
Rice Science ›› 2017, Vol. 24 ›› Issue (2): 109-118.DOI: 10.1016/j.rsci.2016.06.009
• Orginal Article • Previous Articles Next Articles
Yan He, Yong-feng Shi, Xiao-bo Zhang, Hui-mei Wang, Xia Xu, Jian-li Wu( )
)
Received:2016-04-10
Accepted:2016-06-06
Online:2017-02-10
Published:2016-12-28
Yan He, Yong-feng Shi, Xiao-bo Zhang, Hui-mei Wang, Xia Xu, Jian-li Wu. Identification of a Gravitropism-Deficient Mutant in Rice[J]. Rice Science, 2017, 24(2): 109-118.
Add to citation manager EndNote|Ris|BibTeX
| Hormone type | Treatment | |||||
| 1 | 2 | 3 | 4 | 5 | 6 | |
| 2,4-dichlorophenoxyacetic acid | 0.1 | 0.1 | 0.5 | 1.0 | 2.0 | 4.0 |
| Gibberellic acid 3 | 0.5 | 1.0 | 2.0 | 4.0 | 10.0 | 20.0 |
Table 1 Concentrations of exogenous hormones. mg/L
| Hormone type | Treatment | |||||
| 1 | 2 | 3 | 4 | 5 | 6 | |
| 2,4-dichlorophenoxyacetic acid | 0.1 | 0.1 | 0.5 | 1.0 | 2.0 | 4.0 |
| Gibberellic acid 3 | 0.5 | 1.0 | 2.0 | 4.0 | 10.0 | 20.0 |

Fig. 1. Phenotypes of wild type (WT) and M96 mutant.A, Sprout of WT (left) and M96 (right); B, WT (left) and M96 (right) at the early tillering stage; C, M96 at the heading stage; D, WT at the heading stage.
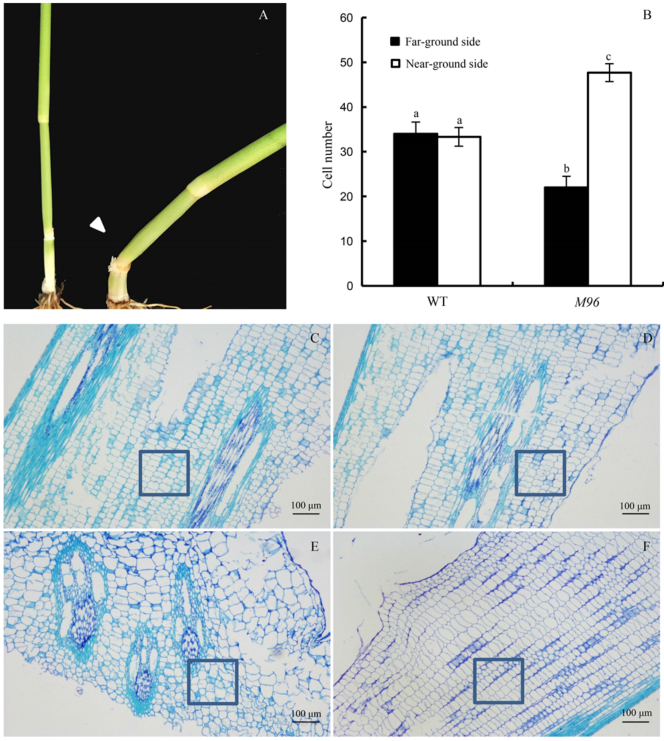
Fig. 3. Paraffin section observation of the second node longitudinal sections.A, Stem of wild type (WT, left) and M96 mutant (right). The arrow indicates the curved node of M96 mutant; B, Longitudinal cell numbers of the sides of the second node in WT and M96. Different lowercase letters indicate significant differences at the 0.01 level by the Duncan’s test. Error bars represent the standard deviation (n = 5); C, Longitudinal section on the far-ground side of the second node in WT; D, Longitudinal section on the near-ground side of the second node in WT; E, Longitudinal section on the far-ground side of the second node in M96; F, Longitudinal section on the near-ground side of the second node in M96.

Fig. 4. Gravity response of wild type (WT) and M96 under light and dark conditions.A, 5-day-old WT (top) and M96 (bottom) seedlings grown under light; B, 5-day-old WT (top) and M96 (bottom) seedlings grown under dark. Arrows indicate gravity direction.
| Marker | Forward primer (5′-3′) | Reverse primer (5′-3′) |
|---|---|---|
| RM536 | TCTCTCCTCTTGTTTGGCTC | ACACACCAACACGACCACAC |
| RM6091 | GCGGACACACCAGAGAATAAGC | GTGCTGTCCTGTCCTTGAATCC |
| RM229 | CACTCACACGAACGACTGAC | CGCAGGTTCTTGTGAAATGT |
| RM27051 | ACCTGGCTACCATCCAAACACG | GCTTTAGGGAGTTCCTGATGTGC |
| RM287 | GGCTACACCTACACGCGAGAACC | AGATGCATGGAATGCCTGTTTGG |
| RM5960 | CGAGCAGCACTGGAGAACACC | CTCCTAGGTGCAGCGGACTACC |
| InDel8 | AACACCACCCGATTCCCT | CAGATTGGATGAGCAGCAAC |
| M96 | ATCATTGCCGTTGTCATCATCT | CAGCACATTCAAGCCCTTCTAT |
| M96RT-2 | AAAGTCTACCCCGAGAACAC | CTCTTGTTGCCGTTCATCTC |
| Ubiquitin | CCCTCCACCTCGTCCTCAG | AGATAACAACGGAAGCATAAAAGTC |
Table 2 Part of primers used in this study.
| Marker | Forward primer (5′-3′) | Reverse primer (5′-3′) |
|---|---|---|
| RM536 | TCTCTCCTCTTGTTTGGCTC | ACACACCAACACGACCACAC |
| RM6091 | GCGGACACACCAGAGAATAAGC | GTGCTGTCCTGTCCTTGAATCC |
| RM229 | CACTCACACGAACGACTGAC | CGCAGGTTCTTGTGAAATGT |
| RM27051 | ACCTGGCTACCATCCAAACACG | GCTTTAGGGAGTTCCTGATGTGC |
| RM287 | GGCTACACCTACACGCGAGAACC | AGATGCATGGAATGCCTGTTTGG |
| RM5960 | CGAGCAGCACTGGAGAACACC | CTCCTAGGTGCAGCGGACTACC |
| InDel8 | AACACCACCCGATTCCCT | CAGATTGGATGAGCAGCAAC |
| M96 | ATCATTGCCGTTGTCATCATCT | CAGCACATTCAAGCCCTTCTAT |
| M96RT-2 | AAAGTCTACCCCGAGAACAC | CTCTTGTTGCCGTTCATCTC |
| Ubiquitin | CCCTCCACCTCGTCCTCAG | AGATAACAACGGAAGCATAAAAGTC |
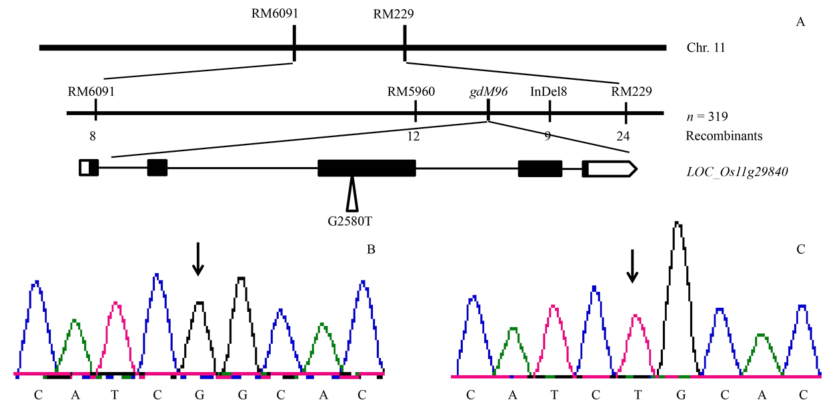
Fig. 5. Gene mapping and candidate gene analysis.A, Gene mapping of M96 mutant; B, Sequence of the mutational region in LOC_Os11g29840 from the wild type; C, Sequence of the mutational region in LOC_Os11g29840 from M96. Arrows indicate the position at 2 580 bp.
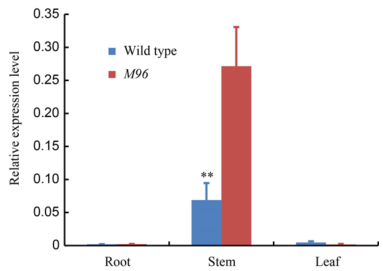
Fig. 6. Expression analysis of target gene in root, stem and leaf of the wild type and M96 by real-time PCR.Error bars represent the standard deviation (n = 5). ** represents the significant difference at the 0.01 level by the Duncan’s test.
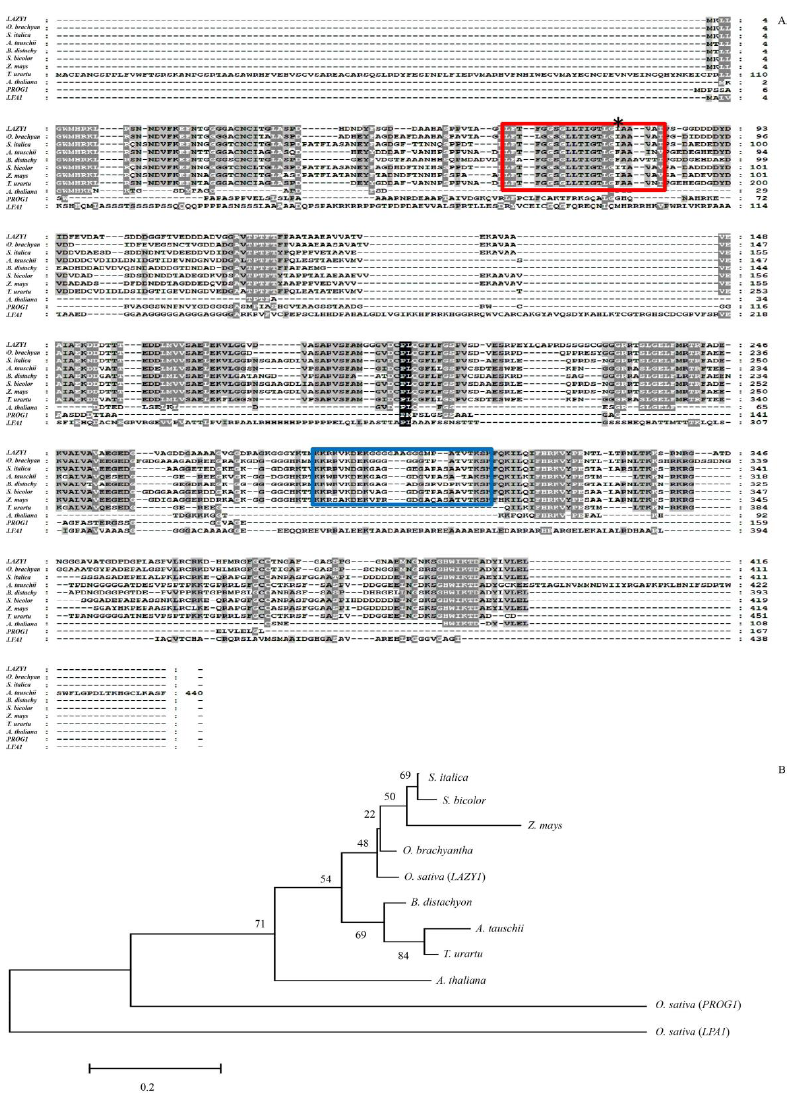
Fig. 7. Structural and phylogenetic analysis of gdM96 homologs.A, Comparison of amino acid sequences of gdM96 homologs. Accession numbers for the respective protein sequences are as follows: O. sativa (LAZY1) (LOC_Os11g29840), O. sativa (PROG1) (LOC_Os07g05900), O. sativa (LPA1) (LOC_Os03g13400), A. thaliana (NP_196913.2); O. brachyantha (XP_015697926), S. italic (XP_004979290), A. tauschii (EMT22503), B. distachyon (XP_010237715), S. bicolor (XP_002449512), Z. mays (AEM59513) and T. urartu (EMS62694). The asterisk indicates the mutant site in M96, the red squared box indicates the predicted transmembrane domain and the blue squared box indicates the putative nuclear localization sequence domain. B, Dendrogram of gdM96 homologs.
| 1 | Azuma T, Hatanaka T, Uchida N, Yasuda T.2003. Interactions between abscisic acid, ethylene and gibberellin in internodal elongation in floating rice: The promotive effect of abscisic acid at low humidity.Plant Growth Regul, 41(2): 105-109. |
| 2 | Buer C S, Wasteneys G O, Masle J.2003. Ethylene modulates root-wave responses in Arabidopsis.Plant Physiol, 132(2): 1085-1096. |
| 3 | Digby J, Firn R D.2002. Light modulation of the gravitropic set-point angle (GSA).J Exp Bot, 53: 377-381. |
| 4 | Dong Z B.2014. Map-based cloning and functional analysis on the maize LAZY1 gene. Beijing: China Agricultural University. (in Chinese with English abstract) |
| 5 | Fukaki H, Fujisawa H, Tasaka M.1996. How do plant shoots bend up? The initial step to elucidate the molecular mechanisms of shoot gravitropism using Arabidopsis thaliana.J Plant Res, 109(2): 129-137. |
| 6 | Gaiser J C, Lomax T L.1993. The altered gravitropic response of the lazy-2 mutant of tomato is phytochrome regulated.Plant Physiol, 102(2): 339-344. |
| 7 | Hangarter R P.1997. Gravity, light and plant form.Plant Cell Environ, 20(6): 796-800. |
| 8 | Hasenstein K H, Kuznetsov O A.1999. The response of lazy-2 tomato seedlings to curvature-inducing magnetic gradients is modulated by light.Planta, 208(1): 59-65. |
| 9 | Jin J, Huang W, Gao J P, Yang J, Shi M, Zhu M Z, Luo D, Lin H X.2008. Genetic control of rice plant architecture under domestication.Nat Genet, 40(11): 1365-1369. |
| 10 | Knauer T, Dümmer M, Landgraf F, Forreiter C.2011. A negative effector of blue light-induced and gravitropic bending in Arabidopsis.Plant Physiol, 156(1): 439-447. |
| 11 | Kovach M J, Sweeney M T, McCouch S R.2007. New insights into the history of rice domestication.Trends Genet, 23(11): 578-587. |
| 12 | Li P J, Wang Y H, Qian Q, Fu Z M, Wang M, Zeng D L, Li B H, Wang X J, Li J Y.2007. LAZY1 controls rice shoot gravitropism through regulating polar auxin transport.Cell Res, 17(5): 402-410. |
| 13 | Liu J M, Park S J, Huang J, Lee E J, Xuan Y H, Je B I, Kumar V, Priatama R A, Raj K V, Kim S H, Min M K, Cho J H, Kim T H, Chandran A K N, Jung K H, Takatsuto S, Fujioka S, Han C D.2016. Loose Plant Architecture1 (LPA1) determines lamina joint bending by suppressing auxin signaling that interacts with C-22-hydroxylated and 6-deoxo brassinosteroids in rice.J Exp Bot, 67(3): 1883-1895. |
| 14 | Lu Y J, Zheng K L.1992. A simple method for isolation of rice DNA.Chin J Rice Sci, 6(1): 47-48. (in Chinese with English abstract) |
| 15 | Miyata M, Komori T, Yamamoto T, Ueda T, Yano M, Nitta N.2005. Fine scale and physical mapping of Spk(t) controlling spreading stub in rice.Breeding Sci, 55(2): 237-239. |
| 16 | Pennazio S.2002. The discovery of the chemical nature of the plant hormone auxin.Rivista Di Biol, 95(2): 289-308. |
| 17 | Saitou N, Nei M.1987. The neighbor-joining method: A new method for reconstructing phylogenetic trees.Mol Biol Evol, 4(4): 406-425. |
| 18 | Sang D J, Chen D Q, Liu G F, Liang Y, Huang L Z, Meng X B, Chu J F, Sun X H, Dong G J, Yuan Y D, Qian Q, Li J Y, Wang Y H.2014. Strigolactones regulate rice tiller angle by attenuating shoot gravitropism through inhibiting auxin biosynthesis.Proc Natl Acad Sci USA, 111: 11199-11204. |
| 19 | Shi Y F, Chen J, Liu W Q, Huang Q N, Shen B, Leung H, Wu J L.2009. Genetic analysis and gene mapping of a new rolled-leaf mutant in rice (Oryza sativa L.).Sci China C: Life Sci, 52(9): 885-890. |
| 20 | Song C B, Yang W T, Du D L, Li D.2006. Progress on plant gravitropism mechanism and its related functional genes. In: Proceedings of the Workshop on Research & Development of Biotechnology in Hainan. |
| 21 | Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S.2011. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods.Mol Biol Evol, 28(10): 2731-2739. |
| 22 | Tan L B, Li X R, Liu F X, Sun X Y, Li C G, Zhu Z F, Fu Y C, Cai H W, Wang X K, Xie D X, Sun C Q.2008. Control of a key transition from prostrate to erect growth in rice domestication.Nat Genet, 40: 1360-1364. |
| 23 | Tasaka M, Kato T, Fukaki H.1999. The endodermis and shoot gravitropism. Trends Plant Sci, 4(3): 103-107. |
| 24 | Vandenbussche F, Smalle J, Le J, Saibo N J M, de Paepe A, Chaerle L, Tietz O, Smets R, Laarhoven L J J, Harren F J M, van Onckelen H, Palme K, Verbelen J P, van Der Straeten D.2003. The Arabidopsis mutant alh1 illustrates a cross talk between ethylene and auxin.Plant Physiol, 131(3): 1228-1238. |
| 25 | Wu X R, Tang D, Li M, Wang K J, Cheng Z K.2013. Loose Plant Architecture 1, an INDETERMINATE DOMAIN protein involved in shoot gravitropism, regulates plant architecture in rice.Plant Physiol, 161(1): 317-329. |
| 26 | Yoshihara T, Iino M.2007. Identification of the gravitropism- related rice gene LAZY1 and elucidation of LAZY1-dependent and -independent gravity signaling pathways.Plant Cell Physiol, 48(5): 678-688. |
| 27 | Zhang J, Ku L X, Han Z P, Guo S L, Liu H J, Zhang Z Z, Cao L R, Cui X J, Chen Y H.2014. The ZmCLA4 gene in the qLA4-1 QTL controls leaf angle in maize (Zea mays L.).J Exp Bot, 65: 5063-5076. |
| 28 | Zhang Q F, Xu J D, Li Y, Xu P Z, Zhang H Y, Wu X J.2007. Morphological, anatomical and genetic analysis for a rice mutant with abnormal hull.J Genet Genomics, 34(6): 519-526. |
| [1] | XIA Xiaodong, ZHANG Xiaobo, WANG Zhonghao, CHENG Benyi, Sun Huifeng, XU Xia, GONG Junyi, YANG Shihua, WU Jianli, SHI Yongfeng, XU Rugen. Mapping and Functional Analysis of LE Gene in a Lethal Etiolated Rice Mutant at Seedling Stage [J]. Rice Science, 2023, 30(6): 13-. |
| [2] | Yong Yang, Qiujun Lin, Xinyu Chen, Weifang Liang, Yuwen Fu, Zhengjin Xu, Yuanhua Wu, Xuming Wang, Jie Zhou, Chulang Yu, Chengqi Yan, Qiong Mei, Jianping Chen. Characterization and Proteomic Analysis of Novel Rice Lesion Mimic Mutant with Enhanced Disease Resistance [J]. Rice Science, 2021, 28(5): 466-478. |
| [3] | Xiaobo Zhu, Mu Ze, Mawsheng Chern, Xuewei Chen, Jing Wang. Deciphering Rice Lesion Mimic Mutants to Understand Molecular Network Governing Plant Immunity and Growth [J]. Rice Science, 2020, 27(4): 278-288. |
| [4] | Yuexiong Zhang, Gang Qin, Qianqian Ma, Minyi Wei, Xinghai Yang, Zengfeng Ma, Haifu Liang, Chi Liu, Zhenjing Li, Fang Liu, Dahui Huang, Rongbai Li. Identification of Major Locus Bph35 Resistance to Brown Planthopper in Rice [J]. Rice Science, 2020, 27(3): 237-245. |
| [5] | Yaling Chen, Yuehan Pang, Jinsong Bao. Expression Profiles and Protein Complexes of Starch Biosynthetic Enzymes from White-Core and Waxy Mutants Induced from High Amylose Indica Rice [J]. Rice Science, 2020, 27(2): 152-161. |
| [6] | Cheabu Sulaiman, Panichawong Nat, Rattanametta Prisana, Wasuri Boonthong, Kasemsap Poonpipope, Arikit Siwaret, Vanavichit Apichart, Malumpong Chanate. Screening for Spikelet Fertility and Validation of Heat Tolerance in a Large Rice Mutant Population [J]. Rice Science, 2019, 26(4): 229-238. |
| [7] | Ran Qin, Akhter Delara, Chengcong Yang, Kumar Nath Ujjal, Eshag Jamal, Xiaoli Jin, Chunhai Shi. SRG1, Encoding a Kinesin-4 Protein, Is an Important Factor for Determining Grain Shape in Rice [J]. Rice Science, 2018, 25(6): 297-307. |
| [8] | Hua Zhang, Xu Liu, Yongyi Yang, Ning Xuan, Fangyin Yao. Mapping of Hd-6-2 for Heading Date Using Two Secondary Segregation Populations in Rice [J]. Rice Science, 2018, 25(3): 161-168. |
| [9] | Zhongkang Wang, Dongdong Zeng, Ran Qin, Jialin Liu, Chunhai Shi, Xiaoli Jin. A Novel and Pleiotropic Factor SLENDER GRAIN3 Is Involved in Regulating Grain Size in Rice [J]. Rice Science, 2018, 25(3): 132-141. |
| [10] | Shu-hua Liu, Jian Tang, Ju Luo, Bao-jun Yang, Ai-ying Wang, Jin-cai Wu. Cloning and Characterization of karmoisin Homologue Gene (Nlka) in Two Brown Planthopper Strains with Different Eye Colors [J]. Rice Science, 2016, 23(2): 104-110. |
| [11] | Daniel Abacar Jose, Zhao-miao Lin, Xin-cheng Zhang, Cheng-qiang Ding, She Tang, Zheng-hui Liu, Shao-hua Wang, Yan-feng Ding. Variation in Yield and Physicochemical Quality Traits among Mutants of Japonica Rice Cultivar Wuyujing 3 [J]. Rice Science, 2016, 23(1): 33-41. |
| [12] | Xia Xu, Xiao-bo Zhang, Yong-feng Shi, Hui-mei Wang, Bao-hua Feng, Xiao-hong Li, Qi-na Huang, Li-xin Song, Dan Guo, Yan He, Jian-li Wu. A Point Mutation in an F-Box Domain-Containing Protein Is Responsible for Brown Hull Phenotype in Rice [J]. Rice Science, 2016, 23(1): 1-8. |
| [13] | Lian-ping Sun, Ying-xin Zhang, Pei-pei Zhang, Zheng-fu Yang, Xiao-deng Zhan, Xi-hong Shen, Zhen-hua Zhang, Xia Hu, Dan-dan Xuan, Wei-xun Wu, Zi-he Li, Li-yong Cao, Shi-hua Cheng. K-Domain Splicing Factor OsMADS1 Regulates Open Hull Male Sterility in Rice [J]. Rice Science, 2015, 22(5): 207-216. |
| [14] | Zhong-hua Sheng, Peng-fei Fang, San-feng Li, Gui-ai Jiao, Li-hong Xie, Pei-song Hu, Shao-qing Tang, Xiang-jin Wei. Phenotype of Rice Floury Endosperm Mutant flo7 and Fine Mapping of Mutated Gene [J]. Rice Science, 2015, 22(4): 162-170. |
| [15] | Min Guo, Rong-de Li, Jian Yao, Juan Zhu, Xiang-yun Fan, Wei Wang, Shu-zhu Tang, Ming-hong Gu, Chang-jie Yan. RL3(t), Responsible for Leaf Shape Formation, Delimited to a 46-kb DNA Fragment in Rice [J]. Rice Science, 2015, 22(1): 44-48. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||