
Rice Science ›› 2019, Vol. 26 ›› Issue (4): 239-247.DOI: 10.1016/j.rsci.2019.01.004
• Orginal Article • Previous Articles Next Articles
Donde Ravindra1, Kumar Jitendra1, Gouda Gayatri1, Kumar Gupta Manoj3, Mukherjee Mitadru1, Yasin Baksh Sk1, Mahadani Pradosh, Kumar Sahoo Khirod2, Behera Lambodar1, Kumar Dash Sushanta1( )
)
Received:2018-11-14
Accepted:2019-01-14
Online:2019-07-28
Published:2019-04-04
Donde Ravindra, Kumar Jitendra, Gouda Gayatri, Kumar Gupta Manoj, Mukherjee Mitadru, Yasin Baksh Sk, Mahadani Pradosh, Kumar Sahoo Khirod, Behera Lambodar, Kumar Dash Sushanta. Assessment of Genetic Diversity of Drought Tolerant and Susceptible Rice Genotypes Using Microsatellite Markers[J]. Rice Science, 2019, 26(4): 239-247.
Add to citation manager EndNote|Ris|BibTeX
| No. | Genotypes | Eco Types | Tolerance to stresses | Other information |
| 1 | Swarna sub1 | Shallow low land | Tolerant to submergence and susceptible to drought | Popular variety |
| 2 | IR64 sub1 | Irrigated | Tolerant to Submergence and susceptible to drought | - |
| 3 | FR13A | Shallow low land | Tolerant to Submergence and susceptible to drought | International submergence tolerance donor |
| 4 | CR143-2-2 | Upland | Drought Tolerance | Drought tolerant check |
| 5 | N 22 | Upland | Drought Tolerance | Drought and heat stress check |
| 6 | Brahamanakhi | Upland/Medium land | Drought Tolerance | Landrace |
| 7 | Satyabhama | Upland | Drought Tolerance | Released variety |
| 8 | IR 20 | Irrigated | Drought susceptible | International Check |
| 9 | Nerica1 | Upland/ Aerobic | Drought Tolerance | Suitable for Africa |
| 10 | Azucena | Upland | Drought Tolerance | Tropical japonica |
| 11 | Curinga | Upland | Drought Tolerance | Tropical japonica |
| 12 | MER20 | Upland | Drought Tolerance | CSSLs/BC4F4 of O. meridionalis/ Curinga |
| 13 | RUF44 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| 14 | RUF16 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| 15 | RUF48 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| 16 | RUF13 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
Supplemental Table 1 The list of drought tolerant and susceptible rice genotypes used for the assessment of genetic diversity.
| No. | Genotypes | Eco Types | Tolerance to stresses | Other information |
| 1 | Swarna sub1 | Shallow low land | Tolerant to submergence and susceptible to drought | Popular variety |
| 2 | IR64 sub1 | Irrigated | Tolerant to Submergence and susceptible to drought | - |
| 3 | FR13A | Shallow low land | Tolerant to Submergence and susceptible to drought | International submergence tolerance donor |
| 4 | CR143-2-2 | Upland | Drought Tolerance | Drought tolerant check |
| 5 | N 22 | Upland | Drought Tolerance | Drought and heat stress check |
| 6 | Brahamanakhi | Upland/Medium land | Drought Tolerance | Landrace |
| 7 | Satyabhama | Upland | Drought Tolerance | Released variety |
| 8 | IR 20 | Irrigated | Drought susceptible | International Check |
| 9 | Nerica1 | Upland/ Aerobic | Drought Tolerance | Suitable for Africa |
| 10 | Azucena | Upland | Drought Tolerance | Tropical japonica |
| 11 | Curinga | Upland | Drought Tolerance | Tropical japonica |
| 12 | MER20 | Upland | Drought Tolerance | CSSLs/BC4F4 of O. meridionalis/ Curinga |
| 13 | RUF44 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| 14 | RUF16 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| 15 | RUF48 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| 16 | RUF13 | Upland | Drought Tolerance | CSSLs /BC4F4 of O. rufipogon/ Curinga |
| Marker | Na | Np | Nu | Fla | Fha | PIC | Marker | Na | Np | Nu | Fla | Fha | PIC |
| RM315 | 2 | 2 | 0 | 1 | 1 | 0.49 | RM596 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM472 | 2 | 2 | 1 | 0 | 1 | 0.12 | RM512 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM302 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM179 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM431 | 2 | 2 | 0 | 1 | 1 | 0.34 | RM277 | 1 | 0 | 0 | 0 | 2 | 0.75 |
| RM212 | 2 | 2 | 0 | 0 | 2 | 0.75 | RM313 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM543 | 2 | 2 | 0 | 0 | 2 | 0.68 | RM83 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM259 | 2 | 2 | 0 | 1 | 1 | 0.53 | RM101 | 2 | 2 | 0 | 1 | 1 | 0.53 |
| RM488 | 2 | 2 | 1 | 0 | 1 | 0.12 | RM309 | 2 | 2 | 0 | 0 | 2 | 0.75 |
| RM521 | 2 | 2 | 0 | 0 | 2 | 0.75 | RM28130 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM526 | 2 | 2 | 0 | 0 | 2 | 0.75 | RM28050 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM555 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28089 | 2 | 2 | 1 | 0 | 1 | 1.00 |
| RM530 | 2 | 2 | 0 | 1 | 1 | 0.96 | RM244 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM279 | 3 | 3 | 0 | 2 | 1 | 0.98 | RM28067 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM416 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28088 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM22 | 2 | 2 | 0 | 1 | 1 | 0.96 | RM28099 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM16030 | 2 | 2 | 0 | 1 | 1 | 0.90 | RM28078 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM60 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM3349 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM15780 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28079 | 2 | 2 | 0 | 1 | 1 | 0.96 |
| RM537 | 2 | 2 | 1 | 0 | 1 | 0.12 | RM28082 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM252 | 3 | 3 | 0 | 0 | 3 | 0.86 | RM28048 | 2 | 2 | 0 | 0 | 2 | 0.75 |
| RM136 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28069 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM527 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28088 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM528 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM1261 | 2 | 2 | 0 | 1 | 1 | 0.96 |
| RM5371 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28075 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM541 | 2 | 2 | 1 | 0 | 1 | 1.00 | RM28051 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM276 | 3 | 3 | 1 | 0 | 2 | 1.00 | RM28050 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM210 | 3 | 3 | 0 | 1 | 2 | 0.34 | RM12091 | 2 | 2 | 0 | 1 | 1 | 0.23 |
| RM339 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28090 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM337 | 2 | 2 | 0 | 0 | 2 | 0.61 | RM28059 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM25 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28095 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM464 | 2 | 2 | 0 | 0 | 2 | 0.00 | Total | 95 | 60 | 6 | 13 | 77 | 41.76 |
| RM566 | 2 | 2 | 0 | 0 | 2 | 0.81 | Mean | 1.5 | 1.0 | 0.1 | 0.2 | 1.2 | 0.66 |
| RM24390 | 1 | 0 | 0 | 0 | 1 | 0.00 |
Table 1 Number of alleles (Na), number of polymorphic alleles (Np), unique allele (Nu), low-frequency allele (Fla), high-frequency allele (Fha) and polymorphism information content (PIC) for 63 simple sequence repeats (SSRs) in 16 rice genotypes.
| Marker | Na | Np | Nu | Fla | Fha | PIC | Marker | Na | Np | Nu | Fla | Fha | PIC |
| RM315 | 2 | 2 | 0 | 1 | 1 | 0.49 | RM596 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM472 | 2 | 2 | 1 | 0 | 1 | 0.12 | RM512 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM302 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM179 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM431 | 2 | 2 | 0 | 1 | 1 | 0.34 | RM277 | 1 | 0 | 0 | 0 | 2 | 0.75 |
| RM212 | 2 | 2 | 0 | 0 | 2 | 0.75 | RM313 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM543 | 2 | 2 | 0 | 0 | 2 | 0.68 | RM83 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM259 | 2 | 2 | 0 | 1 | 1 | 0.53 | RM101 | 2 | 2 | 0 | 1 | 1 | 0.53 |
| RM488 | 2 | 2 | 1 | 0 | 1 | 0.12 | RM309 | 2 | 2 | 0 | 0 | 2 | 0.75 |
| RM521 | 2 | 2 | 0 | 0 | 2 | 0.75 | RM28130 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM526 | 2 | 2 | 0 | 0 | 2 | 0.75 | RM28050 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM555 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28089 | 2 | 2 | 1 | 0 | 1 | 1.00 |
| RM530 | 2 | 2 | 0 | 1 | 1 | 0.96 | RM244 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM279 | 3 | 3 | 0 | 2 | 1 | 0.98 | RM28067 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM416 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28088 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM22 | 2 | 2 | 0 | 1 | 1 | 0.96 | RM28099 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM16030 | 2 | 2 | 0 | 1 | 1 | 0.90 | RM28078 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM60 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM3349 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM15780 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28079 | 2 | 2 | 0 | 1 | 1 | 0.96 |
| RM537 | 2 | 2 | 1 | 0 | 1 | 0.12 | RM28082 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM252 | 3 | 3 | 0 | 0 | 3 | 0.86 | RM28048 | 2 | 2 | 0 | 0 | 2 | 0.75 |
| RM136 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28069 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM527 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28088 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM528 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM1261 | 2 | 2 | 0 | 1 | 1 | 0.96 |
| RM5371 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28075 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM541 | 2 | 2 | 1 | 0 | 1 | 1.00 | RM28051 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM276 | 3 | 3 | 1 | 0 | 2 | 1.00 | RM28050 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM210 | 3 | 3 | 0 | 1 | 2 | 0.34 | RM12091 | 2 | 2 | 0 | 1 | 1 | 0.23 |
| RM339 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28090 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM337 | 2 | 2 | 0 | 0 | 2 | 0.61 | RM28059 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM25 | 1 | 0 | 0 | 0 | 1 | 0.00 | RM28095 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| RM464 | 2 | 2 | 0 | 0 | 2 | 0.00 | Total | 95 | 60 | 6 | 13 | 77 | 41.76 |
| RM566 | 2 | 2 | 0 | 0 | 2 | 0.81 | Mean | 1.5 | 1.0 | 0.1 | 0.2 | 1.2 | 0.66 |
| RM24390 | 1 | 0 | 0 | 0 | 1 | 0.00 |
| Microsatellite marker | Amp. range (bp) | Markers positions (cM) | Chr. | QTLs | Reference |
| RM315 | 120-133 | 165.0 | 1 | qDTY1.1 | Dixit et al, 2012 |
| RM472 | 296-300 | 168.2-172.0 | 1 | qDTY1.1 | Venuprasad et al, 2012 |
| RM431 | 296-250 | 178.3 | 1 | qDTY1.1 | Gimhani et al, 2016; Vikram et al, 2011; Kumar et al, 2014 |
| RM212 | 136-150 | 163.1 | 1 | qDTY1.1 | Vikram et al, 2011 |
| RM488 | 177-200 | 101 | 1 | qDTY1.1 | Kumar et al, 2014 |
| RM555 | 223 | 20.3 | 2 | qDTY2.2 | Kumar et al, 2014 |
| RM279 | 140-174 | 134 | 2 | qDTY2.2 | Sandhu et al, 2018 |
| RM60 | 165 | - | 3 | qDTF3.2 | Vikram et al, 2011; Awasthi, 2014 |
| RM22 | 180-194 | 7.2 | 3 | qDTF3.2 | Vikram et al, 2011; Awasthi, 2014 |
| RM541 | 150-158 | 75.5 | 6 | qDTY 6.2 | Dixit et al, 2012 |
| RM28048 | 80-93 | - | 12 | qDTY12.1 | Awasthi, 2014;Bernier et al, 2009 |
| RM1261 | 160-167 | 61.6 | 12 | qDTY12.1 | Dixit et al, 2012 |
Supplemental Table 2 List of microsatellite markers associated with drought-tolerant QTLs used for assessment of genetic diversity study.
| Microsatellite marker | Amp. range (bp) | Markers positions (cM) | Chr. | QTLs | Reference |
| RM315 | 120-133 | 165.0 | 1 | qDTY1.1 | Dixit et al, 2012 |
| RM472 | 296-300 | 168.2-172.0 | 1 | qDTY1.1 | Venuprasad et al, 2012 |
| RM431 | 296-250 | 178.3 | 1 | qDTY1.1 | Gimhani et al, 2016; Vikram et al, 2011; Kumar et al, 2014 |
| RM212 | 136-150 | 163.1 | 1 | qDTY1.1 | Vikram et al, 2011 |
| RM488 | 177-200 | 101 | 1 | qDTY1.1 | Kumar et al, 2014 |
| RM555 | 223 | 20.3 | 2 | qDTY2.2 | Kumar et al, 2014 |
| RM279 | 140-174 | 134 | 2 | qDTY2.2 | Sandhu et al, 2018 |
| RM60 | 165 | - | 3 | qDTF3.2 | Vikram et al, 2011; Awasthi, 2014 |
| RM22 | 180-194 | 7.2 | 3 | qDTF3.2 | Vikram et al, 2011; Awasthi, 2014 |
| RM541 | 150-158 | 75.5 | 6 | qDTY 6.2 | Dixit et al, 2012 |
| RM28048 | 80-93 | - | 12 | qDTY12.1 | Awasthi, 2014;Bernier et al, 2009 |
| RM1261 | 160-167 | 61.6 | 12 | qDTY12.1 | Dixit et al, 2012 |
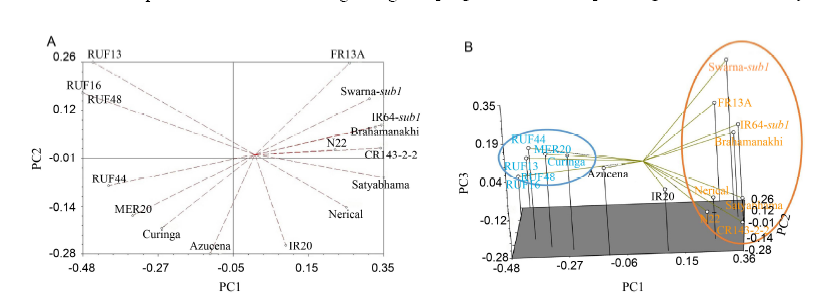
Fig. 1. Two-dimensional plot (A) and three-dimensional plot (B) from the principal component analysis (PCA) for 16 rice genotypes based on 63 simple sequence repeat markers.
| Genotype | Swarna-sub1 | IR64-sub1 | FR13A | CR143-2-2 | Bra | Sat | N22 | Nerical | IR20 | Azucena | Curinga | MER20 | RUF44 | RUF16 | RUF48 |
| IR64-sub1 | 0.85 | ||||||||||||||
| FR13A | 0.79 | 0.80 | |||||||||||||
| CR143-2-2 | 0.73 | 0.83 | 0.75 | ||||||||||||
| Bra | 0.77 | 0.78 | 0.75 | 0.78 | |||||||||||
| Sat | 0.79 | 0.90 | 0.78 | 0.93 | 0.82 | ||||||||||
| N22 | 0.72 | 0.77 | 0.81 | 0.85 | 0.75 | 0.86 | |||||||||
| Nerical | 0.78 | 0.84 | 0.77 | 0.86 | 0.81 | 0.93 | 0.87 | ||||||||
| IR20 | 0.73 | 0.74 | 0.68 | 0.76 | 0.73 | 0.82 | 0.77 | 0.83 | |||||||
| Azucena | 0.68 | 0.71 | 0.65 | 0.68 | 0.66 | 0.74 | 0.72 | 0.81 | 0.83 | ||||||
| Curinga | 0.65 | 0.68 | 0.66 | 0.65 | 0.63 | 0.71 | 0.73 | 0.76 | 0.76 | 0.86 | |||||
| MER20 | 0.61 | 0.64 | 0.63 | 0.62 | 0.62 | 0.67 | 0.69 | 0.73 | 0.75 | 0.84 | 0.96 | ||||
| RUF44 | 0.56 | 0.59 | 0.62 | 0.57 | 0.57 | 0.62 | 0.64 | 0.67 | 0.71 | 0.80 | 0.89 | 0.93 | |||
| RUF16 | 0.54 | 0.56 | 0.59 | 0.58 | 0.54 | 0.59 | 0.65 | 0.64 | 0.68 | 0.77 | 0.82 | 0.86 | 0.87 | ||
| RUF48 | 0.54 | 0.56 | 0.59 | 0.58 | 0.54 | 0.59 | 0.65 | 0.64 | 0.68 | 0.77 | 0.82 | 0.86 | 0.87 | 1.00 | |
| RUF13 | 0.56 | 0.57 | 0.58 | 0.57 | 0.55 | 0.58 | 0.62 | 0.61 | 0.65 | 0.73 | 0.78 | 0.82 | 0.83 | 0.96 | 0.96 |
| Bra, Brahamanakhi; Sat, Satyabhama. | |||||||||||||||
Table 2 Genetic similarity coefficient among 16 rice genotypes.
| Genotype | Swarna-sub1 | IR64-sub1 | FR13A | CR143-2-2 | Bra | Sat | N22 | Nerical | IR20 | Azucena | Curinga | MER20 | RUF44 | RUF16 | RUF48 |
| IR64-sub1 | 0.85 | ||||||||||||||
| FR13A | 0.79 | 0.80 | |||||||||||||
| CR143-2-2 | 0.73 | 0.83 | 0.75 | ||||||||||||
| Bra | 0.77 | 0.78 | 0.75 | 0.78 | |||||||||||
| Sat | 0.79 | 0.90 | 0.78 | 0.93 | 0.82 | ||||||||||
| N22 | 0.72 | 0.77 | 0.81 | 0.85 | 0.75 | 0.86 | |||||||||
| Nerical | 0.78 | 0.84 | 0.77 | 0.86 | 0.81 | 0.93 | 0.87 | ||||||||
| IR20 | 0.73 | 0.74 | 0.68 | 0.76 | 0.73 | 0.82 | 0.77 | 0.83 | |||||||
| Azucena | 0.68 | 0.71 | 0.65 | 0.68 | 0.66 | 0.74 | 0.72 | 0.81 | 0.83 | ||||||
| Curinga | 0.65 | 0.68 | 0.66 | 0.65 | 0.63 | 0.71 | 0.73 | 0.76 | 0.76 | 0.86 | |||||
| MER20 | 0.61 | 0.64 | 0.63 | 0.62 | 0.62 | 0.67 | 0.69 | 0.73 | 0.75 | 0.84 | 0.96 | ||||
| RUF44 | 0.56 | 0.59 | 0.62 | 0.57 | 0.57 | 0.62 | 0.64 | 0.67 | 0.71 | 0.80 | 0.89 | 0.93 | |||
| RUF16 | 0.54 | 0.56 | 0.59 | 0.58 | 0.54 | 0.59 | 0.65 | 0.64 | 0.68 | 0.77 | 0.82 | 0.86 | 0.87 | ||
| RUF48 | 0.54 | 0.56 | 0.59 | 0.58 | 0.54 | 0.59 | 0.65 | 0.64 | 0.68 | 0.77 | 0.82 | 0.86 | 0.87 | 1.00 | |
| RUF13 | 0.56 | 0.57 | 0.58 | 0.57 | 0.55 | 0.58 | 0.62 | 0.61 | 0.65 | 0.73 | 0.78 | 0.82 | 0.83 | 0.96 | 0.96 |
| Bra, Brahamanakhi; Sat, Satyabhama. | |||||||||||||||
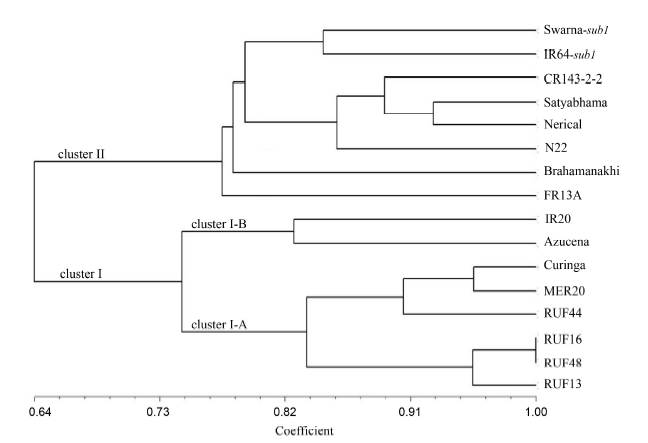
Fig. 2. Unweighted pair-group method with arithmetic means (UPGMA) dendrogram for 16 rice genotypes based on genetic similarity by 63 simple sequence repeat markers.
| [1] | Anandan A, Anumalla M, Pradhan S K, Ali J.2016. Population structure, diversity and trait association analysis in rice (Oryza sativa L.) germplasm for early seedling vigor (ESV) using trait linked SSR markers. PLoS One, 11(3): e0152406. |
| [2] | Anderson J A, Churchill G A, Autrique J E, Tanksley S D, Sorrells M E.1993. Optimizing parental selection for genetic linkage maps.Genome, 36(1): 181-186. |
| [3] | Arbelaez J D, Moreno L T, Singh N, Tung C W, Maron L G, Ospina Y, Martinez C P, Grenier C, Lorieux M, McCouch S.2015. Development and GBS-genotyping of introgression lines (ILs) using two wild species of rice,O. meridionalis and O. rufipogon, in a common recurrent parent, O. sativa cv. Curinga. Mol Breeding, 35: 81. |
| [4] | Barik M, Dash S K, Padhi S, Swain P.2017. Effect of drought on morpho-physiological, yield and yield traits of chromosome segment substitution lines (CSSLs) derived from wild species of rice.ORYZA-Int J Rice, 54(1): 65-72. |
| [5] | Behera L, Patra B C, Sahu R K, Nanda A, Sahu S C, Patnaik A, Rao G J N, Singh O N.2012. Assessment of genetic diversity in medicinal rices using microsatellite markers.Aust J Crop Sci, 6(9): 1369-1376. |
| [6] | Behera L, Mohanty S, Pradhan S, Singh S, Singh O, Sahu R, Sahu S, Dash S, Mohapatra T.2013. Assessment of genetic diversity of rainfed lowland rice genotypes using microsatellite markers.Indian J Genet, 73: 142-152. |
| [7] | Charcosset A, Moreau L.2004. Use of molecular markers for the development of new cultivars and the evaluation of genetic diversity.Euphytica, 137(1): 81-94. |
| [8] | Choudhary G, Ranjitkumar N, Surapaneni M, Deborah D A, Vipparla A, Anuradha G, Siddiq E A, Vemireddy L R.2013. Molecular genetic diversity of major Indian rice cultivars over decadal periods.PLoS One, 8(6): e66197. |
| [9] | Dellaporta S L, Wood J, Hicks J B.1983. A plant DNA minipreparation: Version II.Plant Mol Biol Rep, 1: 19-21. |
| [10] | Flint-Garcia S A, Thuillet A C, Yu J, Pressoir G, Romero S M, Mitchell S E, Doebley J, Kresovich S, Goodman M M, Buckler E S.2005. Maize association population: A high-resolution platform for quantitative trait locus dissection.Plant J, 44: 1054-1064. |
| [11] | Gaikwad K B, Singh N, Bhatia D, Kaur R, Bains N S, Bharaj T S, Singh K.2014. Yield-enhancing heterotic QTL transferred from wild species to cultivated riceOryza sativa L. PLoS One, 9(6): e96939. |
| [12] | Ganie S A, Borgohain M J, Kritika K, Talukdar A, Pani D R, Mondal T K.2016. Assessment of genetic diversity ofSaltol QTL among the rice(Oryza sativa L.) genotypes. Physiol Mol Biol Pla, 22(1): 107-114. |
| [13] | Gawenda I, Schrӧder-Lorenz A, Debener T.2012. Markers for ornamental traits inPhalaenopsis orchids: Population structure, linkage disequilibrium and association mapping. Mol Breeding, 30(1): 305-316. |
| [14] | Ismail A M, Singh U S, Singh S, Dar M H, Mackill D J.2013. The contribution of submergence-tolerant (sub1) rice varieties to food security in flood-prone rainfed lowland areas in Asia. Field Crops Res, 152: 83-93. |
| [15] | Jasim Aljumaili S, Rafii M Y, Latif M, Sakimin S Z, Arolu I W, Miah G.2018. Genetic diversity of aromatic rice germplasm revealed by SSR markers.BioMed Res Int, 2018: 1-11. |
| [16] | Khush G S.2005. What it will take to feed 5.0 billion rice consumers in 2030.Plant Mol Biol, 59: 1-6. |
| [17] | Kumbhar S D, Kulwal P L, Patil J V, Sarawate C D, Gaikwad A P, Jadhav A S.2015. Genetic diversity and population structure in landraces and improved rice varieties from India.Rice Sci, 22(3): 99-107. |
| [18] | Marathi B, Guleria S, Mohapatra T, Parsad R, Mariappan N, Kurungara V K, Atwal S S, Prabhu K V, Singh N K, Singh A K.2012. QTL analysis of novel genomic regions associated with yield and yield related traits in new plant type based recombinant inbred lines of rice (Oryza sativa L.). BMC Plant Boil, 12: 137. |
| [19] | Molden D, Oweis T, Steduto P, Bindraban P, Hanjra M A, Kijne J.2010. Improving agricultural water productivity: Between optimism and caution.Agr Water Manage, 97(4): 528-535. |
| [20] | Nachimuthu V V, Muthurajan R, Duraialaguraja S, Sivakami R, Pandian B A, Ponniah G, Gunasekaran K, Swaminathan M, Suji K K, Sabariappan R.2015. Analysis of population structure and genetic diversity in rice germplasm using SSR markers: An initiative towards association mapping of agronomic traits inOryza sativa. Rice, 8: 30. |
| [21] | Patel S, Ravikiran R, Chakraborty S, Macwana S, Sasidharan N, Trivadi R, Aher B.2014. Genetic diversity analysis of colored and white rice genotypes using microsatellite (SSR) and insertion-deletion (INDEL) markers.Emir J Food Agr, 26(6): 497-507. |
| [22] | Pradhan S K, Barik S R, Sahoo A, Mohapatra S, Nayak D K, Mahender A, Meher J, Anandan A, Pandit E.2016. Population structure, genetic diversity and molecular marker-trait association analysis for high temperature stress tolerance in rice.PLoS One, 11: e0160027. |
| [23] | Rohlf F J.1988. NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System. Exeter Publishing. |
| [24] | Roy P S, Rao G J N, Jena S, Samal R, Patnaik A, Patnaik S S C, Jambhulkar N N, Sharma S, Mohapatra T.2016. Nuclear and chloroplast DNA variation provides insights into population structure and multiple origin of native aromatic rices of Odisha, India.PLoS One, 11(9): e0162268. |
| [25] | Roy S, Banerjee A, Mawkhlieng B, Misra1 A K, Pattanayak A, Harish G D, Singh S K, Ngachan S V, Bansal K C, 2015. Genetic diversity and population structure in aromatic and quality rice (Oryza sativa L.) landraces from North-Eastern India. PLoS One, 10: e0129607. |
| [26] | Saini N, Jain N, Jain S, Jain R K.2004. Assessment of genetic diversity within and among Basmati and non-Basmati rice varieties using AFLP, ISSR and SSR markers.Euphytica, 140(3): 133-146. |
| [27] | Sajib A M, Hossain M, Mosnaz A, Hossain H, Islam M, Ali M, Prodhan S H.2012. SSR marker-based molecular characterization and genetic diversity analysis of aromatic landreces of rice (Oryza sativa L.). J BioSci Biotechnol, 1: 107-116. |
| [28] | Septiningsih E M, Pamplona A M, Sanchez D L, Neeraja C N, Vergara G V, Heuer S, Ismail A M, Mackill D J.2009. Development of submergence-tolerant rice cultivars: Thesub1 locus and beyond. Ann Bot, 103(2): 151-160. |
| [29] | Shah S M, Arif M, Aslam K, Shabir G, Thomson M J.2016. Genetic diversity analysis of Pakistan rice (Oryza sativa) germplasm using multiplexed single nucleotide polymorphism markers. Genet Res Crop Ev, 63(7): 1113-1126. |
| [30] | Singh N, Dang T T M, Vergara G V, Pandey D M, Sanchez D, Neeraja C N, Septiningsih E M, Mendioro M, Tecson-Mendoza E M, Ismail A M, Mackill D J.2010. Molecular marker survey and expression analyses of the rice submergence-tolerance geneSUB1A. Theor Appl Genet, 121(8): 1441-1453. |
| [31] | Singh N, Choudhury D R, Tiwari G, Singh A K, Kumar S, Srinivasan K, Tyagi R K, Sharma A D, Singh N K, Singh R.2016. Genetic diversity trend in Indian rice varieties: An analysis using SSR markers.BMC Genet, 17: 127. |
| [32] | Sun J, Qian Q, Ma D R, Xu Z J, Liu D, Du H B, Chen W F.2013. Introgression and selection shaping the genome and adaptive loci of weedy rice in northern China.New Phytol, 197(1): 290-299. |
| [33] | Sweeney M, McCouch S.2007. The complex history of the domestication of rice.Ann Bot, 100(5): 951-957. |
| [34] | Tian F, Li D J, Fu Q, Zhu Z F, Fu Y C, Wang X K, Sun C Q.2006. Construction of introgression lines carrying wild rice (Oryza rufipogon Griff.) segments in cultivated rice(Oryza sativa L.) background and characterization of introgressed segments associated with yield-related traits. Theor Appl Genet, 112(3): 570-580. |
| [35] | Upadhyay P, Singh V K, Neeraja C.2011. Identification of genotype specific alleles and molecular diversity assessment of popular rice (Oryza sativa L.) varieties of India. Int J Plant Breeding Genet, 5(2): 130-140. |
| [36] | Varshney R K, Bansal K C, Aggarwal P K, Datta S K, Craufurd P Q.2011. Agricultural biotechnology for crop improvement in a variable climate: Hope or hype?Trends Plant Sci, 16(7): 363-371. |
| [37] | Venuprasad R, Bool M, Dalid C O, Bernier J, Kumar A, Atlin G N.2009a. Genetic loci responding to two cycles of divergent selection for grain yield under drought stress in a rice breeding population.Euphytica, 167(2): 261-269. |
| [38] | Venuprasad R, Dalid C O, Del Valle M, Zhao D, Espiritu M, Sta Cruz M T, Amante M, Kumar A, Atlin G N.2009b. Identification and characterization of large-effect quantitative trait loci for grain yield under lowland drought stress in rice using bulk-segregant analysis.Theor Appl Genet, 120(1): 177-190. |
| [39] | Venuprasad R, Bool M E, Quiatchon L, Sta Cruz M T, Amante M, Atlin G.2012. A large-effect QTL for rice grain yield under upland drought stress on chromosome 1.Mol Breeding, 30(1): 535-547. |
| [40] | Xu Q, Yuan X P, Wang S, Feng Y, Yu H Y, Wang Y P, Yang Y L, Wei X H, Li X M.2016. The genetic diversity and structure ofindica rice in China as detected by single nucleotide polymorphism analysis. BMC Genet, 17: 53. |
| [41] | Yang F, Chen Y L, Tong C, Huang Y, Xu F F, Li K H, Corke H, Sun M, Bao J S.2014. Association mapping of starch physicochemical properties with starch synthesis-related gene markers in nonwaxy rice (Oryza sativa L.). Mol Breeding, 34(4): 1747-1763. |
| [42] | Zhao Y L, Wang H M, Chen W, Li Y H.2014. Genetic structure, linkage disequilibrium and association mapping of Verticillium wilt resistance in elite cotton (Gossypium hirsutum L.) germplasm population. PLoS One, 9(1): e86308. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [13] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [14] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| [15] | Zhang Guomei, Li Han, Liu Shanshan, Zhou Xuming, Lu Mingyang, Tang Liang, Sun Lihua. Water Extract of Rice False Smut Balls Activates Nrf2/HO-1 and Apoptosis Pathways, Causing Liver Injury [J]. Rice Science, 2023, 30(5): 473-485. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||