
Rice Science ›› 2019, Vol. 26 ›› Issue (3): 178-188.DOI: 10.1016/j.rsci.2018.09.001
• Research Papers • Previous Articles Next Articles
F. Aala Jr Wilson( ), B. Gregorio Glenn
), B. Gregorio Glenn
Received:2018-05-08
Accepted:2018-09-28
Online:2019-05-28
Published:2019-01-25
F. Aala Jr Wilson, B. Gregorio Glenn. Morphological and Molecular Characterization of Novel Salt-Tolerant Rice Germplasms from the Philippines and Bangladesh[J]. Rice Science, 2019, 26(3): 178-188.
Add to citation manager EndNote|Ris|BibTeX
| Score | Description | Remark |
|---|---|---|
| 1 | Normal growth, only the old leaves show white tips | Highly tolerant |
| 3 | Near normal growth, only leaf tips burn, few older leaves become whitish partially | Tolerant |
| 5 | Growth severely retarded; most old leaves severely injured, few young leaves are elongated | Moderately tolerant |
| 7 | Complete cessation of growth; most leaves are dried; only few young leaves still green | Susceptible |
| 9 | Almost all plants dead or dying | Highly susceptible |
Table 1 Modified standard evaluation score for assessing the visual symptoms of salt injury at the seedling stage.
| Score | Description | Remark |
|---|---|---|
| 1 | Normal growth, only the old leaves show white tips | Highly tolerant |
| 3 | Near normal growth, only leaf tips burn, few older leaves become whitish partially | Tolerant |
| 5 | Growth severely retarded; most old leaves severely injured, few young leaves are elongated | Moderately tolerant |
| 7 | Complete cessation of growth; most leaves are dried; only few young leaves still green | Susceptible |
| 9 | Almost all plants dead or dying | Highly susceptible |
| CN | ACCNO | Designation | Source | SES | SL (cm) | SFW (g) | SDW (g) | RSLD (%) | RSWC (%) | RSDR (%) | Na-K ratio |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 6 | 36968 | Akundo | BD | 3.00 d | 60.67 a | 1.708 a | 0.413 a | 27.200 e-o | 75.85 a | 41.769 a-d | 0.39 no |
| 7 | 37179 | Kuplod | BD | 3.00 d | 57.17 a-c | 1.178 bc | 0.286 b-g | 28.243 e-n | 75.65 a | 35.072 a-f | 0.76 k-o |
| 24 | 64772 | Chondoni | BD | 5.67 a | 33.67 o-s | 0.213 tu | 0.100 s-u | 48.205 a-c | 50.41 l | 61.740 a-b | 4.41 bc |
| 52 | 66765 | Asha | BD | 3.67 cd | 46.67 f-k | 0.612 g-s | 0.204 e-p | 35.180 a-k | 66.69 a-i | 41.040 a-d | 2.42 d-i |
| 85 | 66798 | Jakor | BD | 3.00 d | 47.50 f-j | 0.666 e-r | 0.191 h-r | 40.625 a-f | 71.25 a-f | 55.478 a-c | 1.93 e-l |
| 102 | 66816 | Moisdol | BD | 3.00 d | 49.67 d-h | 0.703 e-q | 0.204 f-p | 19.677 i-o | 70.58 a-g | 28.370 a-f | 0.65 l-o |
| 117 | 77204 | Naptasa | BD | 3.00 d | 45.00 g-l | 0.465 n-u | 0.127 o-u | 37.063 a-j | 72.21 a-d | 58.874 a-c | 1.11 h-o |
| 119 | 77213 | Aguni Kartiksail | BD | 3.00 d | 52.83 b-f | 0.827 c-n | 0.230 c-m | 31.236 d-m | 71.91 a-e | 46.429 a-f | 0.96 j-o |
| 127 | 77226 | Chakol | BD | 3.00 d | 51.17 c-g | 0.807 d-o | 0.235 c-k | 28.438 e-n | 70.09 a-g | 40.756 a-d | 1.22 h-o |
| 131 | 77230 | Chorua Kartiksail | BD | 3.00 d | 49.67 d-h | 0.859 c-m | 0.242 c-k | 38.046 a-i | 68.68 a-h | 57.210 a-c | 0.92 j-o |
| 154 | 77273 | Lal Moni | BD | 3.00 d | 49.67 d-h | 0.912 c-j | 0.242 c-k | 33.184 c-k | 73.71 ab | 45.543 a-d | 1.29 e-o |
| 155 | 77276 | Modhu Bash | BD | 5.00 ab | 43.17 h-m | 0.914 c-i | 0.290 b-e | 37.590 a-j | 62.75 d-k | 18.692 a-g | 2.66 de |
| 157 | 77278 | Mridom | BD | 4.33 bc | 41.17 j-n | 0.416 p-u | 0.166 j-u | 38.710 a-h | 57.63 i-l | 35.371 a-e | 1.86 e-m |
| 174 | 77297 | Roa | BD | 3.00 d | 40.50 j-o | 0.623 f-s | 0.206 e-o | 27.246 f-o | 66.74 a-i | 4.341 f-h | 1.45 e-o |
| 176 | 77299 | Sada Rupa | BD | 3.67 cd | 26.67 s-w | 0.384 p-u | 0.125 p-u | 34.959 c-k | 67.01 a-i | 32.855 a-f | 2.10 e-k |
| 190 | 69809 | Pilet Lubang | PH | 3.67 cd | 26.17 t-x | 0.319 r-u | 0.116 q-u | 34.583 b-k | 60.02 h-l | 28.454 a-g | 1.14 h-o |
| 193 | 69812 | Binutiti | PH | 3.67 cd | 35.17 n-r | 0.296 s-u | 0.106 r-u | 47.250 a-d | 64.48 b-j | 56.198 a-c | 2.65 d-f |
| 198 | 69818 | Londran | PH | 3.00 d | 49.33 e-h | 0.532 k-u | 0.186 h-s | 27.451 f-o | 61.06 g-k | 15.175 b-g | 1.60 e-o |
| 199 | 69819 | Macapuno | PH | 3.67 cd | 38.00 l-q | 0.428 p-u | 0.158 k-u | 43.284 a-e | 62.26 e-k | 37.101 a-f | 1.30 e-o |
| 206 | 69826 | Reppeng | PH | 3.67 cd | 35.83 n-r | 0.529 l-u | 0.153 k-u | 36.578 a-k | 67.13 a-i | 37.207 b-g | 2.60 d-g |
| 207 | 69827 | Ricorico | PH | 3.00 d | 37.67 m-q | 0.630 e-s | 0.194 h-r | 52.017 a | 69.04 a-h | 54.786 a-c | 1.81 e-m |
| 272 | 19502 | Ikogan | PH | 3.00 d | 49.00 f-h | 0.866 c-l | 0.258 c-i | 0.000 p | 70.21 a-g | -2.520 e-h | 0.91 j-o |
| 317 | 24230 | Betalga | PH | 3.00 d | 56.33 a-e | 1.011 b-f | 0.287 b-f | 12.208 n-p | 71.58 a-f | 11.408 c-g | 0.43 no |
| 318 | 24231 | Maranao | PH | 3.00 d | 51.17 c-g | 0.824 d-o | 0.231 c-l | 10.234 op | 71.43 a-f | 5.075 d-h | 0.97 j-o |
| 319 | 24232 | Mori | PH | 3.00 d | 59.50 ab | 0.881 c-k | 0.246 c-j | 16.589 l-p | 71.68 a-f | 28.461 a-f | 0.65 l-o |
| 327 | 24484 | IR5494 | PH | 3.67 cd | 21.33 wx | 0.186 u | 0.083 u | 36.318 a-k | 54.87 j-l | 18.627 a-g | 3.68 cd |
| 365 | 40546 | IR4493-5-5-3 | PH | 3.00 d | 23.67 v-x | 0.308 s-u | 0.117 p-u | 39.574 a-g | 62.05 f-k | 33.206 a-f | 5.13 ab |
| 366 | 44265 | AC | PH | 3.00 d | 56.50 a-d | 0.981 b-h | 0.262 b-i | 22.955 g-o | 73.31 a-c | 29.884 a-f | 0.33 o |
| 369 | 44273 | Ampipit | PH | 3.00 d | 45.33 g-k | 0.486 m-u | 0.166 j-u | 36.300 a-k | 64.60 b-j | 47.357 a-d | 2.48 d-h |
| 370 | 44274 | Anangka | PH | 3.00 d | 57.33 a-c | 0.967 b-h | 0.263 b-i | 15.892 m-p | 72.57 a-d | 15.054 b-g | 0.50 m-o |
| 391 | 44323 | Binalasang | PH | 3.00 d | 48.50 f-i | 1.024 b-e | 0.274 b-h | 34.459 c-k | 73.26 a-c | 32.486 a-f | 1.43 e-o |
| 403 | 44380 | Casibon | PH | 3.00 d | 29.33 r-v | 0.403 p-u | 0.142 l-u | 22.467 h-o | 64.49 b-j | -34.700 h | 1.28 f-o |
| 413 | 44409 | Dukab | PH | 3.00 d | 46.50 f-k | 0.750 d-p | 0.226 c-m | 28.46 2e-n | 70.07 a-g | 33.627 a-f | 2.20 e-j |
| 420 | 44421 | Gallano | PH | 3.00 d | 43.00 h-m | 0.377 q-u | 0.141 m-u | 32.813 c-k | 60.10 h-l | 59.012 a-c | 1.26 g-o |
| 469 | 44651 | Motit Motit (469) | PH | 3.00 d | 43.17 h-m | 0.553 i-u | 0.201 g-q | 27.042 f-o | 63.50 c-k | 37.318 a-e | 1.77 e-n |
| 470 | 44652 | Motit Motit (470) | PH | 3.00 d | 46.00 f-k | 0.629 f-s | 0.189 h-s | 37.273 a-k | 69.76 a-h | 44.585 a-d | 1.36 e-o |
| 509 | 44753 | Sarujao | PH | 3.00 d | 59.50 ab | 1.095 b-d | 0.299 b-d | 25.934 f-o | 72.36 a-d | 33.383 a-f | 1.53 e-o |
| 529 | 47241 | Kalagnon | PH | 3.00 d | 25.00 u-x | 0.505 m-u | 0.156 k-u | 35.897 a-k | 69.00 a-h | -31.921 gh | 1.64 e-o |
| 553 | 52867 | Binangahon | PH | 3.67 cd | 33.17 p-t | 0.279 s-u | 0.109 q-u | 50.620 ab | 60.98 g-k | 65.687 a | 2.22 e-j |
| 554 | 52869 | Binog | PH | 3.00 d | 40.00 k-p | 0.432 p-u | 0.135 n-u | 36.340 a-k | 66.88 a-i | 41.954 a-d | 1.07 i-o |
| 573 | 52944 | Pilit | PH | 3.00 d | 41.83 i-n | 0.609 i-s | 0.175 i-t | 40.521 a-f | 70.73 a-g | 48.678 a-d | 1.05 i-o |
| 577 | 52960 | Tjeremas | PH | 3.00 d | 25.67 u-x | 0.585 i-t | 0.177 i-t | 53.474 a | 69.46 a-h | 34.724 a-f | 1.20 h-o |
| 596 | 52988 | Sinan-Pablo | PH | 3.00 d | 50.00 d-h | 0.997 b-g | 0.307 bc | 31.663 d-m | 69.24 a-h | 35.393 a-e | 1.41 e-o |
| 597 | 52989 | Veronica | PH | 3.00 d | 45.50 g-k | 0.725 e-q | 0.221 d-n | 28.534 e-n | 69.59 a-h | 31.579 a-f | 1.82 e-m |
| FL478 (Check) | PH | 3.00 d | 31.00 q-u | 1.303 b | 0.340 ab | 32.364 d-l | 73.93 a | 44.954 a-d | 1.75 e-n | ||
| IR29 (Check) | PH | 9.00 e | 19.50 x | 0.205 tu | 0.087 tu | 48.684 a-c | 54.04 kl | 61.218 a-c | 6.08 a | ||
| Normal | 1.00 a | 63.49 a | 1.550 a | 0.322 a | - | 78.09 a | - | 0.13 a | |||
| Salinized | 3.24 b | 42.72 b | 0.674 b | 0.201 b | - | 67.28 b | - | 1.73 b | |||
Table 2 Physio-morphometric data for the 44 salt-tolerant accessions under salinized conditions (EC = 18 dSm-1).
| CN | ACCNO | Designation | Source | SES | SL (cm) | SFW (g) | SDW (g) | RSLD (%) | RSWC (%) | RSDR (%) | Na-K ratio |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 6 | 36968 | Akundo | BD | 3.00 d | 60.67 a | 1.708 a | 0.413 a | 27.200 e-o | 75.85 a | 41.769 a-d | 0.39 no |
| 7 | 37179 | Kuplod | BD | 3.00 d | 57.17 a-c | 1.178 bc | 0.286 b-g | 28.243 e-n | 75.65 a | 35.072 a-f | 0.76 k-o |
| 24 | 64772 | Chondoni | BD | 5.67 a | 33.67 o-s | 0.213 tu | 0.100 s-u | 48.205 a-c | 50.41 l | 61.740 a-b | 4.41 bc |
| 52 | 66765 | Asha | BD | 3.67 cd | 46.67 f-k | 0.612 g-s | 0.204 e-p | 35.180 a-k | 66.69 a-i | 41.040 a-d | 2.42 d-i |
| 85 | 66798 | Jakor | BD | 3.00 d | 47.50 f-j | 0.666 e-r | 0.191 h-r | 40.625 a-f | 71.25 a-f | 55.478 a-c | 1.93 e-l |
| 102 | 66816 | Moisdol | BD | 3.00 d | 49.67 d-h | 0.703 e-q | 0.204 f-p | 19.677 i-o | 70.58 a-g | 28.370 a-f | 0.65 l-o |
| 117 | 77204 | Naptasa | BD | 3.00 d | 45.00 g-l | 0.465 n-u | 0.127 o-u | 37.063 a-j | 72.21 a-d | 58.874 a-c | 1.11 h-o |
| 119 | 77213 | Aguni Kartiksail | BD | 3.00 d | 52.83 b-f | 0.827 c-n | 0.230 c-m | 31.236 d-m | 71.91 a-e | 46.429 a-f | 0.96 j-o |
| 127 | 77226 | Chakol | BD | 3.00 d | 51.17 c-g | 0.807 d-o | 0.235 c-k | 28.438 e-n | 70.09 a-g | 40.756 a-d | 1.22 h-o |
| 131 | 77230 | Chorua Kartiksail | BD | 3.00 d | 49.67 d-h | 0.859 c-m | 0.242 c-k | 38.046 a-i | 68.68 a-h | 57.210 a-c | 0.92 j-o |
| 154 | 77273 | Lal Moni | BD | 3.00 d | 49.67 d-h | 0.912 c-j | 0.242 c-k | 33.184 c-k | 73.71 ab | 45.543 a-d | 1.29 e-o |
| 155 | 77276 | Modhu Bash | BD | 5.00 ab | 43.17 h-m | 0.914 c-i | 0.290 b-e | 37.590 a-j | 62.75 d-k | 18.692 a-g | 2.66 de |
| 157 | 77278 | Mridom | BD | 4.33 bc | 41.17 j-n | 0.416 p-u | 0.166 j-u | 38.710 a-h | 57.63 i-l | 35.371 a-e | 1.86 e-m |
| 174 | 77297 | Roa | BD | 3.00 d | 40.50 j-o | 0.623 f-s | 0.206 e-o | 27.246 f-o | 66.74 a-i | 4.341 f-h | 1.45 e-o |
| 176 | 77299 | Sada Rupa | BD | 3.67 cd | 26.67 s-w | 0.384 p-u | 0.125 p-u | 34.959 c-k | 67.01 a-i | 32.855 a-f | 2.10 e-k |
| 190 | 69809 | Pilet Lubang | PH | 3.67 cd | 26.17 t-x | 0.319 r-u | 0.116 q-u | 34.583 b-k | 60.02 h-l | 28.454 a-g | 1.14 h-o |
| 193 | 69812 | Binutiti | PH | 3.67 cd | 35.17 n-r | 0.296 s-u | 0.106 r-u | 47.250 a-d | 64.48 b-j | 56.198 a-c | 2.65 d-f |
| 198 | 69818 | Londran | PH | 3.00 d | 49.33 e-h | 0.532 k-u | 0.186 h-s | 27.451 f-o | 61.06 g-k | 15.175 b-g | 1.60 e-o |
| 199 | 69819 | Macapuno | PH | 3.67 cd | 38.00 l-q | 0.428 p-u | 0.158 k-u | 43.284 a-e | 62.26 e-k | 37.101 a-f | 1.30 e-o |
| 206 | 69826 | Reppeng | PH | 3.67 cd | 35.83 n-r | 0.529 l-u | 0.153 k-u | 36.578 a-k | 67.13 a-i | 37.207 b-g | 2.60 d-g |
| 207 | 69827 | Ricorico | PH | 3.00 d | 37.67 m-q | 0.630 e-s | 0.194 h-r | 52.017 a | 69.04 a-h | 54.786 a-c | 1.81 e-m |
| 272 | 19502 | Ikogan | PH | 3.00 d | 49.00 f-h | 0.866 c-l | 0.258 c-i | 0.000 p | 70.21 a-g | -2.520 e-h | 0.91 j-o |
| 317 | 24230 | Betalga | PH | 3.00 d | 56.33 a-e | 1.011 b-f | 0.287 b-f | 12.208 n-p | 71.58 a-f | 11.408 c-g | 0.43 no |
| 318 | 24231 | Maranao | PH | 3.00 d | 51.17 c-g | 0.824 d-o | 0.231 c-l | 10.234 op | 71.43 a-f | 5.075 d-h | 0.97 j-o |
| 319 | 24232 | Mori | PH | 3.00 d | 59.50 ab | 0.881 c-k | 0.246 c-j | 16.589 l-p | 71.68 a-f | 28.461 a-f | 0.65 l-o |
| 327 | 24484 | IR5494 | PH | 3.67 cd | 21.33 wx | 0.186 u | 0.083 u | 36.318 a-k | 54.87 j-l | 18.627 a-g | 3.68 cd |
| 365 | 40546 | IR4493-5-5-3 | PH | 3.00 d | 23.67 v-x | 0.308 s-u | 0.117 p-u | 39.574 a-g | 62.05 f-k | 33.206 a-f | 5.13 ab |
| 366 | 44265 | AC | PH | 3.00 d | 56.50 a-d | 0.981 b-h | 0.262 b-i | 22.955 g-o | 73.31 a-c | 29.884 a-f | 0.33 o |
| 369 | 44273 | Ampipit | PH | 3.00 d | 45.33 g-k | 0.486 m-u | 0.166 j-u | 36.300 a-k | 64.60 b-j | 47.357 a-d | 2.48 d-h |
| 370 | 44274 | Anangka | PH | 3.00 d | 57.33 a-c | 0.967 b-h | 0.263 b-i | 15.892 m-p | 72.57 a-d | 15.054 b-g | 0.50 m-o |
| 391 | 44323 | Binalasang | PH | 3.00 d | 48.50 f-i | 1.024 b-e | 0.274 b-h | 34.459 c-k | 73.26 a-c | 32.486 a-f | 1.43 e-o |
| 403 | 44380 | Casibon | PH | 3.00 d | 29.33 r-v | 0.403 p-u | 0.142 l-u | 22.467 h-o | 64.49 b-j | -34.700 h | 1.28 f-o |
| 413 | 44409 | Dukab | PH | 3.00 d | 46.50 f-k | 0.750 d-p | 0.226 c-m | 28.46 2e-n | 70.07 a-g | 33.627 a-f | 2.20 e-j |
| 420 | 44421 | Gallano | PH | 3.00 d | 43.00 h-m | 0.377 q-u | 0.141 m-u | 32.813 c-k | 60.10 h-l | 59.012 a-c | 1.26 g-o |
| 469 | 44651 | Motit Motit (469) | PH | 3.00 d | 43.17 h-m | 0.553 i-u | 0.201 g-q | 27.042 f-o | 63.50 c-k | 37.318 a-e | 1.77 e-n |
| 470 | 44652 | Motit Motit (470) | PH | 3.00 d | 46.00 f-k | 0.629 f-s | 0.189 h-s | 37.273 a-k | 69.76 a-h | 44.585 a-d | 1.36 e-o |
| 509 | 44753 | Sarujao | PH | 3.00 d | 59.50 ab | 1.095 b-d | 0.299 b-d | 25.934 f-o | 72.36 a-d | 33.383 a-f | 1.53 e-o |
| 529 | 47241 | Kalagnon | PH | 3.00 d | 25.00 u-x | 0.505 m-u | 0.156 k-u | 35.897 a-k | 69.00 a-h | -31.921 gh | 1.64 e-o |
| 553 | 52867 | Binangahon | PH | 3.67 cd | 33.17 p-t | 0.279 s-u | 0.109 q-u | 50.620 ab | 60.98 g-k | 65.687 a | 2.22 e-j |
| 554 | 52869 | Binog | PH | 3.00 d | 40.00 k-p | 0.432 p-u | 0.135 n-u | 36.340 a-k | 66.88 a-i | 41.954 a-d | 1.07 i-o |
| 573 | 52944 | Pilit | PH | 3.00 d | 41.83 i-n | 0.609 i-s | 0.175 i-t | 40.521 a-f | 70.73 a-g | 48.678 a-d | 1.05 i-o |
| 577 | 52960 | Tjeremas | PH | 3.00 d | 25.67 u-x | 0.585 i-t | 0.177 i-t | 53.474 a | 69.46 a-h | 34.724 a-f | 1.20 h-o |
| 596 | 52988 | Sinan-Pablo | PH | 3.00 d | 50.00 d-h | 0.997 b-g | 0.307 bc | 31.663 d-m | 69.24 a-h | 35.393 a-e | 1.41 e-o |
| 597 | 52989 | Veronica | PH | 3.00 d | 45.50 g-k | 0.725 e-q | 0.221 d-n | 28.534 e-n | 69.59 a-h | 31.579 a-f | 1.82 e-m |
| FL478 (Check) | PH | 3.00 d | 31.00 q-u | 1.303 b | 0.340 ab | 32.364 d-l | 73.93 a | 44.954 a-d | 1.75 e-n | ||
| IR29 (Check) | PH | 9.00 e | 19.50 x | 0.205 tu | 0.087 tu | 48.684 a-c | 54.04 kl | 61.218 a-c | 6.08 a | ||
| Normal | 1.00 a | 63.49 a | 1.550 a | 0.322 a | - | 78.09 a | - | 0.13 a | |||
| Salinized | 3.24 b | 42.72 b | 0.674 b | 0.201 b | - | 67.28 b | - | 1.73 b | |||
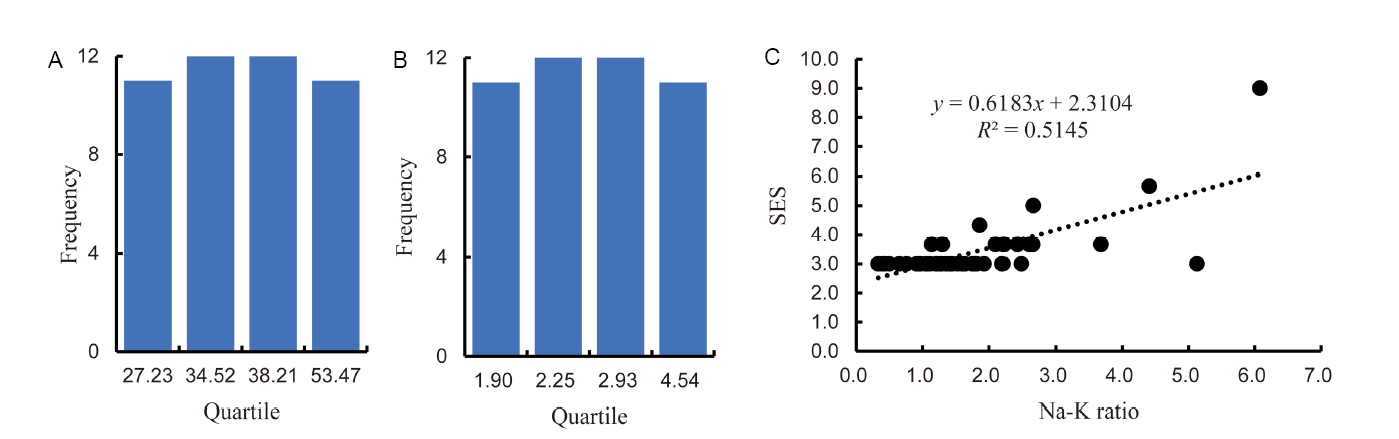
Fig. 1. General distributions of selected accessions for relative shoot length difference (A), relative shoot dry weight reduction (B) and scatter plot of Na-K ratio to standard evaluation score (SES) (C).
| Source of error | SES | RSWC | RSDR | SDW | SFW | SL | RSLD | Shoot Na-K ratio |
|---|---|---|---|---|---|---|---|---|
| Replicate | 3.52** | 30.6 | 654.71 | 0.015 | 0.225 | 54.49 | 189.82 | 0.28 |
| Genotype | 0.91* | 103.15** | 538.61* | 0.016** | 0.304** | 365.30** | 373.38** | 4.12** |
| Total error | 0.57 | 8 012.594 | 55 547.23 | 0.903 | 17.125 | 18.66 | 24 089.63 | 0.72 |
Table 3 Mean square values of each physiomorphometric data.
| Source of error | SES | RSWC | RSDR | SDW | SFW | SL | RSLD | Shoot Na-K ratio |
|---|---|---|---|---|---|---|---|---|
| Replicate | 3.52** | 30.6 | 654.71 | 0.015 | 0.225 | 54.49 | 189.82 | 0.28 |
| Genotype | 0.91* | 103.15** | 538.61* | 0.016** | 0.304** | 365.30** | 373.38** | 4.12** |
| Total error | 0.57 | 8 012.594 | 55 547.23 | 0.903 | 17.125 | 18.66 | 24 089.63 | 0.72 |
| Source of error | RSWC | SDW | SFW | SL | Shoot Na-K ratio |
|---|---|---|---|---|---|
| Combined treatment | 8 060.26** | 1.001** | 52.950** | 29 760.71** | 176.58** |
| Total error | 22.71 | 4.619 | 147.187 | 27.23 | 0.36 |
Table 4 Mean square values using the combined treatment as an error term in each of the physio-morphometric data.
| Source of error | RSWC | SDW | SFW | SL | Shoot Na-K ratio |
|---|---|---|---|---|---|
| Combined treatment | 8 060.26** | 1.001** | 52.950** | 29 760.71** | 176.58** |
| Total error | 22.71 | 4.619 | 147.187 | 27.23 | 0.36 |
| Marker | MAF | GN | AN | GD | H | PIC | Marker | MAF | GN | AN | GD | H | PIC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AP3206f | 0.9783 | 2 | 2 | 0.0416 | 0 | 0.0416 | RM433 | 0.7174 | 2 | 2 | 0.3967 | 0 | 0.3233 | |
| RM490 | 0.6413 | 6 | 4 | 0.5317 | 0.0652 | 0.5037 | RM455 | 0.9783 | 2 | 2 | 0.0416 | 0 | 0.0416 | |
| RM1287 | 0.413 | 5 | 5 | 0.6741 | 0 | 0.6337 | RM493 | 0.413 | 6 | 6 | 0.6999 | 0 | 0.6724 | |
| RM3412b | 0.413 | 6 | 6 | 0.7138 | 0 | 0.6898 | RM536 | 0.4348 | 4 | 4 | 0.6611 | 0 | 0.6182 | |
| RM10694 | 0.5 | 2 | 2 | 0.4891 | 0 | 0.375 | RM3843 | 0.2391 | 10 | 10 | 0.8497 | 0 | 0.8555 | |
| RM10793 | 0.4565 | 5 | 5 | 0.675 | 0 | 0.6421 | RM10748 | 0.587 | 4 | 4 | 0.5613 | 0 | 0.5155 | |
| RM315 | 0.8261 | 3 | 3 | 0.2876 | 0 | 0.2617 | RM10764 | 0.3913 | 6 | 4 | 0.7013 | 0.0652 | 0.666 | |
| RM6329 | 0.8261 | 3 | 3 | 0.2922 | 0 | 0.2728 | RM105 | 0.6087 | 5 | 4 | 0.5469 | 0.0435 | 0.5038 | |
| RM7075 | 0.4022 | 6 | 4 | 0.6849 | 0.0435 | 0.6455 | RM24330 | 0.3696 | 5 | 5 | 0.7323 | 0 | 0.7095 | |
| RM13197 | 0.8152 | 3 | 2 | 0.295 | 0.0217 | 0.2559 | RM140 | 0.8478 | 3 | 3 | 0.258 | 0 | 0.2386 | |
| RM20224 | 0.4783 | 3 | 3 | 0.5881 | 0 | 0.5181 | RM144 | 0.587 | 5 | 5 | 0.5714 | 0 | 0.5333 | |
| RM3867 | 0.4565 | 6 | 6 | 0.6694 | 0 | 0.635 | RM171 | 0.8261 | 4 | 3 | 0.2954 | 0.0435 | 0.2795 | |
| RM19 | 0.5652 | 4 | 4 | 0.589 | 0 | 0.5487 | RM178 | 0.8478 | 3 | 3 | 0.2635 | 0 | 0.2523 | |
| RM125 | 0.8913 | 3 | 3 | 0.1951 | 0 | 0.1897 | RM10772 | 0.913 | 2 | 2 | 0.1553 | 0 | 0.1462 | |
| RM127 | 0.6304 | 2 | 2 | 0.4558 | 0 | 0.3574 | RM13332 | 0.6739 | 3 | 3 | 0.4762 | 0 | 0.432 | |
| RM152 | 0.4674 | 7 | 5 | 0.6893 | 0.0652 | 0.666 | RM495 | 0.7065 | 5 | 4 | 0.4508 | 0.0217 | 0.4199 | |
| RM413 | 0.3804 | 8 | 6 | 0.7432 | 0.0435 | 0.7252 | RM10745 | 0.7391 | 4 | 4 | 0.4189 | 0 | 0.4 |
Table 5 Summary statistics for the 34 markers used to screen the selected rice accessions.
| Marker | MAF | GN | AN | GD | H | PIC | Marker | MAF | GN | AN | GD | H | PIC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AP3206f | 0.9783 | 2 | 2 | 0.0416 | 0 | 0.0416 | RM433 | 0.7174 | 2 | 2 | 0.3967 | 0 | 0.3233 | |
| RM490 | 0.6413 | 6 | 4 | 0.5317 | 0.0652 | 0.5037 | RM455 | 0.9783 | 2 | 2 | 0.0416 | 0 | 0.0416 | |
| RM1287 | 0.413 | 5 | 5 | 0.6741 | 0 | 0.6337 | RM493 | 0.413 | 6 | 6 | 0.6999 | 0 | 0.6724 | |
| RM3412b | 0.413 | 6 | 6 | 0.7138 | 0 | 0.6898 | RM536 | 0.4348 | 4 | 4 | 0.6611 | 0 | 0.6182 | |
| RM10694 | 0.5 | 2 | 2 | 0.4891 | 0 | 0.375 | RM3843 | 0.2391 | 10 | 10 | 0.8497 | 0 | 0.8555 | |
| RM10793 | 0.4565 | 5 | 5 | 0.675 | 0 | 0.6421 | RM10748 | 0.587 | 4 | 4 | 0.5613 | 0 | 0.5155 | |
| RM315 | 0.8261 | 3 | 3 | 0.2876 | 0 | 0.2617 | RM10764 | 0.3913 | 6 | 4 | 0.7013 | 0.0652 | 0.666 | |
| RM6329 | 0.8261 | 3 | 3 | 0.2922 | 0 | 0.2728 | RM105 | 0.6087 | 5 | 4 | 0.5469 | 0.0435 | 0.5038 | |
| RM7075 | 0.4022 | 6 | 4 | 0.6849 | 0.0435 | 0.6455 | RM24330 | 0.3696 | 5 | 5 | 0.7323 | 0 | 0.7095 | |
| RM13197 | 0.8152 | 3 | 2 | 0.295 | 0.0217 | 0.2559 | RM140 | 0.8478 | 3 | 3 | 0.258 | 0 | 0.2386 | |
| RM20224 | 0.4783 | 3 | 3 | 0.5881 | 0 | 0.5181 | RM144 | 0.587 | 5 | 5 | 0.5714 | 0 | 0.5333 | |
| RM3867 | 0.4565 | 6 | 6 | 0.6694 | 0 | 0.635 | RM171 | 0.8261 | 4 | 3 | 0.2954 | 0.0435 | 0.2795 | |
| RM19 | 0.5652 | 4 | 4 | 0.589 | 0 | 0.5487 | RM178 | 0.8478 | 3 | 3 | 0.2635 | 0 | 0.2523 | |
| RM125 | 0.8913 | 3 | 3 | 0.1951 | 0 | 0.1897 | RM10772 | 0.913 | 2 | 2 | 0.1553 | 0 | 0.1462 | |
| RM127 | 0.6304 | 2 | 2 | 0.4558 | 0 | 0.3574 | RM13332 | 0.6739 | 3 | 3 | 0.4762 | 0 | 0.432 | |
| RM152 | 0.4674 | 7 | 5 | 0.6893 | 0.0652 | 0.666 | RM495 | 0.7065 | 5 | 4 | 0.4508 | 0.0217 | 0.4199 | |
| RM413 | 0.3804 | 8 | 6 | 0.7432 | 0.0435 | 0.7252 | RM10745 | 0.7391 | 4 | 4 | 0.4189 | 0 | 0.4 |
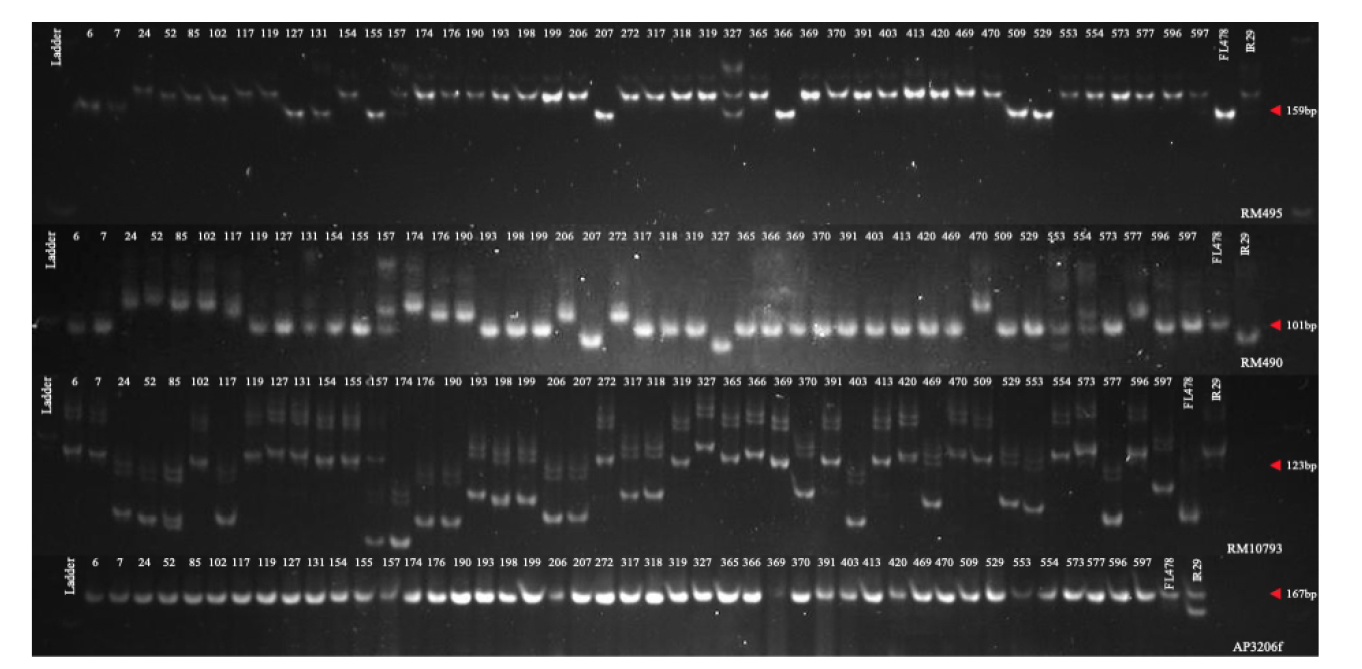
Supplmental Fig. 1 Representative gel documentation and banding pattern of AP3206f, RM3421b and RM10793 used to detect the genetic polymorphisms in the 44 test accessions.
| [1] | Asfaw K G.2011. Effects of salinity on seedling biomass production and relative water content of twenty sorghum (Sorghum bicolor L. Moench) accessions. Asian J Agric Sci, 3(3): 242-249. |
| [2] | Bhowmik S K, Titov S, Islam M M, Siddika A, Sultana S, Haquel M D S.2009. Phenotypic and genotypic screening of rice genotypes at seedling stage for salt tolerance.Afr J Biotechnol, 8(23): 6490-6494. |
| [3] | Bonilla P S, Dvorak J, Mackill D, Deal K, Gregorio G.2002. RFLP and SSLP mapping of salinity tolerance genes in chromosome 1 of rice (Oryza sativa L.) using recombinant inbred lines. Phil Agric Sci, 65(1): 68-76. |
| [4] | Delaporta S L, Wood J, Hicks J B.1983. A plant DNA minipreparation: Version II. Plant Mol Biol Rep, 1(4): 19-21. |
| [5] | Demiral T, Türkan I.2005. Comparative lipid peroxidation, antioxidant defence systems and proline content in roots of two rice cultivars in salt tolerance.Environ Exp Bot, 53: 247-257. |
| [6] | Felsentein J.1989. PHYLIP: Phylogeny inference package (version 3.2).Cladistics, 5: 164-166. |
| [7] | Flowers T, Flowers S.2005. Why does salinity pose such a difficult problem for plant breeders?Agric Water Manag, 78(1): 15-24. |
| [8] | Ganie S A, Karmakar J, Roychowdhury R, Mondal T K, Dey N.2014. Assessment of genetic diversity in salt-tolerant rice and its wild relative for ten SSR loci and one allele mining primer of salT gene located on 1st chromosome. Plant Syst Evol, 300: 1741-1747. |
| [9] | Ghosh N, Adak M K, Ghosh P D, Gupta S, Sen Gupta D N, Mandal C.2011. Differential responses of two rice varieties to salt stress.Plant Biotechnol Rep, 5: 89-103. |
| [10] | Gregorio G B, Senadhira D.1993. Genetic analysis of salinity tolerance in rice (Oryza sativa L.). Theor Appl Genet, 86: 333-338. |
| [11] | Gregorio G B, Senadhira D, Mendoza R D.1997. Screening rice for salinity tolerance. In: IRRI Discussion Paper Series No. 22 . Los Baños, Laguna, the Philippines:International Rice Research Institute. |
| [12] | Gregorio G B, Senadhira D, Mendoza R D, Manigbas N L, Roxas J P, Guerta C Q.2002. Progress in breeding for salinity tolerance and associated abiotic stresses in rice.Field Crops Res, 76(2/3): 91-101. |
| [13] | Guevarra E, Loresto G C, Jackson M T.2001. Use of conserved rice germplasm.Plant Genet Resour Newslett, 124: 51-56. |
| [14] | Ha S, Vankova R, Yamaguchi-Shinozaki K, Shinozaki K, Tran L S.2012. Cytokinins: Metabolism and function in plant adaptation to environmental stresses.Trends Plant Sci, 17(3): 172-179. |
| [15] | Hamid M A, Islam M R.1984. Rice cultivation in the tidal swamps of Bangladesh. In: Workshop on Research Priorities in Tidal Swamp Rice. Los Baños, the Philippines: International Rice Research Institute: 69-77. |
| [16] | Hoang T M L, Tran T N, Nguyen T K T, Williams B, Wurm P, Bellairs S, Mundree S.2016. Improvement of salinity stress tolerance in rice: Challenges and opportunities.Agronomy, 6(54): 1-23. |
| [17] | Hoisington D, Khairallah M, Reeves T, Ribaut J M, Skovmand B, Taba S, Warburton M.1999. Plant genetic resources: What can they contribute toward increased crop productivity?Proc Natl Acad Sci USA, 96: 5937-5943. |
| [18] | Horie T, Karahara I, Katsuhara M.2012. Salinity tolerance mechanisms in glycophytes: An overview with the central focus on rice plants.Rice, 5(1): 1-18. |
| [19] | Iqbal A, Shoajib J.1993. Evaluation of soil and water salinity for cropping in the coastal region of Bangladesh.J Noami, 10: 12-18. |
| [20] | Islam M R, Salam M A, Hassan L, Collard B C Y, Singh R K, Gregorio G B.2011. QTL mapping for salinity tolerance at seedling stage in rice (Oryza sativa L.). Emir J Food Agric, 23(2): 137-146. |
| [21] | James R A, Rivelli A R, Munns R, Caemmerer S V.2002. Factors affecting CO2 assimilation, leaf injury and growth in salt-stressed durum wheat.Funct Plant Biol, 29: 1393-1403. |
| [22] | Janaguiraman D M, Ramadass R, Devi D D.2003. Effect of salt stress on germination and seedling growth in rice genotypes.Madras Agric J, 90(1/2/3): 50-53. |
| [23] | Javed A S, Khan M F A.1975. Effect of sodium chloride and sodium sulphate on IRRI rice.J Agric Res, 13: 705-710. |
| [24] | Kader M A, Lindberg S.2005. Uptake of sodium in protoplasts of salt-sensitive and salt-tolerant cultivars of rice,Oryza sativa L. determined by the fluorescent dye SBFI. J Exp Bot, 56: 3149-3158. |
| [25] | Khan M S A, Hamid A, Karim M A.1997. Effect of sodium chloride on germination and seedling characters of different types of rice (Oryza sativa L.). J Agron Crop Sci, 179: 163-169. |
| [26] | Kumar K, Kumar M, Kim S R, Ryu H, Cho Y G.2013. Insights into genomics of salt stress response in rice.Rice, 6(1): 27. |
| [27] | Lafitte H, Ismail A, Bennett J.2004. Abiotic stress tolerance in rice for Asia: Progress and the future. In: Proceedings of a Symposium on the New Directions for a Diverse Planet at the 4th Meeting of the International Crop Science Congress. September 26 to October 1, 2004. Brisbane,Australia: International Crop Science Congress: 1-17. |
| [28] | Lapitan V C, Brar D S, Abe T, Redoña E D.2007. Assessment of genetic diversity of Philippine rice cultivars carrying good quality traits using SSR markers.Breeding Sci, 57(4): 263-270. |
| [29] | Lisa L A, Seraj Z I, Elahi C M F, Das K C, Biswas K, Islam M R, Salam M A, Gomosta A R.2004. Genetic variation in microsatellite DNA, physiology and morphology of coastal saline rice (Oryza sativa L.) landraces of Bangladesh. Plant Soil, 263: 213-228. |
| [30] | Lee K S, Choi W Y, Ko J C, Kim T S, Gregorio G B.2003. Salinity tolerance of japonica and indica rice(Oryza sativa L.) at the seedling stage. Planta, 216: 1043-1046. |
| [31] | Liu K, Muse S V.2005. Powermarker: An integrated analysis environment for genetic marker analysis.Bioinformatics, 21(9): 2128-2129. |
| [32] | Mazid M A, Malek M A, Hossain M.2016. The BRAC approach to small farmer innovations. In: Technological and Institutional Innovations for Marginalized Smallholders in Agricultural Development. Switzerland: Springer International Publishing: 99-112. |
| [33] | Mishra B, Singh R, Senadhira D.2003. Advances in breeding salt-tolerant rice varieties. In: Khush G S, Brar D S, Hardy B. Advances in Rice Genetics. Proceedings of a Symposium on Rice Genetics at the 4th meeting of the International Rice Genetics Symposium; October 22-27, 2000. Los Baños, the Philippines: International Rice Research Institute: 5-7. |
| [34] | Mohamed A M, Natarajan S, Vanathi D, Ramasamy S, Sathyamoorthi K.2007. Lowland rice in costal saline soils: A review.Agric Rev, 28(4): 245-253. |
| [35] | Mohammadi-Nejad G, Arzani A, Rezai A M, Singh R K, Gregorio G B.2008. Assessment of rice genotypes for salt tolerance using microsatellite markers associated with the Saltol QTL. Afr J Biotechnol, 7(6): 730-736. |
| [36] | Mohammadi-Nejad G, Singh R K, Arzani A, Rezaie A M, Sabouri H, Gregorio G B.2009. Evaluation of salinity tolerance in rice (Oryza sativa L.) genotypes. Int J Plant Prod, 4: 199-208. |
| [37] | Moradi F, Ismail A M.2007. Responses of photosynthesis, chlorophyll fluorescence and ros-scavenging systems to salt stress during seedling and reproductive stages in rice.Ann Bot, 99: 1161-1173. |
| [38] | Moreno L, Ordoñez S, Dela Cruz I, Redoña E.2003. Relationship of parental genetic diversity with heterosis in two-line and three-line Philippine rice hybrids. In: Khush G S, Brar D S, Hardy B. Advances in Rice Genetics. Proceedings of a Symposium on Rice Genetics at the 4th Meeting of the International Rice Research Institute: 12-14. |
| [39] | Munns R, Tester M.2008. Mechanisms of salinity tolerance.Annu Rev Plant Biol, 59: 651-681. |
| [40] | Natarajan S, Ganapthy M, Krishnakumar S, Dhanalakshmi R, Saliha B.2005. Grouping of rice genotypes for salinity tolerance based upon grain yield and Na:K ratio under coastal environment. Res J Agric Biol Sci, 1(2): 162-165. |
| [41] | PhilRice (Philippine Rice Research Institute). 2011. Management of salt-affected soils for rice production.Department Agric, 77: 1-28. |
| [42] | Qin F, Kodaira K S, Maruyama K, Mizoi J, Tran L S, Fujita Y, Morimoto K, Shinozaki K, Yamaguchi-Shinozaki K.2011. SPINDLY, a negative regulator of gibberellic acid signaling, is involved in the plant abiotic stress response.Plant Physiol, 157: 1900-1913. |
| [43] | Rahman M A, Thomson M J, Shah-E-Alam M, de Ocampo M, Egdane J, Ismail A M.2016. Exploring novel genetic sources of salinity tolerance in rice through molecular and physiological characterization.Ann Bot, 117(6): 1083-1097. |
| [44] | Rajendran K, Tester M, Roy S J.2009. Quantifying the three main components of salinity tolerance in cereals. Plant Cell Environ, 32(3): 237-249. |
| [45] | Rashid MM, Imran S, Islam MA, Hassan L.2018. Genetic diversity analysis of rice landraces (Oryza sativa L.) for salt tolerance using SSR markers in Bangladesh. Fund Appl Agric, 3(2): 460-466. |
| [46] | Reddy I N B L, Kim B K, Yoon I S, Kim K H.2017. Salt tolerance in rice: Focus on mechanisms and approaches. Rice Sci, 24(3): 123-144. |
| [47] | Rines H W, Molnar S J, Tinker N A, Phillips R L.2006. Cereals and millets. In: Kole C. Genome Mapping and Molecular Breeding in Plants. Berlin: Springer: 211-242. |
| [48] | Roy S K, Patra S K, Sarkar K K.2002. Studies on the effect of salinity stress on rice (Oryza sativa L.) at seedling stage. J Interac, 6(3): 254-259. |
| [49] | Nei M, Chesser R K.1983. Estimation of fication indices and gene diversities.Ann Human Genet, 47(3): 253-259. |
| [50] | Saxena M K, Pandey U K.1981. Physiological studies on salt tolerance of ten rice varieties growth and yield aspect.Ind J Plant Physiol, 24: 61-68. |
| [51] | Schmidt R, Caldanab C, Mueller-Roeberab B, Schippers J H M.2014. The contribution of SERF1 to root-to-shoot signaling during salinity stress in rice.Plant Sig Behav, 9(1): 1-10. |
| [52] | Sexcion F S H, Egdane J A, Ismail A M, Dionisio-Sese M L.2009. Morpho-physiological traits associated with tolerance of salinity during seedling stage in rice (Oryza sativa L.). Phil J Crop Sci, 34(2): 27-37. |
| [53] | Sirault X R R, James R A, Furbank R T.2009. A new screening method for osmotic component of salinity tolerance in cereals using infrared thermography.Funct Plant Biol, 36(11): 970-977. |
| [54] | Tahjib-Ul-Arif M D, Sayed M A, Islam M M, Siddiqui M N, Begum S N, Hossain M A.2018. Screening of rice landraces (Oryza sativa L.) for seedling stage salinity tolerance using morpho-physiological and molecular markers. Acta Physiol Plant, 40(70): 1-12. |
| [55] | Thomson M J, de Ocampo M, Egdane J, Rahman M A, Sajise A G, Adorada D L, Tumimbang-Raiz E, Blumwald E, Seraj Z I, Singh R K, Gregorio G B, Ismail A M.2010. Characterizing the Saltol quantitative trait locus for salinity tolerance in rice. Rice, 3(2/3): 148-160. |
| [56] | Türkana I, Demiral T.2009. Recent developments in understanding salinity tolerance.Environ Exp Bot, 67: 2-9. |
| [57] | Umali D L.1993. Irrigation-Induced Salinity a Growing Problem for Development and the Environment . Washington DC, USA: the Word Bank. |
| [58] | Verslues P E, Agarwal M, Katiyar-Agarwal S, Zhu J H, Zhu J K.2006. Methods and concepts in quantifying resistance to drought, salt and freezing, abiotic stresses that affect plant water status.Plant J, 45(4): 523-539. |
| [59] | Wahhab A.1961. Salt tolerance of various varieties of agricultural crops at germination stage.In: Salinity Problems in the Arid Zone, Proc. Teheran Symp. Arid Zone Res, 14: 185-192. |
| [60] | Walia H, Wilson C, Condamine P, Liu X, Ismail A M, Zeng L, Wanamaker S I, Mandal J, Xu J, Cui X, Close T J.2005. Comparative transcriptional profiling of two contrasting rice genotypes under salinity stress during the vegetative growth stage.Plant Physiol, 139: 822-835. |
| [61] | Xu Y, Crouch J.2008. Marker-assisted selection in plant breeding: From publications to practice.Crop Sci, 48: 391-407. |
| [62] | Yeo A R, Yeo M E, Flowers S A, Flowers T J.1990. Screening of rice (Oryza sativa) genotypes for physiological characters contributing to salinity resistance, and their relationship to overall performance. Theor Appl Genet, 79(3): 377-384. |
| [63] | Yoshida S, Forno D A, Cock J H, Gomez K A.1976. Laboratory Manual for Physiological Studies of Rice. 3rd edn . Los Banos,the Philippines: International Rice Research Institute: 83. |
| [64] | Zeng L H, Shannon M C.2000. Salinity effects on seedling growth and yield components rice.Crop Sci, 40(4): 996-1003. |
| [1] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [2] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [3] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [4] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [5] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [6] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [7] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [8] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [9] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [10] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [14] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [15] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||