
Rice Science ›› 2022, Vol. 29 ›› Issue (1): 76-88.DOI: 10.1016/j.rsci.2021.12.007
• Research Paper • Previous Articles Next Articles
Zhang Chengming1, Nobuhiro Tanaka2, Maria Stefanie Dwiyanti1, Matthew Shenton2, Hayato Maruyama1, Takuro Shinano1, Chu Qingnan3, Xie Jun4, Toshihiro Watanabe1( )
)
Received:2021-01-01
Accepted:2021-05-13
Online:2022-01-28
Published:2022-01-01
Contact:
Toshihiro Watanabe
Zhang Chengming, Nobuhiro Tanaka, Maria Stefanie Dwiyanti, Matthew Shenton, Hayato Maruyama, Takuro Shinano, Chu Qingnan, Xie Jun, Toshihiro Watanabe. Ionomic Profiling of Rice Genotypes and Identification of Varieties with Elemental Covariation Effects[J]. Rice Science, 2022, 29(1): 76-88.
Add to citation manager EndNote|Ris|BibTeX
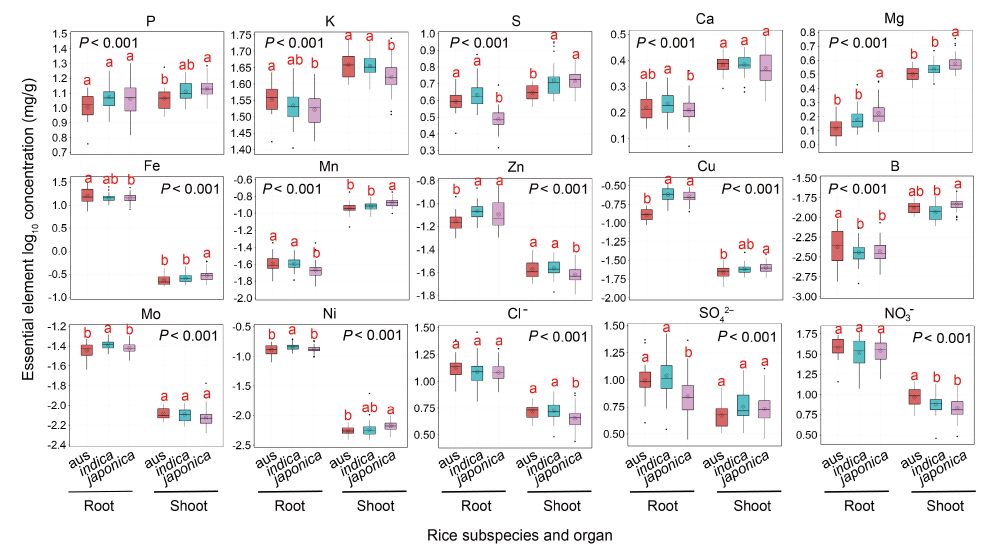
Fig. 1. Boxplot showing log 10 concentrations of 15 essential elements (P, K, S, Ca, Mg, Fe, Mn, Zn, Cu, B, Mo, Ni, Cl?, SO42? and NO3?) in rice shoots and roots.
| Element | Shoot | Root | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| F value a | Contribution (%) b | CV of japonica | CV of indica | CV of aus | F value a | Contribution (%) b | CV of japonica | CV of indica | CV of aus | ||
| P | 3.31** | 63.57 | 0.14 | 0.16 | 0.19 | 1.08 | 37.24 | 0.27 | 0.29 | 0.19 | |
| K | 2.58** | 56.71 | 0.09 | 0.07 | 0.09 | 3.60** | 64.69 | 0.10 | 0.11 | 0.10 | |
| S | 2.76** | 58.33 | 0.13 | 0.23 | 0.10 | 8.10** | 80.47 | 0.14 | 0.21 | 0.15 | |
| Ca | 2.65** | 57.34 | 0.14 | 0.08 | 0.08 | 1.16 | 37.05 | 0.10 | 0.10 | 0.12 | |
| Mg | 4.91** | 71.38 | 0.14 | 0.12 | 0.11 | 12.11** | 86.03 | 0.20 | 0.19 | 0.17 | |
| Fe | 1.29 | 39.48 | 0.30 | 0.21 | 0.28 | 1.53* | 44.64 | 0.20 | 0.20 | 0.37 | |
| Mn | 3.13** | 61.29 | 0.11 | 0.11 | 0.19 | 3.56** | 64.44 | 0.22 | 0.16 | 0.26 | |
| Zn | 1.54* | 43.90 | 0.17 | 0.16 | 0.19 | 1.32 | 40.24 | 0.30 | 0.17 | 0.20 | |
| Cu | 3.74** | 65.46 | 0.15 | 0.17 | 0.21 | 9.13** | 82.27 | 0.15 | 0.20 | 0.18 | |
| B | 2.86** | 59.19 | 0.16 | 0.25 | 0.17 | 0.81 | 29.09 | 0.31 | 0.30 | 0.47 | |
| Mo | 2.96** | 59.99 | 0.22 | 0.16 | 0.14 | 3.07** | 60.99 | 0.13 | 0.11 | 0.16 | |
| Ni | 1.05 | 34.73 | 0.17 | 0.51 | 0.16 | 1.63* | 45.36 | 0.12 | 0.13 | 0.19 | |
| Cl‒ | 1.24 | 40.21 | 0.22 | 0.19 | 0.16 | 1.08 | 41.28 | 0.21 | 0.30 | 0.26 | |
| SO42‒ | 2.13** | 53.69 | 0.30 | 0.38 | 0.28 | 2.19** | 59.04 | 0.47 | 0.43 | 0.44 | |
| NO3‒ | 1.57* | 45.91 | 0.28 | 0.24 | 0.27 | 1.39 | 47.32 | 0.29 | 0.35 | 0.26 | |
| Al | 0.85 | 30.21 | 0.49 | 0.53 | 0.36 | 1.15 | 37.07 | 0.25 | 0.21 | 0.24 | |
| Ba | 2.70** | 57.80 | 0.18 | 0.14 | 0.14 | 3.55** | 64.35 | 0.14 | 0.11 | 0.19 | |
| Na | 2.33** | 54.19 | 0.29 | 0.34 | 0.30 | 3.03** | 60.61 | 0.26 | 0.32 | 0.30 | |
| Sr | 2.98** | 60.18 | 0.17 | 0.16 | 0.16 | 2.60** | 56.92 | 0.14 | 0.12 | 0.17 | |
| As | 4.11** | 67.56 | 0.12 | 0.09 | 0.10 | 4.53** | 69.71 | 0.15 | 0.18 | 0.24 | |
| Cd | 8.76** | 81.61 | 0.30 | 0.19 | 0.32 | 6.77** | 77.50 | 0.19 | 0.20 | 0.23 | |
| Co | 2.99** | 60.19 | 0.27 | 0.21 | 0.26 | 4.56** | 69.85 | 0.17 | 0.17 | 0.20 | |
| Cr | 1.67** | 46.07 | 0.52 | 0.68 | 0.51 | 1.47* | 42.76 | 0.20 | 0.14 | 0.22 | |
| Cs | 4.21** | 68.07 | 0.13 | 0.16 | 0.20 | 9.19** | 82.37 | 0.10 | 0.16 | 0.18 | |
| Li | 1.87** | 48.69 | 0.28 | 0.21 | 0.21 | 1.15 | 36.97 | 0.20 | 0.17 | 0.19 | |
| Se | 6.11** | 75.56 | 0.14 | 0.21 | 0.35 | 8.83** | 81.78 | 0.18 | 0.20 | 0.28 | |
Table 1. Variations in elemental concentrations in shoots and roots among 120 rice varieties.
| Element | Shoot | Root | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| F value a | Contribution (%) b | CV of japonica | CV of indica | CV of aus | F value a | Contribution (%) b | CV of japonica | CV of indica | CV of aus | ||
| P | 3.31** | 63.57 | 0.14 | 0.16 | 0.19 | 1.08 | 37.24 | 0.27 | 0.29 | 0.19 | |
| K | 2.58** | 56.71 | 0.09 | 0.07 | 0.09 | 3.60** | 64.69 | 0.10 | 0.11 | 0.10 | |
| S | 2.76** | 58.33 | 0.13 | 0.23 | 0.10 | 8.10** | 80.47 | 0.14 | 0.21 | 0.15 | |
| Ca | 2.65** | 57.34 | 0.14 | 0.08 | 0.08 | 1.16 | 37.05 | 0.10 | 0.10 | 0.12 | |
| Mg | 4.91** | 71.38 | 0.14 | 0.12 | 0.11 | 12.11** | 86.03 | 0.20 | 0.19 | 0.17 | |
| Fe | 1.29 | 39.48 | 0.30 | 0.21 | 0.28 | 1.53* | 44.64 | 0.20 | 0.20 | 0.37 | |
| Mn | 3.13** | 61.29 | 0.11 | 0.11 | 0.19 | 3.56** | 64.44 | 0.22 | 0.16 | 0.26 | |
| Zn | 1.54* | 43.90 | 0.17 | 0.16 | 0.19 | 1.32 | 40.24 | 0.30 | 0.17 | 0.20 | |
| Cu | 3.74** | 65.46 | 0.15 | 0.17 | 0.21 | 9.13** | 82.27 | 0.15 | 0.20 | 0.18 | |
| B | 2.86** | 59.19 | 0.16 | 0.25 | 0.17 | 0.81 | 29.09 | 0.31 | 0.30 | 0.47 | |
| Mo | 2.96** | 59.99 | 0.22 | 0.16 | 0.14 | 3.07** | 60.99 | 0.13 | 0.11 | 0.16 | |
| Ni | 1.05 | 34.73 | 0.17 | 0.51 | 0.16 | 1.63* | 45.36 | 0.12 | 0.13 | 0.19 | |
| Cl‒ | 1.24 | 40.21 | 0.22 | 0.19 | 0.16 | 1.08 | 41.28 | 0.21 | 0.30 | 0.26 | |
| SO42‒ | 2.13** | 53.69 | 0.30 | 0.38 | 0.28 | 2.19** | 59.04 | 0.47 | 0.43 | 0.44 | |
| NO3‒ | 1.57* | 45.91 | 0.28 | 0.24 | 0.27 | 1.39 | 47.32 | 0.29 | 0.35 | 0.26 | |
| Al | 0.85 | 30.21 | 0.49 | 0.53 | 0.36 | 1.15 | 37.07 | 0.25 | 0.21 | 0.24 | |
| Ba | 2.70** | 57.80 | 0.18 | 0.14 | 0.14 | 3.55** | 64.35 | 0.14 | 0.11 | 0.19 | |
| Na | 2.33** | 54.19 | 0.29 | 0.34 | 0.30 | 3.03** | 60.61 | 0.26 | 0.32 | 0.30 | |
| Sr | 2.98** | 60.18 | 0.17 | 0.16 | 0.16 | 2.60** | 56.92 | 0.14 | 0.12 | 0.17 | |
| As | 4.11** | 67.56 | 0.12 | 0.09 | 0.10 | 4.53** | 69.71 | 0.15 | 0.18 | 0.24 | |
| Cd | 8.76** | 81.61 | 0.30 | 0.19 | 0.32 | 6.77** | 77.50 | 0.19 | 0.20 | 0.23 | |
| Co | 2.99** | 60.19 | 0.27 | 0.21 | 0.26 | 4.56** | 69.85 | 0.17 | 0.17 | 0.20 | |
| Cr | 1.67** | 46.07 | 0.52 | 0.68 | 0.51 | 1.47* | 42.76 | 0.20 | 0.14 | 0.22 | |
| Cs | 4.21** | 68.07 | 0.13 | 0.16 | 0.20 | 9.19** | 82.37 | 0.10 | 0.16 | 0.18 | |
| Li | 1.87** | 48.69 | 0.28 | 0.21 | 0.21 | 1.15 | 36.97 | 0.20 | 0.17 | 0.19 | |
| Se | 6.11** | 75.56 | 0.14 | 0.21 | 0.35 | 8.83** | 81.78 | 0.18 | 0.20 | 0.28 | |
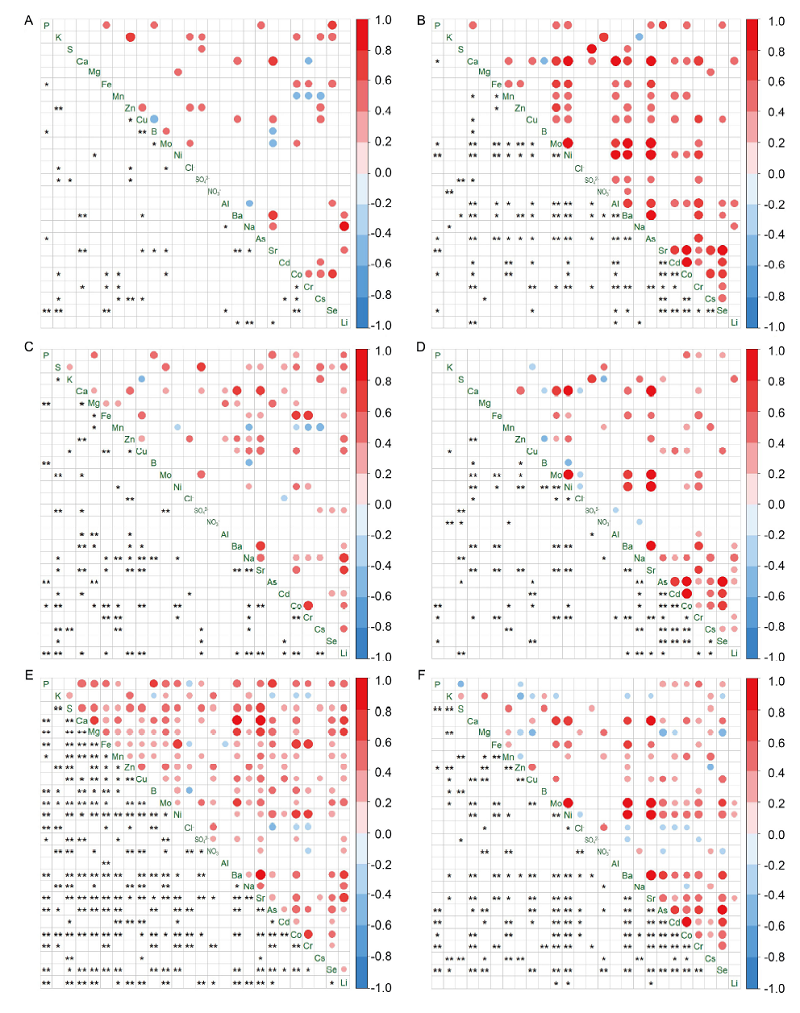
Fig. 3. Correlation heatmaps for aus shoots (A), aus roots (B), indica shoots (C), indica roots (D), japonica shoots (E) and japonica roots (F). nly significant (P < 0.05) correlations are displayed in the Pearson’s correlation analysis. The circle in upper triangle matrix represents significant correlation. A larger circle indicates a stronger correlation between the row and the column variables. The red circle indicates a positive correlation coefficient, and the blue indicates a negative one. * and ** in lower triangle matrix represent P < 0.05 and P < 0.01, respectively.
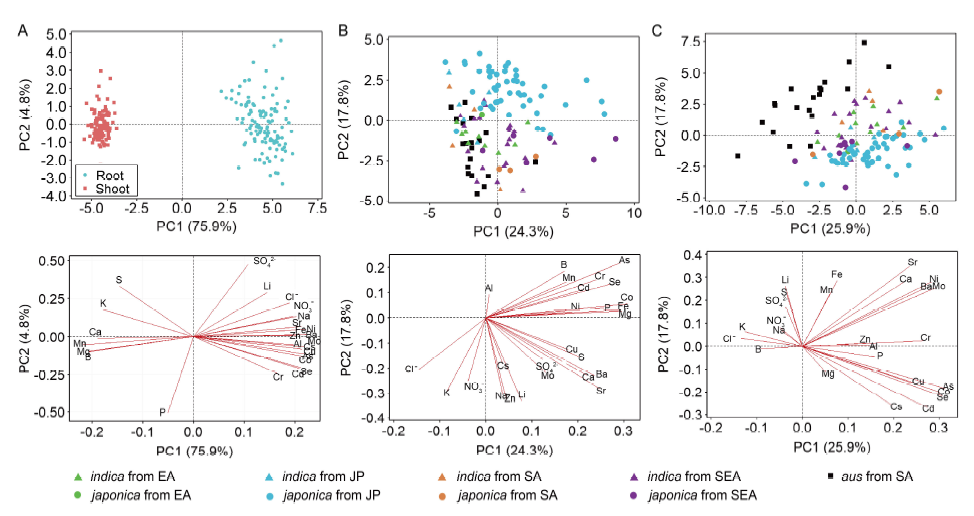
Fig. 4. Combination of principal component analysis (PCA) score plots and loading plots for all samples (A), shoots (B) and roots (C). EA, JP, SA and SEA represent East Asia (except Japan), Japan, South Asia and Southeast Asia, respectively.
| ID | Variety | Subspecies | High | Low | |||
|---|---|---|---|---|---|---|---|
| Essential element | Nonessential element | Essential element | Nonessential element | ||||
| JRC06 | Kaneko B | japonica | Mn, B | As, Cd, Co, Cr, Se | K | ‒ | |
| JRC12 | Oiran | japonica | Ca, Mg, Mn | Sr, Co, Se | Mo | ‒ | |
| JRC13 | Bouzu Mochi | japonica | Mg, Fe, B, Ni | As, Cd, Se | ‒ | ‒ | |
| JRC36 | Sekiyama | japonica | ‒ | ‒ | K, Zn, Cu, Mo | Ba, Sr | |
| JRC37 | Shinyamadaho 2 | japonica | ‒ | Al | S, Ca, Mo | Ba, Sr, Co | |
| JRC40 | Akamai | indica | ‒ | ‒ | Ca, Zn | Ba, Sr | |
| JRC49 | Rikutou Rikuu 2 | japonica | ‒ | ‒ | Zn, Mo, SO42‒ | Na, Li | |
| JRC51 | Shinshuu | japonica | ‒ | ‒ | NO3‒ | Na, Sr, Li | |
| WRC04 | Jena 035 | aus | ‒ | Li | P | Na, As, Cd, Se | |
| WRC10 | Shuusoushu | indica | Cl‒ | ‒ | ‒ | Co, Cs | |
| WRC11 | Jinguoyin | indica | Zn, Cl‒ | ‒ | NO3‒ | Cd | |
| WRC14 | IR58 | indica | Mg, Mo | Al | ‒ | Cs | |
| WRC18 | Qingyu (Seiyu) | indica | ‒ | ‒ | P, Ni | Co, Cs | |
| WRC19 | Deng Pao Zhai | indica | ‒ | ‒ | Ni | Co, Cr | |
| WRC20 | Tadukan | indica | ‒ | ‒ | Ni | Co, Cr | |
| WRC21 | Shwe Nang Gyi | indica | Mo, SO42‒ | Se | ‒ | ‒ | |
| WRC23 | Lebed | japonica | ‒ | ‒ | Ni | Cr, Cs | |
| WRC24 | Pinulupot 1 | indica | ‒ | ‒ | S, B | Cs | |
| WRC25 | Muha | aus | ‒ | ‒ | S, SO42‒ | Cd, Se | |
| WRC26 | Jhona 2 | aus | K, Zn, Cu | ‒ | ‒ | Cr | |
| WRC27 | Nepal 8 | aus | ‒ | Li | ‒ | Cd, Se | |
| WRC29 | Kalo Dhan | aus | ‒ | Na, Li | P, Mg, Mn | Cd, Se | |
| WRC30 | Anjana Dhan | aus | K, Zn | Co, Cs | Mg, Mn | ‒ | |
| WRC32 | Tupa 121-3 | aus | Mn | ‒ | Cu, Ni | As | |
| WRC33 | Surjamukhi | aus | ‒ | ‒ | Mg, Cu, Ni | Ba, Sr | |
| WRC38 | Arc 11094 | aus | ‒ | ‒ | S, Ca, Cu | Sr, As, Se | |
| WRC42 | Local Basmati | aus | P, K, NO3‒ | Se | ‒ | ‒ | |
| WRC47 | Jaguary | japonica | Ca, Zn, Cu | Al | ‒ | ‒ | |
| WRC48 | Khau Mac Kho | japonica | P, Ca, Mg, B, Mo | Ba, Na, Sr, As | ‒ | ‒ | |
| WRC49 | Padi Perak | japonica | P, S, Fe, B, Mo, Ni | Ba, Sr, Co, Cr, Se | ‒ | ‒ | |
| WRC50 | Rexmont | japonica | P, Fe, Ni | Co, Cr | Cl-, NO3‒ | ‒ | |
| WRC51 | Urasan 1 | japonica | Ca, Mg, Fe | Ba, Sr, As, Co, Cr | ‒ | ‒ | |
| WRC52 | Khau Tan Chiem | japonica | Cu, NO3‒ | Cd | Ca | Al | |
| WRC53 | Tima | japonica | S, Mn, SO42‒ | Cs | ‒ | ‒ | |
| WRC55 | Tupa 729 | japonica | K, NO3‒ | Cs | ‒ | ‒ | |
| WRC57 | Milyang 23 | indica | Fe, Ni | Cr | ‒ | ‒ | |
| WRC58 | Neang Menh | indica | S, SO42‒ | Na | Mn, B | As | |
| WRC60 | Hakphaynhay | indica | S, SO42‒ | ‒ | P, Fe, B | Al | |
| WRC63 | Bleiyo | indica | Zn, Cl‒ | Sr | ‒ | ‒ | |
| WRC65 | Rambhog | indica | K, S | Cs | Mn | ‒ | |
| WRC67 | Phulba | japonica | Ca, Mn, Cl‒ | Li | ‒ | ‒ | |
| WRC68 | Khao Nam Jen | japonica | Mg, Cu | Cd, Li | ‒ | ‒ | |
Table 2. Highest and lowest multi-element accumulation varieties for elements in shoots.
| ID | Variety | Subspecies | High | Low | |||
|---|---|---|---|---|---|---|---|
| Essential element | Nonessential element | Essential element | Nonessential element | ||||
| JRC06 | Kaneko B | japonica | Mn, B | As, Cd, Co, Cr, Se | K | ‒ | |
| JRC12 | Oiran | japonica | Ca, Mg, Mn | Sr, Co, Se | Mo | ‒ | |
| JRC13 | Bouzu Mochi | japonica | Mg, Fe, B, Ni | As, Cd, Se | ‒ | ‒ | |
| JRC36 | Sekiyama | japonica | ‒ | ‒ | K, Zn, Cu, Mo | Ba, Sr | |
| JRC37 | Shinyamadaho 2 | japonica | ‒ | Al | S, Ca, Mo | Ba, Sr, Co | |
| JRC40 | Akamai | indica | ‒ | ‒ | Ca, Zn | Ba, Sr | |
| JRC49 | Rikutou Rikuu 2 | japonica | ‒ | ‒ | Zn, Mo, SO42‒ | Na, Li | |
| JRC51 | Shinshuu | japonica | ‒ | ‒ | NO3‒ | Na, Sr, Li | |
| WRC04 | Jena 035 | aus | ‒ | Li | P | Na, As, Cd, Se | |
| WRC10 | Shuusoushu | indica | Cl‒ | ‒ | ‒ | Co, Cs | |
| WRC11 | Jinguoyin | indica | Zn, Cl‒ | ‒ | NO3‒ | Cd | |
| WRC14 | IR58 | indica | Mg, Mo | Al | ‒ | Cs | |
| WRC18 | Qingyu (Seiyu) | indica | ‒ | ‒ | P, Ni | Co, Cs | |
| WRC19 | Deng Pao Zhai | indica | ‒ | ‒ | Ni | Co, Cr | |
| WRC20 | Tadukan | indica | ‒ | ‒ | Ni | Co, Cr | |
| WRC21 | Shwe Nang Gyi | indica | Mo, SO42‒ | Se | ‒ | ‒ | |
| WRC23 | Lebed | japonica | ‒ | ‒ | Ni | Cr, Cs | |
| WRC24 | Pinulupot 1 | indica | ‒ | ‒ | S, B | Cs | |
| WRC25 | Muha | aus | ‒ | ‒ | S, SO42‒ | Cd, Se | |
| WRC26 | Jhona 2 | aus | K, Zn, Cu | ‒ | ‒ | Cr | |
| WRC27 | Nepal 8 | aus | ‒ | Li | ‒ | Cd, Se | |
| WRC29 | Kalo Dhan | aus | ‒ | Na, Li | P, Mg, Mn | Cd, Se | |
| WRC30 | Anjana Dhan | aus | K, Zn | Co, Cs | Mg, Mn | ‒ | |
| WRC32 | Tupa 121-3 | aus | Mn | ‒ | Cu, Ni | As | |
| WRC33 | Surjamukhi | aus | ‒ | ‒ | Mg, Cu, Ni | Ba, Sr | |
| WRC38 | Arc 11094 | aus | ‒ | ‒ | S, Ca, Cu | Sr, As, Se | |
| WRC42 | Local Basmati | aus | P, K, NO3‒ | Se | ‒ | ‒ | |
| WRC47 | Jaguary | japonica | Ca, Zn, Cu | Al | ‒ | ‒ | |
| WRC48 | Khau Mac Kho | japonica | P, Ca, Mg, B, Mo | Ba, Na, Sr, As | ‒ | ‒ | |
| WRC49 | Padi Perak | japonica | P, S, Fe, B, Mo, Ni | Ba, Sr, Co, Cr, Se | ‒ | ‒ | |
| WRC50 | Rexmont | japonica | P, Fe, Ni | Co, Cr | Cl-, NO3‒ | ‒ | |
| WRC51 | Urasan 1 | japonica | Ca, Mg, Fe | Ba, Sr, As, Co, Cr | ‒ | ‒ | |
| WRC52 | Khau Tan Chiem | japonica | Cu, NO3‒ | Cd | Ca | Al | |
| WRC53 | Tima | japonica | S, Mn, SO42‒ | Cs | ‒ | ‒ | |
| WRC55 | Tupa 729 | japonica | K, NO3‒ | Cs | ‒ | ‒ | |
| WRC57 | Milyang 23 | indica | Fe, Ni | Cr | ‒ | ‒ | |
| WRC58 | Neang Menh | indica | S, SO42‒ | Na | Mn, B | As | |
| WRC60 | Hakphaynhay | indica | S, SO42‒ | ‒ | P, Fe, B | Al | |
| WRC63 | Bleiyo | indica | Zn, Cl‒ | Sr | ‒ | ‒ | |
| WRC65 | Rambhog | indica | K, S | Cs | Mn | ‒ | |
| WRC67 | Phulba | japonica | Ca, Mn, Cl‒ | Li | ‒ | ‒ | |
| WRC68 | Khao Nam Jen | japonica | Mg, Cu | Cd, Li | ‒ | ‒ | |
| [1] | Affholder M C, Flöhr A, Kirchmann H. 2019. Can Cd content in crops be controlled by Se fertilization? A meta-analysis and outline of Cd sequestration mechanisms. Plant Soil, 440(1/2): 369-380. |
| [2] |
Amtmann A, Troufflard S, Armengaud P. 2008. The effect of potassium nutrition on pest and disease resistance in plants. Physiol Plant, 133(4): 682-691.
PMID |
| [3] | Arao T, Ishikawa S, Murakami M, Abe K, Maejima Y, Makino T. 2010. Heavy metal contamination of agricultural soil and countermeasures in Japan. Paddy Water Environ, 8(3): 247-257. |
| [4] | Balonov M I, Bruk G Y, Golikov V Y, Barkovsky A N, Kravtsova E M, Kravtosova O S, Mubasarov A A, Shutov V N, Travnikova I G, Howard B J, Brown J E, Strand P. 2007. Assessment of current exposure of the population living in the Techa River Basin from radioactive releases of the Mayak facility. Health Phys, 92(2): 134-147. |
| [5] |
Baxter I, Dilkes B P. 2012. Elemental profiles reflect plant adaptations to the environment. Science, 336: 1661-1663.
PMID |
| [6] | Baxter I R, Vitek O, Lahner B, Muthukumar B, Borghi M, Morrissey J, Guerinot M L, Salt D E. 2008. The leaf ionome as a multivariable system to detect a plant’s physiological status. Proc Natl Acad Sci USA, 105(33): 12081-12086. |
| [7] | Becker M, Asch F. 2005. Iron toxicity in rice: Conditions and management concepts. J Plant Nutr Soil Sci, 168(4): 558-573. |
| [8] |
Campbell M T, Du Q, Liu K, Sharma S, Zhang C, Walia H. 2020. Characterization of the transcriptional divergence between the subspecies of cultivated rice (Oryza sativa). BMC Genomics, 21(1): 394.
PMID |
| [9] | Cao Y, Sun D, Ai H, Mei H Y, Liu X, Sun S B, Xu G H, Liu Y G, Chen Y S, Ma L Q. 2017. Knocking out OsPT4 gene decreases arsenate uptake by rice plants and inorganic arsenic accumulation in rice grains. Environ Sci Technol, 51(21): 12131-12138. |
| [10] | Chen H P, Tang Z, Wang P, Zhao F J. 2018. Geographical variations of cadmium and arsenic concentrations and arsenic speciation in Chinese rice. Environ Pollut, 238: 482-490. |
| [11] | Chen H R, Yang Y, Ye Y F, Tao L Z, Fu X D, Liu B M, Wu Y J. 2019. Differences in cadmium accumulation between indica and japonica rice cultivars in the reproductive stage. Ecotoxicol Environ Saf, 186: 109795. |
| [12] | Chen Z, Watanabe T, Shinano T, Okazaki K, Osaki M. 2009. Rapid characterization of plant mutants with an altered ion-profile: A case study using Lotus japonicus. New Phytol, 181: 795-801. |
| [13] | Chu Q N, Watanabe T, Sha Z M, Osaki M, Shinano T. 2015. Interactions between Cs, Sr, and other nutrients and trace element accumulation in Amaranthus shoot in response to variety effect. J Agric Food Chem, 63(8): 2355-2363. |
| [14] | Civáň P, Craig H, Cox C J, Brown T A. 2015. Three geographically separate domestications of Asian rice. Nat Plants, 1: 15164. |
| [15] | Clemens S. 2014. Zn and Fe biofortification: The right chemical environment for human bioavailability. Plant Sci, 225: 52-57. |
| [16] | Curie C, Cassin G, Couch D, Divol F, Higuchi K, Le Jean M, Misson J, Schikora A, Czernic P, Mari S. 2009. Metal movement within the plant: Contribution of nicotianamine and yellow stripe 1-like transporters. Ann Bot, 103(1): 1-11. |
| [17] | Du F, Liu P, Wang K, Yang Z G, Wang L. 2020. Ionomic responses of rice plants to the stresses of different arsenic species in hydroponics. Chemosphere, 243: 125398. |
| [18] | El Azhari A, Rhoujjati A, El Hachimi M L, Ambrosi J. 2017. Pollution and ecological risk assessment of heavy metals in the soil-plant system and the sediment-water column around a former Pb/Zn-mining area in NE Morocco. Ecotoxicol Environ Saf, 144: 464-474. |
| [19] |
El Mehdawi A F, Jiang Y, Guignardi Z S, Esmat A, Pilon M, Pilon-Smits E A H, Schiavon M. 2018. Influence of sulfate supply on selenium uptake dynamics and expression of sulfate/ selenate transporters in selenium hyperaccumulator and nonhyperaccumulator Brassiaceae. New Phytol, 217(1): 194-205.
PMID |
| [20] | Feng X M, Han L, Chao D Y, Liu Y, Zhang Y J, Wang R G, Guo J K, Feng R W, Xu Y M, Ding Y Z, Huang B Y, Zhang G L. 2017. Ionomic and transcriptomic analysis provides new insight into the distribution and transport of cadmium and arsenic in rice. J Hazard Mater, 331: 246-256. |
| [21] | Gu J F, Zhou H, Tang H L, Yang W T, Zeng M, Liu Z M, Peng P Q, Liao B H. 2019. Cadmium and arsenic accumulation during the rice growth period under in situ remediation. Ecotoxicol Environ Saf, 171: 451-459. |
| [22] | Huang X H, Wei X H, Sang T, Zhao Q, Feng Q, Zhao Y, Li C Y, Zhu C R, Lu T T, Zhang Z W, Li M, Fan D L, Guo Y L, Wang A H, Wang L, Deng L W, Li W J, Lu Y Q, Weng Q J, Liu K Y, Huang T, Zhou T Y, Jing Y F, Li W, Lin Z, Buckler E S, Qian Q, Zhang Q F, Li J Y, Han B. 2010. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet, 42(11): 961-967. |
| [23] | Huang X Y, Salt D E. 2016. Plant ionomics: From elemental profiling to environmental adaptation. Mol Plant, 9(6): 787-797. |
| [24] | Ishikawa J, Fujimura S, Kondo M, Murai-Hatano M, Goto A, Shinano T. 2018. Dynamic changes in the Cs distribution throughout rice plants during the ripening period, and effects of the soil-K level. Plant Soil, 429(1/2): 503-518. |
| [25] | Kabir A H, Begum M C, Haque A, Amin R, Swaraz A M, Haider S A, Paul N K, Hossain M M. 2016. Genetic variation in Fe toxicity tolerance is associated with the regulation of translocation and chelation of iron along with antioxidant defence in shoots of rice. Funct Plant Biol, 43(11): 1070-1081. |
| [26] | Khush G S. 2005. What it will take to feed 5.0 billion rice consumers in 2030. Plant Mol Biol, 59(1): 1-6. |
| [27] | Kubo K, Kobayashi H, Nitta M, Takenaka S, Nasuda S, Fujimura S, Takagi K, Nagata O, Ota T, Shinano T. 2020. Variations in radioactive cesium accumulation in wheat germplasm from fields affected by the 2011 Fukushima nuclear power plant accident. Sci Rep, 10: 3744. |
| [28] | Li Z Y, Ma Z W, van der Kuijp T J, Yuan Z W, Huang L. 2014. A review of soil heavy metal pollution from mines in China: Pollution and health risk assessment. Sci Total Environ, 468-469: 843-853. |
| [29] | Muthayya S, Sugimoto J D, Montgomery S, Maberly G F. 2014. An overview of global rice production, supply, trade, and consumption. Ann N Y Acad Sci, 1324(1): 7-14. |
| [30] | Ozaki T, Ambe S, Abe T, Francis A J. 2005. Competitive inhibition and selectivity enhancement by Ca in the uptake of inorganic elements (Be, Na, Mg, K, Ca, Sc, Mn, Co, Zn, Se, Rb, Sr, Y, Zr, Ce, Pm, Gd, Hf) by carrot (Daucus carota cv. U. S. harumakigosun). Biol Trace Elem Res, 103(1): 69-82. |
| [31] |
Palmgren M G, Clemens S, Williams L E, Krämer U, Borg S, Schjørring J K, Sanders D. 2008. Zinc biofortification of cereals: Problems and solutions. Trends Plant Sci, 13(9): 464-473.
PMID |
| [32] | Pinto E, Ferreira I M P L V O. 2015. Cation transporters/channels in plants: Tools for nutrient biofortification. J Plant Physiol, 179: 64-82. |
| [33] | Qiao K, Wang F H, Liang S, Wang H, Hu Z L, Chai T Y. 2019. New biofortification tool: Wheat TaCNR5 enhances zinc and manganese tolerance and increases zinc and manganese accumulation in rice grains. J Agric Food Chem, 67(35): 9877-9884. |
| [34] | Rai H, Yokoyama S, Satoh-Nagasawa N, Furukawa J, Nomi T, Ito Y, Fujimura S, Takahashi H, Suzuki R, Yousra E, Goto A, Fuji S, Nakamura S I, Shinano T, Nagasawa N, Wabiko H, Hattori H. 2017. Cesium uptake by rice roots largely depends upon a single gene, HAK1, which encodes a potassium transporter. Plant Cell Physiol, 58(9): 1486-1493. |
| [35] | Rennenberg H. 1984. The fate of excess sulfur in higher plants. Annu Rev Plant Physiol, 35(1): 121-153. |
| [36] | Römheld V, Kirkby E A. 2010. Research on potassium in agriculture: Needs and prospects. Plant Soil, 335(1): 155-180. |
| [37] | Salt D E, Baxter I, Lahner B. 2008. Ionomics and the study of the plant ionome. Annu Rev Plant Biol, 59(1): 709-733. |
| [38] |
Schiavon M, Pilon-Smits E A H, Wirtz M, Hell R, Malagoli M. 2008. Interactions between chromium and sulfur metabolism in Brassica juncea. J Environ Qual, 37(4): 1536-1545.
PMID |
| [39] | Singh A, Septiningsih E M, Balyan H S, Singh N K, Rai V. 2017. Genetics, physiological mechanisms and breeding of flood- tolerant rice (Oryza sativa L.). Plant Cell Physiol, 58(2): 185-197. |
| [40] | Spielmann J, Ahmadi H, Scheepers M, Weber M, Nitsche S, Carnol M, Bosman B, Kroymann J, Motte P, Clemens S, Hanikenne M. 2020. The two copies of the zinc and cadmium ZIP6 transporter of Arabidopsis halleri have distinct effects on cadmium tolerance. Plant Cell Environ, 43(9): 2143-2157. |
| [41] | Stein J C, Yu Y, Copetti D, Zwickl D J, Zhang L, Zhang C J, Chougule K, Gao D Y, Iwata A, Goicoechea J L, Wei S, Wang J, Liao Y, Wang M H, Jacquemin J, Becker C, Kudrna D, Zhang J W, Londono C E M, Song X, Lee S, Sanchez P, Zuccolo A, Ammiraju J S S, Talag J, Danowitz A, Rivera L F, Gschwend A R, Noutsos C, Wu C C, Kao S M, Zeng J W, Wei F J, Zhao Q, Feng Q, Baidouri M E, Carpentier M C, Lasserre E, Cooke R, da Rosa Farias D, da Maia L C, dos Santos R S, Nyberg K G, McNally K L, Mauleon R, Alexandrov N, Schmutz J, Flowers D, Fan C Z, Weigel D, Jena K K, Wicker T, Chen M S, Han B, Henry R, Hsing Y I C, Kurata N, de Oliveira A C, Panaud O, Jackson S A, Machado C A, Sanderson M J, Long M Y, Ware D, Wing R A. 2018. Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza. Nat Genet, 50(2): 285-296. |
| [42] | Suriyagoda L D B, Dittert K, Lambers H. 2018. Mechanism of arsenic uptake, translocation and plant resistance to accumulate arsenic in rice grains. Agric Ecosyst Environ, 253: 23-37. |
| [43] | Takahashi R, Ishimaru Y, Shimo H, Ogo Y, Senoura T, Nishizawa N K, Nakanishi H. 2012. The OsHMA2 transporter is involved in root-to-shoot translocation of Zn and Cd in rice. Plant Cell Environ, 35(11): 1948-1957. |
| [44] | Tan Y J, Sun L, Song Q N, Mao D H, Zhou J Q, Jiang Y R, Wang J R, Fan T, Zhu Q H, Huang D Y, Xiao H, Chen C Y. 2020. Genetic architecture of subspecies divergence in trace mineral accumulation and elemental correlations in the rice grain. Theor Appl Genet, 133(2): 529-545. |
| [45] | Tanaka N, Shenton M, Kawahara Y, Kumagai M, Sakai H, Kanamori H, Yonemaru J, Fukuoka S, Sugimoto K, Ishimoto M, Wu J, Ebana K. 2020a. Whole-genome sequencing of the NARO World Rice Core Collection (WRC) as the basis for diversity and association studies. Plant Cell Physiol, 61(5): 922-932. |
| [46] | Tanaka N, Shenton M, Kawahara Y, Kumagai M, Sakai H, Kanamori H, Yonemaru J I, Fukuoka S, Sugimoto K, Ishimoto M, Wu J Z, Ebana K. 2020b. Investigation of the genetic diversity of a core collection of Japanese landraces using whole- genome sequencing. Plant Cell Physiol, 61(12): 2087-2096. |
| [47] | Watanabe T, Kouho R, Katayose T, Kitajima N, Sakamoto N, Yamaguchi N, Shinano T, Yurimoto H, Osaki M. 2014. Arsenic alters uptake and distribution of sulphur in Pteris vittata. Plant Cell Environ, 37(1): 45-53. |
| [48] | Watanabe T, Maejima E, Yoshimura T, Urayama M, Yamauchi A, Owadano M, Okada R, Osaki M, Kanayama Y, Shinano T. 2016. The ionomic study of vegetable crops. PLoS One, 11(8): e0160273. |
| [49] | White P J, Broadley M R. 2009. Biofortification of crops with seven mineral elements often lacking in human diets: Iron, zinc, copper, calcium, magnesium, selenium and iodine. New Phytol, 182: 49-84. |
| [50] | White P J, Brown P H. 2010. Plant nutrition for sustainable development and global health. Ann Bot, 105(7): 1073-1080. |
| [51] | White P J, Broadley M R, Thompson J A, McNicol J W, Crawley M J, Poulton P R, Johnston A E. 2012. Testing the distinctness of shoot ionomes of angiosperm families using the Rothamsted Park Grass Continuous Hay Experiment. New Phytol, 196(1): 101-109. |
| [52] | Wu Q, Zhu X F, Zhao X S, Shen R F. 2020. Potassium affects cadmium resistance in Arabidopsis through facilitating root cell wall Cd retention in a nitric oxide dependent manner. Environ Exp Bot, 178: 104175. |
| [53] | Xu Q, Wang C Q, Li S G, Li B, Li Q Q, Chen G D, Chen W L, Wang F. 2017. Cadmium adsorption, chelation and compartmentalization limit root-to-shoot translocation of cadmium in rice (Oryza sativa L.). Environ Sci Pollut Res, 24(12): 11319-11330. |
| [54] | Yang M, Lu K, Zhao F J, Xie W B, Ramakrishna P, Wang G Y, Du Q Q, Liang L M, Sun C J, Zhao H, Zhang Z Y, Liu Z H, Tian J J, Huang X Y, Wang W S, Dong H X, Hu J T, Ming L C, Xing Y Z, Wang G W, Xiao J H, Salt D E, Lian X M. 2018. Genome-wide association studies reveal the genetic basis of ionomic variation in rice. Plant Cell, 30(11): 2720-2740. |
| [55] | Zhang C M, Zhao W Y, Gao A X, Su T T, Wang Y K, Zhang Y Q, Zhou X B, He X H. 2018. How could agronomic biofortification of rice be an alternative strategy with higher cost-effectiveness for human iron and zinc deficiency in China? Food Nutr Bull, 39(2): 246-259. |
| [56] | Zhang L H, Hu B, Li W, Che R H, Deng K, Li H, Yu F Y, Ling H Q, Li Y J, Chu C C. 2014. OsPT2, a phosphate transporter, is involved in the active uptake of selenite in rice. New Phytol, 201(4): 1183-1191. |
| [57] | Zhou H, Zhu W, Yang W T, Gu J F, Gao Z X, Chen L W, Du W Q, Zhang P, Peng P Q, Liao B H. 2018. Cadmium uptake, accumulation, and remobilization in iron plaque and rice tissues at different growth stages. Ecotoxicol Environ Saf, 152: 91-97. |
| [1] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [2] | Sheikh Faruk Ahmed, Hayat Ullah, May Zun Aung, Rujira Tisarum, Suriyan Cha-Um, Avishek Datta. Iron Toxicity Tolerance of Rice Genotypes in Relation to Growth, Yield and Physiochemical Characters [J]. Rice Science, 2023, 30(4): 321-334. |
| [3] | Guan-fu Fu, Cai-xia Zhang, Yong-jie Yang, Jie Xiong, Xue-qin Yang, Xiu-fu Zhang, Qian-yu Jin, Long-xing Tao. Male Parent Plays More Important Role in Heat Tolerance in Three-Line Hybrid Rice [J]. Rice Science, 2015, 22(3): 116-122. |
| [4] | Challagulla Vineela, Bhattarai Surya, J. Midmore David. In-vitro vs in-vivo Inoculation: Screening for Resistance of Australian Rice Genotypes Against Blast Fungus [J]. Rice Science, 2015, 22(3): 132-137. |
| [5] | M. PUNITHAVALLI, N. M. MUTHUKRISHNAN, M. B. RAJKUMAR. Influence of Rice Genotypes on Folding and Spinning Behaviour of Leaffolder (Cnaphalocrocis medinalis) and Its Interaction with Leaf Damage [J]. RICE SCIENCE, 2013, 20(6): 442-450. |
| [6] | M. PUNITHAVALLI, N. M. MUTHUKRISHNAN, M. BALAJI RAJKUMAR. Defensive Responses of Rice Genotypes for Resistance Against Rice Leaffolder Cnaphalocrocis medinalis [J]. RICE SCIENCE, 2013, 20(5): 363-370. |
| [7] | LIU Li1, WANG Li1, DENG Fei1, HUANG Yun1, LIU Dai-yin2, REN Wan-jun1, YANG Wen-yu1. Response of Osmotic Regulation Substance Content and Protective Enzyme Activities to Shading in Leaves of Different Rice Genotypes [J]. RICE SCIENCE, 2013, 20(4): 276-283. |
| [8] | ZHANG Tao, NI Xian-lin, JIANG Kai-feng, DENG Hua-feng, HE Qing, YANG Qian-hua, YANG Li, WAN Xian-Qi, CAO Ying-jiang, ZHENG Jia-kui, . Relationship Between Heterosis and Parental Genetic Distance Based on Molecular Markers for Functional Genes Related to Yield Traits in Rice [J]. RICE SCIENCE, 2010, 17(4): 288-295 . |
| [9] | HAO Xian-bin, MA Xiu-fang, HU Pei-song, ZHANG Zhong-xu, SUI Guo-min, HUA Ze-tian. Relationship Between Plant Type and Grain Quality of Japonica Hybrid Rice in Northern China [J]. RICE SCIENCE, 2010, 17(1): 43-50 . |
| [10] | CHEN Yu-min, FAN Cheng-ming, YANG Yan, HE Yue-qiu, . Relationship Between Resistance Gene Analogue and Blast Resistance in Rice [J]. RICE SCIENCE, 2009, 16(2): 99-105 . |
| [11] | Keykhosrow KEYMANESH, Mohammad Hassan DARVISHI, Soroush SARDARI. Metabolome Comparison of Transgenic and Non-transgenic Rice by Statistical Analysis of FTIR and NMR Spectra [J]. RICE SCIENCE, 2009, 16(2): 119-123 . |
| [12] | WANG Xiu-hong, SHI Xiang-yuan, WU Xian-jun. Tissue Culture Responses from Different Explants of Rice [J]. RICE SCIENCE, 2005, 12(3): 229-232 . |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||