
Rice Science ›› 2021, Vol. 28 ›› Issue (4): 344-357.DOI: 10.1016/j.rsci.2021.05.005
• Research Papers • Previous Articles Next Articles
Yan Wang#, Xiaoqin Zeng#, Lu Lu, Qinglan Cheng, Fayu Yang, Mingjiang Huang, Mao Xiong, Yunfeng Li( )
)
Received:2020-08-27
Accepted:2020-11-17
Online:2021-07-28
Published:2021-07-28
About author:#These authors contributed equally to this work
Yan Wang, Xiaoqin Zeng, Lu Lu, Qinglan Cheng, Fayu Yang, Mingjiang Huang, Mao Xiong, Yunfeng Li. MULTI-FLORET SPIKELET 4 (MFS4) Regulates Spikelet Development and Grain Size in Rice[J]. Rice Science, 2021, 28(4): 344-357.
Add to citation manager EndNote|Ris|BibTeX

Fig. 1. Phenotypes of spikelet in wild type (WT) and mfs4 mutants. A, Spikelet of the WT. A1, WT spikelet; A2, The lemma and palea were removed in A1; A3 to A6, The WT spikelet surface characters of sl, pa and le; A7, Transverse section of WT spikelet. B?E, Spikelets of the mfs4 mutants. B1 and B2, The Type I mfs4 mutant spikelet; B3?B5, The Type I mfs4 mutant spikelet surface characters of spikelet, le and ele; B6, Transverse section of the Type I mfs4 mutant. C1 and C2, The Type II mfs4 mutant spikelet; C2, The lemma and extra lemma were removed; C3?C6, The Type II mfs4 mutant spikelet surface characters of sl, pa, le and ele; C7, Transverse section of the Type II mfs4 mutant. D1 and D2, The Type II mfs4 mutant spikelet; D2, The lemma, palea and elongate sterile lemma were removed; D3?D6, The Type II mfs4 mutant spikelet surface characters of sl, pa, le and mrp; D7, Transverse section of the Type II mfs4 mutant. E1, The Type III mfs4 mutant spikelet; E2?E4, The Type III mfs4 mutant spikelet surface characters of sl, pa and le; E5, Transverse section of the Type III mfs4 mutant. F, Percentage of mutant organs in mfs4 spikelets.
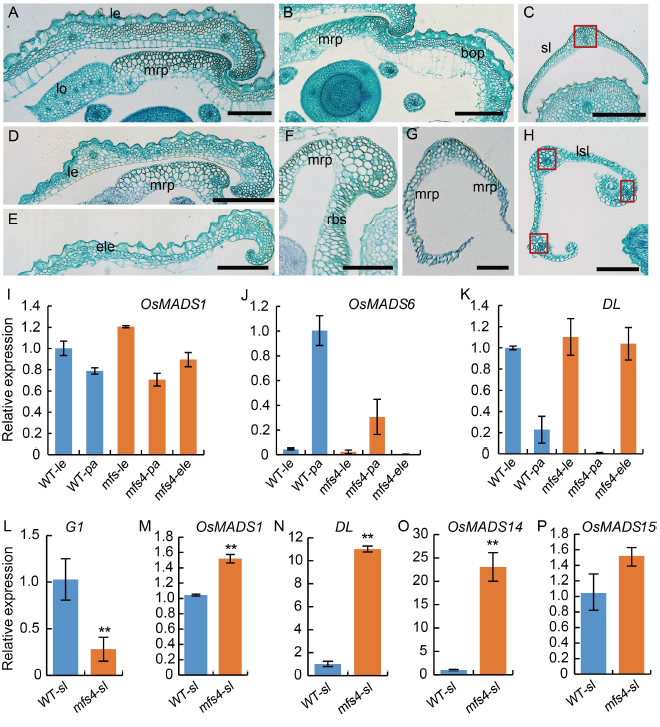
Fig. 2. Histological and qRT-PCR analysis of lateral organs in spikelets of wild type (WT) and mfs4 mutants. A?C, Transverse sections of lateral organs in the WT spikelet. A and B, The lemma and palea of WT. C, The sterile lemma of WT. Red box indicates the vascular bundle. Scale bars are 500 μm. D?H, Transverse sections of lateral organs in the mfs4 mutant spikelets. D?G, The lemma, palea and extra lemma of the mfs4 mutant spikelets. H, The elongate sterile lemma of the mfs4 mutant spikelets. Red boxes indicate the vascular bundles. Scale bars are 500 μm. I?P, qRT-PCR analysis of OsMADS1, OsMADS6, DL, G1, OsMADS14 and OsMADS15 expression. Actin was used as a control. RNA was isolated from the ?ower organs of the WT and mfs4 mutant spikelets. Data are Mean ± SE (n = 3). *, P? ≤ 0.05 and **, P? ≤ 0.01 by the Student’s t-test. bop, Body of palea; ele, Extra lemma; le, Lemma; lo, Lodicule; lsl, Lemma- like sterile lemma; mrp, Marginal region of palea; Pa, Palea; bs, Residual bop-like structure; sl, Sterile lemma. bop, Body of palea; ele, Extra lemma; le, Lemma; lo, Lodicule; mrp, Marginal region of palea; lsl, Lemma-like sterile lemma; ov, Ovule; pa, Palea; pi, Pistil; rbs, Residual bop-like structure; rg, Rudimentary glume; sl, Sterile lemma; st, Stamen. Scale bars are 1 000 μm in A1, A2, B1, B2, C1, C2, D1, D2 and E1; Scale bars are 500 μm in A3?A6, B3?B5, C3?C6, D3?D6 and E2?E4; Scale bars are 100 μm in A7, B6, C7, D7 and E5.
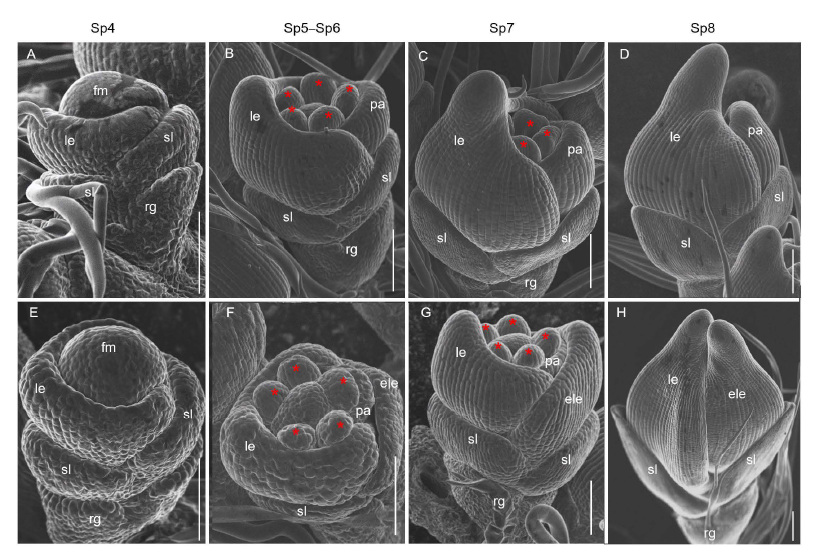
Fig. 3. Scanning electron micrographs of spikelets at early developmental stages in wild type (WT) and mfs4 mutant. A-D, WT. E-H, mfs4 with ele. Asterisks indicate the stamens. Scale bars are 100 μm. ele, Extra lemma; fm, Flower meristem; le, Lemma; pa, Palea; rg, Rudimentary glume; sl, Sterile lemma.
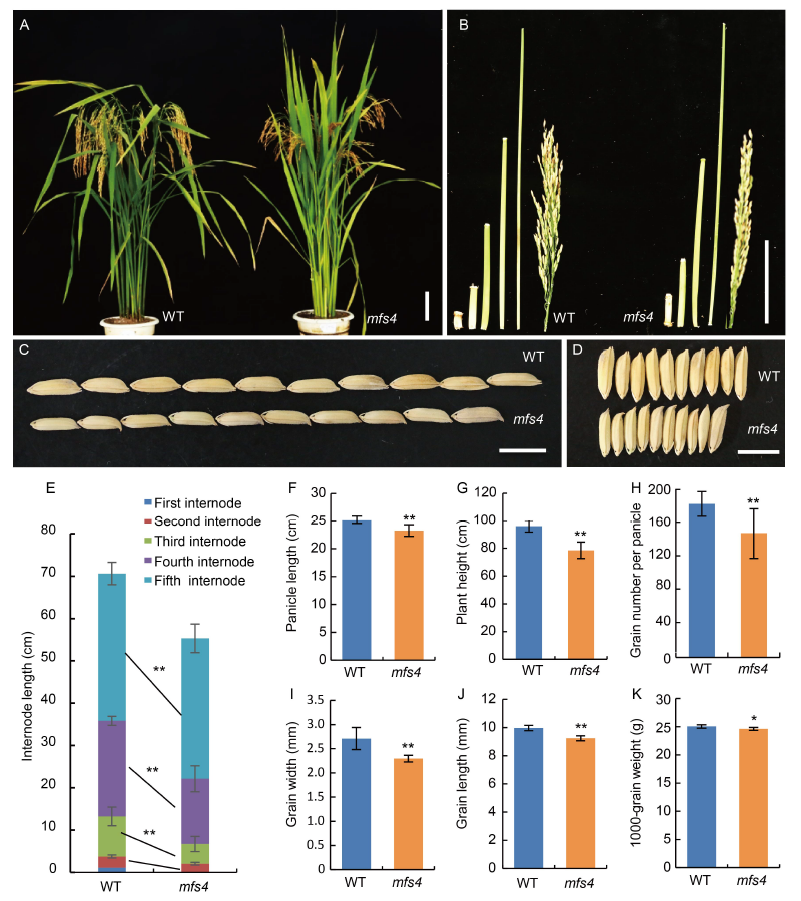
Fig. 4. Comparison of morphological characters between wild type (WT) and mfs4 mutant. A, Plant morphology of WT and mfs4 mutant at the heading stage. B, Internodes and panicles of WT and mfs4 mutant. C and D, Comparisons of grain length (C) and width (D) in the WT and mfs4 mutant.E?K, Comparisons of internode length (E), panicle length (F), plant height (G), grain number per panicle (H), grain width (I), grain length (J) and 1000-grain weight (K) between the WT and mfs4 mutant. Scale bars are 10 cm in A and B, and 1 cm in C and D. *, P ≤ 0.05 and **, P ≤ 0.01 by the Student’s t-test.
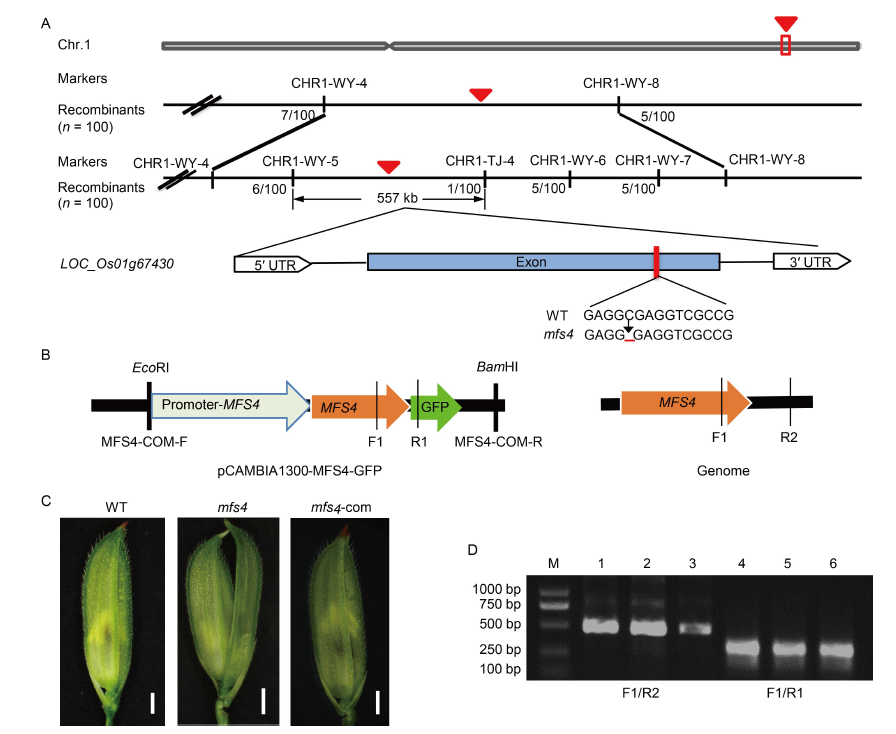
Fig. 5. Fine mapping and sequencing analysis of MFS4 gene. A, Gene mapping of MFS4 on rice chromosome 1. The red triangles indicate the target gene. The red line in the exon indicates the mutational site of candidate gene. WT, Wild type. B, Schematic structure of the complementary vector pCAMBIA1300-MFS4-GFP and MFS4 genome. MFS4-COM-F/MFS4-COM-R are complementary vector primers, F1/R2 are primers for amplifying endogenous MFS4, and F1/R1 are primers for amplifying exogenous MFS4-GFP. C, Defect in mfs4 flowers were completely rescued by introduction of pCAMBIA1300-MFS4. mfs4-com, mfs4 complementary flower. Scale bars are 1 mm. D, Identification of transgenic plants. M, DNA marker; Lanes 1?3, Detection of positive transgenic plants by F1/R2 primers; Lanes 4?6, Detection of positive transgenic plants by F1/R1 primers.
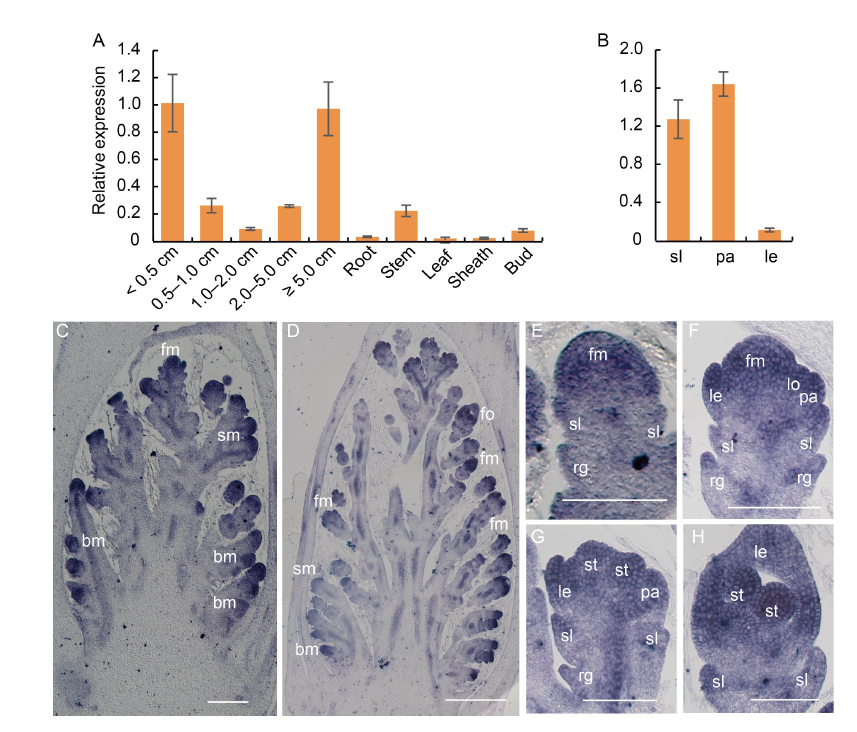
Fig. 6. Spatiotemporal expression pattern of MFS4 gene. A and B, qRT-PCR of MFS4. Actin was used as a control. RNA was isolated from young panicles < 0.5 cm, 0.5?1.0 cm, 1.0?2.0 cm, 2.0?5.0 cm and ≥ 5.0 cm, as well as vegetative organ and floral organ of wild type plants. Data are Mean ± SD (n = 3). C?H, In situ hybridization expression in panicles of wild type (C and D), and spikelets at Sp2?Sp3 (E), Sp4?Sp5 (F), Sp5?Sp6 (G) and Sp7?Sp8 (H). bm, Branch meristem; fm, Floral meristem; fo, Floral organ; le, Lemma; lo, Lodicule; pa, Palea; rg, Rudimentary glume; sl, Sterile lemma; sm, Spikelet meristem; st, Stamen. Scale bars are 500 μm in C and D, and 100 μm in E?H.
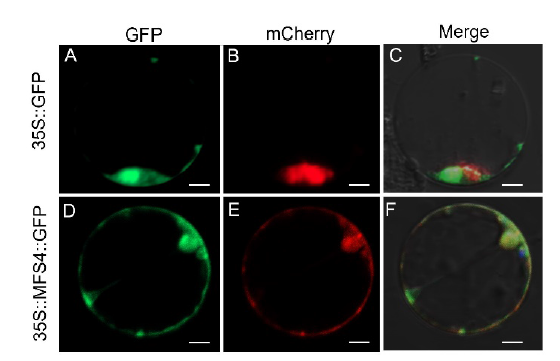
Fig. 7. Subcellular localization of MFS4 protein. A and D, GFP illuminant. B and E, mCherry illuminant which indicates the mitochondria marker. C and F, Merge illuminant. Scale bars are 50 μm.
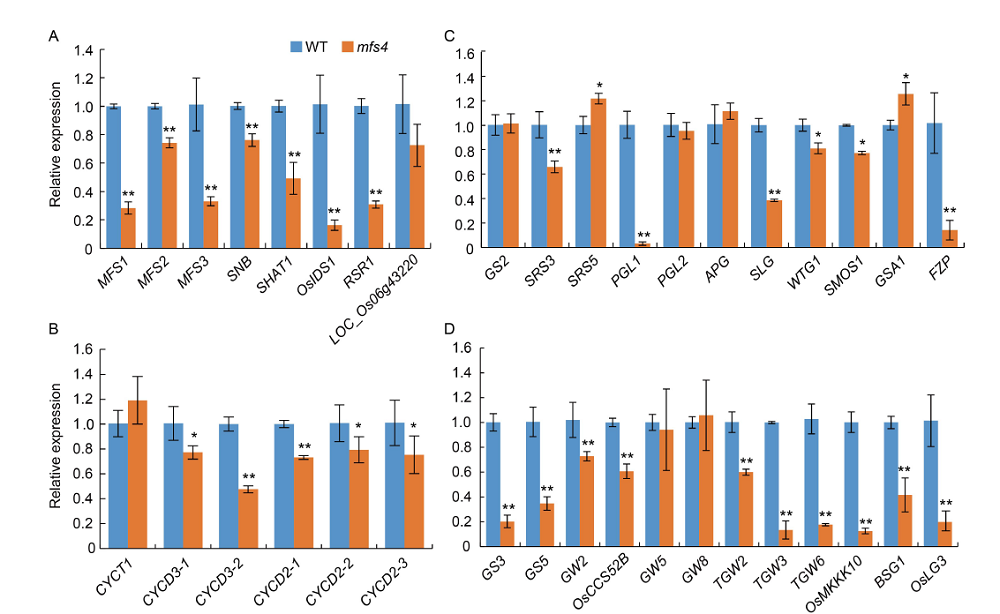
Fig. 8. Expression levels of related genes in wild type (WT) and mfs4 mutant. A, Expression analysis of related genes influencing spikelet meristem determination in young panicles. B, Expression analysis of related genes influencing grain size by cell cycle-related genes in panicles. C, Expression analysis of related genes influencing grain size by hull cell expansion-related genes in panicles. D, Expression analysis of related genes influencing grain size by hull cell proliferation-related genes in panicles. Actin was used as a control. RNA was isolated from panicles of WT and mfs4 mutant. At least three replicates were performed and the mean value was used. Error bars mean standard error. *, P?≤ 0.05 and **, P?≤ 0.01 by the Student’s t-test.
| Primer | Forward sequence (5′→3′) | Reverse sequence (3′→5′) |
|---|---|---|
| CHR1-WY-4 | GGATGAACCATACACGCCCATT | GCATGAGAATGCCTGACTGGATG |
| CHR1-TJ-4 | CATGAACAAATCTCAAGTTGTTGAG | TCTGCTTTCTTGGTTACGAGGT |
| CHR1-WY-5 | CGGAGAACTCTGGTTGTGCTTTG | TGCCGTTGAGCCAAAGACATCT |
| CHR1-WY-6 | ACTGAGCTAGCTATGCGGGCTG | AGCAGAGTCATCGACGAGTCGA |
| CHR1-WY-7 | TCAGACCCCATTCTTCACTCCG | GTCGACTTGTTGCCCTCCTCTT |
| CHR1-WY-8 | GCTCTCTCGTCGATCTTAGCAGC | CGTTCGTTGCATGAAGAGGAG |
Table S1. Sequences of primers for gene mapping.
| Primer | Forward sequence (5′→3′) | Reverse sequence (3′→5′) |
|---|---|---|
| CHR1-WY-4 | GGATGAACCATACACGCCCATT | GCATGAGAATGCCTGACTGGATG |
| CHR1-TJ-4 | CATGAACAAATCTCAAGTTGTTGAG | TCTGCTTTCTTGGTTACGAGGT |
| CHR1-WY-5 | CGGAGAACTCTGGTTGTGCTTTG | TGCCGTTGAGCCAAAGACATCT |
| CHR1-WY-6 | ACTGAGCTAGCTATGCGGGCTG | AGCAGAGTCATCGACGAGTCGA |
| CHR1-WY-7 | TCAGACCCCATTCTTCACTCCG | GTCGACTTGTTGCCCTCCTCTT |
| CHR1-WY-8 | GCTCTCTCGTCGATCTTAGCAGC | CGTTCGTTGCATGAAGAGGAG |
| Primer | Forward sequence (5′→3′) | Reverse sequence (3′→5′) |
|---|---|---|
| OsMADS1 | ATCACCATCAGGGTCTTCTC | CAACCATGTCTGCTGCTTCA |
| OsMADS6 | CCAACAATGCACTTTCTGAAAC | GGAGGCTTGCTGCATGGC |
| DL | CCCATCTGCTTACAACCGCTT | GTTGGAGGTGGAAACCGTCG |
| MFS4 | AACGTACACGACCCGATCAC | CCGACCTTGAAGAAGTCGAG |
| G1 | GGCGTCTACTTGCCATTTCTG | TCGATCAGCATCAAAGCACAG |
| OsMADS14 | CCATTAACGAGCTTCAACGG | TGGTATGGATCTGAAGCCTCC |
| OsMADS15 | AGTACGCCACTGACTCCAGG | TGCTGGCCCCTCACATTC |
| MFS1 | AACGTACACGACCCGATCAC | CCGACCTTGAAGAAGTCGAG |
| MFS2 | GGAGGGCCTCAGATGGCAGAGG | GCTTCAACACGGGACCAAGATGG |
| MFS3 | AGCGAGCCGTGCATGGTGTA | GCTGCTGCTTAATTGCAGTGATG |
| SNB | AACGTACACGACCCGATCAC | CCGACCTTGAAGAAGTCGAG |
| SHAT1 | GCGATGGGAATCGCACATATGG | AAACCCGTCCATGCTGCTCTTC |
| OsIDS1 | GATGGGTGGCTCTCAGCATTCT | CAGCTTTGCAAGCCAACACGTC |
| RSR1 | AGCTGCCTACTGATGCTGCTGCT | GGTTCAGATCCACATCGGCTCCT |
| Os06g43220 | GTGGAGTGGAGGCAGACATCA | CCGGAGAACCTGAACGAACTC |
| CYCT1 | AGACAGCAGGTGATCAGGCCAC | CCTTGCTGGGCAAACAGTTGA |
| CYCD3-1 | TTGCTGGTTTACCTGTCATGCC | AGGAACCGGTCCAGGTAGTTCA |
| CYCD3-2 | GATGCCCCCAAGGTCAGTAGTA | TCGGAGATGATCTTGGCGCA |
| CYCD2-1 | GATGGAGGAAACCCTAGTTCCC | CTACCTGGCAACGAACAATGCA |
| CYCD2-2 | TCAGTGATCGCGCGATGATCT | ATGGGGTCACAGCTTGCAGC |
| CYCD2-3 | AGCACCATGAAAGGTACCATCG | ACACCGCCAAAGTCAAGAACG |
| GS2 | GTTGGGTTCTCAGCTGCACATG | ACTGCATCCACCTGACACCAAT |
| SRS3 | ACCAGGCGACAAAGAGATCGAG | GCATTCCTTCAGAGCGAGCAG |
| SRS5 | CTCGACATTGAGCGCCCAAC | TTCAGAGCACCATCGAACCTCA |
| PGL1 | GCTCGACTCCGGAGCAATGATA | CCGATCATGTTCTACCCCCAT |
| PGL2 | CGAGCTCATCTCCAAGCTCCAG | ACCACCACCCACACACACGACT |
| APG | CAAGAGCTCATCCCAAACTGCA | CGTCTTGAGGTACTCGATGGCT |
| SLG | TGGAGCACATCACATCCACCTC | AGGTGATGTAGGGGAGGTCGAAT |
| WTG1 | TAGTGCATGCTTTGTCATGGCA | TGCCAGTGCCAGGTTATTCACA |
| SMOS1 | ATCCTGCCAGCAAGTCAACCTG | AAGCTCGTACGGTCACTCTTGA |
| GSA1 | TGCAAGTCCCCAAGCTGATCC | GCATCGTGGGTCTCAATTGGAT |
| FZP | ACACAAGTGCAGTGCATGCATG | GCCTCGAGTGTTCATGACCAAT |
| GS3 | GTTCTACCGAAATGGCCGGAA | AGGAGAGGTAGCTGAGGCAGCA |
| GS5 | TGCTTGGCCATCTACAGGTGAT | GCATAGCTCTCCCCTGAGATGT |
| OsCCS5 | TGGAACGTGTTTCCATCTCCC | ATGATGTGGCCCCGATGCTA |
| GW2 | CAGGTCTGCACAGGCTGCAA | TGCTAGTCTGGGCAGAGCATGA |
| GW5 | CGTTGTGTGTTGGCGATGGAT | ATCTTGGGGCTCCGGTCGTA |
| GW8 | GGTGCAAGGAGGACCTGAGCAAG | AATTTGGCGGGAAGGAAGGAGAC |
| TGW2 | GGTAGGTGCAAGCACTGAAGCA | TTCGTCTACCTCATCATGCCGA |
| TGW3 | TGTGTGGCTGGAAATCCGGAT | AATGCCGTTTCCTCCAGCATC |
| TGW6 | AACGAGAGCACCAGCACGAGA | GAGAGCAGCACGAGGAAGACAA |
| OsMKKK10 | GATGCCGAAAATGCAACACTTG | GACACCCCTTGGTGATGTGGAT |
| OsMKK4 | ACGGCTTCACTTGGCCTCGT | GCGTGTGCGTTTTCTGGTGG |
| OsLG3 | ACACTTCGTGTTCGCGTCCAA | ACGACTGCACGAACAAAACCAA |
| BSG1 | AGATCGGCTGCTACCTGACCTAG | GTGGTGATGGTGACGATCCATC |
Table S2. Sequences of primers for RT-qPCR.
| Primer | Forward sequence (5′→3′) | Reverse sequence (3′→5′) |
|---|---|---|
| OsMADS1 | ATCACCATCAGGGTCTTCTC | CAACCATGTCTGCTGCTTCA |
| OsMADS6 | CCAACAATGCACTTTCTGAAAC | GGAGGCTTGCTGCATGGC |
| DL | CCCATCTGCTTACAACCGCTT | GTTGGAGGTGGAAACCGTCG |
| MFS4 | AACGTACACGACCCGATCAC | CCGACCTTGAAGAAGTCGAG |
| G1 | GGCGTCTACTTGCCATTTCTG | TCGATCAGCATCAAAGCACAG |
| OsMADS14 | CCATTAACGAGCTTCAACGG | TGGTATGGATCTGAAGCCTCC |
| OsMADS15 | AGTACGCCACTGACTCCAGG | TGCTGGCCCCTCACATTC |
| MFS1 | AACGTACACGACCCGATCAC | CCGACCTTGAAGAAGTCGAG |
| MFS2 | GGAGGGCCTCAGATGGCAGAGG | GCTTCAACACGGGACCAAGATGG |
| MFS3 | AGCGAGCCGTGCATGGTGTA | GCTGCTGCTTAATTGCAGTGATG |
| SNB | AACGTACACGACCCGATCAC | CCGACCTTGAAGAAGTCGAG |
| SHAT1 | GCGATGGGAATCGCACATATGG | AAACCCGTCCATGCTGCTCTTC |
| OsIDS1 | GATGGGTGGCTCTCAGCATTCT | CAGCTTTGCAAGCCAACACGTC |
| RSR1 | AGCTGCCTACTGATGCTGCTGCT | GGTTCAGATCCACATCGGCTCCT |
| Os06g43220 | GTGGAGTGGAGGCAGACATCA | CCGGAGAACCTGAACGAACTC |
| CYCT1 | AGACAGCAGGTGATCAGGCCAC | CCTTGCTGGGCAAACAGTTGA |
| CYCD3-1 | TTGCTGGTTTACCTGTCATGCC | AGGAACCGGTCCAGGTAGTTCA |
| CYCD3-2 | GATGCCCCCAAGGTCAGTAGTA | TCGGAGATGATCTTGGCGCA |
| CYCD2-1 | GATGGAGGAAACCCTAGTTCCC | CTACCTGGCAACGAACAATGCA |
| CYCD2-2 | TCAGTGATCGCGCGATGATCT | ATGGGGTCACAGCTTGCAGC |
| CYCD2-3 | AGCACCATGAAAGGTACCATCG | ACACCGCCAAAGTCAAGAACG |
| GS2 | GTTGGGTTCTCAGCTGCACATG | ACTGCATCCACCTGACACCAAT |
| SRS3 | ACCAGGCGACAAAGAGATCGAG | GCATTCCTTCAGAGCGAGCAG |
| SRS5 | CTCGACATTGAGCGCCCAAC | TTCAGAGCACCATCGAACCTCA |
| PGL1 | GCTCGACTCCGGAGCAATGATA | CCGATCATGTTCTACCCCCAT |
| PGL2 | CGAGCTCATCTCCAAGCTCCAG | ACCACCACCCACACACACGACT |
| APG | CAAGAGCTCATCCCAAACTGCA | CGTCTTGAGGTACTCGATGGCT |
| SLG | TGGAGCACATCACATCCACCTC | AGGTGATGTAGGGGAGGTCGAAT |
| WTG1 | TAGTGCATGCTTTGTCATGGCA | TGCCAGTGCCAGGTTATTCACA |
| SMOS1 | ATCCTGCCAGCAAGTCAACCTG | AAGCTCGTACGGTCACTCTTGA |
| GSA1 | TGCAAGTCCCCAAGCTGATCC | GCATCGTGGGTCTCAATTGGAT |
| FZP | ACACAAGTGCAGTGCATGCATG | GCCTCGAGTGTTCATGACCAAT |
| GS3 | GTTCTACCGAAATGGCCGGAA | AGGAGAGGTAGCTGAGGCAGCA |
| GS5 | TGCTTGGCCATCTACAGGTGAT | GCATAGCTCTCCCCTGAGATGT |
| OsCCS5 | TGGAACGTGTTTCCATCTCCC | ATGATGTGGCCCCGATGCTA |
| GW2 | CAGGTCTGCACAGGCTGCAA | TGCTAGTCTGGGCAGAGCATGA |
| GW5 | CGTTGTGTGTTGGCGATGGAT | ATCTTGGGGCTCCGGTCGTA |
| GW8 | GGTGCAAGGAGGACCTGAGCAAG | AATTTGGCGGGAAGGAAGGAGAC |
| TGW2 | GGTAGGTGCAAGCACTGAAGCA | TTCGTCTACCTCATCATGCCGA |
| TGW3 | TGTGTGGCTGGAAATCCGGAT | AATGCCGTTTCCTCCAGCATC |
| TGW6 | AACGAGAGCACCAGCACGAGA | GAGAGCAGCACGAGGAAGACAA |
| OsMKKK10 | GATGCCGAAAATGCAACACTTG | GACACCCCTTGGTGATGTGGAT |
| OsMKK4 | ACGGCTTCACTTGGCCTCGT | GCGTGTGCGTTTTCTGGTGG |
| OsLG3 | ACACTTCGTGTTCGCGTCCAA | ACGACTGCACGAACAAAACCAA |
| BSG1 | AGATCGGCTGCTACCTGACCTAG | GTGGTGATGGTGACGATCCATC |
| Primer | Sequence | Purpose |
|---|---|---|
| MFS4-com-GFP-F-EcoRI | CTATGACCATGATTACGAAGATAGCTCGATCAGCCACAACCAT | Complementary test |
| MFS4-com-GFP-R-BamHI | CATGTCGACTCTAGAGGATCCGATTAGCCCGAGCGTCTGCAC | Complementary test |
| 580-MFS4-XbalI-F | AAGTCCGGAGCTAGCTCTAGAATGACGCTCCCGAGGCAATG | Subcellular localization |
| 580-MFS4-BamHI-R | GCCCTTGCTCACCATGGATCCGATTAGCCCGAGCGTCTGCA | Subcellular localization |
| MFS4-F | GCTCAAGATGGATTGGGAAGGTC | Gene test |
| MFS4-R | TGCCAGATCGTCCAACTGGTGT | Gene test |
| MFS4-COM-F1 | AAGGTGATGGACTTCGTGGGTC | Complementary plant test |
| MFS4-COM-R1 | CGGTGGTGCAGATGAACTTCAG | Complementary plant test |
| MFS4-COM-R2 | CTGCTCAGATTTGGGCGTTCTG | Complementary plant test |
Table S3. Primers used in this study.
| Primer | Sequence | Purpose |
|---|---|---|
| MFS4-com-GFP-F-EcoRI | CTATGACCATGATTACGAAGATAGCTCGATCAGCCACAACCAT | Complementary test |
| MFS4-com-GFP-R-BamHI | CATGTCGACTCTAGAGGATCCGATTAGCCCGAGCGTCTGCAC | Complementary test |
| 580-MFS4-XbalI-F | AAGTCCGGAGCTAGCTCTAGAATGACGCTCCCGAGGCAATG | Subcellular localization |
| 580-MFS4-BamHI-R | GCCCTTGCTCACCATGGATCCGATTAGCCCGAGCGTCTGCA | Subcellular localization |
| MFS4-F | GCTCAAGATGGATTGGGAAGGTC | Gene test |
| MFS4-R | TGCCAGATCGTCCAACTGGTGT | Gene test |
| MFS4-COM-F1 | AAGGTGATGGACTTCGTGGGTC | Complementary plant test |
| MFS4-COM-R1 | CGGTGGTGCAGATGAACTTCAG | Complementary plant test |
| MFS4-COM-R2 | CTGCTCAGATTTGGGCGTTCTG | Complementary plant test |
| [1] | Arora R, Agarwal P, Ray S, Singh A K, Singh V P, Tyagi A K, Kapoor S. 2007. MADS-box gene family in rice: Genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genom, 8(1): 242. |
| [2] | Ashikari M, Sakakibara H, Lin S Y, Yamamoto T, Takashi T, Nishimura A, Angeles E R, Qian Q, Kitano H, Matsuoka M. 2005. Cytokinin oxidase regulates rice grain production. Science, 309: 741‒745. |
| [3] | Aya K, Hobo T, Sato-Izawa K, Ueguchi-Tanaka M, Kitano H, Matsuoka M. 2014. A novel AP2-type transcription factor, SMALL ORGAN SIZE1, controls organ size downstream of an auxin signaling pathway. Plant Cell Physiol, 55(5): 897-912. |
| [4] | Cai Q, Yuan Z, Chen M J, Yin C S, Luo Z J, Zhao X X, Liang W Q, Hu J P, Zhang D B. 2014. Jasmonic acid regulates spikelet development in rice. Nat Commun, 5: 3476. |
| [5] | Chuck G, Meeley R, Hake S. 2008. Floral meristem initiation and meristem cell fate are regulated by the maize AP2 genes ids1 and sid1. Development, 135(18): 3013-3019. |
| [6] | Deng Z Y, Liu L T, Li T, Yan S, Kuang B J, Huang S J, Yan C J, Wang T. 2015. OsKinesin-13A is an active microtubule depolymerase involved in glume length regulation via affecting cell elongation. Sci Rep, 5: 9457. |
| [7] | Dong N Q, Sun Y W, Guo T, Shi C L, Zhang Y M, Kan Y, Xiang Y H, Zhang H, Yang Y B, Li Y C, Zhao H Y, Yu H X, Lu Z Q, Wang Y, Ye W W, Shan J X, Lin H X. 2020. UDP-glucosyltransferase regulates grain size and abiotic stress tolerance associated with metabolic flux redirection in rice. Nat Commun, 11(1): 2629. |
| [8] | Duan P G, Ni S, Wang J M, Zhang B L, Xu R, Wang Y X, Chen H Q, Zhu X D, Li Y H. 2016. Regulation of OsGRF4 by OsmiR396 controls grain size and yield in rice. Nat Plants, 2(1): 1-5. |
| [9] | Feng Z M, Wu C Y, Wang C M, Roh J, Zhang L, Chen J, Zhang S Z, Zhang H, Yang C Y, Hu J L, You Xi M, Liu X, Yang X M, Guo X P, Zhang X, Wu F Q, Terzaghi W, Kim S K, Jiang L, Wan J M. 2016. SLG controls grain size and leaf angle by modulating brassinosteroid homeostasis in rice. J Exp Bot, 67(14): 4241-4253. |
| [10] | Guo T, Lu Z Q, Shan J X, Ye W W, Dong N Q, Lin H X. 2020. ERECTA1 acts upstream of the OsMKKK10-OsMKK4- OsMPK6 cascade to control spikelet number by regulating cytokinin metabolism in rice. Plant Cell, 32(9): 2763-2779. |
| [11] | Heang D, Sassa H. 2012a. Antagonistic actions of HLH/bHLH proteins are involved in grain length and weight in rice. PLoS One, 7(2): e31325. |
| [12] | Heang D, Sassa H. 2012b. An atypical bHLH protein encoded by POSITIVE REGULATOR OF GRAIN LENGTH 2 is involved in controlling grain length and weight of rice through interaction with a typical bHLH protein APG. Breeding Sci, 62(2): 133-141. |
| [13] | Hu J, Wang Y X, Fang Y X, Zeng L J, Xu J, Yu H P, Shi Z Y, Pan J J, Zhang D, Kang S J, Zhu L, Dong G J, Guo L B, Zeng D L, Zhang G H, Xie L H, Xiong G S, Li J Y, Qian Q. 2015. A rare allele of GS2 enhances grain size and grain yield in rice. Mol Plant, 8(10): 1455-1465. |
| [14] | Huang K, Wang D K, Duan P G, Zhang B L, Xu R, Li N, Li Y H. 2017. WIDE AND THICK GRAIN 1, which encodes an otubain- like protease with deubiquitination activity, influences grain size and shape in rice. Plant J, 91(5): 849-860. |
| [15] | Huang X Z, Qian Q, Liu Z B, Sun H Y, He S Y, Luo D, Xia G M, Chu C C, Li J Y, Fu X D. 2009. Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet, 41(4): 494-497. |
| [16] | Ikeda K, Sunohara H, Nagato Y. 2004. Developmental course of inflorescence and spikelet in rice. Breeding Sci, 54(2): 147-156. |
| [17] | Ishimaru K, Hirotsu N, Madoka Y, Murakami N, Hara N, Onodera H, Kashiwagi T, Ujiie K, Shimizu B, Onishi A, Miyagawa H, Katoh E. 2013. Loss of function of the IAA-glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nat Genet, 45(6): 707-713. |
| [18] | Jeon J S, Jang S, Lee S, Nam J, Kim C, Lee S H, Chung Y Y, Kim S R, Lee Y H, Cho Y G, An G. 2000. leafy hull sterile1 is a homeotic mutation in a rice MADS box gene affecting rice flower development. Plant Cell, 12(6): 871-884. |
| [19] | Jin Y, Luo Q, Tong H N, Wang A J, Cheng Z J, Tang J F, Li D Y, Zhao X F, Li X B, Wan J M, Jiao Y L, Chu C C, Zhu L H. 2011. An AT-hook gene is required for palea formation and floral organ number control in rice. Develop Biol, 359(2): 277-288. |
| [20] | Kitagawa K, Kurinami S, Oki K, Abe Y, Ando T, Kono I, Yano M, Kitano H, Iwasaki Y. 2010. A novel kinesin 13 protein regulating rice seed length. Plant Cell Physiol, 51(8): 1315-1329. |
| [21] | La H G, Li J, Ji Z D, Cheng Y J, Li X L, Jiang S Y, Venkatesh P N, Ramachandran S. 2006. Genome-wide analysis of cyclin family in rice (Oryza sativa L.). Mol Genet Genom, 275(4): 374-386. |
| [22] | Lee D Y, An G. 2012. Two AP2 family genes, SUPERNUMERARY BRACT (SNB) and OsINDETERMINATE SPIKELET 1 (OsIDS1), synergistically control inflorescence architecture and floral meristem establishment in rice. Plant J, 69(3): 445-461. |
| [23] | Lee D Y, Lee J, Moon S, Park S Y, An G. 2007. The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem. Plant J, 49(1): 64-78. |
| [24] | Li H F, Liang W Q, Jia R D, Yin C S, Zong J, Kong H Z, Zhang D B. 2010. The AGL6-like gene OsMADS6 regulates floral organ and meristem identities in rice. Cell Res, 20(3): 299-313. |
| [25] | Li H G, Xue D W, Gao Z Y, Yan M X, Xu W Y, Xing Z, Huang D N, Qian Q, Xue Y B. 2009. A putative lipase gene EXTRA GLUME1 regulates both empty-glume fate and spikelet development in rice. Plant J, 57(4): 593-605. |
| [26] | Li X J, Sun L J, Tan L B, Liu F, Zhu Z F, Fu Y C, Sun X Y, Sun X W, Xie D X, Sun C Q. 2012. TH1, a DUF640 domain-like gene controls lemma and palea development in rice. Plant Mol Biol, 78(4/5): 351-359. |
| [27] | Li Y B, Fan C C, Xing Y Z, Jiang Y H, Luo L J, Sun L, Shao D, Xu C J, Li X H, Xiao J H, He Y Q, Zhang Q F. 2011. Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet, 43(12): 1266-1270. |
| [28] | Li Y F, Zeng X Q, Li Y, Wang L, Zhuang H, Wang Y, Tang J, Wang H L, Xiong M, Yang F Y, Yuan X Z, He G H. 2020. MULTI-FLORET SPIKELET 2, a MYB transcription factor, determines spikelet meristem fate and floral organ identity in rice. Plant Physiol, 184: 988‒1003. |
| [29] | Ma X D, Cheng Z J, Wu F Q, Jin M G, Zhang L G, Zhou F, Wang J L, Zhou K N, Ma J, Lin Q B, Lei C L, Wan J M. 2013. BEAK LIKE SPIKELET1 is required for lateral development of lemma and palea in rice. Plant Mol Biol Rep, 31(1): 98-108. |
| [30] | Malcomber S T, Preston J C, Reinheimer R, Kossuth J, Kellogg E A. 2006. Developmental gene evolution and the origin of grass inflorescence diversity. Adv Bot Res, 44: 425‒481. |
| [31] | Nagasawa N, Miyoshi M, Sano Y, Satoh H, Hirano H, Sakai H, Nagato Y. 2003. SUPERWOMAN1 and DROOPING LEAF genes control floral organ identity in rice. Development, 130(4): 705-718. |
| [32] | Ren D Y, Li Y F, Zhao F M, Sang X C, Shi J Q, Wang N, Guo S, Ling Y H, Zhang C W, Yang Z L, He G H. 2013. MULTI- FLORET SPIKELET1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice. Plant Physiol, 162(2): 872-884. |
| [33] | Ren D Y, Rao Y C, Wu L W, Xu Q K, Li Z Z, Yu H P, Zhang Y, Leng Y J, Hu J, Zhu L, Gao Z Y, Dong G J, Zhang G H, Guo L B, Zeng D L, Qian Q. 2016. The pleiotropic ABNORMAL FLOWER AND DWARF1 affects plant height, floral development and grain yield in rice. J Integr Plant Biol, 58(6): 529-539. |
| [34] | Ren D Y, Hu J, Xu Q K, Cui Y J, Zhang Y, Zhou T T, Rao Y C, Xue D W, Zeng D L, Zhang G H, Gao Z Y, Zhu L, Shen L, Chen G, Guo L B, Qian Q. 2018a. FZP determines grain size and sterile lemma fate in rice. J Exp Bot, 69(20): 4853-4866. |
| [35] | Ren D Y, Yu H P, Rao Y C, Xu Q K, Zhou T T, Hu J, Zhang Y, Zhang G H, Zhu L, Gao Z Y, Chen G, Guo L B, Zeng D L, Qian Q. 2018b. ‘Two-floret spikelet’ as a novel resource has the potential to increase rice yield. Plant Biotechnol J, 16(2): 351-353. |
| [36] | Ren D Y, Rao Y C, Yu H P, Xu Q K, Cui Y J, Xia S S, Yu X Q, Liu H, Hu H T, Xue D W, Zeng D L, Hu J, Zhang G H, Gao Z Y, Zhu L, Zhang Q, Shen L, Guo L B, Qian Q. 2020. MORE FLORET 1 encodes a MYB transcription factor that regulates spikelet development in rice. Plant Physiol, 184(1): 251-265. |
| [37] | Ruan B P, Shang L G, Zhang B, Hu J, Wang Y X, Lin H, Zhang A P, Liu C L, Peng Y L, Zhu L, Ren D Y, Shen L, Dong G J, Zhang G H, Zeng D L, Guo L B, Qian Q, Gao Z Y. 2020. Natural variation in the promoter of TGW2 determines grain width and weight in rice. New Phytol, 227(2): 629-640. |
| [38] | Segami S, Kono I, Ando T, Yano M, Kitano H, Miura K, Iwasaki Y. 2012. Small and round seed 5 gene encodes alpha-tubulin regulating seed cell elongation in rice. Rice, 5: 4. |
| [39] | Song X J, Huang W, Shi M, Zhu M Z, Lin H X. 2007. A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet, 39(5): 623-630. |
| [40] | Su’udi M, Cha J Y, Ahn I P, Kwak Y S, Woo Y M, Son D Y. 2012. Functional characterization of a B-type cell cycle switch 52 in rice (OsCCS52B). Plant Cell Tissue Organ Cult, 111(1): 101-111. |
| [41] | Wan X Y, Weng J F, Zhai H Q, Wang J K, Lei C L, Liu X L, Guo T, Jiang L, Su N, Wan J M. 2008. Quantitative trait loci (QTL) analysis for rice grain width and fine mapping of an identified QTL allele gw-5 in a recombination hotspot region on chromosome 5. Genetics, 179(4): 2239-2252. |
| [42] | Wang K J, Tang D, Hong L L, Xu W Y, Huang J, Li M, Gu M H, Xue Y B, Cheng Z K. 2010. DEP and AFO regulate reproductive habit in rice. PLoS Genet, 6(1): e1000818. |
| [43] | Wang S K, Wu K, Yuan Q B, Liu X Y, Liu Z B, Lin X Y, Zeng R Z, Zhu H T, Dong G J, Qian Q, Zhang G Q, Fu X D. 2012. Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet, 44(8): 1-6. |
| [44] | Wu T, Shen Y Y, Zheng M, Yang C Y, Chen Y L, Feng Z M, Liu X, Liu S J, Chen Z J, Lei C L, Wang J L, Jiang L, Wan J M. 2014. Gene SGL, encoding a kinesin-like protein with transactivation activity, is involved in grain length and plant height in rice. Plant Cell Rep, 33(2): 235-244. |
| [45] | Xia D, Zhou H, Liu R J, Dan W H, Li P B, Wu B, Chen J X, Wang L Q, Gao G J, Zhang Q L, He Y Q. 2018. GL3.3, a novel QTL encoding a GSK3/SHAGGY-like kinase, epistatically interacts with GS3 to produce extra-long grains in rice. Mol Plant, 11(5): 754-756. |
| [46] | Xiong H Y, Yu J P, Miao J L, Li J J, Zhang H L, Wang X, Liu P L, Zhao Y, Jiang C H, Yin Z G, Li Y, Guo Y, Fu B Y, Wang W S, Li Z K, Ali J, Li Z C. 2018. Natural variation in OsLG3 increases drought tolerance in rice by inducing ROS scavenging. Plant Physiol, 178(1): 451-467. |
| [47] | Xue W Y, Xing Y Z, Weng X Y, Zhao Y, Tang W J, Wang L, Zhou H J, Yu S B, Xu C G, Li X H, Zhang Q F. 2008. Natural variation inGhd7 is an important regulator of heading date and yield potential in rice. Nat Genet, 40: 761. |
| [48] | Yan S, Zou G H, Li S J, Wang H, Liu H Q, Zhai G W, Guo P, Song H M, Yan C J, Tao Y Z. 2011. Seed size is determined by the combinations of the genes controlling different seed characteristics in rice. Theor Appl Genet, 123(7): 1173-1181. |
| [49] | Ying J Z, Ma M, Bai C, Huang X H, Liu J L, Fan Y Y, Song X J. 2018. TGW3, a major QTL that negatively modulates grain length and weight in rice. Mol Plant, 11(5): 750-753. |
| [50] | Yoshida A, Suzaki T, Tanaka W, Hirano H Y. 2009. The homeotic gene long sterile lemma (G1) specifies sterile lemma identity in the rice spikelet. Proc Natl Acad Sci USA, 106(47): 20103-20108. |
| [51] | Zhang B Y, Wu S H, Zhang Y E, Xu T, Guo F F, Tang H S, Li X, Wang P F, Qian W F, Xue Y B. 2016. A high temperature- dependent mitochondrial lipase EXTRA GLUME1 promotes floral phenotypic robustness against temperature fluctuation in rice (Oryza sativa L.). PLoS Genet, 12(7): e1006152. |
| [52] | Zhang L, Yu H, Ma B, Liu G F, Wang J J, Wang J M, Gao R C, Li J J, Liu J Y, Xu J, Zhang Y Y, Li Q, Huang X H, Xu J L, Li J M, Qian Q, Han B, He Z H, Li J Y. 2017. A natural tandem array alleviates epigenetic repression of IPA1 and leads to superior yielding rice. Nat Commun, 8(1): 14789. |
| [53] | Zheng H, Zhang J, Zhuang H, Zeng X Q, Tang J, Wang H L, Chen H, Li Y, Ling Y H, He G H, Li Y F. 2019. Gene mapping and candidate gene analysis of multi-floret spikelet 3 (mfs3) in rice (Oryza sativa L.). J Integr Agric, 18(12): 2673-2681. |
| [54] | Zhu Q H, Upadhyaya N M, Gubler F, Helliwell C A. 2009. Over- expression of miR172 causes loss of spikelet determinacy and floral organ abnormalities in rice (Oryza sativa). BMC Plant Biol, 9: 149. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [13] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [14] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| [15] | Zhang Guomei, Li Han, Liu Shanshan, Zhou Xuming, Lu Mingyang, Tang Liang, Sun Lihua. Water Extract of Rice False Smut Balls Activates Nrf2/HO-1 and Apoptosis Pathways, Causing Liver Injury [J]. Rice Science, 2023, 30(5): 473-485. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||