
Rice Science ›› 2022, Vol. 29 ›› Issue (5): 462-472.DOI: 10.1016/j.rsci.2022.07.006
• Research Paper • Previous Articles Next Articles
Amrit Kumar Nayak3, Anilkumar C1, Sasmita Behera1, Rameswar Prasad Sah1, Gera Roopa Lavanya3, Awadhesh Kumar2, Lambodar Behera1, Muhammed Azharudheen Tp1( )
)
Received:2021-08-26
Accepted:2022-01-26
Online:2022-09-28
Published:2022-07-14
Contact:
Muhammed Azharudheen Tp
Amrit Kumar Nayak, Anilkumar C, Sasmita Behera, Rameswar Prasad Sah, Gera Roopa Lavanya, Awadhesh Kumar, Lambodar Behera, Muhammed Azharudheen Tp. Genetic Dissection of Grain Size Traits Through Genome-Wide Association Study Based on Genic Markers in Rice[J]. Rice Science, 2022, 29(5): 462-472.
Add to citation manager EndNote|Ris|BibTeX
| Trait | Phenotype | Skewness | Kurtosis | Shapiro-Wilks ‘p’ | |||
|---|---|---|---|---|---|---|---|
| Min | Max | Mean ± SE | PV | ||||
| TGW | 10.6 | 31.9 | 21.50 ± 0.04 | 0.18 | 0.07 | 0.41 | 0.197 |
| GL | 5.2 | 10.6 | 8.39 ± 0.14 | 1.82 | -0.09 | -0.51 | 0.754 |
| GW | 1.7 | 3.3 | 2.62 ± 0.04 | 0.14 | -0.22 | -0.74 | 0.094 |
| LWR | 2.01 | 5.59 | 3.31 ± 2.01 | 0.52 | 0.67 | 0.24 | 0.016 |
Table 1. Phenotype variation and distribution pattern of four grain size-related traits.
| Trait | Phenotype | Skewness | Kurtosis | Shapiro-Wilks ‘p’ | |||
|---|---|---|---|---|---|---|---|
| Min | Max | Mean ± SE | PV | ||||
| TGW | 10.6 | 31.9 | 21.50 ± 0.04 | 0.18 | 0.07 | 0.41 | 0.197 |
| GL | 5.2 | 10.6 | 8.39 ± 0.14 | 1.82 | -0.09 | -0.51 | 0.754 |
| GW | 1.7 | 3.3 | 2.62 ± 0.04 | 0.14 | -0.22 | -0.74 | 0.094 |
| LWR | 2.01 | 5.59 | 3.31 ± 2.01 | 0.52 | 0.67 | 0.24 | 0.016 |
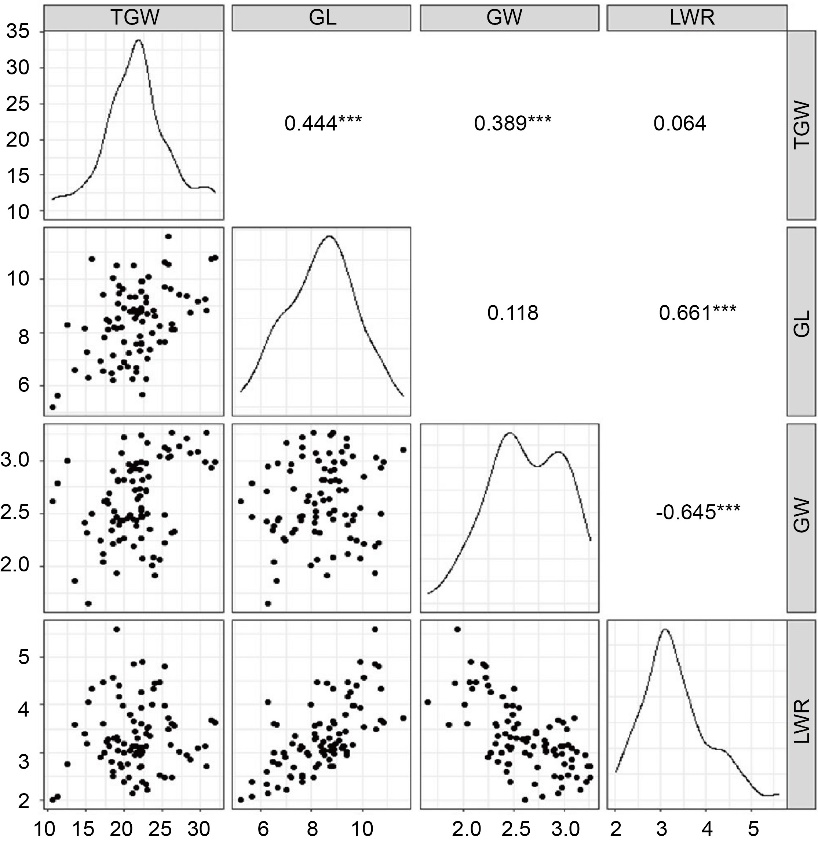
Fig. 2. Correlation coefficients and trend of distribution among grain size characters estimated based on across season best linear unbiased predictor values of phenotypes. TGW, 1000-grain weight; GL, Grain length; GW, Grain width; LWR, Length-width ratio. ***, P < 0.001 by Pearson’s correlation approach.
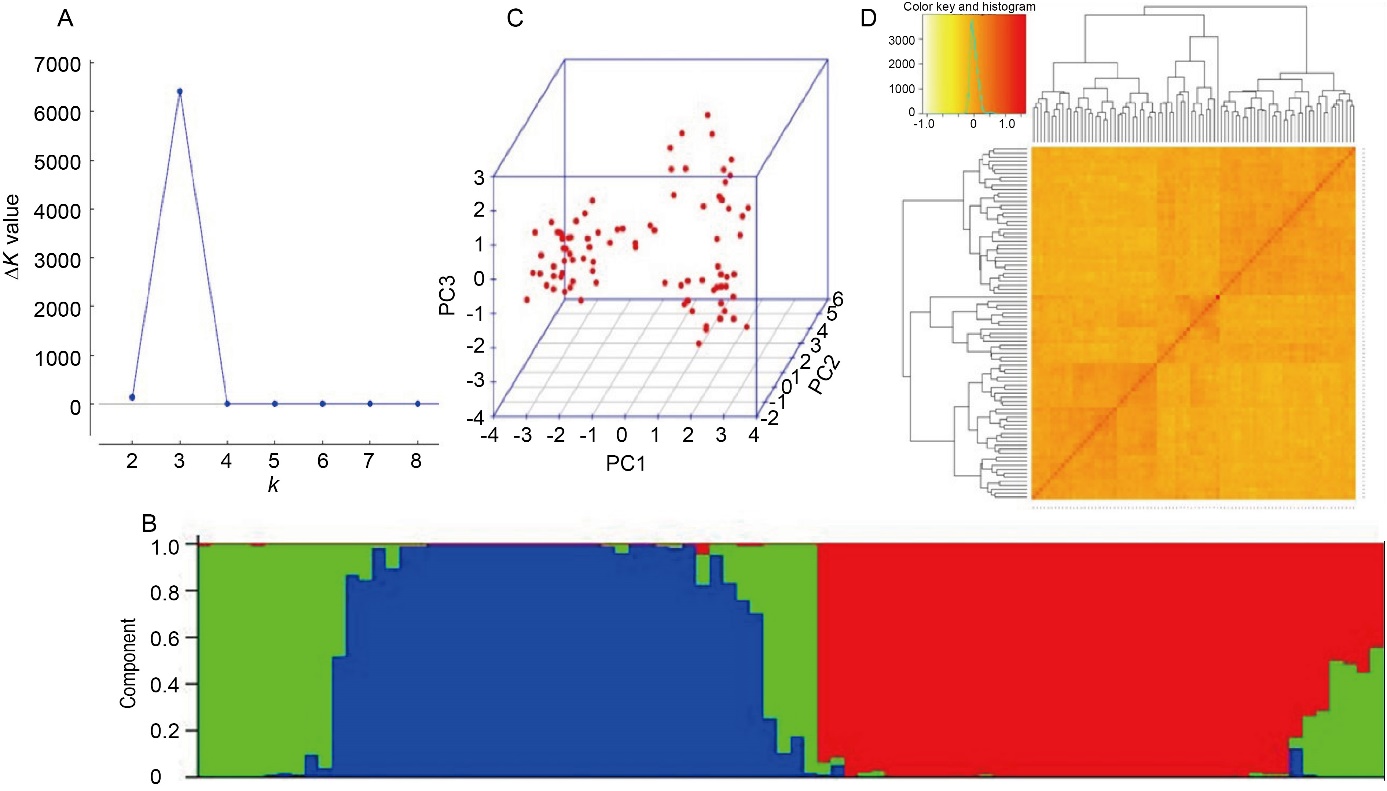
Fig. 3. Population structure analysis. A, Magnitude of ∆K values with k ranging from 2 to 8 (x-axis) in association mapping panel. B, Population structure of association panel based on 142 new candidate gene based SSR markers at K = 3. Different color columns represent different sub-populations. C, 3D representation of principle component (PC) analysis showing three sub-populations. D, Heat map of kinship matrix. The heat map shows the level of relatedness among the population. The darker areas show the level of relatedness between varieties and the dendrogram depicts clustering of sub-populations.
| Trait | Marker | Chr | Position (bp) | P-value | PVE (%) | Known gene |
|---|---|---|---|---|---|---|
| TGW | M69 | 5 | 18 724 905 | 0.01 | 11.01 | OSBC1L4 |
| Sd14 | 6 | 5 315 178 | 0.02 | 10.23 | OsC1 | |
| M55 | 4 | 25 489 003 | 0.02 | 10.00 | SHO1 | |
| Sdi21 | 5 | 1 160 267 | 0.04 | 9.54 | RSR1 | |
| GL | M55 | 4 | 25 489 003 | 0.04 | 6.34 | SHO1 |
| GW | M35 | 8 | 26 439 584 | 0.01 | 13.25 | NPP1 |
| Sdi1 | 1 | 5 236 623 | 0.01 | 13.07 | OsD2 | |
| M99 | 1 | 25 382 698 | 0.02 | 11.00 | Rd | |
| M69 | 5 | 18 724 905 | 0.02 | 10.56 | OSBC1L4 | |
| LWR | Sdi1 | 1 | 5 236 623 | 0.02 | 8.00 | OsD2 |
Table 2. Significant marker-trait associations identified for four grain size-related traits based on mixed line model.
| Trait | Marker | Chr | Position (bp) | P-value | PVE (%) | Known gene |
|---|---|---|---|---|---|---|
| TGW | M69 | 5 | 18 724 905 | 0.01 | 11.01 | OSBC1L4 |
| Sd14 | 6 | 5 315 178 | 0.02 | 10.23 | OsC1 | |
| M55 | 4 | 25 489 003 | 0.02 | 10.00 | SHO1 | |
| Sdi21 | 5 | 1 160 267 | 0.04 | 9.54 | RSR1 | |
| GL | M55 | 4 | 25 489 003 | 0.04 | 6.34 | SHO1 |
| GW | M35 | 8 | 26 439 584 | 0.01 | 13.25 | NPP1 |
| Sdi1 | 1 | 5 236 623 | 0.01 | 13.07 | OsD2 | |
| M99 | 1 | 25 382 698 | 0.02 | 11.00 | Rd | |
| M69 | 5 | 18 724 905 | 0.02 | 10.56 | OSBC1L4 | |
| LWR | Sdi1 | 1 | 5 236 623 | 0.02 | 8.00 | OsD2 |
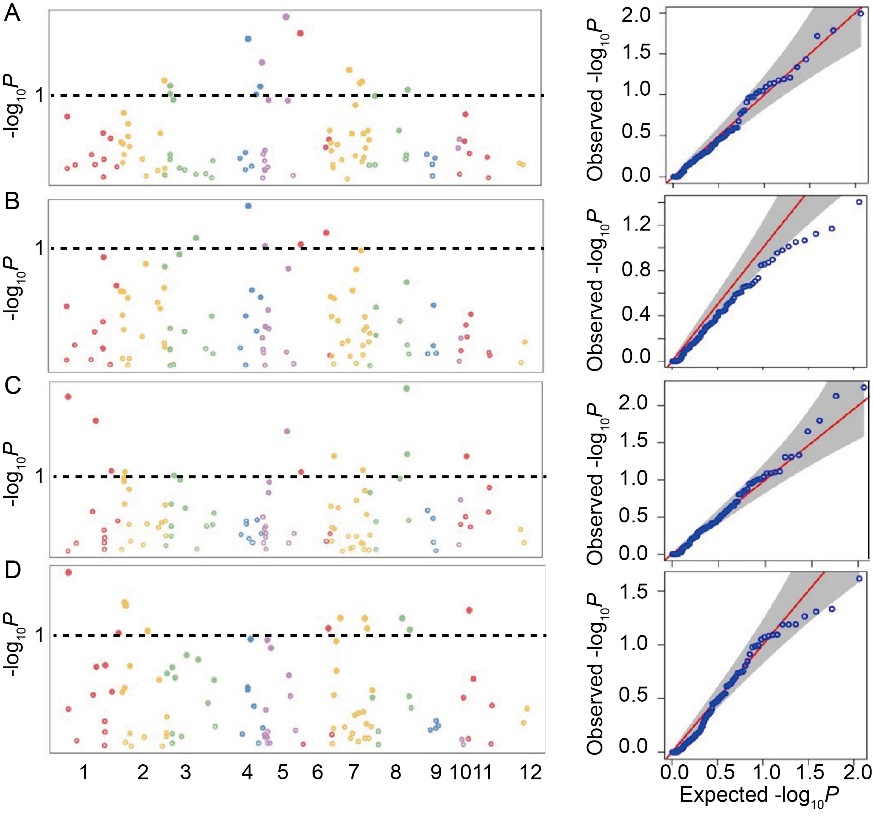
Fig. 4. Manhattan plots and Quantile-quantile plots for markers associated with grain traits across the genome. A, 1000-grain weight; B, Grain length; C, Grain width; D, Length-width ratio. In Manhattan plots, x-axis represents chromosomes and explains chromosome-wise marker distribution, and -log10P values on y-axis indicates significant associations. Quantile-quantile plots show deviation of observed -log10P values and expected -log10P values indicating the significant marker trait associations.
| [1] | Agrama H A, Eizenga G C, Yan W. 2007. Association mapping of yield and its components in rice cultivars. Mol Breed, 19(4): 341-356. |
| [2] |
Alqudah A M, Sallam A, Stephen Baenziger P, Börner A. 2020. GWAS: Fast-forwarding gene identification and characterization in temperate cereals: Lessons from barley: A review. J Adv Res, 22: 119-135.
PMID |
| [3] | Alvarado G, Rodríguez F M, Pacheco A, Burgueño J, Crossa J, Vargas M, Pérez-Rodríguez P, Lopez-Cruz M A. 2020. META-R: A software to analyze data from multi-environment plant breeding trials. Crop J, 8(5): 745-756. |
| [4] | Anandan A, Mahender A, Sah R P, Haque S, Pradhan S K, Roy P S, Singh O N, Ali J. 2021. Genetic diversity and population structure among an assorted group of genotypes pertinent to reproductive stage drought stress in rice (Oryza sativa L.). Acta Sci Agric, 5(3): 77-89. |
| [5] | Atwell S, Huang Y S, Vilhjálmsson B J, Willems G, Horton M, Li Y, Meng D, Platt A, Tarone A M, Hu T T, Jiang R, Muliyati N W, Zhang X, Amer M A, Baxter I, Brachi B, Chory J, Dean C, Debieu M, de Meaux J, Ecker J R, Faure N, Kniskern J M, Jones J D, Michael T, Nemri A, Roux F, Salt D E, Tang C, Todesco M, Traw M B, Weigel D, Marjoram P, Borevitz J O, Bergelson J, Nordborg M. 2010. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature, 465: 627-631. |
| [6] | Azharudheen T P M, Nayak A K, Behera S, Anilkumar C, Marndi B C, Moharana D, Singh L K, Upadhyay S, Sah R P. 2022. Genome-wide association analysis for plant type characters and yield using cgSSR markers in rice (Oryza sativa L.). Euphytica, 218(6): 1-13. |
| [7] | Bai X F, Luo L J, Yan W H, Kovi M R, Zhan W, Xing Y Z. 2010. Genetic dissection of rice grain shape using a recombinant inbred line population derived from two contrasting parents and fine mapping a pleiotropic quantitative trait locus qGL7. BMC Genet, 11: 16. |
| [8] | Chakraborti M, Anilkumar C, Verma R L, Abdul Fiyaz R, Reshmi Raj K R, Patra B C, Balakrishnan D, Sarkar S, Mondal N P, Kar M K, Meher J, Sundaram R M, Rao L S. 2021. Rice breeding in India: Eight decades of journey towards enhancing the genetic gain for yield, nutritional quality, and commodity value. Oryza, 58: 69-88. |
| [9] |
Ching A, Caldwell K S, Jung M, Dolan M, Smith O S, Tingey S, Morgante M, Rafalski A J. 2002. SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet, 3: 19.
PMID |
| [10] | Cho Y G, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch S R, Park W D, Ayres N, Cartinhour S. 2000. Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet, 100(5): 713-722. |
| [11] | Collard B C Y, Vera Cruz C M, McNally K L, Virk P S, MacKill D J. 2008. Rice molecular breeding laboratories in the genomics era: Current status and future considerations. Int J Plant Genomics, 2008: 524847. |
| [12] | Dai X, You C, Chen G, Li X, Zhang Q, Wu C. 2011. OsBC1L4 encodes a COBRA-like protein that affects cellulose synthesis in rice. Plant Mol Biol, 75(4): 333-345. |
| [13] | Duan P G, Xu J S, Zeng D L, Zhang B L, Geng M F, Zhang G Z, Huang K, Huang L J, Xu R, Ge S, Qian Q, Li Y H. 2017. Natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol Plant, 10(5): 685-694. |
| [14] | Earl D A, vonHoldt B M. 2012. STRUCTURE HARVESTER: A website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour, 4(2): 359-361. |
| [15] |
Evanno G, Regnaut S, Goudet J. 2005. Detecting the number of clusters of individuals using the software STRUCTURE: A simulation study. Mol Ecol, 14(8): 2611-2620.
PMID |
| [16] | Evans L T. 1972. Storage Capacity as a Limitation on Grain Yield in Rice Breeding. Manila, the Philippines: International Rice Research Institute. |
| [17] | Fu F F, Xue H W. 2010. Coexpression analysis identifies Rice Starch Regulator1, a rice AP2/EREBP family transcription factor, as a novel rice starch biosynthesis regulator. Plant Physiol, 154(2): 927-938. |
| [18] | Fu F H, Wang F, Huang W J, Peng H P, Wu Y Y, Huang D J. 1994. Genetic analysis on grain characters in hybrid rice. Acta Agron Sin, 20(1): 39-45. |
| [19] |
Furukawa T, Maekawa M, Oki T, Suda I, Iida S, Shimada H, Takamure I, Kadowaki K I. 2007. The Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp. Plant J, 49(1): 91-102.
PMID |
| [20] |
Gao D W, Sun W Q, Wang D W, Dong H L, Zhang R, Yu S B. 2020. A xylan glucuronosyltransferase gene exhibits pleiotropic effects on cellular composition and leaf development in rice. Sci Rep, 10(1): 3726.
PMID |
| [21] | Garris A J, Tai T H, Coburn J, Kresovich S, McCouch S. 2005. Genetic structure and diversity in Oryza sativa L. Genetics, 169(3): 1631-1638. |
| [22] | Gobu R, Shiv A, Anilkumar C, Basavaraj P S, Harish D, Adhikari S, Vinita R, Umesh H, Sujatha M. 2020. Accelerated crop breeding towards development of climate resilient varieties. Climate change and Indian Agriculture: Challenges and Adaptation Strategies. In: Rao C S, Srinivas T, Rao R V S, Rao N S, Vinayagam S S, Krishnan P. Climate Change and Indian Agriculture: Challenges and Adaptation Strategies, ICAR-National Academy of Agricultural Research Management, Hyderabad, Telangana, India: 49-69. |
| [23] |
Guo X, Elston R C. 1999. Linkage information content of polymorphic genetic markers. Hum Hered, 49(2): 112-118.
PMID |
| [24] | Hill Jr R R, Rosenberger J L. 1985. Methods for combining data from gemrplasm evaluation trials 1. Crop Sci, 25(3): 467-470. |
| [25] |
Hu X X, Wang C, Fu Y P, Liu Q, Jiao X Z, Wang K J. 2016. Expanding the range of CRISPR/Cas9 genome editing in rice. Mol Plant, 9(6): 943-945.
PMID |
| [26] | Hu Z J, Lu S J, Wang M J, He H H, Sun L, Wang H R, Liu X H, Jiang L, Sun J L, Xin X Y, Kong W, Chu C C, Xue H W, Yang J S, Luo X J, Liu J X. 2018. A novel QTL qTGW3 encodes the GSK3/SHAGGY-like kinase OsGSK5/OsSK41 that interacts with OsARF4 to negatively regulate grain size and weight in rice. Mol Plant, 11(5): 736-749. |
| [27] |
Huang R Y, Jiang L R, Zheng J S, Wang T S, Wang H C, Huang Y M, Hong Z L. 2013. Genetic bases of rice grain shape: So many genes, so little known. Trends Plant Sci, 18(4): 218-226.
PMID |
| [28] |
Huang X H, Han B. 2014. Natural variations and genome-wide association studies in crop plants. Annu Rev Plant Biol, 65: 531-551.
PMID |
| [29] | Hussain K, Zhang Y X, Anley W, Riaz A, Abbas A, Rani M H, Wang H, Shen X H, Cao L Y, Cheng S H. 2020. Association mapping of quantitative trait loci for grain size in introgression line derived from Oryza rufipogon. Rice Sci, 27(3): 246-254. |
| [30] | Jan R, Khan M A, Asaf S, Lee I J, Kim K M. 2020. Overexpression of OsF3H modulates WBPH stress by alteration of phenylpropanoid pathway at a transcriptomic and metabolomic level in Oryza sativa. Sci Rep, 10(1): 14685. |
| [31] | Kaneko K, Inomata T, Masui T, Koshu T, Umezawa Y, Itoh K, Pozueta-Romero J, Mitsui T. 2014. Nucleotide pyrophosphatase/ phosphodiesterase 1 exerts a negative effect on starch accumulation and growth in rice seedlings under high temperature and CO2 concentration conditions. Plant Cell Physiol, 55(2): 320-332. |
| [32] |
Kang H M, Zaitlen N A, Wade C M, Kirby A, Heckerman D, Daly M J, Eskin E. 2008. Efficient control of population structure in model organism association mapping. Genetics, 178(3): 1709-1723.
PMID |
| [33] | Kassambara A, Mundt F. 2017. Factoextra: Extract and visualize the results of multivariate data analyses. [2021-7-25]. https://mirrors.sjtug.sjtu.edu.cn/cran/web/packages/factoextra/index.html. |
| [34] | Katara J L, Parameswaran C, Devanna B N, Verma R L, Anil Kumar C, Patra B C, Samantaray S. 2021. Genomics assisted breeding: The need and current perspective for rice improvement in India. Oryza, 58: 61-68. |
| [35] |
Korte A, Farlow A. 2013. The advantages and limitations of trait analysis with GWAS: A review. Plant Methods, 9: 29.
PMID |
| [36] |
Lipka A E, Tian F, Wang Q S, Peiffer J, Li M, Bradbury P J, Gore M A, Buckler E S, Zhang Z W. 2012. GAPIT: Genome association and prediction integrated tool. Bioinformatics, 28(18): 2397-2399.
PMID |
| [37] |
Liu K J, Muse S V. 2005. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics, 21(9): 2128-2129.
PMID |
| [38] | Lu H, Redus M A, Coburn J R, Rutger J N, McCouch S R, Tai T H. 2005. Population structure and breeding patterns of 145 US rice cultivars based on SSR marker analysis. Crop Sci, 45(1): 66-76. |
| [39] | Ma X S, Feng F J, Zhang Y, Elesawi I E, Xu K, Li T F, Mei H W, Liu H Y, Gao N N, Chen C L, Luo L J, Yu S W. 2019. A novel rice grain size gene OsSNB was identified by genome-wide association study in natural population. PLoS Genet, 15(5): e1008191. |
| [40] | Mather D E, Hyes P M, Chalmers K J, Eglinton J, Matus I, Richardson K, Von Zitzewitz J, Marquez-Cedillo L, Hearnden P, Pal N. 2004. Use of SSR marker data to study linkage disequilibrium and population structure in Hordeum vulgare: Prospects for association mapping in barley. In: Jaroslav S, Jarmila J. 9th International Barley Genetics Symposium. Brno, Czech Republic: International barley genetics symposium: 302-307. |
| [41] | Meng L J, Zhao X Q, Ponce K, Ye G Y, Leung H. 2016. QTL mapping for agronomic traits using multi-parent advanced generation inter-cross (MAGIC) populations derived from diverse elite indica rice lines. Field Crops Res, 189: 19-42. |
| [42] | Mohanty S. 2013. Trends in global rice consumption. Rice Today, 12: 44-45. |
| [43] | Molla K A, Debnath A B, Ganie S A, Mondal T K. 2015. Identification and analysis of novel salt responsive candidate gene based SSRs (cgSSRs) from rice (Oryza sativa L.). BMC Plant Biol, 15: 122. |
| [44] | Molla K A, Azharudheen T P M, Ray S, Sarkar S, Swain A, Chakraborti M, Vijayan J, Singh O N, Baig M J, Mukherjee A K. 2019. Novel biotic stress responsive candidate gene based SSR (cgSSR) markers from rice. Euphytica, 215(2): 17. |
| [45] | Nanjo Y, Oka H, Ikarashi N, Kaneko K, Kitajima A, Mitsui T, Muñoz F J, Rodríguez-López M, Baroja-Fernández E, Pozueta- Romero J. 2006. Rice plastidial N-glycosylated nucleotide pyrophosphatase/phosphodiesterase is transported from the ER- Golgi to the chloroplast through the secretory pathway. Plant Cell, 18(10): 2582-2592. |
| [46] |
Norton G J, Travis A J, Douglas A, Fairley S, Alves E D, Ruang- Areerate P, Naredo M, Elizabeth B, McNally K L, Hossain M, Islam M. 2018. Genome wide association mapping of grain and straw biomass traits in the rice Bengal and Assam Aus panel (BAAP) grown under alternate wetting and drying and permanently flooded irrigation. Front Plant Sci, 9: 1223.
PMID |
| [47] | Pahlich E, Gerlitz C. 1980. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemistry, 19: 11-13. |
| [48] | Patra B C, Anilkumar C, Chakraborti M. 2020. Rice breeding in India: A journey from phenotype based pure-line selection to genomics assisted breeding. Agric Res J, 57(6): 816-825. |
| [49] | Piepho H P, Möhring J, Melchinger A E, Büchse A. 2008. BLUP for phenotypic selection in plant breeding and variety testing. Euphytica, 161: 209-228. |
| [50] | Ponce K, Zhang Y, Guo L B, Leng Y J, Ye G Y. 2020. Genome- wide association study of grain size traits in indica rice multiparent advanced generation intercross (MAGIC) population. Front Plant Sci, 11: 395. |
| [51] |
Pritchard J K, Stephens M, Donnelly P. 2000. Inference of population structure using multilocus genotype data. Genetics, 155(2): 945-959.
PMID |
| [52] | Qiu X J, Pang Y L, Yuan Z H, Xing D Y, Xu J L, Dingkuhn M, Li Z K, Ye G Y. 2015. Genome-wide association study of grain appearance and milling quality in a worldwide collection of indica rice germplasm. PLoS One, 10(12): e0145577. |
| [53] | R Core Team. 2021. R, A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. [2021-11-25]. |
| [54] |
Rafalski J A. 2010. Association genetics in crop improvement. Curr Opin Plant Biol, 13(2): 174-180.
PMID |
| [55] | Rahman S N, Islam M S, Alam M S, Nasiruddin K M. 2007. Genetic polymorphism in rice (Oryza sativa L.) through RAPD analysis. Indian J Biotechnol, 6: 224-229. |
| [56] | Raju B R, Mohankumar M V, Sumanth K K, Rajanna M P, Udayakumar M, Prasad T G, Sheshshayee M S. 2016. Discovery of QTLs for water mining and water use efficiency traits in rice under water-limited condition through association mapping. Mol Breeding, 36(3): 35. |
| [57] | Sahu R K, Patnaik S S C, Sah R P. 2020. Quality seed production in rice. Odisha, India: ICAR-National Rice Research Institute: 58. |
| [58] | Sakamoto T, Ohnishi T, Fujioka S, Watanabe B, Mizutani M. 2012. Rice CYP90D2 and CYP90D3 catalyze C-23 hydroxylation of brassinosteroids in vitro. Plant Physiol Biochem, 58: 220-226. |
| [59] | Sanghamitra P, Sah R P, Bagchi T B, Sharma S G, Kumar A, Munda S, Sahu R K. 2018. Evaluation of variability and environmental stability of grain quality and agronomic parameters of pigmented rice (O. sativa L.). J Food Sci Technol, 55(3): 879-890. |
| [60] | Seo H, Kim S H, Lee B D, Lim J H, Lee S J, An G, Paek N C. 2020. The rice basic helix-loop-helix 79 (OsbHLH079) determines leaf angle and grain shape. Int J Mol Sci, 21(6): 2090. |
| [61] | Shapiro S S, Wilk M B. 1965. An analysis of variance test for normality (complete samples). Biometrika, 52: 591-611. |
| [62] | Song X W, Li P C, Zhai J X, Zhou M, Ma L J, Liu B, Jeong D H, Nakano M, Cao S Y, Liu C Y, Chu C C, Wang X J, Green P J, Meyers B C, Cao X F. 2012. Roles of DCL4 and DCL3b in rice phased small RNA biogenesis. Plant J, 69(3): 462-474. |
| [63] | Tan Y F, Xing Y Z, Li J X, Yu S B, Xu C G, Zhang Q F. 2000. Genetic bases of appearance quality of rice grains in Shanyou 63, an elite rice hybrid. Theor Appl Genet, 101: 823-829. |
| [64] |
Upadhyaya G, Das A, Ray S. 2021. A rice R2R3-MYB (OsC1) transcriptional regulator improves oxidative stress tolerance by modulating anthocyanin biosynthesis. Physiol Plant, 173(4): 2334-2349.
PMID |
| [65] |
VanRaden P M. 2008. Efficient methods to compute genomic predictions. J Dairy Sci, 91(11): 4414-4423.
PMID |
| [66] |
Varshney R K, Graner A, Sorrells M E. 2005. Genic microsatellite markers in plants: Features and applications. Trends Biotechnol, 23(1): 48-55.
PMID |
| [67] |
Vieira M L C, Santini L, Diniz A L, de Freitas Munhoz C. 2016. Microsatellite markers: What they mean and why they are so useful. Genet Mol Biol, 39(3): 312-328.
PMID |
| [68] | Wang C H, Yang Y L, Yuan X P, Xu Q, Feng Y, Yu H Y, Wang Y P, Wei X H. 2014. Genome-wide association study of blast resistance in indica rice. BMC Plant Biol, 14: 311. |
| [69] | Wang Y H, Zheng Y M, Cai Q H, Liao C J, Mao X H, Xie H G, Zhu Y S, Lian L, Luo X, Xie H A, Zhang J F. 2016. Population structure and association analysis of yield and grain quality traits in hybrid rice primal parental lines. Euphytica, 212(2): 261-273. |
| [70] | Wei T, Simko V. 2021. R package ‘corrplot’: Visualization of a Correlation Matrix. Version 0.88. https://github.com/taiyun/corrplot. |
| [71] |
Wu J H, Feng F J, Lian X M, Teng X Y, Wei H B, Yu H H, Xie W B, Yan M, Fan P Q, Li Y, Ma X S, Liu H Y, Yu S B, Wang G W, Zhou F S, Luo L J, Mei H W. 2015. Genome-wide association study (GWAS) of mesocotyl elongation based on re-sequencing approach in rice. BMC Plant Biol, 15: 218.
PMID |
| [72] |
Xing Y Z, Zhang Q F. 2010. Genetic and molecular bases of rice yield. Annu Rev Plant Biol, 61: 421-442.
PMID |
| [73] | Xu J L, Xue Q Z, Luo L J, Li Z K. 2002. Genetic dissection of grain weight and its related traits in rice (Oryza sativa L.). Chin J Rice Sci, 16: 6-10. (in Chinese with English abstract) |
| [74] |
Yu J M, Pressoir G, Briggs W H, Vroh Bi I, Yamasaki M, Doebley J F, McMullen M D, Gaut B S, Nielsen D M, Holland J B, Kresovich S, Buckler E S. 2006. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet, 38(2): 203-208.
PMID |
| [75] |
Yu J P, Xiong H Y, Zhu X Y, Zhang H L, Li H H, Miao J L, Wang W S, Tang Z S, Zhang Z Y, Yao G X, Zhang Q, Pan Y H, Wang X, Rashid M A R, Li J J, Gao Y M, Li Z K, Yang W C, Fu X D, Li Z C. 2017. OsLG3 contributing to rice grain length and yield was mined by Ho-LAMap. BMC Biol, 15(1): 28.
PMID |
| [76] | Zhang D L, Zhang H L, Qi Y W, Wang M X, Sun J L, Ding L, Li Z C. 2013. Genetic structure and eco-geographical differentiation of cultivated Hsien rice (Oryza sativa L. subsp. indica) in China revealed by microsatellites. Chin Sci Bull, 58(3): 344-352. |
| [77] | Zhang P, Liu X D, Tong H H, Lu Y G, Li J Q. 2014. Association mapping for important agronomic traits in core collection of rice (Oryza sativa L.) with SSR markers. PLoS One, 9(10): e111508. |
| [78] | Zhao D S, Li Q F, Zhang C Q, Zhang C, Yang Q Q, Pan L X, Ren X Y, Lu J, Gu M H, Liu Q Q. 2018. GS9 acts as a transcriptional activator to regulate rice grain shape and appearance quality. Nat Commun, 9(1): 1240. |
| [79] | Zhou Q Y, An H, Zhang Y, Shen F C. 2000. Study on heredity of morphological characters of rice grain. J Southwest Agric Univ, 22(2): 102-104. |
| [1] | Rakotoson Tatiana, Dusserre Julie, Letourmy Philippe, Frouin Julien, Ramonta Ratsimiala Isabelle, Victorine Rakotoarisoa Noronirina, Cao Tuong-Vi, Vom Brocke Kirsten, Ramanantsoanirina Alain, Ahmadi Nourollah, Raboin Louis-Marie. Genome-Wide Association Study of Nitrogen Use Efficiency and Agronomic Traits in Upland Rice [J]. Rice Science, 2021, 28(4): 379-390. |
| [2] | Yang Lv, Yueying Wang, Jahan Noushin, Haitao Hu, Ping Chen, Lianguang Shang, Haiyan Lin, Guojun Dong, Jiang Hu, Zhenyu Gao, Qian Qian, Yu Zhang, Longbiao Guo. Genome-Wide Association Analysis and Allelic Mining of Grain Shape-Related Traits in Rice [J]. Rice Science, 2019, 26(6): 384-392. |
| [3] | Zongxiang Chen, Zhiming Feng, Houxiang Kang, Jianhua Zhao, Tianxiao Chen, Qianqian Li, Hongbing Gong, Yafang Zhang, Xijun Chen, Xuebiao Pan, Wende Liu, Guoliang Wang, Shimin Zuo. Identification of New Resistance Loci Against Sheath Blight Disease in Rice Through Genome-Wide Association Study [J]. Rice Science, 2019, 26(1): 21-31. |
| [4] | Lee Jae-Sung, Wissuwa Matthias, B. Zamora Oscar, M. Ismail Abdelbagi. Novel Sources of aus Rice for Zinc Deficiency Tolerance Identified Through Association Analysis Using High-Density SNP Array [J]. Rice Science, 2018, 25(5): 293-296. |
| [5] | Ya-fang Zhang, Yu-yin Ma, Zong-xiang Chen, Jie Zou, Tian-xiao Chen, Qian-qian Li, Xue-biao Pan, Shi-min Zuo. Genome-Wide Association Studies Reveal New Genetic Targets for Five Panicle Traits of International Rice Varieties [J]. Rice Science, 2015, 22(5): 217-226. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||