | < 11% | Highly susceptible |
| Cultivar | Origin | Grain type | Special trait |
| Dawk Pa-yawm | Native rice | Long | Upland rice |
| Homcholasit | RGDU* | Long | submergence tolerance |
| Jao Hom Nil | RGDU | Medium | Wild Type |
| Pathum Thani 1 | Rice Department. | Long | Good quality |
| Pin Kaset 1 | RGDU | Long | High yield |
| Pin Kaset 2 | RGDU | Medium | High yield |
| Pin Kaset 3 | RGDU | Long | High yield |
| PSL2 | Rice Department. | Medium | Commercial cultivar |
| RD15 | Rice Department. | Medium | Adaptable to drought |
| RD33 | Rice Department. | Long | Blast resistance |
| Suphan Buri 1 | Rice Department. | Medium | Commercial cultivar |
| Thanyasirin | RGDU | Medium | Waxy rice |
| A5029 | RGDU | Long | Very long grain |
| Sinlek | RGDU | Long | Control for high temperature sensitivity |
| * RGDU; Rice gene discovery unit | |||
Supplemental Table 1 Origin, grain type and special traits of various rice cultivars used during the wet season of 2013.
| Cultivar | Origin | Grain type | Special trait |
| Dawk Pa-yawm | Native rice | Long | Upland rice |
| Homcholasit | RGDU* | Long | submergence tolerance |
| Jao Hom Nil | RGDU | Medium | Wild Type |
| Pathum Thani 1 | Rice Department. | Long | Good quality |
| Pin Kaset 1 | RGDU | Long | High yield |
| Pin Kaset 2 | RGDU | Medium | High yield |
| Pin Kaset 3 | RGDU | Long | High yield |
| PSL2 | Rice Department. | Medium | Commercial cultivar |
| RD15 | Rice Department. | Medium | Adaptable to drought |
| RD33 | Rice Department. | Long | Blast resistance |
| Suphan Buri 1 | Rice Department. | Medium | Commercial cultivar |
| Thanyasirin | RGDU | Medium | Waxy rice |
| A5029 | RGDU | Long | Very long grain |
| Sinlek | RGDU | Long | Control for high temperature sensitivity |
| * RGDU; Rice gene discovery unit | |||
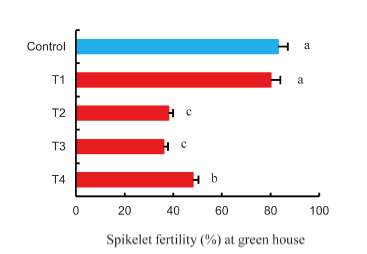
Fig. 1. Sensitivity of spikelet fertility to heat stress at 40 ºC to 45 ºC during the reproductive stages.T1, High temperature from panicle initiation to booting stages; T2, High temperature from panicle initiation to harvesting stages; T3, High temperature from booting to harvesting stages; T4, High temperature from flowering to harvesting stages. For the control, the plants were grown outside of the heat chamber under normal conditions. Different lowercase letters indicate significant difference at the 0.05 level. Bars are SE (n =15).
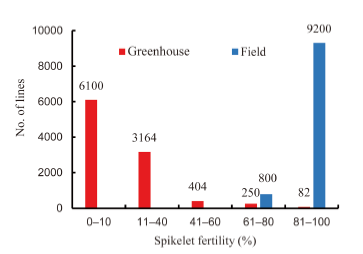
Fig. 2. Frequency distribution of 10 000 mutant plants and lines selected for spikelet fertility at 40 ºC to 45 ºC daytime temperature during the reproductive stage until harvesting.
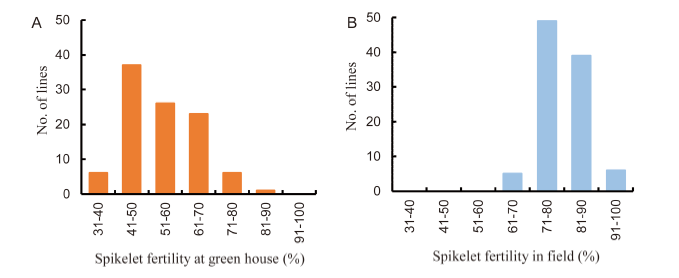
Fig. 3. Selection of rice mutants for heat tolerance based on spikelet fertility. A, Frequency of spikelet fertility in the greenhouse at high temperature (40 ºC to 45 ºC). B, Frequency of spikelet fertility in the field.
| Genotype | Spikelet fertility (%) | Panicle length (cm) | Filled grains per panicle | Total florets per panicle | |||||||
| Field | Greenhouse | Field | Greenhouse | Field | Greenhouse | Field | Greenhouse | ||||
| M9962 | 84.3 aA | 82.2 abA | 20.4 deA | 19.9 eA | 105.4 aA | 75.1 cdB | 119.7 aA | 104.4 bcB | |||
| M3181 | 79.5 abcdA | 75.1 cdeA | 22.3 cdA | 20.6 deA | 73.1 cdeA | 53.6 hiB | 92.4 cdefA | 83.7 fgA | |||
| M7988 | 75.4 deA | 74.3 deA | 20.5 deB | 23.5 bcA | 81.7 bcA | 53.1 hiB | 109.8 abA | 83.5 fgB | |||
| M8269 | 76.1 bcdA | 73.7 deA | 20.3 deA | 19.6 eA | 54.3 ghjA | 50.9 ijA | 84.6 efggA | 85.7 defgA | |||
| M8281 | 83.0 abA | 72.2 deB | 20.6 deA | 19.6 eA | 64.3 efgA | 54.4 ghiA | 100.4 bcdA | 85.7 defgA | |||
| M8372 | 76.6 bcdA | 69.8 eB | 20.2 deA | 19.8 eA | 62.7 fghA | 40.7 jB | 94.9 bcdefA | 77.1 gB | |||
| M7766 | 69.6 eA | 16.2 hB | 24.7 abA | 23.3 bcA | 73.1 cdeA | 9.2 kB | 107.9 abA | 101.8 bcA | |||
| JHN (wild type) | 82.3 abcA | 39.3 gB | 24.9 abA | 21.9 cdeA | 85.6 bA | 40.7 jB | 103.0 bcA | 99.2 bcdeA | |||
| Sinlek (control) | 62.1 fA | 10.4 hB | 25.9 aA | 24.1 abcA | 70.1 defA | 17.8 kB | 109.5 abA | 108.6 abA | |||
| Genotype (G) | 2.50* | 0.44* | 4.37* | 4.68* | |||||||
| Treatment (T) | 4.94* | 1.67* | 7.26* | 10.54* | |||||||
| G × T | 6.98* | 2.37 ns | 10.27* | 14.91 | |||||||
| Different lowercase letters followed the values in the column are significantly different at P < 0.05 by the Tukey’s least significant difference (LSD) test. Different uppercase letters in each row for the same trait indicate significant difference at P < 0.05 by the Tukey’s LSD test. | |||||||||||
Table 3 Effects of heat stress on panicle traits in the candidate heat tolerant lines.
| Genotype | Spikelet fertility (%) | Panicle length (cm) | Filled grains per panicle | Total florets per panicle | |||||||
| Field | Greenhouse | Field | Greenhouse | Field | Greenhouse | Field | Greenhouse | ||||
| M9962 | 84.3 aA | 82.2 abA | 20.4 deA | 19.9 eA | 105.4 aA | 75.1 cdB | 119.7 aA | 104.4 bcB | |||
| M3181 | 79.5 abcdA | 75.1 cdeA | 22.3 cdA | 20.6 deA | 73.1 cdeA | 53.6 hiB | 92.4 cdefA | 83.7 fgA | |||
| M7988 | 75.4 deA | 74.3 deA | 20.5 deB | 23.5 bcA | 81.7 bcA | 53.1 hiB | 109.8 abA | 83.5 fgB | |||
| M8269 | 76.1 bcdA | 73.7 deA | 20.3 deA | 19.6 eA | 54.3 ghjA | 50.9 ijA | 84.6 efggA | 85.7 defgA | |||
| M8281 | 83.0 abA | 72.2 deB | 20.6 deA | 19.6 eA | 64.3 efgA | 54.4 ghiA | 100.4 bcdA | 85.7 defgA | |||
| M8372 | 76.6 bcdA | 69.8 eB | 20.2 deA | 19.8 eA | 62.7 fghA | 40.7 jB | 94.9 bcdefA | 77.1 gB | |||
| M7766 | 69.6 eA | 16.2 hB | 24.7 abA | 23.3 bcA | 73.1 cdeA | 9.2 kB | 107.9 abA | 101.8 bcA | |||
| JHN (wild type) | 82.3 abcA | 39.3 gB | 24.9 abA | 21.9 cdeA | 85.6 bA | 40.7 jB | 103.0 bcA | 99.2 bcdeA | |||
| Sinlek (control) | 62.1 fA | 10.4 hB | 25.9 aA | 24.1 abcA | 70.1 defA | 17.8 kB | 109.5 abA | 108.6 abA | |||
| Genotype (G) | 2.50* | 0.44* | 4.37* | 4.68* | |||||||
| Treatment (T) | 4.94* | 1.67* | 7.26* | 10.54* | |||||||
| G × T | 6.98* | 2.37 ns | 10.27* | 14.91 | |||||||
| Different lowercase letters followed the values in the column are significantly different at P < 0.05 by the Tukey’s least significant difference (LSD) test. Different uppercase letters in each row for the same trait indicate significant difference at P < 0.05 by the Tukey’s LSD test. | |||||||||||
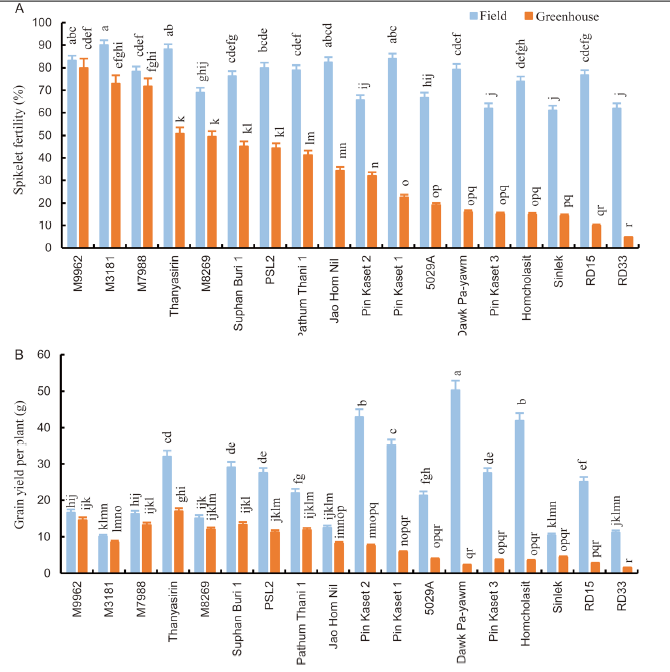
Fig. 4. Effects of high temperature stress during the booting to harvesting stages on the spikelet fertility (A) and grain yield per plant (B). Values (Mean ± SE, n = 15) with the same lowercase letters are not significantly different at the 0.05 level.
| Genotype | 100-grain weight (g) | DF (%) | No. of filled grains per panicle | DF (%) | Panicle weight (g) | DF (%) | |||
| In field | Greenhouse | In field | Greenhouse | In field | Greenhouse | ||||
| M9962 | 2.10 uA | 1.93 vB | 8 | 101.0 defgA | 103.0 defgA | - | 3.45 fghiA | 2.82 jklmB | 18.3 |
| M3181 | 2.49 noA | 2.31 qrB | 7 | 68.3 jklmnA | 50.9 mnoA | 25 | 2.42 klmnopA | 1.65 rsB | 31.8 |
| M7988 | 2.27 rstA | 2.09 uB | 8 | 77.3 hijklA | 79.9 ghijklA | - | 2.85 ijklA | 2.43 klmnoA | 14.7 |
| Thanyasirin | 2.57 mnA | 2.22 rstB | 14 | 143.2 abA | 104.1 defB | 27 | 5.76 aA | 3.82 efgB | 33.7 |
| M8269 | 2.44 opA | 2.31 qrB | 5 | 74.1 ijklmA | 58.2 lmnoA | 37 | 2.75 jklmA | 2.34 lmnopqA | 14.9 |
| Suphan Buri 1 | 2.67 jkA | 2.43 opB | 9 | 115.0 cdeA | 66.2 klmnoB | 42 | 4.80 bcA | 2.88 ijklB | 40.0 |
| PSL2 | 3.17 cA | 2.30 qrsB | 27 | 92.4 efghiA | 60.8 lmoB | 33 | 4.40 cdeA | 2.64 jklmnB | 40.0 |
| Pathum Thani 1 | 2.85 ghA | 2.46 opB | 14 | 91.0 efghiA | 53.1 mnoB | 41 | 3.62 fghA | 2.11 nopqrsB | 41.7 |
| Jao Hom Nil | 2.52 mnoA | 2.21 stB | 12 | 85.3 fghijkA | 46.6 nopB | 45 | 2.99 hijkA | 2.08 nopqrsB | 30.4 |
| Pin Kaset 2 | 2.91 fgA | 2.67 jkB | 8 | 134.0 abcA | 44.3 opqB | 67 | 5.03 bA | 2.54 klmnB | 49.5 |
| Pin Kaset 1 | 3.04 deA | 2.86 ghB | 6 | 114.1 cdeA | 26.9 pqrB | 76 | 4.50 bcdA | 2.06 nopqrsB | 54.2 |
| A5029 | 3.45 aA | 2.98 efB | 14 | 87.2 fghikA | 20.2 rB | 77 | 4.83 bcA | 1.83 pqrsB | 62.1 |
| Dawk Payawm | 2.77 hiA | 2.37 pqB | 14 | 154.3 aA | 24.8 pqrB | 83 | 4.90 bcA | 2.21 mnopqrB | 54.8 |
| Pin Kaset 3 | 3.12 cdA | 2.78 hiB | 11 | 120.0 bcdA | 15.4 rB | 87 | 4.71 bcA | 1.74 qrsB | 63.1 |
| Homcholasit | 3.28 bA | 2.61 klmB | 20 | 99.0 defghA | 22.3 qrB | 77 | 3.70 efgA | 1.89 opqrsB | 48.9 |
| Sinlek | 2.98 efA | 2.66 jklB | 11 | 69.6 ijklmnA | 17.8 rB | 74 | 3.19 ghijA | 1.79 pqrsB | 43.9 |
| RD15 | 2.64 klA | 2.48 noB | 6 | 122.0 bcdA | 15.2 rB | 88 | 3.90 defA | 1.53 sB | 60.8 |
| RD33 | 2.74 ijA | 2.18 tuB | 20 | 73.4 ijklmA | 7.4 rB | 89 | 2.43 klmnoA | 1.59 sB | 34.5 |
| Genotype (G) | 0.07* | 19.9* | 2.04* | ||||||
| Treatment (T) | 0.02* | 5.6* | 0.14* | ||||||
| G × T | 0.09* | 23.9* | 0.62* | ||||||
| DF, Percentage decrease from the field. Different lowercase letters followed the values in the column are significantly different at P < 0.05 by the Tukey’s least significant difference (LSD) test. Different uppercase letters in the each row for the same trait indicate significant difference at P < 0.05 by the Tukey’s LSD test. | |||||||||
Table 4 Effects of heat stress at the booting stages on the 100-grain weight, filled grains per panicle and panicle weight in different rice cultivars in wet season 2013.
| Genotype | 100-grain weight (g) | DF (%) | No. of filled grains per panicle | DF (%) | Panicle weight (g) | DF (%) | |||
| In field | Greenhouse | In field | Greenhouse | In field | Greenhouse | ||||
| M9962 | 2.10 uA | 1.93 vB | 8 | 101.0 defgA | 103.0 defgA | - | 3.45 fghiA | 2.82 jklmB | 18.3 |
| M3181 | 2.49 noA | 2.31 qrB | 7 | 68.3 jklmnA | 50.9 mnoA | 25 | 2.42 klmnopA | 1.65 rsB | 31.8 |
| M7988 | 2.27 rstA | 2.09 uB | 8 | 77.3 hijklA | 79.9 ghijklA | - | 2.85 ijklA | 2.43 klmnoA | 14.7 |
| Thanyasirin | 2.57 mnA | 2.22 rstB | 14 | 143.2 abA | 104.1 defB | 27 | 5.76 aA | 3.82 efgB | 33.7 |
| M8269 | 2.44 opA | 2.31 qrB | 5 | 74.1 ijklmA | 58.2 lmnoA | 37 | 2.75 jklmA | 2.34 lmnopqA | 14.9 |
| Suphan Buri 1 | 2.67 jkA | 2.43 opB | 9 | 115.0 cdeA | 66.2 klmnoB | 42 | 4.80 bcA | 2.88 ijklB | 40.0 |
| PSL2 | 3.17 cA | 2.30 qrsB | 27 | 92.4 efghiA | 60.8 lmoB | 33 | 4.40 cdeA | 2.64 jklmnB | 40.0 |
| Pathum Thani 1 | 2.85 ghA | 2.46 opB | 14 | 91.0 efghiA | 53.1 mnoB | 41 | 3.62 fghA | 2.11 nopqrsB | 41.7 |
| Jao Hom Nil | 2.52 mnoA | 2.21 stB | 12 | 85.3 fghijkA | 46.6 nopB | 45 | 2.99 hijkA | 2.08 nopqrsB | 30.4 |
| Pin Kaset 2 | 2.91 fgA | 2.67 jkB | 8 | 134.0 abcA | 44.3 opqB | 67 | 5.03 bA | 2.54 klmnB | 49.5 |
| Pin Kaset 1 | 3.04 deA | 2.86 ghB | 6 | 114.1 cdeA | 26.9 pqrB | 76 | 4.50 bcdA | 2.06 nopqrsB | 54.2 |
| A5029 | 3.45 aA | 2.98 efB | 14 | 87.2 fghikA | 20.2 rB | 77 | 4.83 bcA | 1.83 pqrsB | 62.1 |
| Dawk Payawm | 2.77 hiA | 2.37 pqB | 14 | 154.3 aA | 24.8 pqrB | 83 | 4.90 bcA | 2.21 mnopqrB | 54.8 |
| Pin Kaset 3 | 3.12 cdA | 2.78 hiB | 11 | 120.0 bcdA | 15.4 rB | 87 | 4.71 bcA | 1.74 qrsB | 63.1 |
| Homcholasit | 3.28 bA | 2.61 klmB | 20 | 99.0 defghA | 22.3 qrB | 77 | 3.70 efgA | 1.89 opqrsB | 48.9 |
| Sinlek | 2.98 efA | 2.66 jklB | 11 | 69.6 ijklmnA | 17.8 rB | 74 | 3.19 ghijA | 1.79 pqrsB | 43.9 |
| RD15 | 2.64 klA | 2.48 noB | 6 | 122.0 bcdA | 15.2 rB | 88 | 3.90 defA | 1.53 sB | 60.8 |
| RD33 | 2.74 ijA | 2.18 tuB | 20 | 73.4 ijklmA | 7.4 rB | 89 | 2.43 klmnoA | 1.59 sB | 34.5 |
| Genotype (G) | 0.07* | 19.9* | 2.04* | ||||||
| Treatment (T) | 0.02* | 5.6* | 0.14* | ||||||
| G × T | 0.09* | 23.9* | 0.62* | ||||||
| DF, Percentage decrease from the field. Different lowercase letters followed the values in the column are significantly different at P < 0.05 by the Tukey’s least significant difference (LSD) test. Different uppercase letters in the each row for the same trait indicate significant difference at P < 0.05 by the Tukey’s LSD test. | |||||||||
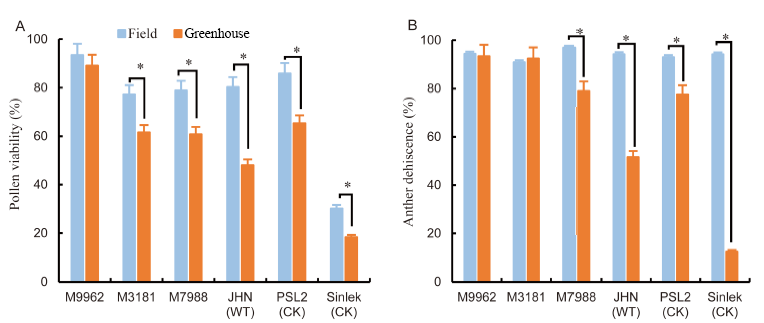
Fig. 5. Pollen viability (A) and anther dehiscence (B) under field conditions and at high temperature for the six rice genotypes. *, Significant at the 0.05 level.
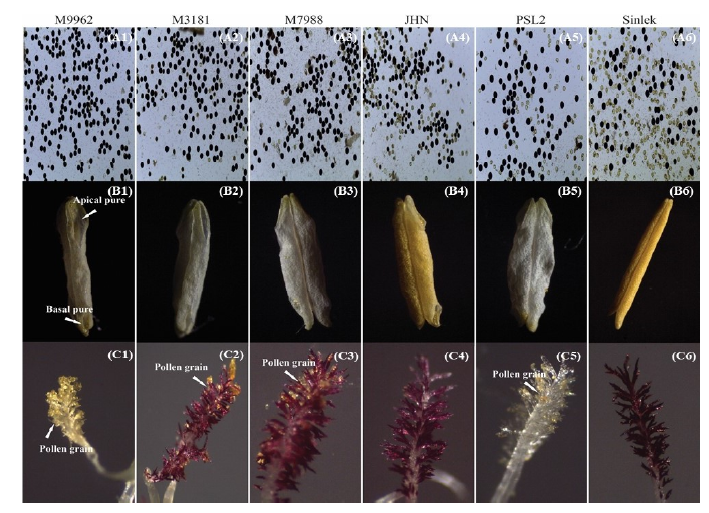

Fig. 6. Images of six cultivars showing the pollen viability (A1-A6), anther dehiscence (apical and basal pore) (B1-B6) and pollen number on the stigma (C1-C6) under high temperature conditions.
| [1] | Ahloowalia B S, Maluszynski M, Nichterlein K.2004. Global impact of mutation-derived varieties.Euphytica, 135(2): 187-204. |
| [2] | Baker J T, Allen Jr L H, Boote K J.1992. Temperature effects on rice at elevated CO2 concentration.J Exp Bot, 43: 959-964. |
| [3] | Chakhonkaen S, Pitnjam K, Saisuk W, Ukoskit K, Muangprom A.2012. Genetic structure of Thai rice and rice accessions obtained from the International Rice Research Institute.Rice, 5(1): 19. |
| [4] | Chumpolsri W, Wijit N, Boontakham P, Nimmanpipug P, Sookwong P, Luangkamin S, Wongpornchai S.2015. Variation of terpenoid flavor odorants in bran of some black and white rice varieties analyzed by GC × GC-MS.J Food Nutr Res, 3(2): 114-120. |
| [5] | Counce P A, Keisling T C, Mitchell A J.2000. A uniform, objective, and adaptive system for expressing rice development.Crop Sci, 40(2): 436-443. |
| [6] | Endo M, Tsuchiya T, Hamada K, Kawamura S, Yano K, Ohshima M, Higashitani A, Watanabe M, Kawagishi-Kobayashi M.2009. High temperatures cause male sterility in rice plants with transcriptional alterations during pollen development.Plant Cell Physiol, 50(11): 1911-1922. |
| [7] | González-Schain N, Dreni L, Lawas L M F, Galbiati M, Colombo L, Heuer S, Jagadish K S V, Kater M M.2015. Genome-wide transcriptome analysis during anthesis reveals new insights into the molecular basis of heat stress responses in tolerant and sensitive rice varieties.Plant Cell Physiol, 57(1): 57-68. |
| [8] | Huang L Y, Sun Y, Peng S B, Wang F.2016. Genotypic differences ofjaponica rice responding to high temperature in China. Agron J, 108(2): 626-636. |
| [9] | IPCC. 2014. Climate Change 2013: The Physical Science Basis. Cambridge, and New York: Cambridge University Press. |
| [10] | IRRI.2013. Standard Evaluation System for Rice (SES). Manila, the Philippines:International Rice Research Institute. 5: 35-36. |
| [11] | Ishimaru T, Hirabayashi H, Ida M, Takai T, San-Oh Y A, Yoshinaga S, Ando I, Ogawa T, Kondo M.2010. A genetic resource for early-morning flowering trait of wild riceOryza officinalis to mitigate high temperature-induced spikelet sterility at anthesis. Ann Bot, 106(3): 515-520. |
| [12] | Jagadish S V K, Craufurd P Q, Wheeler T R.2007. High temperature stress and spikelet fertility in rice (Oryza sativa L.). J Exp Bot, 58(7): 1627-1635. |
| [13] | Jagadish S V K, Craufurd P Q, Wheeler T R.2008. Phenotyping parents of mapping populations of rice for heat tolerance during anthesis.Crop Sci, 48(3): 1140-1146. |
| [14] | Jagadish S V K, Muthurajan R, Oane R, Wheeler T R, Heuer S, Bennett J, Craufurd P Q.2010. Physiological and proteomic approaches to address heat tolerance during anthesis in rice (Oryza sativa L.). J Exp Bot, 61(1): 143-156. |
| [15] | Kobayashi K, Matsui T, Murata Y, Yamamoto M.2011. Percentage of dehisced thecae and length of dehiscence control pollination stability of rice cultivars at high temperatures.Plant Prod Sci, 14(2): 89-95. |
| [16] | Lee I S, Kim D S, Lee S J, Song H S, Lim Y P, Lee Y I.2003. Selection and characterizations of radiation-induced salinity- tolerant lines in rice.Breeding Sci, 53(4): 313-318. |
| [17] | Manigbas N L, Lambio L A F, Madrid L B, Cardenas C C.2014. Germplasm innovation of heat tolerance in rice for irrigated lowland conditions in the Philippines.Rice Sci, 21(3): 162-169. |
| [18] | Maruyama A, Weerakoon W M W, Wakiyama Y, Ohba K.2013. Effects of increasing temperatures on spikelet fertility in different rice cultivars based on temperature gradient chamber experiments.J Agron Crop Sci, 199(6): 416-423. |
| [19] | Masuduzzaman A S M, Ahmad H U, Haque M, Ahmed M M E.2016. Evaluation of rice lines tolerant to heat during flowering stage.J Rice Res, 4: 170. |
| [20] | Matsui T, Namuco O S, Ziska L H, Horie T.1997. Effects of high temperature and CO2 concentration on spikelet sterility inindica rice. Field Crops Res, 51(3): 213-219. |
| [21] | Matsui T, Omasa K, Horie T.2000. High temperature at flowering inhibits swelling of pollen grains, a driving force for thecae dehiscence in rice (Oryza sativa L.). Plant Prod Sci, 3(4): 430-434. |
| [22] | Matsui T, Omasa K, Horie T.2001. The difference in sterility due to high temperatures during the flowering period amongjaponica-rice varieties. Plant Prod Sci, 4(2): 90-93. |
| [23] | Matsui T, Omasa K.2002. Rice (Oryza sativa L.) cultivars tolerant to high temperature at flowering: Anther characteristics. Ann Bot, 89(6): 683-687. |
| [24] | Miyagawa S.2001. Dynamics of rainfed lowland rice varieties in north-east Thailand. In: Saxena K G. Small-Scale Livelihoods and Natural Resource Management in Marginal Areas ofMonsoon Asia. Cornell University. |
| [25] | Mohapatra T, Robin S, Sarla N, Sheshashayee M, Singh A K, Singh K, Singh N K, Amitha Mithra S V, Sharma R P.2014. EMS induced mutants of upland rice variety Nagina 22: Generation and characterization.Proc Ind Nat Sci Acad, 80(1): 163-172. |
| [26] | Moung-ngam P.2016. Evaluation of Heat Tolerance in Rice Germplasm. [Master thesis]. Bangkok, Thailand:Kasetsart University. |
| [27] | Moya T B, Ziska L H, Namuco O S, Olszyk D.1998. Growth dynamics and genotypic variation in tropical, field-grown paddy rice (Oryza sativa L.) in response to increasing carbon dioxide and temperature. Global Change Biol, 4(6): 645-656. |
| [28] | Nakagawa H, Horie T, Matsui T.2003. Effects of Climate Change on Rice Production and Adaptive Technologies. Manila,the Philippines: International Rice Research Institute: 635-658. |
| [29] | Napasintuwong O.2012. Survey of recent innovations in aromatic rice. In: The 131st EAAE Seminar ‘Innovation for Agricultural Competitiveness and Sustainability of Rural Areas’. September 18-19, 2012. Prague,Czech Republic. |
| [30] | Peng S, Huang J, Sheehy J E, Laza R C, Visperas R M, Zhong X, Centeno G S, Khush G S, Cassman K G.2004. Rice yields decline with higher night temperature from global warming.Proc Natl Acad Sci USA, 101: 9971-9975. |
| [31] | Poli Y, Basava R K, Panigrahy M, Vinukonda V P, Dokula N R, Voleti S R, Desiraju S, Neelamraju S.2013. Characterization of a Nagina 22 rice mutant for heat tolerance and mapping of yield traits.Rice, 6(1): 36. |
| [32] | Prasad P V V, Boote K J, Allen L H, Sheehy J E, Thomas J M G.2006. Species, ecotype and cultivar differences in spikelet fertility and harvest index of rice in response to high temperature stress.Field Crops Res, 95: 398-411. |
| [33] | Prasanth V V, Basava K R, Babu M S, Venkata Tripura V G N, Rama Devi S J S, Mangrauthia S K, Voleti S R, Sarla N.2016. Field level evaluation of rice introgression lines for heat tolerance and validation of markers linked to spikelet fertility.Physiol Mol Biol Plants, 22(2): 179-192. |
| [34] | Rang Z W, Jagadish S V K, Zhou Q M, Craufurd P Q, Heuer S.2011. Effect of high temperature and water stress on pollen germination and spikelet fertility in rice.Environ Exp Bot, 70(1): 58-65. |
| [35] | Ruengphayak S, Ruanjaichon V, Saensuk C, Phromphan S, Tragoonrung S, Kongkachuichai R, Vanavichit A.2015. Forward screening for seedling tolerance to Fe toxicity reveals a polymorphic mutation in ferric chelate reductase in rice.Rice, 8(1): 3. |
| [36] | Saleem M Y, Mukhtar Z, Cheema A A, Atta B M.2005. Induced mutation andin vitro techniques as a method to induce salt tolerance in Basmati rice(Oryza sativa L.). Int J Environ Sci Technol, 2(2): 141-145. |
| [37] | Satake T, Yoshida S.1978. High temperature-induced sterility inindica rices at flowering. Jpn J Crop Sci, 47(1): 6-17. |
| [38] | Shah F, Huang J L, Cui K H, Nie L M, Shah T, Chen C, Wang K.2011. Impact of high-temperature stress on rice plant and its traits related to tolerance.J Agric Sci, 149(5): 545-556. |
| [39] | Shi W J, Ishimaru T, Gannaban R B, Oane W, Jagadish S V K.2015. Popular rice (L.) cultivars show contrasting responses to heat stress at gametogenesis and anthesis.Crop Sci, 55(2): 589-596. |
| [40] | Sukkeoa S, Rerkasemb B, Jamjoda S.2017. Heat tolerance in Thai rice varieties.Sci Asia, 43(2): 61-69. |
| [41] | Tenorio F A, Ye C, Redoña E, Sierra S, Laza M, Argayoso M A2013. Screening rice genetic resources for heat tolerance.SABRAO J Breeding Genet, 45(3): 371-381. |
| [42] | Wassmann R, Jagadish S V K, Sumfleth K, Pathak H, Howell G, Ismail A, Serraj R, Redona E, Singh R K, Heuer S.2009. Regional vulnerability of climate change impacts on Asian rice production and scope for adaptation.Adv Agron, 102: 91-133. |
| [43] | Wu Y J, Chen Y, Wang J, Zhu C X, Xu B L.2006. RAPD analysis of jasmine rice-specific genomic structure.Genome, 49(6): 716-719. |
| [44] | Ye C R, Argayoso M A, Redona E D, Sierra S N, Laza M A, Dilla C J, Mo Y J, Thomson M J, Chin J, Delaviña C B, Diaz G Q, Hernandez J E.2012. Mapping QTL for heat tolerance at flowering stage in rice using SNP markers.Plant Breeding, 131(1): 33-41. |
| [45] | Ziska L H, Manalo P A, Ordonez R A.1996. Intraspecific variation in the response of rice (Oryza sativa L.) to increased CO2 and temperature: Growth and yield response of 17 cultivars. J Exp Bot, 47(9): 1353-1359. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | XIA Xiaodong, ZHANG Xiaobo, WANG Zhonghao, CHENG Benyi, Sun Huifeng, XU Xia, GONG Junyi, YANG Shihua, WU Jianli, SHI Yongfeng, XU Rugen. Mapping and Functional Analysis of LE Gene in a Lethal Etiolated Rice Mutant at Seedling Stage [J]. Rice Science, 2023, 30(6): 13-. |
| [7] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [8] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [9] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [10] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [11] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [12] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [13] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [14] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [15] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||