
Rice Science ›› 2017, Vol. 24 ›› Issue (3): 163-172.DOI: 10.1016/j.rsci.2016.10.001
• Orginal Article • Previous Articles Next Articles
Jannoey Panatda( ), Channei Duangdao, Kotcharerk Jate, Pongprasert Weerathep, Nomura Mika
), Channei Duangdao, Kotcharerk Jate, Pongprasert Weerathep, Nomura Mika
Received:2016-08-01
Accepted:2016-10-17
Online:2017-05-28
Published:2017-03-03
Jannoey Panatda, Channei Duangdao, Kotcharerk Jate, Pongprasert Weerathep, Nomura Mika. Expression Analysis of Genes Related to Rice Resistance Against Brown Planthopper, Nilaparvata lugens[J]. Rice Science, 2017, 24(3): 163-172.
Add to citation manager EndNote|Ris|BibTeX
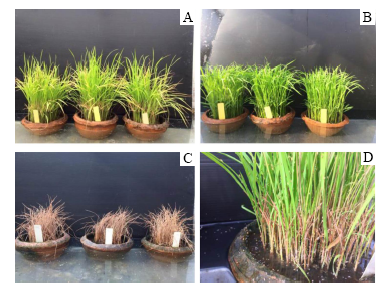

Fig. 1. Hopperburn symptom of rice plants.( A, PSL2 (resistant variety) attacked by brown planthopper (BPH) for 14 d; B, Control without BPH inoculation; C, TN1 (susceptible variety) in the cage condition; D, BPH nymph behaviors settled on the rice seedling stem.)
| Function | Gene | Protein name | Forward primer | Reverse primer |
|---|---|---|---|---|
| Pathogenesis related-gene | PR1a | SCP-like extracellular protein | GGAAGTACGGCGAGAACATC | GGTCGTACCACTGCTTCTCC |
| PR2 | Glucanase | TGCTATGTTCGACGAGAACG | GTTGAACAGCCCAAAGTGCT | |
| PR3 | Chitinase | CTCACCACGAACATCCTCAC | GTCGCAGTAGCGCTTGTAGA | |
| PR4 | Chitinase | CCGTCTTCTCCAAGATCGAC | TGGTAGTCGACGATGAGGTG | |
| PR6 | Protease inhibitor | GTGCATCTTGCATGCTTTGT | TTTTCCTCATGGTCCACACA | |
| PR15 | Oxalate oxidase4 | AGGCCTTCTGCAACAAGATG | CACTCCTTCACCTCGTCCAT | |
| PR9 | Lignin-forming peroxidase | GATGGTGAAGATGGGGAACA | ACGGAGCACTTGATCCTGAC | |
| PR10a | Ribonuclease | GCCGAATACGCCTAAGATGA | ACATTTCTGCGGCTCTCATT | |
| PR13 | Thionin | CCGCTTCTGTACCAAGGAAG | ATGTGTGAAGCCCCTTATGC | |
| PRpha | Phenylalanine ammonia lyase | AGAATCACCGAGTGCAGGTC | GCCGGTCAGGTACTTTGTTC | |
| Lignin synthesis | CHS | Chalcone synthase | AGGGAAGAATGGGGACTGAT | TGCCTCGAACTAGCATTCCT |
| CHI | Chalcone isomerase | AGCTCCTGAAGGCGGAAT | GATTTTCACGCGGACACC | |
| C4H | Cinnamate-4-hydroxylase | CTCGTCCAGAGCTTCGACCT | GGATCTGGTTGCTGAACTGG | |
| Signaling pathway | LOX | Lipoxygenase | GGAGGTTCAACGAGAGGATG | GATCCTTGTTCCGGCAGTC |
| AOS | Allene oxide synthase | GGAGGAAGCTGCTGCAATAC | TGCTTGTTGTCAACGCTAGG | |
| AXR | Auxin responsive protein | TGTTCCATGGGAGATGTTCA | CCAATTGCATCTGAGCCTTT | |
| ACO | ACC oxidase | CCTACCCGAGGTTCGTGTT | CTCCTTGGCCTCGAACTTGT | |
| Oxidative stress | SOD | Superoxide dismutase | CGATCCTGATGATCTTGGAAA | CAGCCTTGAAGTCCGATGAT |
| CAT | Catalase | AGGCAAGATCGTTTTCTCCA | GCGACCAGTAGGAGATCCAG | |
| TRX | Thioredoxin1 | GACAGCTGCATGGAGTTCCT | CCCTGATGAAGAGGAAGGTG | |
| GST | Glutathione transferase | GTAGGCTCGCCGAGTACG | CAGCTGCTGCCCACTCTG | |
| Control | Actin | ATCACCATCGGAGCAGAAAG | AAAAGATGGCTGGAAGAGCA |
Table 1 Primers for qRT-PCR amplification designed by primer3 software
| Function | Gene | Protein name | Forward primer | Reverse primer |
|---|---|---|---|---|
| Pathogenesis related-gene | PR1a | SCP-like extracellular protein | GGAAGTACGGCGAGAACATC | GGTCGTACCACTGCTTCTCC |
| PR2 | Glucanase | TGCTATGTTCGACGAGAACG | GTTGAACAGCCCAAAGTGCT | |
| PR3 | Chitinase | CTCACCACGAACATCCTCAC | GTCGCAGTAGCGCTTGTAGA | |
| PR4 | Chitinase | CCGTCTTCTCCAAGATCGAC | TGGTAGTCGACGATGAGGTG | |
| PR6 | Protease inhibitor | GTGCATCTTGCATGCTTTGT | TTTTCCTCATGGTCCACACA | |
| PR15 | Oxalate oxidase4 | AGGCCTTCTGCAACAAGATG | CACTCCTTCACCTCGTCCAT | |
| PR9 | Lignin-forming peroxidase | GATGGTGAAGATGGGGAACA | ACGGAGCACTTGATCCTGAC | |
| PR10a | Ribonuclease | GCCGAATACGCCTAAGATGA | ACATTTCTGCGGCTCTCATT | |
| PR13 | Thionin | CCGCTTCTGTACCAAGGAAG | ATGTGTGAAGCCCCTTATGC | |
| PRpha | Phenylalanine ammonia lyase | AGAATCACCGAGTGCAGGTC | GCCGGTCAGGTACTTTGTTC | |
| Lignin synthesis | CHS | Chalcone synthase | AGGGAAGAATGGGGACTGAT | TGCCTCGAACTAGCATTCCT |
| CHI | Chalcone isomerase | AGCTCCTGAAGGCGGAAT | GATTTTCACGCGGACACC | |
| C4H | Cinnamate-4-hydroxylase | CTCGTCCAGAGCTTCGACCT | GGATCTGGTTGCTGAACTGG | |
| Signaling pathway | LOX | Lipoxygenase | GGAGGTTCAACGAGAGGATG | GATCCTTGTTCCGGCAGTC |
| AOS | Allene oxide synthase | GGAGGAAGCTGCTGCAATAC | TGCTTGTTGTCAACGCTAGG | |
| AXR | Auxin responsive protein | TGTTCCATGGGAGATGTTCA | CCAATTGCATCTGAGCCTTT | |
| ACO | ACC oxidase | CCTACCCGAGGTTCGTGTT | CTCCTTGGCCTCGAACTTGT | |
| Oxidative stress | SOD | Superoxide dismutase | CGATCCTGATGATCTTGGAAA | CAGCCTTGAAGTCCGATGAT |
| CAT | Catalase | AGGCAAGATCGTTTTCTCCA | GCGACCAGTAGGAGATCCAG | |
| TRX | Thioredoxin1 | GACAGCTGCATGGAGTTCCT | CCCTGATGAAGAGGAAGGTG | |
| GST | Glutathione transferase | GTAGGCTCGCCGAGTACG | CAGCTGCTGCCCACTCTG | |
| Control | Actin | ATCACCATCGGAGCAGAAAG | AAAAGATGGCTGGAAGAGCA |
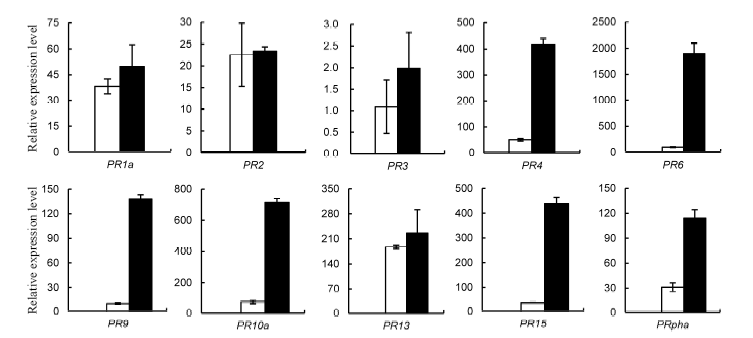
Fig. 2. Differential expression of pathogenesis-related genes in rice response to brown planthopper (BPH) (Mean ± SD, n = 3).( Data were analyzed at the 0.05 level. The blank columns are control rice plants and the black ones are rice plants infested with BPH.)
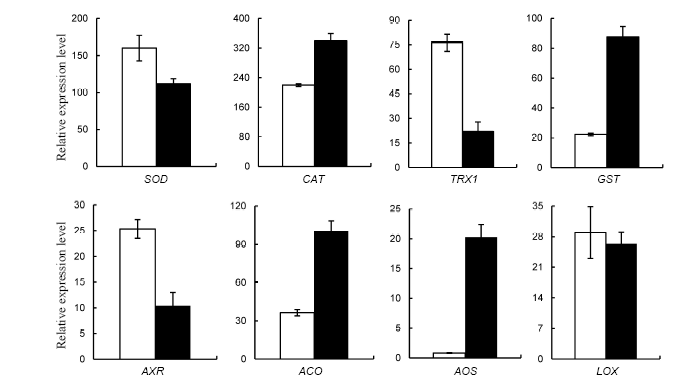
Fig. 3. Differential expression of antioxidant enzyme-related genes (SOD, CAT, TXR1 and GST) and signaling compound biosynthesis-related genes (AXR, ACO, AOS and LOX) in rice response to brown planthopper (BPH) (Mean ± SD, n = 3).( Data were analyzed at the 0.05 level. The blank columns are control rice plants and the black ones are rice plants infested with BPH.)
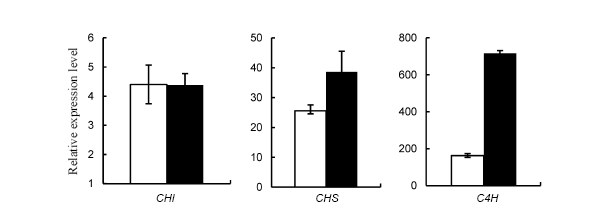
Fig. 4. Differential expression of lignin biosynthesis-related genes in rice response to brown planthopper (BPH) (Mean ± SD, n = 3).( Data were analyzed at the 0.05 level. The blank columns are control rice plants and the black ones are rice plants infested with BPH.)
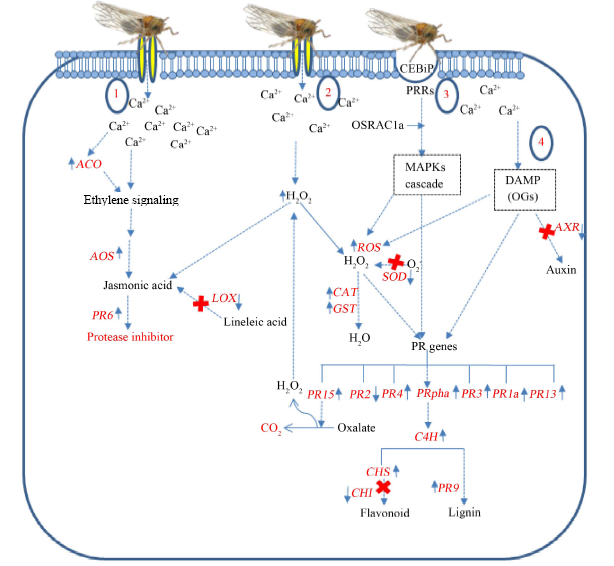
Fig. 5. Schematic representation of brown planthopper (BPH) triggered rice defense mechanism (Cheng et al, 2013 with modifications). (BPH induced the expression of wound-response genes in rice represented by following mechanism: (1) Wound-induced Ca2+ fluxes occur in rice as a second messenger. The rice CBL-interacting protein kinases (CIPK14 and CIPK15) are Ca2+-related-protein sensor and involve in induced accumulation of signaling molecule (ethylene and jasmonic acid). (2) H2O2 is emerging as signal molecules after wounding and subsequently Ca2+ accumulates. H2O2 activates jasmonic acid, and controls the expression of defense related-gene and metabolite biosynthesis. (3) Rice chitin elicitor-binding protein (CEBiP) is essential for chitin recognition for chitin-trigged immunity induction. Plants recognize pathogen by cell surface localized pattern-recognition receptor (PRRs). PRRs also interact with OSRAC1 for regulation of the final step of signaling pathway including the production of reactive oxygen species (ROS), lignin, PR proteins and mitogen-activated protein kinase (MAPK) cascade. (4) Damage-associate molecular pattern (DAMP) molecules are release from wound tissue, and activate the plant innate immunity. Oligogalactoronides (OGs), one of the DAMP molecules, involve in plant response to wounding. OGs can induce the protease inhibitor, ROS, nitrix oxide, phytoalexin, PR2 and PR3 accumulation, and callose deposition. However, OGs also regulate auxin-antagonistic activity.)
| 1 | Akiyama T, Jin S, Yoshida M, Hoshino T, Opassiri R, Cairns J R K.2009. Expression of an endo-(1,3;1,4)-beta-glucanase in response to wounding, methyl jasmonate, abscisic acid and ethephon in rice seedlings.J Plant Physiol, 166(16): 1814-1825. |
| 2 | Alagar M, Suresh S, Samiyappan R, Saravanakuma D.2007. Reaction of resistant and susceptible rice genotypes against brown planthopper (Nilaparvata lugens). Phytoparasitica, 35: 346. |
| 3 | Anand A, Zhou T, Trick H N, Gill B S, Bockus W W, Muthukrishnan S.2003. Greenhouse and field testing of transgenic wheat plants stably expressing genes for thaumatin-like protein, chitinase and glucanase againstFusarium graminearum. J Exp Bot, 54: 1101-1111. |
| 4 | Cheng X Y, Zhu L L, He G C.2013. Towards understanding of molecular interactions between rice and the brown planthopper.Mol Plant, 6(3): 621-634. |
| 5 | Du B, Zhang W L, Liu B F, Hu J, Wei Z, Shi Z Y, He R F, Zhu L L, Chen R Z, Han B, He G C.2009. Identification and characterization ofBph14, a gene conferring resistance to brown planthopper in rice. Proc Natl Acad Sci USA, 106(52): 22163-22168. |
| 6 | Duan C X, Yu J J, Bai J Y, Zhu Z D, Wang X M.2014. Induced defense responses in rice plants against small brown planthopper infestation.Crop J, 2(1): 55-62. |
| 7 | Ebrahim S, Usha K, Singh B.2011. Pathogenesis related (PR) proteins in plant defense mechanism: Science against microbial pathogens: Communicating current research and technological advances.Sci Against Microb Pathog, 2: 1043-1054. |
| 8 | Gill S S, Tuteja N.2010. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants.Plant Physiol Biochem, 48(12): 909-930. |
| 9 | Hao Z N, Wang L P, Tao R X.2009. Expression patterns of defence genes and antioxidant defence responses in a rice variety that is resistant to leaf blast but susceptible to neck blast.Physiol Mol Plant Pathol, 74(2): 167-174. |
| 10 | Hao Z N, Wang L P, He Y P, Liang J G, Tao R X.2011. Expression of defense genes and activities of antioxidant enzymes in rice resistance to rice stripe virus and small brown planthopper.Plant Physiol Biochem, 49(7): 744-751. |
| 11 | Hu T Z.2014. A glutathione S-transferase confers herbicide tolerance in rice.Crop Breeding Appl Biotechnol, 14(2): 76-81. |
| 12 | International Rice Research Institute (IRRI). 1996. Standard Evaluation System for Rice. Manila, the Philippines. |
| 13 | Iwamoto M, Baba-Kasai A, Kiyota S, Hara N, Takano M.2010. ACO1, a gene for aminocyclopropane-1-carboxylateoxidase: Effects on internode elongation at the heading stage in rice.Plant Cell Environ, 33(5): 805-815. |
| 14 | Jach G, Görnhardt B, Mundy J, Logemann J, Pinsdorf E, Leah R, Schell J, Maas C.1995. Enhanced quantitative resistance against fungal disease by combinatorial expression of different barley antifungal proteins in transgenic tobacco.Plant J, 8(1): 97-109. |
| 15 | Jamal F, Pandey P K, Singh D, Ahmed W.2015. A Kunitz-type serine protease inhibitor fromButea monosperma seed and its influence on developmental physiology of Helicoverpa armigera. Proc Biochem, 50(2): 311-316. |
| 16 | Jannoey P, Weerathep P, Lumyong S, Roytrakul S, Nomura M.2015. Comparative proteomic analysis of two rice cultivars (Oryza sativa L.) contrasting in brown planthopper (BPH) stress resistance. Plant Omic J, 8(2): 96-105. |
| 17 | Jongedijk E, Tigelaar H, van Roekel J S C, Bres-Vloemans S A, Dekker I, van den Elzen P J M, Cornelissen B J C, Melchers L S.1995. Synergistic activity of chitinases and β-1,3-glucanases enhances fungal resistance in transgenic tomato plants.Euphytica, 85(1): 173-180. |
| 18 | Jwa N S, Agrawal G K, Rakwal R, Park C H, Agrawal V P.2011. Molecular cloning and characterization of a novel jasmonate inducible pathogenesis-related class 10 protein gene,JIOsPR10, from rice(Oryza sativa L.) seedling leaves. Biochem Biophys Res Commun, 286(5): 973-983. |
| 19 | Kuwar S S, Pauchet Y, Vogel H, Hecke D G.2015. Adaptive regulation of digestive serine proteases in the larval midgut ofHelicoverpa armigera in response to a plant protease inhibitor. Insect Biochem Mol Biol, 59: 18-29. |
| 20 | Li X C, Liao Y Y, Leung D W M, Wang H Y, Chen B L, Peng X X, Liu E E.2015. Divergent biochemical and enzymatic properties of oxalate oxidase isoforms encoded by four similar genes in rice.Phytochem, 118: 216-223. |
| 21 | Melchers L S, Stuiver M H.2000. Novel genes for disease-resistance breeding.Curr Opin Plant Biol, 3(2): 147-152. |
| 22 | Molla K A, Karmakar S, Chanda P K, Ghosh S, Sarkar S N, Datta S K, Datta K.2013. Riceoxalate oxidase gene driven by green tissue-specific promoter increases tolerance to sheath blight pathogen(Rhizoctonia solani) in transgenic rice. Mol Plant Pathol, 14(9): 910-922. |
| 23 | Mott G A, Middleton M A, Desveaux D, Guttman D S.2014. Peptides and small molecules of the plant-pathogen apoplastic arena.Front Plant Sci, 5: 677. |
| 24 | Muthamilarasan M, Prasad M.2013. Plant innate immunity: An updated insight into defense mechanism.J Biosci, 38(2): 433-449. |
| 25 | Nishizawa Y, Saruta M, Nakazono K, Nishio Z, Soma M, Yoshida T, Nakajima E, Hibi T.2003. Characterization of transgenic rice plants over-expressing the stress-inducible β-1,3-glucanase geneGns1. Plant Mol Biol, 51(1): 143-152. |
| 26 | Opassiri R, Maneesan J, Akiyama T, Pomthong B, Jin S, Kimura A, Ketudat Cairns J R.2010. Rice Os4BGlu12 is a wound-induced β-glucosidase that hydrolyzes cell wall-β-glucan-derived oligosaccharides and glycosides.Plant Sci, 179(3): 273-278. |
| 27 | Ponciano G, Yoshikawa M, Lee J L, Ronald P C, Whalen M C.2006. Pathogenesis-related gene expression in rice is correlated with developmentally controlledXa21-mediated resistance against Xanthomonas oryzae pv. Oryzae. Physiol Mol Plant Pathol, 69: 131-139. |
| 28 | Qiu Z B, Guo J L, Zhu A J, Zhang L, Zhang M M.2014. Exogenous jasmonic acid can enhance tolerance of wheat seedlings to salt stress.Ecotox Environ Safe, 104: 202-208. |
| 29 | Rezaei M K, Shobbar Z S, Shahbazi M, Abedini R, Zare S.2013. GlutathioneS-transferase (GST) family in barley: Identification of members, enzyme activity, and gene expression pattern. J Plant Physiol, 170(14): 1277-1284. |
| 30 | Senthil-Nathan S, Kalaivani K, Choi M Y, Paik C H.2009. Effects of jasmonic acid-induced resistance in rice on the plant brownhopper,Nilaparvata lugens Stål (Homoptera: Delphacidae). Pest Biochem Physiol, 95(2): 77-84. |
| 31 | Sridevi G, Parameswari C, Sabapathi N, Raghupathy V, Veluthambi K.2008. Combined expression of chitinase and β-1,3-glucanase genes in indica rice (Oryza sativa L.) enhances resistance against Rhizoctonia solani. Plant Sci, 175(3): 283-290. |
| 32 | Stael S, Kmiecik P, Willems P, van der Kelen K, Coll N S, Teige M, van Breusegem F.2015. Plant innate immunity-sunny side up?Trends Plant Sci, 20(1): 1-11. |
| 33 | Sun H Y, Li Y, Feng S Q, Zou W H, Guo K, Fan C F, Si S L, Peng L C.2013. Analysis of five rice 4-coumarate: Coenzyme A ligase enzyme activity and stress response for potential roles in lignin and flavonoid biosynthesis in rice.Biochem Biophys Res Commun, 430(3): 1151-1156. |
| 34 | Ting A S Y, Chai J Y.2015. Chitinase and β-1,3-glucanase activities ofTrichoderma harzianum in response towards pathogenic and non-pathogenic isolates: Early indications of compatibility in consortium. Biocatal Agric Biotechnol, 4(1): 109-113. |
| 35 | Turner J G, Ellis C, Devoto A.2002. The jasmonate signal pathway.Plant Cell, 14(S1): S153-S164. |
| 36 | van Loon L C, van Strien E A.1999. The families of pathogenesis-related proteins, their activities, and comparative analysis of PR-1 type proteins.Physiol Mol Plant Pathol, 55(2): 85-97. |
| 37 | Vieira Dos Santos C, Rey P.2006. Plant thioredoxins are key actors in the oxidative stress response.Trends Plannt Sci, 11(7): 329-334. |
| 38 | Watkins D, Nuruddin M D, Hosur M, Tcherbi-Narteh A, Jeelan S.2015. Extraction and characterization of lignin from different biomass resources.J Mater Res Technol, 4(1): 26-32. |
| 39 | Weng Q M, Huang Z, Wang X L, Zhu L L, He G C.2003. In situ localization of proteinase inhibitor mRNA in rice plant challenged by brown planthopper.Chin Sci Bull, 48(10): 979-982. |
| 40 | Wu S W, Wang H W, Yang Z D, Kong L R.2014. Expression comparisons of pathogenesis-related (PR) genes in wheat in response to infection/infestation by Fusarium, Yellow dwarf virus (YDV) aphid-transmitted and hessian fly. J Integr Agric, 13(5): 926-936. |
| 41 | Xayphakatsa K, Tsukiyama T, Inouye K, Okumoto Y, Nakazaki T, Tanisaka T.2008. Gene cloning, expression, purification and characterization of rice (Oryza sativa L.) class II chitinase CHT11. Enzyme Microb Technol, 43(1): 19-24. |
| 42 | Xi Y, Pan P L, Ye Y X, Yu B, Xu H J, Zhang C X.2015. Chitinase-like gene family in the brown planthopper,Nilaparvata lugens. Insect Mol Biol, 24(1): 29-40. |
| 43 | Xu J, Xing X J, Tian Y S, Peng R H, Xue Y, Zhao W, Yao Q H.2015. TransgenicArabidopsis plants expressing tomato glutathione S-transferase showed enhanced resistance to salt and drought stress. PLoS One, 10(9): e0136960. |
| 44 | You L X, Wang P, Kong C H.2011. The levels of jasmonic acid and salicylic acid in a rice-barnyardgrass coexistence system and their relation to rice allelochemicals.Biochem Syst Ecol, 39: 491-497. |
| 45 | Zhang F T, Zhu L L, He G C.2004. Differential gene expression in response to brown planthopper feeding in rice.J Plant Physiol, 161(1): 53-62. |
| 46 | Zhang X Y, Nie Z H, Wang W J, Leung D W M, Xu D G, Chen B L, Chen Z, Zeng L X, Liu E E.2013. Relationship between disease resistance and rice oxalate oxidases in transgenic rice.PLoS One, 8(10): e78348. |
| 47 | Zheng W J, Ma L, Zhao J M, Li Z Q, Sun F Y, Lu X C.2013. Comparative transcriptome analysis of two rice varieties in response to rice stripe virus and small brown planthoppers during early interaction.PLoS One, 8(12): e82126. |
| 48 | Zhu Q, Maher E A, Masoud S, Dixon R A, Lamb C J.1994. Enhanced protection against fungal attack by constitutive co-expression of chitinase and glucanase genes in transgenic tobacco.Biol Technol, 12: 807-812. |
| 49 | (Managing Editor: Li Guan) |
| [1] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [2] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [3] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [4] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [5] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [6] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [7] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [8] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [9] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [10] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [14] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [15] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||