
Rice Science ›› 2017, Vol. 24 ›› Issue (3): 123-144.DOI: 10.1016/j.rsci.2016.09.00
• Orginal Article • Next Articles
Naga Bheema Lingeswara Reddy Inja, Kim Beom-Ki, Yoon In-Sun, Kim Kyung-Hwan, Kwon Taek-Ryoun( )
)
Received:2016-07-25
Accepted:2016-09-23
Online:2017-05-28
Published:2017-03-03
Naga Bheema Lingeswara Reddy Inja, Kim Beom-Ki, Yoon In-Sun, Kim Kyung-Hwan, Kwon Taek-Ryoun. Salt Tolerance in Rice: Focus on Mechanisms and Approaches[J]. Rice Science, 2017, 24(3): 123-144.
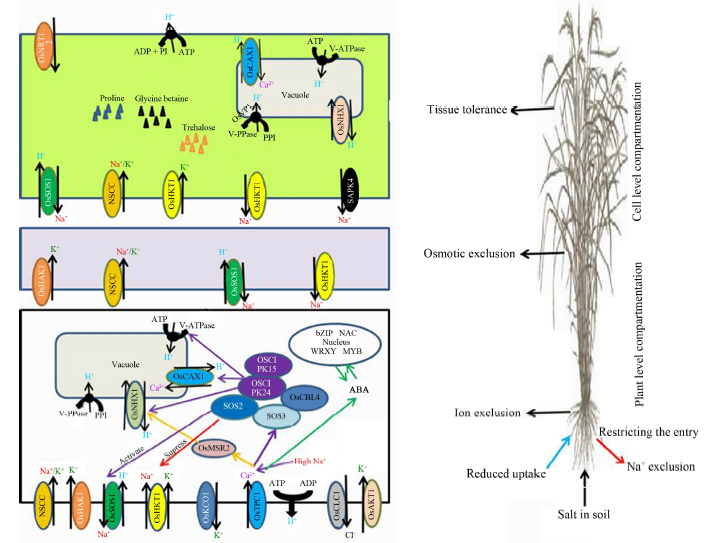
Fig. 1. Rice salt tolerance mechanism-overview of important genes involved at root, shoot and leaf levels. (Tissue tolerance, osmotic exclusion and ion exclusion prevent the accumulation of toxic concentrations of Na+ and Cl-. Mechanisms include genes such as OsSOS1, which reduces Na+ content in cell compartmentation of ions in vacuoles and/or efflux of ions back to the soil etc.)
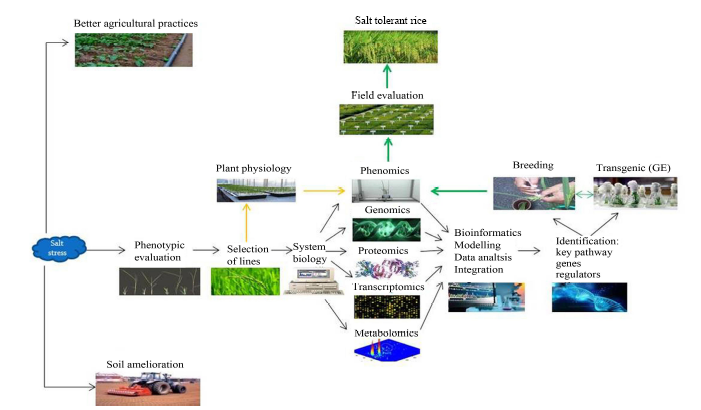
Fig. 2. Approaches for developing of salt tolerant rice plants.( Platforms such as phenomics in tandem with combination of genomics-transcriptomics-proteomics-metabolomics need to be integrated for identifying key genes/regulators which can improve salinity tolerance in rice apart from general practices like usage of better agricultural practices and soil amelioration.)
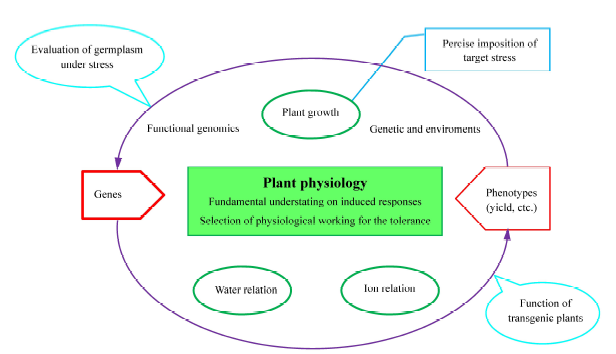
Fig. 3. Schematic flow of plant physiological studies to link between gene and phenotype.( Physiological traits related to stress tolerances are (complex multiple attributes to maintain growth of plants in the presence of abiotic stress condition. Physiological works have a significant role to bridge between gene work and phenotyping in the improvement of the tolerance in plants.))
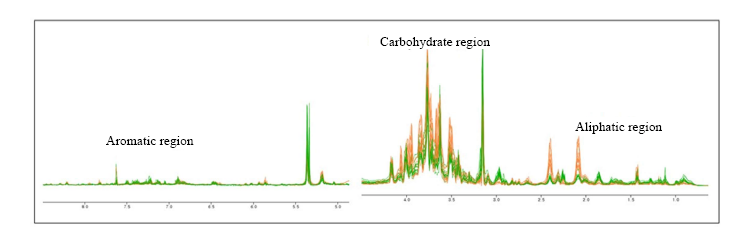
Fig. 5. H-NMR (Proton Nuclear Magnetic Resonance) spectrum of Oryza sativa treated with or without NaCl (45 mmol/L).( Orange line indicates the control, and green line indicates the salt treated one.NMR-based metabolomics was used to differentiate between 20 salt sensitive and 20 salt tolerant varieties. The major factor for this discrimination was due to the change of carbohydrate and aliphatic regions.)
| Salt responsive element | Important gene | Reference |
|---|---|---|
| Receptor-like kinase | OsSIK1, OsSIK2, OsRPK1 | Ouyang et al, 2010; Chen et al, 2013; Zou et al, 2014 |
| Mitogen activated protein | OsEDR1, OsMAPK5, OsMAPK44, OsMAPK33, OsMKK1, OsMPK4, OsMKK6 | Kim et al, 2003; Xiong and Yang, 2003; Jeong et al, 2006; Lee et al, 2011; Kumar et al, 2013; Wang et al, 2014 |
| SNF1-related protein kinase | SAPK4, SnRK2 | Kobayashi et al, 2004; Diedhiou et al, 2008; Nam et al, 2012 |
| Transcription factor | OsABF2, OsbZIP23, OsbZIP71, OsABI5, OsMYB3R-2, OsMYB2, OsMYB91, OsNAC6, OsNAC5, OSISAP1, DST, OsTZF, OsTZF1, OsWRKY13, OsWRKY42, OsERF922, OsNAC5, OsNAC10, OsNAP, ONAC, ONAC022, OsRAB1, MYC/MYB, OsNAC/SNAC, SERF1, OsDREB1A, OsDREB1F, OsDREB2A, OrbHLH001, | Dubouzet et al, 2003; Mukhopadhyay et al, 2004; Hu et al, 2006, 2008; Dai et al, 2007; Nakashima et al, 2007; Qiu et al, 2008; Wang et al, 2008; Xiang et al, 2008; Zou et al, 2008; Huang et al, 2009; Zheng et al, 2009; Hossain et al, 2010; Jeong et al, 2010, 2013; Takasaki et al, 2010; Mallikarjuna et al, 2011; Song et al, 2011; Asano et al, 2012; Liu et al, 2012; Redillas et al, 2012; Yang A et al, 2012; Yang C et al, 2012; Jan et al, 2013; Schmidt et al, 2013; Chen X et al, 2014; Liang et al, 2014; Yoshida et al, 2014; Hong et al, 2016 |
| OrbHLH, OsNAC6, SERF1, DST, OsTZF1 | ||
| MicroRNA | Osa-miR396, Osa-miR820, Osa-miR169, Osa-miR2006, Osa-miR29, MIR393, MIR1425 | Zhao et al, 2009; Jian et al, 2010; Gao et al, 2011; Barrera-Figueroa et al, 2012; Xia et al, 2012; Sharma et al, 2015 |
| ROS-producing/scavenging enzyme | NADP-ME, katE encoding catalase, OsIM1, PDH45, SERF1, OsCatC, OsAPx2 | Cournac et al, 2002; Kong et al, 2003; Liu et al, 2007; Wutipraditkul et al, 2011; Guan et al, 2012; Schmidt et al, 2013; Nath et al, 2016 |
| SOS pathway | OsIM1, OsSOS3, OsSOS2 | Motohashi et al, 2010; Son et al, 2013 |
| Calmodulin (CaM) pathway | OsMSR2, OsCam1-1, OsCML4, OsCML5, OsCML8, OsCML11, OsCaM1, OsCaM2 | Chinpongpanich et al, 2012; Saeng-ngam et al, 2012; Xu et al, 2013 |
| Phospholipid and Ca2+ ion | OsCIPK24, OsCBL4, OsCIPK15 | Xiang et al, 2007; Yang A et al, 2012 |
| Ca-dependent protein kinase | OsCPK4, OsCPK12, OsCDPK7 | Saijo et al, 2000; Asano et al, 2012; Campo et al, 2014 |
| Kinase | SIT1 | Li et al, 2014 |
| Calcium transporter | OsACA6 | Kamrul Huda et al, 2014 |
| Nitrate transporter | OsNRT1;2 | Wang et al, 2012 |
| Helicase | OsSUV3 | Sahoo et al, 2014 |
| Na+/H+ antiporter/exchanger | OsSOS1, OsNHX1 | Bassil et al, 2012; Kumar et al, 2013; Amin et al, 2016 |
| Na+/K+ symporter | OsHKT1;4, OsHKT1;1, OsHKT2:1, OsHKT1;5, HKT2;3, SKC1 | Cotsaftis et al, 2012; Takagi et al, 2015; Mishra et al, 2016; Suzuki et al, 2016 |
| H+-ATPase | OSA3, OsVHA-a1, OSA7 | Sahu and Shaw, 2009; Krebs et al, 2010; Kumar et al, 2013; Son et al, 2013 |
| Channel protein | OsCLC1, OsAKT1, OsHAK1, OsHAK5, | Diedhiou and Golldack, 2006; Yang et al, 2014; Chen et al, 2015; Shen et al, 2015; Ahmad et al, 2016 |
| OsHAK21 | ||
| H+/Ca+ antiporter | OsCAX1 | Kumar et al, 2013 |
| Proline | P5CS, P5CR, SIDP361 | Karthikeyan et al, 2011; Nounjan et al, 2012; Li et al, 2016 |
| Glycine betaine | codA, CMO | Mohanty et al, 2002; Su et al, 2006 |
| Myoinositol | PINO1 | Majee et al, 2004 |
| Trehalose | OsTPS1-11, OsTPS-1 | Li et al, 2011; Zang et al, 2011 |
| Aquaporin | OsTIP1;1, HvPIP2;1 | Katsuhara et al, 2003; Hanba et al, 2004; Sakurai et al, 2005 |
| Abscisic acid | OsAP2-39, OsNCEDs1, OsNCEDs3, | Yaish et al, 2010; Shi et al, 2015; Hong et al, 2016 |
| OsNCEDs4, OsNCEDs5, OsPSY1 | ||
| Jasmonate | OsTIFY1, OsTIFY6, OsTIFY9, OsTIFY10, OsTIFY11 | Ye et al, 2009 |
| Brassinosteroid | OsGSK1(BIN2), OsBRI1, OsDWF4, OsSalT | Koh et al, 2007; Sharma et al, 2013; Zhao et al, 2013 |
Table 1 Various stress responsive elements and their corresponding genes which can lead to development of salinity tolerant rice plants.
| Salt responsive element | Important gene | Reference |
|---|---|---|
| Receptor-like kinase | OsSIK1, OsSIK2, OsRPK1 | Ouyang et al, 2010; Chen et al, 2013; Zou et al, 2014 |
| Mitogen activated protein | OsEDR1, OsMAPK5, OsMAPK44, OsMAPK33, OsMKK1, OsMPK4, OsMKK6 | Kim et al, 2003; Xiong and Yang, 2003; Jeong et al, 2006; Lee et al, 2011; Kumar et al, 2013; Wang et al, 2014 |
| SNF1-related protein kinase | SAPK4, SnRK2 | Kobayashi et al, 2004; Diedhiou et al, 2008; Nam et al, 2012 |
| Transcription factor | OsABF2, OsbZIP23, OsbZIP71, OsABI5, OsMYB3R-2, OsMYB2, OsMYB91, OsNAC6, OsNAC5, OSISAP1, DST, OsTZF, OsTZF1, OsWRKY13, OsWRKY42, OsERF922, OsNAC5, OsNAC10, OsNAP, ONAC, ONAC022, OsRAB1, MYC/MYB, OsNAC/SNAC, SERF1, OsDREB1A, OsDREB1F, OsDREB2A, OrbHLH001, | Dubouzet et al, 2003; Mukhopadhyay et al, 2004; Hu et al, 2006, 2008; Dai et al, 2007; Nakashima et al, 2007; Qiu et al, 2008; Wang et al, 2008; Xiang et al, 2008; Zou et al, 2008; Huang et al, 2009; Zheng et al, 2009; Hossain et al, 2010; Jeong et al, 2010, 2013; Takasaki et al, 2010; Mallikarjuna et al, 2011; Song et al, 2011; Asano et al, 2012; Liu et al, 2012; Redillas et al, 2012; Yang A et al, 2012; Yang C et al, 2012; Jan et al, 2013; Schmidt et al, 2013; Chen X et al, 2014; Liang et al, 2014; Yoshida et al, 2014; Hong et al, 2016 |
| OrbHLH, OsNAC6, SERF1, DST, OsTZF1 | ||
| MicroRNA | Osa-miR396, Osa-miR820, Osa-miR169, Osa-miR2006, Osa-miR29, MIR393, MIR1425 | Zhao et al, 2009; Jian et al, 2010; Gao et al, 2011; Barrera-Figueroa et al, 2012; Xia et al, 2012; Sharma et al, 2015 |
| ROS-producing/scavenging enzyme | NADP-ME, katE encoding catalase, OsIM1, PDH45, SERF1, OsCatC, OsAPx2 | Cournac et al, 2002; Kong et al, 2003; Liu et al, 2007; Wutipraditkul et al, 2011; Guan et al, 2012; Schmidt et al, 2013; Nath et al, 2016 |
| SOS pathway | OsIM1, OsSOS3, OsSOS2 | Motohashi et al, 2010; Son et al, 2013 |
| Calmodulin (CaM) pathway | OsMSR2, OsCam1-1, OsCML4, OsCML5, OsCML8, OsCML11, OsCaM1, OsCaM2 | Chinpongpanich et al, 2012; Saeng-ngam et al, 2012; Xu et al, 2013 |
| Phospholipid and Ca2+ ion | OsCIPK24, OsCBL4, OsCIPK15 | Xiang et al, 2007; Yang A et al, 2012 |
| Ca-dependent protein kinase | OsCPK4, OsCPK12, OsCDPK7 | Saijo et al, 2000; Asano et al, 2012; Campo et al, 2014 |
| Kinase | SIT1 | Li et al, 2014 |
| Calcium transporter | OsACA6 | Kamrul Huda et al, 2014 |
| Nitrate transporter | OsNRT1;2 | Wang et al, 2012 |
| Helicase | OsSUV3 | Sahoo et al, 2014 |
| Na+/H+ antiporter/exchanger | OsSOS1, OsNHX1 | Bassil et al, 2012; Kumar et al, 2013; Amin et al, 2016 |
| Na+/K+ symporter | OsHKT1;4, OsHKT1;1, OsHKT2:1, OsHKT1;5, HKT2;3, SKC1 | Cotsaftis et al, 2012; Takagi et al, 2015; Mishra et al, 2016; Suzuki et al, 2016 |
| H+-ATPase | OSA3, OsVHA-a1, OSA7 | Sahu and Shaw, 2009; Krebs et al, 2010; Kumar et al, 2013; Son et al, 2013 |
| Channel protein | OsCLC1, OsAKT1, OsHAK1, OsHAK5, | Diedhiou and Golldack, 2006; Yang et al, 2014; Chen et al, 2015; Shen et al, 2015; Ahmad et al, 2016 |
| OsHAK21 | ||
| H+/Ca+ antiporter | OsCAX1 | Kumar et al, 2013 |
| Proline | P5CS, P5CR, SIDP361 | Karthikeyan et al, 2011; Nounjan et al, 2012; Li et al, 2016 |
| Glycine betaine | codA, CMO | Mohanty et al, 2002; Su et al, 2006 |
| Myoinositol | PINO1 | Majee et al, 2004 |
| Trehalose | OsTPS1-11, OsTPS-1 | Li et al, 2011; Zang et al, 2011 |
| Aquaporin | OsTIP1;1, HvPIP2;1 | Katsuhara et al, 2003; Hanba et al, 2004; Sakurai et al, 2005 |
| Abscisic acid | OsAP2-39, OsNCEDs1, OsNCEDs3, | Yaish et al, 2010; Shi et al, 2015; Hong et al, 2016 |
| OsNCEDs4, OsNCEDs5, OsPSY1 | ||
| Jasmonate | OsTIFY1, OsTIFY6, OsTIFY9, OsTIFY10, OsTIFY11 | Ye et al, 2009 |
| Brassinosteroid | OsGSK1(BIN2), OsBRI1, OsDWF4, OsSalT | Koh et al, 2007; Sharma et al, 2013; Zhao et al, 2013 |

Fig. 6. Comparison of OsCIPK induced plants to control under salt stress.( Salt response was compared between null, overexpressed (OX), knockout (KO) and control/salt free (Pokkali) lines. OsCIPK could achieve higher biomass and less Na+ through enhanced Na+ exclusion via Na+/H+ antiporter.)
| 1 | Abdallah M M S, Abdelgawad Z A, El-Bassiouny H M S.2016. Alleviation of the adverse effects of salinity stress using trehalose in two rice varieties.South Afr J Bot, 103: 275-282. |
| 2 | Afzal Z, Howton T C, Sun Y, Mukhtar M S.2016. The roles of aquaporins in plant stress responses.J Dev Biol, 4(1): 9. |
| 3 | Ahmad I, Mian A, Maathuis F J M.2016. Overexpression of the rice AKT1 potassium channel affects potassium nutrition and rice drought tolerance.J Exp Bot, 67(9): 2689-2698. |
| 4 | Ahmad P, Azooz M M, Prasad M N V.2013. Salt Stress in Plants: Signalling, Omics and Adaptations. New York, USA: Science, Springer Science & Business Media. |
| 5 | Akbar M, Yabuno T.1977. Breeding for saline-resistant varieties of rice: IV. Inheritance of delayed-type panicle sterility induced by salinity.Jpn J Breeding, 27(3): 237-240. |
| 6 | Amin U S M, Biswas S, Elias S M, Razzaque S, Haque T, Malo R, Seraj Z I.2016. Enhanced salt tolerance conferred by the complete 2.3 kb cDNA of the rice vacuolar Na+/H+ antiporter gene compared to 1.9 kb coding region with 5′-UTR in transgenic lines of rice.Front Plant Sci, 7: 14. |
| 7 | Amtmann A, Sanders D.1998. Mechanisms of Na+ uptake by plant cells.Adv Bot Res, 29: 75-112. |
| 8 | Anuradha S, Rao S S R.2003. Application of brassinosteroids to rice seeds (Oryza sativa L.) reduced the impact of salt stress on growth, prevented photosynthetic pigment loss and increased nitrate reductase activity. Plant Growth Regul, 40(1): 29-32. |
| 9 | Asano T, Hayashi N, Kobayashi M, Aoki N, Miyao A, Mitsuhara I, Ichikawa H, Komatsu S, Hirochika H, Kikuchi S, Ohsugi R.2012. A rice calcium-dependent protein kinase OsCPK12 oppositely modulates salt-stress tolerance and blast disease resistance.Plant J, 69(1): 26-36. |
| 10 | Asch F, Wopereis M C S.2001. Responses of field grown irrigated rice cultivars to varying levels of floodwater salinity in a semi-arid environment.Field Crops Res, 70(2): 127-137. |
| 11 | Ashraf M, Akram N A.2009. Improving salinity tolerance of plants through conventional breeding and genetic engineering: An analytical comparison.Biotechnol Adv, 27(6): 744-752. |
| 12 | Barrera-Figueroa B E, Gao L, Wu Z G, Zhou X F, Zhu J H, Jin H L, Liu R L, Zhu J K.2012. High throughput sequencing reveals novel and abiotic stress-regulated microRNAs in the inflorescences of rice.BMC Plant Biol, 12: 132. |
| 13 | Bassil E, Coku A, Blumwald E.2012. Cellular ion homeostasis: Emerging roles of intracellular NHX Na+/H+ antiporters in plant growth and development.J Exp Bot, 63: 5727-5740. |
| 14 | Blumwald E.1987. Tonoplast vesicles as a tool in the study of ion transport at the plant vacuole.Physiol Plant, 69(4): 731-734. |
| 15 | Blumwald E.2000. Sodium transport and salt tolerance in plants.Curr Opin Cell Biol, 12(4): 431-434. |
| 16 | Bohnert H J, Shen B.1999. Transformation and compatible solutes.Sci Hort, 78: 237-260. |
| 17 | Cai X, Chen T, Zhou Q Y, Xu L, Qu L Q, Hua X J, Lin J X.2011. Development of casparian strip in rice cultivars.Plant Signal Behav, 6(1): 59-65. |
| 18 | Campo S, Baldrich P, Messeguer J, Lalanne E, Coca M, San Segundo B.2014. Overexpression of a calcium-dependent protein kinase confers salt and drought tolerance in rice by preventing membrane lipid peroxidation.Plant Physiol, 165(2): 688-704. |
| 19 | Chen G, Hu Q D, Luo L, Yang T Y, Zhang S, Hu Y B, Yu L, Xu G H.2015. Rice potassium transporter OsHAK1 is essential for maintaining potassium mediated growth and functions in salt tolerance over low and high potassium concentration ranges.Plant Cell Environ, 38(12): 2747-2765. |
| 20 | Chen H D, Xie W B, He H, Yu H H, Chen W, Li J, Yu R B, Yao Y, Zhang W H, He Y Q, Tang X Y, Zhou F S, Deng X W, Zhang Q F.2014. A high-density SNP genotyping array for rice biology and molecular breeding.Mol Plant, 7(3): 541-553. |
| 21 | Chen L J, Wuriyanghan H, Zhang Y Q, Duan K X, Chen H W, Li Q T, Lu X, He S J, Ma B, Zhang W K, Lin Q, Chen S Y, Zhang J S.2013. An S-domain receptor-like kinase, OsSIK2, confers abiotic stress tolerance and delays dark-induced leaf senescence in rice.Plant Physiol, 163(4): 1752-1765. |
| 22 | Chen M, Chen Q J, Niu X G, Zhang R, Lin H Q, Xu C Y, Wang X C, Wang G Y, Chen J.2007. Expression ofOsNHX1 gene in maize confers salt tolerance and promotes plant growth in the field. Plant Soil Environ, 53(11): 490-498. |
| 23 | Chen T, Cai X, Wu X Q, Karahara I, Schreiber L, Lin J X.2011. Casparian strip development and its potential function in salt tolerance.Plant Signal Behav, 6(10): 1499-1502. |
| 24 | Chen T H H, Murata N.2008. Glycinebetaine: An effective protectant against abiotic stress in plants.Trends Plant Sci, 13(9): 499-505. |
| 25 | Chen X, Wang Y F, Lv B, Li J, Luo L Q, Lu S C, Zhang X, Ma H, Ming F.2014. The NAC family transcription factorOsNAP confers abiotic stress response through the ABA pathway. Plant Cell Physiol, 55(3): 604-619. |
| 26 | Chinnusamy V, Jagendorf A, Zhu J K.2005. Understanding and improving salt tolerance in plants.Crop Sci, 45(2): 437-448. |
| 27 | Chinpongpanich A, Limruengroj K, Phean-o-pas S, Limpaseni T, Buaboocha T.2012. Expression analysis of calmodulin and calmodulin-like genes from rice,Oryza sativa L. BMC Res Notes, 5: 625. |
| 28 | Cotsaftis O, Plett D, Johnson A A T, Walia H, Wilson C, Ismail A M, Close T J, Tester M, Baumann U.2011. Root-specific transcript profiling of contrasting rice genotypes in response to salinity stress.Mol Plant, 4(1): 1-17. |
| 29 | Cotsaftis O, Plett D, Shirley N, Tester M, Hormova M.2012. A two-staged model of Na+ exclusion in rice explained by 3D modeling of HKT transporter and alternative splicing.PLoS One, 7(7): e39865. |
| 30 | Cournac L, Latouche G, Cerovic Z, Redding K, Ravenel J, Peltier G.2002. In vivo interactions between photosynthesis, mitorespiration, and chlororespiration inChlamydomonas reinhardtii. Plant Physiol, 129(4): 1921-1928. |
| 31 | Dai X Y, Xu Y Y, Ma Q B, Xu W Y, Wang T, Xue Y B, Chong K.2007. Overexpression of an R1R2R3 MYB gene,OsMYB3R-2, increases tolerance to freezing, drought, and salt stress in transgenic Arabidopsis. Plant Physiol, 143(4): 1739-1751. |
| 32 | Das P, Mishra M, Lakra N, Singla-Pareek S L, Pareek A.2014. Mutation breeding: A powerful approach for obtaining abiotic stress tolerant crops and upgrading food security for human nutrition. In: Tomlekova N, Kojgar M I, Wani M R. Mutagenesis: Exploring Novel Genes and Pathways. Wageningen: Wageningen Academic Publisher: 15-36. |
| 33 | Das P, Nutan K K, Singla-Pareek S L, Pareek A.2015. Oxidative environment and redox homeostasis in plants: Dissecting out significant contribution of major cellular organelles.Front Environ Sci, 2: 70. |
| 34 | Deng P, Jiang D, Dong Y M, Shi X Y, Jing W, Zhang W H.2015. Physiological characterisation and fine mapping of a salt-tolerant mutant in rice (Oryza sativa). Func Plant Biol, 42(11): 1026-1035. |
| 35 | Dhar R, Sägesser R, Weikert C, Yuan J, Wagner A.2011. Adaptation ofSaccharomyces cerevisiae to saline stress through laboratory evolution. J Evol Biol, 24(5): 1135-1153. |
| 36 | Diedhiou C J, Golldack D.2006. Salt-dependent regulation of chloride channel transcripts in rice.Plant Sci, 170(4): 793-800. |
| 37 | Diedhiou C J, Popova O V, Dietz K J, Golldack D.2008. The SNF1-type serine-threonine protein kinase SAPK4 regulates stress-responsive gene expression in rice.BMC Plant Biol, 8: 49. |
| 38 | Dubouzet J G, Sakuma Y, Ito Y, Kasuga M, Dubouzet E G, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K.2003. OsDREB genes in rice,Oryza sativa L. encode transcription activators that function in drought, high salt and cold-responsive gene expression. Plant J, 33(4): 751-763. |
| 39 | Elbein A D, Pan Y T, Pastuszak I, Carroll D.2003. New insights on trehalose: A multifunctional molecule.Glycobiology, 13(4): 17R-27R. |
| 40 | Enstone D E, Peterson C A, Ma F S.2002. Root endodermis and exodermis: Structure, function, and responses to the environment.J Plant Growth Regul, 21(4): 335-351. |
| 41 | Eynard A, Lal R, Wiebe K.2005. Crop response in salt-affected soils.J Sustain Agric, 27: 5-50. |
| 42 | FAO. 2008.. |
| 43 | Ferdose J, Kawasaki M, Taniguchi M, Miyake H.2009. Differential sensitivity of rice cultivars to salinity and its relation to ion accumulation and root tip structure.Plant Prod Sci, 12(4): 453-461. |
| 44 | Flowers T J, Flowers S A.2005. Why does salinity pose such a difficult problem for plant breeders?Agric Water Manag, 78: 15-24. |
| 45 | Fukuda A, Nakamura A, Tagiri A, Tanaka H, Miyao A, Hirochika H, Tanaka Y.2004. Function, intracellular localization and the importance in salt tolerance of a vacuolar Na+/H+ antiporter from rice.Plant Cell Physiol, 45(2): 146-159. |
| 46 | Gao P, Bai X, Yang L, Lv D K, Pan X, Li Y, Cai H, Ji W, Chen Q, Zhu Y M.2011. osa-MIR393: A salinity and alkaline stress- related microRNA gene.Mol Biol Rep, 38(1): 237-242. |
| 47 | Garcia A, Rizzo C A, Ud-Din J, Bartos S L, Senadhira D, Flowers T J, Yeo A R.1997. Sodium and potassium transport to the xylem are inherited independently in rice, and the mechanism of sodium: Potassium selectivity differs between rice and wheat.Plant Cell Environ, 20(9): 1167-1174. |
| 48 | Grattan S R, Zeng L, Shannon M C, Roberts S R.2002. Rice is more sensitive to salinity than previously thought.Calif Agric, 56(6): 189-195. |
| 49 | Greenway H, Munns R.1980. Mechanism of salt tolerance in nonhalophytes.Ann Rev Plant Physiol, 31: 149-190. |
| 50 | Guan Q J, Takano T, Liu S K.2012. Genetic transformation and analysis of riceOsAPx2 gene in Medicago sativa. PLoS One, 7(7): e41233. |
| 51 | Guo Y, Qiu Q S, Quintero F J, Pardo J M, Ohta M, Zhang C, Schumaker K S, Zhu J K.2004. Transgenic evaluation of activated mutant alleles of SOS2 reveals a critical requirement for its kinase activity and C-terminal regulatory domain for salt tolerance inArabidopsis thaliana. Plant Cell, 16(2): 435-449. |
| 52 | Gupta B, Huang B.2014. Mechanism of salinity tolerance in plants: Physiological, biochemical, and molecular characterization.Int J Genom, 2014(1): 701596. |
| 53 | Hairmansis A, Berger B, Tester M, Roy S J.2014. Image-based phenotyping for non-destructive screening of different salinity tolerance traits in rice.Rice, 7: 16. |
| 54 | Hakim M A, Juraimi A S, Begum M, Hanafi M M, Ismail M R, Selamat A.2010. Effect of salt stress on germination and early seedling growth of rice (Oryza sativa L.). Afr J Biotechnol, 9(13): 1911-1918. |
| 55 | Halfter U, Ishitani M, Zhu J K.2000. TheArabidopsis SOS2 protein kinase physically interacts with and is activated by the calcium-binding protein SOS3. Proc Natl Acad Sci USA, 97(7): 3735-3740. |
| 56 | Hanba Y T, Shibasaka M, Hayashi Y, Hayakawa T, Kasamo K, Terashima I, Katsuhara M.2004. Overexpression of the barley aquaporinHvPIP2;1 increases internal CO2 conductance and CO2 assimilation in the leaves of transgenic rice plants. Plant Cell Physiol, 45(5): 521-529. |
| 57 | Hare P D, Cress W A, van Staden J.1998. Dissecting the roles of osmolyte accumulation during stress.Plant Cell Environ, 21(6): 535-553. |
| 58 | Hong Y B, Zhang H J, Huang L, Li D Y, Song F M.2016. Overexpression of a stress-responsive NAC transcription factor geneONAC022 improves drought and salt tolerance in rice. Front Plant Sci, 7: 4. |
| 59 | Hossain M A, Cho J I, Han M, Ahn C H, Jeon J S, An G, Park P B.2010. The ABRE binding bZIP transcription factorOsABF2 is a positive regulator of abiotic stress and ABA signaling in rice. J Plant Physiol, 167(17): 1512-1520. |
| 60 | Hu H H, Dai M Q, Yao J L, Xiao B Z, Li X H, Zhang Q F, Xiong L Z.2006. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice.Proc Natl Acad Sci USA, 103: 12987-12992. |
| 61 | Hu H H, You J, Fang Y J, Zhu X Y, Qi Z Y, Xiong L Z.2008. Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice.Plant Mol Biol, 67: 169-181. |
| 62 | Huang S B, Spielmeyer W, Lagudah E S, Munns R.2008. Comparative mapping of HKT genes in wheat, barley, and rice, key determinants of Na+ transport, and salt tolerance. J Exp Bot, 59(4): 927-937. |
| 63 | Huang X Y, Chao D Y, Gao J P, Zhu M Z, Shi M, Lin H X.2009. A previously unknown zinc finger protein, DST, regulates drought and salt tolerance in rice via stomatal aperture control.Genes Dev, 23: 1805-1817. |
| 64 | Humplik J F, Lazar D, Husickova A, Spichal L.2015. Automated phenotyping of plant shoots using imaging methods for analysis of plant stress responses: A review.Plant Methods, 11: 29. |
| 65 | Ishak N K, Sulaiman Z, Tennakoon K U.2015. Comparative study on growth performance of transgenic (over-expressedOsNHX1) and wild-type Nipponbare under different salinity regimes. Rice Sci, 22(6): 275-282. |
| 66 | Jabnoune M, Espeout S, Mieulet D, Fizames C, Verdeil J L, Conéjéro G, Véry A A.2009. Diversity in expression patterns and functional properties in the rice HKT transporter family.Plant Physiol, 150(4): 1955-1971. |
| 67 | Jan A, Maruyama K, Todaka D, Kidokoro S, Abo M, Yoshimura E, Shinozaki K, Nakashima K, Yamaguchi-Shinozaki K.2013. OsTZF1, a CCCH-tandem zinc finger protein, confers delayed senescence and stress tolerance in rice by regulating stress- related genes.Plant Physiol, 161(3): 1202-1216. |
| 68 | Jangam A P, Pathak R R, Raghuram N.2016. Microarray analysis of riced1 (RGA1) mutant reveals the potential role of G-protein alpha subunit in regulating multiple abiotic stresses such as drought, salinity, heat, and cold. Front Plant Sci, 7: 11. |
| 69 | Jeong J S, Kim Y S, Baek K H, Jung H, Ha S H, Choi Y D, Kim M, Reuzeau C, Kim J K.2010. Root-specific expression ofOsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol, 153(1): 185-197. |
| 70 | Jeong J S, Kim Y S, Redillas M C, Jang G, Jung H, Bang S W, Choi Y D, Ha S H, Reuzeau C, Kim J K.2013. OsNAC5 overexpression enlarges root diameter in rice plants leading to enhanced drought tolerance and increased grain yield in the field.Plant Biotechnol J, 11(1): 101-114. |
| 71 | Jeong M J, Lee S K, Kim B G, Kwon T R, Cho W S, Park Y T, Lee J O, Kwon H B, Byun M O, Park S C.2006. A rice (Oryza sativa L.) MAP kinase gene, OsMAPK44, is involved in response to abiotic stresses. Plant Cell Tiss Organ Cult, 85(2): 151-160. |
| 72 | Jeschke W D.1984. K+-Na+ exchange at cellular membranes, intracellular compartmentation of cations, and salt tolerance. In: Staples R C. Salinity Tolerance in Plants: Strategies for Crop Improvement. New York: Wiley: 37-66. |
| 73 | Jian X Y, Zhang L, Li G L, Zhang L, Wang X J, Cao X F, Fang X H, Chen F.2010. Identification of novel stress-regulated microRNAs fromOryza sativa L. Genomics, 95(1): 47-55. |
| 74 | Kamiya T, Akahori T, Ashikari M, Maesshima M.2005. Expression of the vacuolar Ca2+/H+ exchanger, OsCAX1a, in rice: Cell and age specificity of expression and enhancement by Ca2+.Plant Cell Physiol, 47(1): 96-106. |
| 75 | Kamrul Huda K M, Akhter Banu M S, Yadav S, Sahoo R K, Tuteja R, Tuteja N.2014. Salinity and drought tolerant OsACA6 enhances cold tolerance in transgenic tobacco by interacting with stress-inducible proteins.Plant Physiol Biochem, 82: 229-238. |
| 76 | Karthikeyan A, Pandian S K, Ramesh M.2011. Transgenic indica rice cv. ADT43 expressing aΔ1-pyrroline-5-carboxylate synthetase (P5CS) gene from Vigna aconitifolia demonstrates salt tolerance. Plant Cell, Tiss Organ Cult, 107(3): 383-395. |
| 77 | Kathuria H, Giri J, Nataraja K N, Murata N, Udayakumar M, Tyagi A K.2009. Glycinebetaine-induced water-stress tolerance in codA-expressing transgenic indica rice is associated with up-regulation of several stress responsive genes.Plant Biotech J, 7(6): 512-526. |
| 78 | Katsuhara M, Koshio K, Shibasaka M, Hayashi Y, Hayakawa T, Kasamo K.2003. Over-expression of a barley aquaporin increased the shoot/root ratio and raised salt sensitivity in transgenic rice plants.Plant Cell Physiol, 44(12): 1378-1383. |
| 79 | Khatun S, Flowers T J.1995. Effects of salinity on seed set in rice.Plant Cell Environ, 18(1): 61-67. |
| 80 | Khatun S, Rizzo C A, Flowers T J.1995. Genotypic variation in the effect of salinity on fertility in rice.Plant Soil, 173(2): 239-250. |
| 81 | Kim J A, Agrawal G K, Rakwal R, Han K S, Kim K N, Yun C H, Heu S, Park S Y, Lee Y H, Jwa N S.2003. Molecular cloning and mRNA expression analysis of a novel rice (Oryza sativa L.) MAPK kinase, OsEDR1, an ortholog of Arabidopsis AtEDR1, reveal its role in defense/stress signalling pathways and development. Biochem Biophys Res Commun, 300(4): 868-876. |
| 82 | Kobayashi Y, Yamamoto S, Minami H, Kagaya Y, Hattori T.2004. Differential activation of the rice sucrose nonfermenting1-related protein kinase2 family by hyperosmotic stress and abscisic acid.Plant Cell, 16(5): 1163-1177. |
| 83 | Koh S, Lee S C, Kim M K, Koh J H, Lee S, An G, Choe S, Kim S R.2007. T-DNA tagged knockout mutation of riceOsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses. Plant Mol Biol, 65(4): 453-466. |
| 84 | Kong J, Gong J M, Zhang Z G, Zhang J S, Chen S Y.2003. A new AOX homologous geneOsIM1 from rice(Oryza sativa L.) with an alternative splicing mechanism under salt stress. Theor Appl Genet, 107(2): 326-331. |
| 85 | Krebs M, Beyhl D, Gorlich E, Al-Rasheid K A S, Marten I, Stierhof Y D, Hedrich R, Schumacher K.2010. Arabidopsis V-ATPase activity at the tonoplast is required for efficient nutrient storage but not for sodium accumulation.Proc Natl Acad Sci USA, 107(7): 3251-3256. |
| 86 | Kromdijk J, Long S P.2016. One crop breeding cycle from starvation? How engineering crop photosynthesis for rising CO2 and temperature could be one important route to alleviation.Proc Royal Soc B: Biol Sci, 283: 20152578. |
| 87 | Krishnamurthy P, Ranathunge K, Franke R, Prakash H S, Schreiber L, Mathew M K.2009. The role of root apoplastic transport barriers in salt tolerance of rice (Oryza sativa L.). Planta, 230(1): 119-134. |
| 88 | Kumar K, Kumar M, Kim S R, Ryu H, Cho Y G.2013. Insights into genomics of salt stress response in rice.Rice, 6(1): 27. |
| 89 | Kumar K, Sinha A K.2013. Overexpression of constitutively active mitogen activated protein kinase kinase 6 enhances tolerance to salt stress in rice.Rice, 6(1): 25. |
| 90 | Kumari S, Sabharwal V P N, Khushwaha H R, Sopory S K, Singla-Pareek S L, Pareek A.2009. Transcriptome map for seedling stage specific salinity stress response indicates a specific set of genes as candidate for saline tolerance inOryza sativa L. Funct Integr Genom, 9: 109-123. |
| 91 | Kurokawa Y, Noda T, Yamagata Y, Angeles-Shim R, Sunohara H, Uehara K, Furuta T, Nagai K, Jena K K, Yasui H, Yoshimura A.2016. Construction of a versatile SNP array for pyramiding useful genes of rice.Plant Sci, 242: 131-139. |
| 92 | Kurotani K, Yamanaka K, Toda Y, Ogawa D, Tanaka M, Kozawa H, Nakamura H, Hakata M, Ichikawa H, Hattori T, Takeda S.2015. Stress tolerance profiling of a collection of extant salt-tolerant rice varieties and transgenic plants overexpressing abiotic stress tolerance genes.Plant Cell Physiol, 56(10): 1867-1876. |
| 93 | Kurusu T, Hamada H, Koyano T, Kuchitsu K.2012. Intracellular localization and physiological function of a rice Ca2+-permeable channel OsTPC1.Plant Signal Behav, 7(11): 1428-1430. |
| 94 | Lauchli A, Luttge U.2002. Salinity and Nitrogen Nutrition. Boston: Boston Kluwer Academic Publishers: 229-248. |
| 95 | Lauchli A, Grattan S R.2007. Plant growth and development under salinity stress. In: Jenks M A, Hasegawa P M, Jain S M. Advances in Molecular Breeding Toward Drought and Salt Tolerant Crops. Dordrecht, the Netherlands: Springer: 1-32. |
| 96 | Lee D G, Ahsan N, Lee S H, Lee J J, Bahk J D, Kang K Y, Lee B H.2009. Chilling stress-induced proteomic changes in rice roots.J Plant Physiol, 166(1): 1-11. |
| 97 | Lee S K, Kim B G, Kwon T R, Jeong M J, Park S R, Lee J W, Byun M O, Kwon H B, Matthews B F, Hong C B, Park S C.2011. Overexpression of the mitogen-activated protein kinase geneOsMAPK33 enhances sensitivity to salt stress in rice(Oryza sativa L.). J Biosci, 36(1): 139-151. |
| 98 | Lehmann S, Funck D, Szabados L, Rentsch D.2010. Proline metabolism and transport in plant development.Amino Acids, 39(4): 949-962. |
| 99 | Li C H, Wang G, Zhao J L, Zhang L Q, Ai L F, Han Y F, Sun D Y, Zhang S W, Sun Y.2014. The receptor-like kinase SIT1 mediates salt sensitivity by activating MAPK3/6 and regulating ethylene homeostasis in rice.Plant Cell, 26(6): 2538-2553. |
| 100 | Li H W, Zang B S, Deng X W, Wang X P.2011. Overexpression of the trehalose-6-phosphate synthase geneOsTPS1 enhances abiotic stress tolerance in rice. Planta, 234(5): 1007-1018. |
| 101 | Li M, Guo L J, Guo C M, Wang L J, Chen L.2016. Over-expression of a DUF1644 protein gene,SIDP361, enhances tolerance to salt stress in transgenic rice. J Plant Biol, 59(1): 62-73. |
| 102 | Liang C Z, Wang Y Q, Zhu Y N, Tang J Y, Hu B, Liu L C, Ou S J, Wu H K, Sun X H, Chu J F, Chu C C.2014. OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice.Proc Natl Acad Sci USA, 111: 10013-10018. |
| 103 | Liao Y D, Lin K H, Chen C C, Chang C M.2016. Oryza sativa protein phosphatase 1a (OsPP1a) involved in salt stress tolerance in transgenic rice.Mol Breeding, 36: 22. |
| 104 | Lin C C, Kao C H.2001. Cell wall peroxidase activity, hydrogen peroxidase level and NaCl-inhibited roots of rice seedlings.Plant Soil, 230: 135-143. |
| 105 | Lin H X, Liang Z W, Sasaki T, Yano M.2003. Fine mapping and characterization of quantitative trait lociHd4 and Hd5 controlling heading date in rice. Breeding Sci, 53(1): 51-59. |
| 106 | Lin K H, Pu S F.2010. Tissue- and genotype-specific ascorbate peroxidase expression in sweet potato in response to salt stress.Biol Plant, 54(4): 664-670. |
| 107 | Liu C W, Chang T S, Hsu Y K, Wang A Z, Yen H C, Wu Y P, Wang C S, Lai C C.2014. Comparative proteomic analysis of early salt stress-responsive proteins in roots and leaves of rice.Proteomics, 14: 1759-1775. |
| 108 | Liu D F, Chen X J, Liu J Q, Ye J C, Guo Z J.2012. The rice ERF transcription factorOsERF922 negatively regulates resistance to Magnaporthe oryzae and salt tolerance. J Exp Bot, 63(10): 3899-3911. |
| 109 | Liu J P, Zhu J K.1998. A calcium sensor homolog required for plant salt tolerance.Science, 280: 1943-1945. |
| 110 | Liu S K, Cheng Y X, Zhang X X, Guan Q J, Nishiuchi S, Hase K, Takano T.2007. Expression of an NADP-malic enzyme gene in rice (Oryza sativa L.) is induced by environmental stresses; over-expression of the gene in Arabidopsis confers salt and osmotic stress tolerance. Plant Mol Biol, 64: 49-58. |
| 111 | Lutts S, Kinet J M, Bouharmont J.1995. Changes in plant response to NaCl during development of rice (Oryza sativa L.) varieties differing in salinity resistance. J Exp Bot, 46(12): 1843-1852. |
| 112 | Mahajan S, Tuteja N.2005. Cold, salinity and drought stresses: An overview.Arch Biochem Biophys, 444(2): 139-158. |
| 113 | Majee M, Maitra S, Dastidar K G, Pattnaik S, Chatterjee A, Hait N C, Das K P, Majumder A L.2004. A novel salt-tolerantL-myo-inositol-1-phosphate synthase from Porteresia coarctata(Roxb.) Tateoka, a halophytic wild rice. J Biol Chem, 279: 28539-28552. |
| 114 | Mallikarjuna G, Mallikarjuna K, Reddy M K, Kaul T.2011. Expression ofOsDREB2A transcription factor confers enhanced dehydration and salt stress tolerance in rice(Oryza sativa L.). Biotechnol Lett, 33(8): 1689-1697. |
| 115 | Mantri N, Patade V, Penna S, Ford R, Pang E.2012. Abiotic stress responses in plants: Present and future. In: Ahmad P, Prasad M N V. Abiotic Stress Responses in Plants: Metabolism, Productivity and Sustainability. New York: Springer: 1-19. |
| 116 | Matysik J, Alia A, Bhalu B, Mohanty P.2002. Molecular mechanisms of quenching of reactive oxygen species by proline under stress in plants.Curr Sci, 82(5): 525-532. |
| 117 | McLoughlin F, Galvan-Ampudia C S, Julkowska M M, Caarls L, van der Does D, Lauriere C, Munnik T, Haring M A, Testerink C.2012. The Snf1-related protein kinases SnRK2.4 and SnRK2.10 are involved in maintenance of root system architecture during salt stress.Plant J, 72(3): 436-449. |
| 118 | Mishra S, Singh B, Panda K, Singh B P, Singh N, Misra P, Rai V, Singh N K.2016. Association of SNP haplotypes of HKT family genes with salt tolerance in Indian wild rice germplasm.Rice, 9(1): 15. |
| 119 | Mittal D, Sharma N, Sharma V, Sopory S K, Sanan-Mishra N.2016. Role of microRNAs in rice plant under salt stress.Ann Appl Biol, 168(1): 2-18. |
| 120 | Miyamoto N, Steudle E, Hirasawa T, Lafitte R.2001. Hydraulic conductivity of rice roots.J Exp Bot, 52: 1835-1846. |
| 121 | Miyamoto T, Ochiai K, Nonoue Y, Matsubara K, Yano M, Matoh T.2015. Expression level of the sodium transporter geneOsHKT2;1 determines sodium accumulation of rice cultivars under potassium- deficient conditions. Soil Sci Plant Nutr, 61(3): 481-492. |
| 122 | Mohanty A, Kathuria H, Ferjani A, Sakamoto A, Mohanty P, Murata N, Tyagi A K.2002. Transgenics of an elite indica rice variety Pusa Basmati 1 harboring thecodA gene are highly tolerant to salt stress. Theor Appl Genet, 106(1): 51-57. |
| 123 | Molla K A, Debnath A B, Ganie S A, Mondal T K.2015. Identification and analysis of novel salt responsive candidate gene based SSRs (cgSSRs) from rice (Oryza sativa L.). BMC Plant Biol, 15: 122. |
| 124 | Momayezi M R, Zaharah A R, Hanafi M M, Mohd Razi I.2009. Agronomic characteristics and proline accumulation of Iranian rice genotypes at early seedling stage under sodium salts stress.Malays J Soil Sci, 13: 59-75. |
| 125 | Moradi F, Ismail A M.2007. Responses of photosynthesis, chlorophyll fluorescence and ROS-scavenging systems to salt stress during seedling and reproductive stages in rice.Ann Bot, 99(6): 1161-1173. |
| 126 | Motohashi T, Nagamiya K, Prodhan S H, Nakao K, Shishido T, Yamamoto Y, Moriwaki T, Hattori E, Asada M, Morishima H, Hirose S, Ozawa K, Takabe T, Takabe T, Komamine A.2010. Production of salt stress tolerant rice by overexpression of the catalase gene,katE, derived from Escherichia coli. Asia Pac J Mol Biol Biotechnol, 18(1): 37-41. |
| 127 | Mukhopadhyay A, Vij S, Tyagi A K.2004. Overexpression of a zinc-finger protein gene from rice confers tolerance to cold, dehydration, and salt stress in transgenic tobacco.Proc Natl Acad Sci USA, 101(16): 6309-6314. |
| 128 | Munns R.2002. Comparative physiology of salt and water stress.Plant Cell Environ, 25(2): 239-250. |
| 129 | Munns R, Tester M.2008. Mechanisms of salinity tolerance.Annu Rev Plant Biol, 59: 651-681. |
| 130 | Nakashima K, Tran L S, van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi-Shinozaki K.2007. Functional analysis of a NAC-type transcription factorOsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J, 51(4): 617-630. |
| 131 | Nam M H, Huh S M, Kim K M, Park W W, Seo J B, Cho K, Kim D Y, Kim B G, Yoon I I.2012. Comparative proteomic analysis of early salt stress-responsive proteins in roots of SnRK2 transgenic rice.Prot Sci, 10: 25. |
| 132 | Nam M H, Bang E, Kwon T Y, Kim Y, Kim E H, Cho K, Park W J, Kim B G, Yoon I S.2015. Metabolite profiling of diverse rice germplasm and identification of conserved metabolic markers of rice roots in response to long-term mild salinity stress.Int J Mol Sci, 16(9): 21959-21974. |
| 133 | Nath M, Yadav S, Sahoo R K, Passricha N, Tuteja R, Tuteja N.2016. PDH45 transgenic rice maintain cell viability through lower accumulation of Na+, ROS and calcium homeostasis in roots under salinity stress.J Plant Physiol, 191: 1-11. |
| 134 | Nounjan N, Nghia P T, Theerakulpisut P.2012. Exogenous proline and trehalose promote recovery of rice seedlings from salt-stress and differentially modulate antioxidant enzymes and expression of related genes.J Plant Physiol, 169(6): 596-604. |
| 135 | Ochiai K, Matoh T.2002. Characterization of the Na+ delivery from roots to shoots in rice under saline stress: Excessive salt enhances apoplastic transport in rice plants.Soil Sci Plant Nutr, 48(3): 371-378. |
| 136 | Ouyang S Q, Liu Y F, Liu P, Lei G, He S J, Ma B, Zhang W K, Zhang J S, Chen S Y.2010. Receptor-like kinase OsSIK1 improves drought and salt stress tolerance in rice (Oryza sativa) plants. Plant J, 62(2): 316-329. |
| 137 | Pandit A, Rai V, Bal S, Sinha S, Kumar V, Chauhan M, Gautam R K, Singh R, Sharma P C, Singh A K, Gaikwad K, Sharma T R, Mhoapatra T, Singh N K.2010. Combining QTL mapping and transcriptome profiling of bulked RILs for identification of functional polymorphism for salt tolerance genes in rice (Oryza sativa L.). Mol Genet Genom, 284(2): 121-136. |
| 138 | Pareek A, Sopory S K, Bohnert H J, Govindjee.2010. Abiotic Stress Adaptation in Plants: Physiological, Molecular and Genomic Foundation. Berlin: Springer. |
| 139 | Parida A K, Das A B.2005. Salt tolerance and salinity effect on plants: A review.Ecotox Environ Saf, 60(3): 324-349. |
| 140 | Pearson G A, Bernstein L.1959. Salinity effects at several growth stages of rice.Agron J, 51(11): 654-657. |
| 141 | Peterson C A.1988. Exoderrnal casparian bands: Their significance for ion uptake by toots.Physiol Plant, 72(1): 204-208. |
| 142 | Pitman M G.1984. Transport across the root and shoot/root interactions. In: Staples R C, Toenniessen G H. Salinity Tolerance in Plants: Strategies for Crop Improvement. New York, USA: Wiley: 93-123. |
| 143 | Platten J D, Egdane J A, Ismail A M.2013. Salinity tolerance, Na+ exclusion and allele mining ofHKT1;5 in Oryza sativa and O. glaberrima: Many sources, many genes, one mechanism? BMC Plant Biol, 13: 32. |
| 144 | Qiu D Y, Xiao J, Xie W B, Liu H B, Li X H, Xiong L Z, Wang S P.2008. Rice gene network inferred from expression profiling of plants overexpressingOsWRKY13, a positive regulator of disease resistance. Mol Plant, 1(3): 538-551. |
| 145 | Quintero F J, Ohta M, Shi H Z, Zhu J K, Pardo J M.2002. Reconstitution in yeast of theArabidopsis SOS signaling pathway for Na+ homeostasis. Proc Natl Acad Sci USA, 99(13): 9061-9066. |
| 146 | Rahman M A, Thomson M J, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail A M.2016. Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization.Ann Bot, 117(6): 1083-1097. |
| 147 | Rahman M S, Matsumuro T, Miyake H, Takeoka Y.2001. Effects of salinity stress on the seminal root tip ultrastructures of rice seedlings.Plant Prod Sci, 4(2): 103-111. |
| 148 | Rajendran K, Tester M, Roy S J.2009. Quantifying the three main components of salinity tolerance in cereals.Plant Cell Environ, 32(3): 237-249. |
| 149 | Redillas M C F R, Jeong J S, Kim Y S, Jung H, Bang S W, Choi Y D, Ha S H, Reuzeau C, Kim J K.2012. The overexpression ofOsNAC9 alters the root architecture of rice plants enhancing drought resistance and grain yield under field conditions. Plant Biotechnol J, 10(7): 792-805. |
| 150 | Ren Z H, Gao J P, Li L G, Cai X L, Huang W, Chao D Y, Zhu M Z, Wang Z Y, Luan S, Lin H X.2005. A rice quantitative trait locus for salt tolerance encodes a sodium transporter.Nat Genet, 37: 1141-1146. |
| 151 | Reynolds M P, Ortiz-Monasterio J I, Mc Nab A.2001. Application of Physiology in Wheat Breeding. Mexico: International Maize and Wheat Improvement Center. |
| 152 | Roy S J, Negrao S, Tester M.2014. Salt resistant crop plants.Curr Opin Biotechnol, 26: 115-124. |
| 153 | Royuela M, Gonzalez A, Gonzalez E M, Arrese-Igor C, Aparicio-Tejo P M, Gonzalez-Murua C.2000. Physiological consequences of continuous, sublethal imazethapyr supply to pea plants.J Plant Physiol, 157(3): 345-354. |
| 154 | Saeng-ngam S, Takpirom W, Buaboocha T, Chadchawan S.2012. The role of the OsCam1-1 salt stress sensor in ABA accumulation and salt tolerance in rice.J Plant Biol, 55(3): 198-208. |
| 155 | Sahoo R K, Ansari M W, Tuteja R, Tuteja N.2014. OsSUV3 transgenic rice maintains higher endogenous levels of plant hormones that mitigates adverse effects of salinity and sustains crop productivity.Rice, 7(1): 17. |
| 156 | Sahu B B, Shaw B P.2009. Salt-inducible isoform of plasma membrane H+-ATPase gene in rice remains costitutively expressed in natural halophyte,Suaeda maritima. J Plant Physiol, 166(10): 1077-1089. |
| 157 | Saijo Y, Hata S, Kyozuka J, Shimamoto K, Izui K.2000. Overexpression of a single Ca2+ dependent protein kinase confers both cold and salt/drought tolerance on rice plants.Plant J, 23(3): 319-327. |
| 158 | Sakurai J, Ishikawa F, Yamaguchi T, Uemura M, Maeshima M.2005. Identification of 33 rice aquaporin genes and analysis of their expression and function.Plant Cell Physiol, 46(9): 1568-1577. |
| 159 | Sakurai J, Ahamed A, Murai M, Maeshima M, Uemura M.2008. Tissue and cell-specific localization of rice aquaporins and their water transport activities.Plant Cell Physiol, 49(1): 30-39. |
| 160 | Schmidt R, Mieulet D, Hubberten H M, Obata T, Hoefgen R, Fernie A R, Fisahn J, Segundo B S, Guiderdoni E, Schippers J H M, Mueller-Roeber B.2013. Salt-responsive ERF1 regulates reactive oxygen species-dependent signaling during the initial response to salt stress in rice.Plant Cell, 25(6): 2115-2131. |
| 161 | Sexcion F S H, Egdane J A, Ismail A M, Dionisio-Sese M L.2009. Morpho-physiologicall traits associated with tolerance of salinity during seedling stage in rice (Oryza sativa L.). Phil J Crop Sci, 34(2): 27-37. |
| 162 | Shabala S, Munns R.2012. Salinity stress: Physiological constraints and adaptive mechanisms. In: Shabala S. Plant Stress Physiology. United Kingdom & USA: CAB International: 59-93. |
| 163 | Sharma I, Ching E, Saini S, Bhardwaj R, Pati P K.2013. Exogenous application of brassinosteroid offers tolerance to salinity by altering stress responses in rice variety Pusa Basmati-1.Plant Physiol Biochem, 69: 17-26. |
| 164 | Sharma N, Panchal S, Sanan-Mishra N.2015. Protocol for artificial microRNA mediated over-expression ofmiR820 in indica rice. Am J Plant Sci, 6(12): 1951-1961. |
| 165 | Shen Y, Shen L, Shen Z X, Jing W, Ge H L, Zhao J Z, Zhang W H.2015. The potassium transporter OsHAK21 functions in the maintenance of ion homeostasis and tolerance to salt stress in rice.Plant Cell Environ, 38(12): 2766-2779. |
| 166 | Shi L, Guo M M, Ye N H, Liu Y G, Liu R, Xia Y J, Cui S X, Zhang J H.2015. Reduced ABA accumulation in the root system is caused by ABA exudation in upland rice (Oryza sativa L. var. Gaoshan 1) and this enhanced drought adaptation. Plant Cell Physiol, 56(5): 951-964. |
| 167 | Singh R, Singh Y, Xalaxo S, Verulkar S, Yadav N, Singh S, Singh N, Prasad K S N, Kondayya K, Ramana Rao P V, Rani M G, Anuradha T, Suraynarayana Y, Sharma P C, Krishnamurthy S L, Sharma S K, Dwivedi J L, Singh A K, Singh P K, Nilanjay, Singh N K, Kumar R, Chetia S K, Ahmad T, Rai M, Perraju P, Pande A, Singh D N, Mandal N P, Reddy J N, Singh O N, Katara J L, Marandi B, Swain P, Sarkar R K, Singh D P, Mohapatra T, Padmawathi G, Ram T, Kathiresan R M, Paramsivam K, Nadarajan S, Thirumeni S, Nagarajan M, Singh A K, Vikram P, Kumar A, Septiningshih E, Singh U S, Ismail A M, Mackill D, Singh N K,.2016. From QTL to variety-harnessing the benefits of QTLs for drought, flood and salt tolerance in mega rice varieties of India through a multi-institutional network.Plant Sci, 242: 278-287. |
| 168 | Singh R K, Mishra B, Singh K N.2004. Salt tolerant rice varieties and their role in reclamation programme in Uttar Pradesh. Indian Farming: 6-10. |
| 169 | Sirault X R R, James R A, Furbank R T.2009. A new screening method for osmotic component of salinity tolerance in cereals using infrared thermography.Funct Plant Biol, 36(11): 970-977. |
| 170 | Son Y S, Lm C H, Kim D W, Bahk J D.2013. OsRab11 and OsGAP1 are essential for the vesicle trafficking of the vacuolar H+-ATPase OsVHA-a1 under high salinity conditions.Plant Sci, 198: 58-71. |
| 171 | Song S Y, Chen Y, Chen J, Dai X Y, Zhang W H.2011. Physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress.Planta, 234(2): 331-345. |
| 172 | Staal M, Maathuis F J M, Elzenga J T M, Overbeek J H M, Prins H B A.1991. Na+/H+-antiport activity in tonoplast vesicles from roots of the salt-tolerant plantago maritima and the salt-sensitive plantago media.Physiol Plant, 82(2): 179-184. |
| 173 | Su J, Hirji R, Zhang L, He C K, Selvaraj G, Wu R.2006. Evaluation of the stress-inducible production of choline oxidase in transgenic rice as a strategy for producing the stress-protectant glycinebetaine.J Exp Bot, 57(5): 1129-1135. |
| 174 | Suzuki K, Yamaji N, Costa A, Okuma E, Kobayashi N I, Kashiwagi T, Katsuhara M, Wang C, Tanoi K, Murata Y, Schroeder J I, Ma J F, Horie T.2016. OsHKT1;4-mediated Na+ transport in stems contributes to Na+ exclusion from leaf blades of rice at the reproductive growth stage upon salt stress.BMC Plant, 16: 22. |
| 175 | Takagi H, Tamiru M, Abe A, Yoshida K, Uemura A, Yaegashi H, Obara T, Oikawa K, Utsushi H, Kanzaki E, Mitsuoka C, Natsume S, Kosugi S, Kanzaki H, Matsumura H, Urasaki N, Kamoun S, Terauchi R.2015. MutMap accelerates breeding of a salt-tolerant rice cultivar.Nat Biotechnol, 33: 445-449. |
| 176 | Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguch-Shinozaki K, Nakashima K.2010. The abiotic stress- responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice.Mol Genet Genom, 284(3): 173-183. |
| 177 | Thompson G A, Goggin F L.2006. Transcriptomics and functional genomics of plant defence induction by phloem-feeding insects.J Exp Bot, 57(4): 755-766. |
| 178 | Thomson M J, de Ocampo M, Egdane J, Rahman M A, Sajise A G, Adorada D L, Tumimbang-Raiz E, Blumward E, Seraj Z I, Singh R K, Gregorio G B, Ismail A M.2010. Characterizing theSaltol quantitative trait locus for salinity tolerance in rice. Rice, 3(2): 148-160. |
| 179 | Tiwari S, Krishnamurthy S L, Kumar V, Singh B, Rao A R, Amitha Mithra S V, Rai V, Singh A K, Singh N K.2016. Mapping QTLs for salt tolerance in rice (Oryza sativa L.) by bulked segregant analysis of recombinant inbred lines using 50K SNP chip. PLoS One, 11(4): e0153610. |
| 180 | Todaka D, Nakashima K, Shinozaki K, Yamaguchi-Shinozaki K.2012. Towards understanding transcriptional regulatory networks in abiotic stress responses and tolerance in rice.Rice, 5(1): 6. |
| 181 | Tu Y, Jiang A M, Gan L, Hossain M, Zhang J M, Peng B, Xiong Y G, Song Z J, Cai D T, Xu W F, Zhang J H, He Y C.2014. Genome duplication improves rice root resistance to salt stress.Rice, 7(1): 15. |
| 182 | Tuteja N, Gill S S, Tuteja R.2011. Omics and Plant Stress Tolerance. USA: Bentham Science Publisher: 39-64. |
| 183 | Verslues P E, Agarwal M, Katiyar-Agarwal S, Zhu J H, Zhu J K.2006. Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status.Plant J, 45(4): 523-539. |
| 184 | Wang F Z, Jing W, Zhang W H.2014. The mitogen-activated protein kinase cascade MKK1-MPK4 mediates salt signaling in rice.Plant Sci, 227: 181-189. |
| 185 | Wang H, Zhang M S, Guo R, Shi D C, Liu B, Lin X Y, Yang C W.2012. Effects of salt stress on ion balance and nitrogen metabolism of old and young leaves in rice (Oryza sativa L.). BMC Plant Biol, 12: 194. |
| 186 | Wang Q Y, Guan Y C, Wu Y R, Chen H L, Chen F, Chu C C.2008. Overexpression of a riceOsDREB1F gene increases salt, drought, and low temperature tolerance in both Arabidopsis and rice. Plant Mol Biol, 67(6): 589-602. |
| 187 | Wang R, Jing W, Xiao L Y, Jin Y K, Shen L K, Zhang W H.2015. The rice high-affinity potassium transporter1;1 is involved in salt tolerance and regulated by an MYB-type transcription factor.Plant Physiol, 168(3): 1076-1090. |
| 188 | Wang W S, Zhao X Q, Li M, Huang L Y, Xu J L, Zhang F, Cui Y R, Fu B Y, Li Z K.2016. Complex molecular mechanisms underlying seedling salt tolerance in rice revealed by comparative transcriptome and metabolomic profiling. J Exp Bot, 67(1): 405-419. |
| 189 | Watanabe H, Saigusa M, Morita S.2006. Identification of casparian bands in the mesocotyl and lower internodes of rice (Oryza sativa L.) seedling using fluorescence microscopy. Plant Prod Sci, 9(4): 390-394. |
| 190 | Wutipraditkul N, Boonkomrat S, Buaboocha T.2011. Cloning and characterization of catalases from rice,Oryza sativa L. Biosci Biotechnol Biochem, 75(10): 1900-1906. |
| 191 | Xia K F, Wang R, Ou X J, Fang Z M, Tian C G, Duan J, Wang Y Q, Zhang M Y.2012. OsTIR1 andOsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS One, 7: e30039. |
| 192 | Xiang Y, Huang Y M, Xiong L Z.2007. Characterization of stress-responsive CIPK genes in rice for stress tolerance improvement.Plant Physiol, 144(3): 1416-1428. |
| 193 | Xiang Y, Tang N, Du H, Ye H Y, Xiong L Z.2008. Characterization ofOsbZIP23 as a key player of the basic leucine zipper transcription factor family for conferring abscisic acid sensitivity and salinity and drought tolerance in rice. Plant Physiol, 148(4): 1938-1952. |
| 194 | Xiong L Z, Yang Y N.2003. Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid-inducible mitogen-activated protein kinase.Plant Cell, 15(3): 745-759. |
| 195 | Xu G Y, Cui Y C, Li M J, Wang M L, Yu Y, Zhang B, Huang L F, Xia X J.2013. OsMSR2, a novel rice calmodulin-like gene, confers enhanced salt tolerance in rice (Oryza sativa L.). Aust J Crop Sci, 7(3): 368. |
| 196 | Xu J W, Huang X, Lan H X, Zhang H S, Huang J.2016. Rearrangement of nitrogen metabolism in rice (Oryza sativa L.) under salt stress. Plant Signal Behav, 11(3): e1138194. |
| 197 | Yaish M W, El-Kereamy A, Zhu T, Beatty P H, Good A G, Bi Y M, Rothstein S J.2010. The APETALA-2-like transcription factorOsAP2-39 controls key interactions between abscisic acid and gibberellin in rice. PLoS Genet, 6(9): e1001098. |
| 198 | Yang A, Dai X, Zhang W H.2012. A R2R3-type MYB gene,OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot, 63(7): 2541-2556. |
| 199 | Yang C W, Zhang T Y, Wang H, Zhao N, Liu B.2012. Heritable alteration in salt-tolerance in rice induced by introgression from wild rice (Zizania latifolia). Rice, 5(1): 36. |
| 200 | Yang T Y, Zhang S, Hu Y B, Wu F C, Hu Q D, Chen G, Cai J, Wu T, Moran N, Yu L, Xu G H.2014. The role of a potassium transporter OsHAK5 in potassium acquisition and transport from roots to shoots in rice at low potassium supply levels.Plant Physiol, 166(2): 945-959. |
| 201 | Yasmin F, Biswas S, Jewel G M N A, Elias S M, Seraj Z I.2015. Constitutive overexpression of the plasma membrane Na+/H+ antiporter for conferring salinity tolerance in rice.Plant Tiss Cult Biotechnol, 25(2): 257-272. |
| 202 | Ye H Y, Du H, Tang N, Li X H, Xiong L Z.2009. Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice.Plant Mol Biol, 71(3): 291-305. |
| 203 | Yeo A R, Flowers T J.1986. Salinity resistance in rice (Oryza sativa L.) and a pyramiding approach to breeding for saline soils. Aust J Plant Physiol, 13(1): 161-173. |
| 204 | Yeo A R, Yeo M E, Ftowers T J.1987. The contribution of an apoplastic pathway to sodium uptake by rice roots in saline conditions. J Exp Bot, 38: 1141-1153. |
| 205 | Yeo A R, Yeo M E, Flowers S A, Flowers T J.1990. Screening of rice (Oryza sativa) genotypes for physiological characters contributing to salinity resistance, and their relationship to overall performance. Theor Appl Genet, 79(3): 377-384. |
| 206 | Yoshiba Y, Kiyosue T, Nakashima K, Yamaguchi-Shinozaki K, Shinozak K.1997. Regulation of levels of proline as an osmolyte in plants under water stress.Plant Cell Physiol, 38(10): 1095-1102. |
| 207 | Yoshida T, Mogami J, Yamaguchi-Shinozaki K.2014. ABA- dependent and ABA-independent signaling in response to osmotic stress in plants.Curr Opin Plant Biol, 21: 133-139. |
| 208 | Zang B S, Li H W, Li W J, Deng X W, Wang X P.2011. Analysis of trehalose-6-phosphate synthase (TPS) gene family suggests the formation of TPS complexes in rice.Plant Mol Biol, 76(6): 507-522. |
| 209 | Zeng L H, Shannon M C.2000. Salinity effects on seedling growth and yield components of rice.Crop Sci, 40(4): 996-1003. |
| 210 | Zhao B T, Ge L F, Liang R Q, Li W, Ruan K C, Lin H X, Jin Y X.2009. Members ofmiR-169 family are induced by high salinity and transiently inhibit the NFYA transcription factor. BMC Mol Biol, 10: 29. |
| 211 | Zhao J F, Wu C X, Yuan S J, Yin L, Sun W, Zhao Q L, Zhao B H, Li X Y.2013. Kinase activity of OsBRI1 is essential for brassinosteroids to regulate rice growth and development.Plant Sci, 199/200: 113-120. |
| 212 | Zhang L, Tian L H, Zhao J F, Song Y, Zhang C J, Guo Y.2009. Identification of an apoplastic protein involved in the initial phase of salt stress response in rice root by two-dimensional electrophoresis.Plant Physiol, 149(2): 916-928. |
| 213 | Zheng X N, Chen B, Lu G J, Han B.2009. Overexpression of a NAC transcription factor enhances rice drought and salt tolerance.Biochem Biophys Res Commun, 379(4): 985-989. |
| 214 | Zhou J L, Wang X F, Jiao Y L, Qin Y H, Liu X G, He K, Chen C, Ma L G, Wang J, Xiong L Z, Zhang Q F, Fan L M, Deng X W.2007. Global genome expression analysis of rice in response to drought and high-salinity stresses in shoot, flag leaf, and panicle.Plant Mol Biol, 63(5): 591-608. |
| 215 | Zhou Y, Yang P, Cui F L, Zhang F T, Luo X D, Xie J K.2016. Transcriptome analysis of salt stress responsiveness in the seedlings of Dongxiang wild rice (Oryza rufipogon Griff.). PLoS One, 11(1): e0146242. |
| 216 | Zou M J, Guan Y C, Ren H B, Zhang F, Chen F.2008. A bZIP transcription factor,OsABI5, is involved in rice fertility and stress tolerance. Plant Mol Biol, 66(6): 675-683. |
| 217 | Zou Y, Liu X, Wang Q, Chen Y, Liu C, Qiu Y, Zhang W.2014. OsRPK1, a novel leucine-rich repeat receptor-like kinase, negatively regulates polar auxin transport and root development in rice.Biochim Biophys Acta, 1840(6): 1676-1685. |
| 218 | (Managing Editor: Li Guan) |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | XIA Xiaodong, ZHANG Xiaobo, WANG Zhonghao, CHENG Benyi, Sun Huifeng, XU Xia, GONG Junyi, YANG Shihua, WU Jianli, SHI Yongfeng, XU Rugen. Mapping and Functional Analysis of LE Gene in a Lethal Etiolated Rice Mutant at Seedling Stage [J]. Rice Science, 2023, 30(6): 13-. |
| [7] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [8] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [9] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [10] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [11] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [12] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [15] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||