
Rice Science ›› 2019, Vol. 26 ›› Issue (5): 265-281.DOI: 10.1016/j.rsci.2019.08.001
• Review • Previous Articles Next Articles
Matías Romero Fernando, Gatica-Arias Andrés( )
)
Received:2018-12-15
Accepted:2019-03-07
Online:2019-09-28
Published:2019-05-24
Matías Romero Fernando, Gatica-Arias Andrés. CRISPR/Cas9: Development and Application in Rice Breeding[J]. Rice Science, 2019, 26(5): 265-281.
Add to citation manager EndNote|Ris|BibTeX
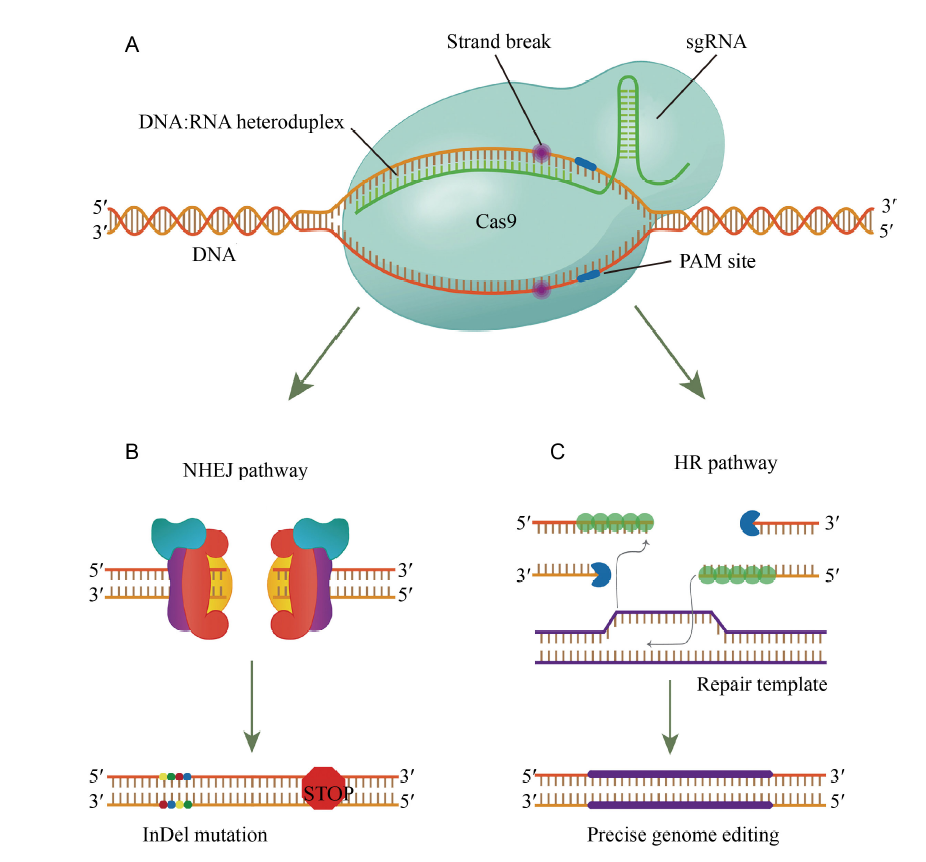
Fig. 1. CRISPR/Cas9 system components and pathways.A, Components required for genome editing using the CRISPR/Cas9 system: a DNA endonuclease (the most commonly used is the Cas9 protein from Streptococcus pyogenes) and a customizable single guide RNA (sgRNA). B, Non-homologous end-joining (NHEJ) repair pathway. C, Homologous recombination (HR) repair pathway. PAM, Protospacer adjacent motif.
| Application | Targeted gene | Strategy | Transformation method/Explant | Reference |
|---|---|---|---|---|
| Response to bacterial blight | SWEET14, SWEET11 | NHEJ | PEG/Protoplast | |
| SWEET13 | NHEJ | Agrobacterium/embriogenic calli | ||
| Rice blast resistance | EFR922 | NHEJ | Agrobacterium/embriogenic calli | |
| SEC3A | NHEJ | Agrobacterium/embriogenic calli | ||
| Response to saline stress | RAV2 | NHEJ | Agrobacterium/embriogenic calli | |
| Response to drought stress | SAPK2 | NHEJ | Agrobacterium/embriogenic calli | |
| Response to cold stress | ANN3 | NHEJ | Agrobacterium/embriogenic calli | |
| Herbicide resistance | EPSPS | Exon replacement by NHEJ | Biobalistic/embriogenic calli | |
| AAC | Base editing by nCas9 | Agrobacterium/embriogenic calli | ||
| ALS | Base editing by nCas9 and/or dCas9 | Agrobacterium/embriogenic calli | ||
| ALS | HR | Biobalistic/embriogenic calli | ||
| Yield improvement | Gn1a, DEP1, GS3, IPA1 | NHEJ | Agrobacterium/embriogenic calli | |
| GS3, Gn1a | NHEJ | Agrobacterium/embriogenic calli | ||
| DEP1 | Gen deletion by NHEJ | Agrobacterium/embriogenic calli | ||
| GW2, GW5, TGW6 | NHEJ | Agrobacterium/embriogenic calli | ||
| GS3, GW2, Gn1a | NHEJ | Agrobacterium/embriogenic calli | ||
| TMS5 | NHEJ | Agrobacterium/embriogenic calli | ||
| Fatty acid metabolism | FAD2-1 | NHEJ | Agrobacterium/embriogenic calli | |
| Flowering time | Hd2, Hd4, Hd5 | NHEJ | Agrobacterium/embriogenic calli | |
| Nitrogen efficiency use | NRT1.1B | Allele replacement by HR | Biobalistic/embriogenic calli | |
| Cadmium accumulation | Nramp5 | NHEJ | Agrobacterium/embriogenic calli | |
| Starch metabolism | SBEI, SBEIIb | NHEJ | Agrobacterium/embriogenic calli | |
| WAXY | NHEJ | Agrobacterium/embriogenic calli | ||
| Transgene excision | GUS | NHEJ | Agrobacterium and biobalistic/embriogenic calli | |
| Comparison between | EPFL9 | NHEJ | Agrobacterium/immature embryos | |
| Cas9 and Cpf1 | ||||
| Genome wide mutant Library | 12 802 genes | NHEJ | Agrobacterium/embriogenic calli | |
| 34 234 genes | NHEJ | Agrobacterium/embriogenic calli |
Table 1 Applications of CRISPR in rice from 2013 to mid-2018.
| Application | Targeted gene | Strategy | Transformation method/Explant | Reference |
|---|---|---|---|---|
| Response to bacterial blight | SWEET14, SWEET11 | NHEJ | PEG/Protoplast | |
| SWEET13 | NHEJ | Agrobacterium/embriogenic calli | ||
| Rice blast resistance | EFR922 | NHEJ | Agrobacterium/embriogenic calli | |
| SEC3A | NHEJ | Agrobacterium/embriogenic calli | ||
| Response to saline stress | RAV2 | NHEJ | Agrobacterium/embriogenic calli | |
| Response to drought stress | SAPK2 | NHEJ | Agrobacterium/embriogenic calli | |
| Response to cold stress | ANN3 | NHEJ | Agrobacterium/embriogenic calli | |
| Herbicide resistance | EPSPS | Exon replacement by NHEJ | Biobalistic/embriogenic calli | |
| AAC | Base editing by nCas9 | Agrobacterium/embriogenic calli | ||
| ALS | Base editing by nCas9 and/or dCas9 | Agrobacterium/embriogenic calli | ||
| ALS | HR | Biobalistic/embriogenic calli | ||
| Yield improvement | Gn1a, DEP1, GS3, IPA1 | NHEJ | Agrobacterium/embriogenic calli | |
| GS3, Gn1a | NHEJ | Agrobacterium/embriogenic calli | ||
| DEP1 | Gen deletion by NHEJ | Agrobacterium/embriogenic calli | ||
| GW2, GW5, TGW6 | NHEJ | Agrobacterium/embriogenic calli | ||
| GS3, GW2, Gn1a | NHEJ | Agrobacterium/embriogenic calli | ||
| TMS5 | NHEJ | Agrobacterium/embriogenic calli | ||
| Fatty acid metabolism | FAD2-1 | NHEJ | Agrobacterium/embriogenic calli | |
| Flowering time | Hd2, Hd4, Hd5 | NHEJ | Agrobacterium/embriogenic calli | |
| Nitrogen efficiency use | NRT1.1B | Allele replacement by HR | Biobalistic/embriogenic calli | |
| Cadmium accumulation | Nramp5 | NHEJ | Agrobacterium/embriogenic calli | |
| Starch metabolism | SBEI, SBEIIb | NHEJ | Agrobacterium/embriogenic calli | |
| WAXY | NHEJ | Agrobacterium/embriogenic calli | ||
| Transgene excision | GUS | NHEJ | Agrobacterium and biobalistic/embriogenic calli | |
| Comparison between | EPFL9 | NHEJ | Agrobacterium/immature embryos | |
| Cas9 and Cpf1 | ||||
| Genome wide mutant Library | 12 802 genes | NHEJ | Agrobacterium/embriogenic calli | |
| 34 234 genes | NHEJ | Agrobacterium/embriogenic calli |
| [1] | Abe K, Araki E, Suzuki Y, Toki S, Saika H.2018. Production of high oleic/low linoleic rice by genome editing.Plant Physiol Biochem, 131: 58-62. |
| [2] | Altpeter F, Springer N M, Bartley L E, Blechl A E, Brutnell T P, Citovsky V, Conrad L J, Gelvin S B, Jackson D P, Kausch A P, Lemaux P G, Medford J I, Orozco-Cárdenas M L, Tricoli D M, Van Eck J, Voytas D F, Walbot V, Wang K, Zhang Z J, Stewart C N Jr.2016. Advancing crop transformation in the era of genome editing.Plant Cell, 28(7): 1510-1520. |
| [3] | Anders C, Bargsten K, Jinek M.2016. Structural plasticity of PAM recognition by engineered variants of the RNA-guided endonuclease Cas9.Mol Cell, 61(6): 895-902. |
| [4] | Bae S, Park J, Kim J S.2014. Cas-OFFinder: A fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases.Bioinformatics, 30(10): 1473-1475. |
| [5] | Belhaj K, Chaparro-Garcia A, Kamoun S, Patron N J, Nekrasov V.2015. Editing plant genomes with CRISPR/Cas9.Curr Opin Biotechnol, 32: 76-84. |
| [6] | Bell C C, Magor G W, Gillinder K R, Perkins A C.2014. A high-throughput screening strategy for detecting CRISPR-Cas9 induced mutations using next-generation sequencing.BMC Genom, 15(1): 1002. |
| [7] | Bertin G, Averbeck D.2006. Cadmium: Cellular effects, modifications of biomolecules, modulation of DNA repair and genotoxic consequences.Biochimie, 88(11): 1549-1559. |
| [8] | Bhaya D, Davison M, Barrangou R.2011. CRISPR-Cas systems in bacteria and archaea: Versatile small RNAs for adaptive defense and regulation.Annu Rev Genet, 45(1): 273-297. |
| [9] | Breviario D, Genga A.2013. Stress response in rice.J Rice Res, 2: e104. |
| [10] | Clemens S, Aarts M G M, Thomine S, Verbruggen N.2013. Plant science: The key to preventing slow cadmium poisoning.Trends Plant Sci, 18(2): 92-99. |
| [11] | Chari R, Yeo N C, Chavez A, Church G M.2017. sgRNA scorer 2.0: A species-independent model to predict CRISPR/Cas9 activity.ACS Synth Biol, 6(5): 902-904. |
| [12] | Chen L Z, Li W, Katin-Grazzini L, Ding J, Gu X B, Li Y J, Gu T T, Wang R, Lin X C, Deng Z N, McAvoy R J, Gmitter F G, Deng Z N, Zhao Y D, Li Y.2018. A method for the production and expedient screening of CRISPR/Cas9-mediated non-transgenic mutant plants.Hort Res, 5: 13. |
| [13] | Cheng S H, Zhuang J Y, Fan Y Y, Du J H, Cao L Y.2007. Progress in research and development on hybrid rice: A super-domesticate in China.Ann Bot, 100(5): 959-966. |
| [14] | Dean R, Van J A L, Pretorius Z A, Hammond-Kosack K E, Di Pietro A, Spanu P D, Rudd J J, Dickman M, Kahmann R, Ellis J, Foster G D.2012. The top 10 fungal pathogens in molecular plant pathology.Mol Plant Pathol, 13(4): 414-430. |
| [15] | Delorge I, Janiak M, Carpentier S, van Dijck P.2014. Fine tuning of trehalose biosynthesis and hydrolysis as novel tools for the generation of abiotic stress tolerant plants.Front Plant Sci, 5: 147. |
| [16] | Doench J G, Fusi N, Sullender M, Hegde M, Vaimberg E W, Donovan K F, Smith I, Tothova Z, Wilen C, Orchard R, Virgin H W, Listgarten J, Root D E.2016. Optimized sgRNA design to maximize activity and minimize off-target effects of CRISPR- Cas9.Nat Biotechnol, 34: 184-191. |
| [17] | Dong Y F, Jin X, Tang Q L, Zhang X, Yang J T, Liu X J, Cai J F, Zhang X B, Wang X J, Wang Z X.2017. Development and event-specific detection of transgenic glyphosate-resistant rice expressing the G2-EPSPS gene. Front Plant Sci, 8: 885. |
| [18] | Duan Y B, Li J, Qin R Y, Xu R F, Li H, Yang Y C, Ma H, Li L, Wei P C, Yang J B.2016. Identification of a regulatory element responsible for salt induction of rice OsRAV2 through ex situ and in situ promoter analysis. Plant Mol Biol, 90(1): 49-62. |
| [19] | Endo M, Nishizawa-Yokoi A, Toki S.2018. Rice genome editing. In: Sasaki T, Ashikari M. Rice Genomics, Genetics and Breeding. Singapore: Springer: 523-539. |
| [20] | Fartyal D, Agarwal A, James D, Borphukan B, Ram B, Sheri V, Agrawal P K, Achary V M M, Reddy M K.2018. Developing dual herbicide tolerant transgenic rice plants for sustainable weed management.Sci Rep, 8(1): 11598. |
| [21] | Feng X P, Peng C, Chen Y, Liu X D, Feng X J, He Y.2017. Discrimination of CRISPR/Cas9-induced mutants of rice seeds using near-infrared hyperspectral imaging.Sci Rep, 7: 15934. |
| [22] | Hamada H, Liu Y L, Nagira Y, Miki R, Taoka N, Imai R.2018. Biolistic-delivery-based transient CRISPR/Cas9 expression enables in planta genome editing in wheat.Sci Rep, 8: 14422. |
| [23] | He Y B, Zhu M, Wang L H, Wu J H, Wang Q Y, Wang R C, Zhao Y D.2018. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants.Mol Plant, 11(9): 1210-1213. |
| [24] | Hermans P W, van Soolingen D, Bik E M, de Haas P E, Dale J W, van Embden J D.1991. Insertion element IS987 from Mycobacterium bovis BCG is located in a hot-spot integration region for insertion elements in Mycobacterium tuberculosis complex strains. Infect Immun, 59(8): 2695-2705. |
| [25] | Huang X Z, Qian Q, Liu Z B, Sun H Y, He S Y, Luo D, Xia G M, Chu C C, Li J Y, Fu X D.2009. Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet, 41: 494-497. |
| [26] | Inui H, Shiota N, Ido Y, Inoue T, Hirose S, Kawahigashi H, Ohkawa Y, Ohkawa H.2001. Herbicide metabolism and tolerance in the transgenic rice plants expressing human CYP2C9 and CYP2C19.Pest Biochem Physiol, 71(3): 156-169. |
| [27] | Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A.1987. Nucleotide sequence of theiap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol, 169(12): 5429-5433. |
| [28] | Jiang W Y, Bikard D, Cox D, Zhang F, Marraffini L A.2013. RNA-guided editing of bacterial genomes using CRISPR-Cas systems.Nat Biotechnol, 31(3): 233-239. |
| [29] | Jiang W Z, Zhou H B, Bi H H, Fromm M, Yang B, Weeks D P.2013. Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucl Acids Res, 41(20): e188. |
| [30] | Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna J A, Charpentier E.2012. A programmable dual-RNA: Guided DNA endonuclease in adaptive bacterial immunity.Science, 337: 816-821. |
| [31] | Jung J H, Seo Y W.2017. Challenges in wide implementation of genome editing for crop improvement.J Crop Sci Biotechnol, 20(2): 129-135. |
| [32] | Kaeppler S M, Kaeppler H F, Rhee Y.2000. Epigenetic aspects of somaclonal variation in plants.Plant Mol Biol, 43: 179-188. |
| [33] | Koonin E V, Makarova K S, Zhang F.2017. Diversity, classification and evolution of CRISPR-Cas systems.Curr Opin Microbiol, 37: 67-78. |
| [34] | Krappmann S.2017. CRISPR-Cas9, the new kid on the block of fungal molecular biology.Med Mycol, 55(1): 16-23. |
| [35] | Li C, Zong Y, Wang Y P, Jin S, Zhang D B, Song Q N, Zhang R, Gao C X.2018. Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion.Genome Biol, 19: 59. |
| [36] | Li J, Meng X B, Zong Y, Chen K L, Zhang H W, Liu J X, Li J Y, Gao C X.2016. Gene replacements and insertions in rice by intron targeting using CRISPR-Cas9.Nat Plants, 2: 16139. |
| [37] | Li J Y, Zhang X, Sun Y W, Zhang J H, Du W M, Guo X P, Li S Y, Zhao Y D, Xia L Q.2018. Efficient allelic replacement in rice by gene editing: A case study of the NRT1.1B gene. J Integr Plant Biol, 60(7): 536-540. |
| [38] | Li M R, Li X X, Zhou Z J, Wu P Z, Fang M C, Pan X P, Lin Q P, Luo W B, Wu G J, Li H Q.2016. Reassessment of the four yield-related genes Gn1a, DEP1, GS3, and IPA1 in rice using a CRISPR/Cas9 system. Front Plant Sci, 7: 377. |
| [39] | Li X F, Liu H Z, Wang M Q, Liu H L, Tian X J, Zhou W J, Lü T X, Wang Z Y, Chu C C, Fang J, Bu Q Y.2015. Combinations of Hd2 and Hd4 genes determine rice adaptability to Heilongjiang Province, northern limit of China. J Integr Plant Biol, 57(8): 698-707. |
| [40] | Li X F, Zhou W J, Ren Y K, Tian X J, Lv T X, Wang Z Y, Fang J, Chu C C, Yang J, Bu Q Y.2017. High-efficiency breeding of early-maturing rice cultivars via CRISPR/Cas9-mediated genome editing.J Genet Genom, 44(3): 175-178. |
| [41] | Li Z S, Liu Z B, Xing A Q, Moon B P, Koellhoffer J P, Huang L X, Ward R T, Clifton E, Falco S C, Cigan A M.2015. Cas9-guide RNA directed genome editing in soybean.Plant Physiol, 169(2): 960-970. |
| [42] | Liang Z, Chen K L, Li T D, Zhang Y, Wang Y P, Zhao Q, Liu J X, Zhang H W, Liu C M, Ran Y D, Gao C X.2017. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes.Nat Commun, 8: 14261. |
| [43] | Liang Z, Chen K L, Zhang Y, Liu J X, Yin K Q, Qiu J L, Gao C X.2018. Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins. Nat Protoc, 13: 413-430. |
| [44] | Liu D F, Chen X J, Liu J Q, Ye J C, Guo Z J.2012. The rice ERF transcription factor OsERF922 negatively regulates resistance to Magnaporthe oryzae and salt tolerance. J Exp Bot, 63(10): 3899-3911. |
| [45] | Liu Q, Wang C, Jiao X Z, Zhang H W, Song L L, Li Y X, Gao C X, Wang K J.2019. Hi-TOM: A platform for high-throughput tracking of mutations induced by CRISPR/Cas systems.Sci China: Life Sci, 62(1): 1-7. |
| [46] | Lou D J, Wang H P, Liang G, Yu D Q.2017. OsSAPK2 confers abscisic acid sensitivity and tolerance to drought stress in rice. Front Plant Sci, 8: 993. |
| [47] | Lu H P, Liu S M, Xu S L, Chen W Y, Zhou X, Tan Y Y, Huang J Z, Shu Q Y.2017. CRISPR-S: An active interference element for a rapid and inexpensive selection of genome-edited, transgene-free rice plants.Plant Biotechnol J, 15(11): 1371-1373. |
| [48] | Lu Y M, Ye X, Guo R M, Huang J, Wang W, Tang J Y, Tan L T, Zhu J K, Chu C C, Qian Y W.2017. Genome-wide targeted mutagenesis in rice using the CRISPR/Cas9 system.Mol Plant, 10(9): 1242-1245. |
| [49] | Ma J, Chen J, Wang M, Ren Y L, Wang S, Lei C L, Cheng Z J, Sodmergen.2018. Disruption of OsSEC3A increases the content of salicylic acid and induces plant defense responses in rice. J Exp Bot, 69(5): 1051-1064. |
| [50] | Matsubara K, Hori K, Ogiso-Tanaka E, Yano M.2014. Cloning of quantitative trait genes from rice reveals conservation and divergence of photoperiod flowering pathways in Arabidopsis and rice. Front Plant Sci, 5: 193. |
| [51] | Meng X B, Yu H, Zhang Y, Zhuang F F, Song X G, Gao S S, Gao C X, Li J Y.2017. Construction of a genome-wide mutant library in rice using CRISPR/Cas9.Mol Plant, 10(9): 1238-1241. |
| [52] | Miao J, Guo D S, Zhang J Z, Huang Q P, Qin G J, Zhang X, Wan J M, Gu H Y, Qu L J.2013. Targeted mutagenesis in rice using CRISPR-Cas system.Cell Res, 23(10): 1233-1236. |
| [53] | Minkenberg B, Xie K B, Yang Y N.2017. Discovery of rice essential genes by characterizing a CRISPR-edited mutation of closely related rice MAP kinase genes.Plant J, 89(3): 636-648. |
| [54] | Mishra R, Joshi R K, Zhao K J.2018. Genome editing in rice: Recent advances, challenges, and future implications.Front Plant Sci, 9: 1361. |
| [55] | Mojica F J M, Juez G, Rodriguez-Valera F.1993. Transcription at different salinities of Haloferax mediterranei sequences adjacent to partially modified PstI sites. Mol Microbiol, 9(3): 613-621. |
| [56] | Moscou M J, Bogdanove A J.2009. A simple cipher governs DNA recognition by TAL effectors.Science, 326: 1501. |
| [57] | Müller M, Munné-Bosch S.2015. Ethylene response factors: A key regulatory hub in hormone and stress signaling.Plant Physiol, 169(1): 32-41. |
| [58] | Nakata A, Amemura M, Makino K.1989. Unusual nucleotide arrangement with repeated sequences in the Escherichia coli K-12 chromosome. J Bacteriol, 171(6): 3553-3556. |
| [59] | Nandy S, Srivastava V.2012. Marker-free site-specific gene integration in rice based on the use of two recombination systems.Plant Biotechnol J, 10(8): 904-912. |
| [60] | Pabo C O, Peisach E, Grant R A.2001. Design and selection of novel Cys2His2 zinc finger proteins.Annu Rev Biochem, 70(1): 313-340. |
| [61] | Peng C, Wang H, Xu X L, Wang X F, Chen X Y, Wei W, Lai Y M, Liu G Q, Godwin I D, Li J Q, Zhang L, Xu J F.2018. High-throughput detection and screening of plants modified by gene editing using quantitative real-time polymerase chain reaction.Plant J, 95(3): 557-567. |
| [62] | Pyott D E, Sheehan E, Molnar A.2016. Engineering of CRISPR/Cas9-mediated potyvirus resistance in transgene-free Arabidopsis plants. Mol Plant Pathol, 17(8): 1276-1288. |
| [63] | Regina A, Bird A, Topping D, Bowden S, Freeman J, Barsby T, Kosar-Hashemi B, Li Z Y, Rahman S, Morell M.2006. High-amylose wheat generated by RNA interference improves indices of large-bowel health in rats.Proc Natl Acad Sci USA, 103(10): 3546-3551. |
| [64] | Ricroch A, Clairand P, Harwood W.2017. Use of CRISPR systems in plant genome editing: Toward new opportunities in agriculture.Emerg Top Life Sci, 1(2): 169-182. |
| [65] | Sarmast M K.2016. Genetic transformation and somaclonal variation in conifers.Plant Biotechnol Rep, 10(6): 309-325. |
| [66] | Shan Q W, Wang Y P, Li J, Zhang Y, Chen K L, Liang Z, Zhang K, Liu J X, Xi J J, Qiu J L, Gao C X.2013. Targeted genome modification of crop plants using a CRISPR-Cas system.Nat Biotechnol, 31(8): 686-688. |
| [67] | Shen C X, Que Q Z, Xia Y M, Tang N, Li D, He R H, Cao M L.2017. Knock out of the annexin gene OsAnn3 via CRISPR/Cas9-mediated genome editing decreased cold tolerance in rice. J Plant Biol, 60(6): 539-547. |
| [68] | Shen L, Wang C, Fu Y P, Wang J J, Liu Q, Zhang X M, Yan C J, Qian Q, Wang K J.2018. QTL editing confers opposing yield performance in different rice varieties.J Integr Plant Biol, 60(2): 89-93. |
| [69] | Shimatani Z, Kashojiya S, Takayama M, Terada R, Arazoe T, Ishii H, Teramura H, Yamamoto T, Komatsu H, Miura K, Ezura H, Nishida K, Ariizumi T, Kondo A.2017. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion.Nat Biotechnol, 35: 441-443. |
| [70] | Shimatani Z, Fujikura U, Ishii H, Matsui Y, Suzuki M, Ueke Y, Taoka K, Terada R, Nishida K, Kondo A.2018. Inheritance of co-edited genes by CRISPR-based targeted nucleotide substitutions in rice.Plant Physiol Biochem, 131: 78-83. |
| [71] | Sprink T, Metje J, Hartung F.2015. Plant genome editing by novel tools: TALEN and other sequence specific nucleases.Curr Opin Biotechnol, 32: 47-53. |
| [72] | Srivastava V, Underwood J L, Zhao S.2017. Dual-targeting by CRISPR/Cas9 for precise excision of transgenes from rice genome.Plant Cell Tiss Org, 129(1): 153-160. |
| [73] | Sun Y W, Zhang X, Wu C Y, He Y B, Ma Y Z, Hou H, Guo X P, Du W M, Zhao Y D, Xia L Q.2016. Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of acetolactate synthase.Mol Plant, 9(4): 628-631. |
| [74] | Sun Y W, Jiao G A, Liu Z P, Zhang X, Li J Y, Guo X P, Du W M, Du J L, Francis F, Zhao Y D, Xia L Q.2017. Generation of high-amylose rice through CRISPR/Cas9-mediated targeted mutagenesis of starch branching enzymes.Front Plant Sci, 8: 298. |
| [75] | Svitashev S, Schwartz C, Lenderts B, Young J K, Mark Cigan A.2016. Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes.Nat Commun, 7: 13274. |
| [76] | Tang L, Mao B G, Li Y K, Lv Q M, Zhang L P, Chen C Y, He H J, Wang W P, Zeng X F, Shao Y, Pan Y L, Hu Y Y, Peng Y, Fu X Q, Li H Q, Xia S T, Zhao B R.2017. Knockout of OsNramp5 using the CRISPR/Cas9 system produces low Cd-accumulating indica rice without compromising yield. Sci Rep, 7: 14438. |
| [77] | Tang X, Liu G Q, Zhou J P, Ren Q R, You Q, Tian L, Xin X H, Zhong Z H, Liu B L, Zheng X L, Zhang D W, Malzahn A, Gong Z Y, Qi Y P, Zhang T, Zhang Y.2018. A large-scale whole-genome sequencing analysis reveals highly specific genome editing by both Cas9 and Cpf1 (Cas12a) nucleases in rice.Genome Biol, 19: 84. |
| [78] | Wang F J, Wang C L, Liu P Q, Lei C L, Hao W, Gao Y, Liu Y G, Zhao K J.2016. Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor geneOsERF922. PLoS One, 11(4): e0154027. |
| [79] | Wang Y, Geng L Z, Yuan M L, Wei J, Jin C, Li M, Yu K, Zhang Y, Jin H B, Wang E, Chai Z J, Fu X D, Li X G.2017. Deletion of a target gene in indica rice via CRISPR/Cas9. Plant Cell Rep, 36(8): 1333-1343. |
| [80] | Woo J W, Kim J, Kwon S I, Corvalán C, Cho S W, Kim H, Kim S G, Kim S T, Choe S, Kim J S.2015. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins.Nat Biotechnol, 33: 1162-1164. |
| [81] | Xie K B, Yang Y N.2013. RNA-guided genome editing in plants using a CRISPR-Cas system.Mol Plant, 6(6): 1975-1983. |
| [82] | Xie K B, Minkenberg B, Yang Y N.2015. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA- processing system.Proc Natl Acad Sci USA, 112(11): 3570-3575. |
| [83] | Xu R F, Yang Y C, Qin R Y, Li H, Qiu C H, Li L, Wei P C, Yang J B.2016. Rapid improvement of grain weight via highly efficient CRISPR/Cas9-mediated multiplex genome editing in rice.J Genet Genom, 43(8): 529-532. |
| [84] | Yin X J, Biswal A K, Dionora J, Perdigon K M, Balahadia C P, Mazumdar S, Chater C, Lin H C, Coe R A, Kretzschmar T, Gray J E, Quick P W, Bandyopadhyay A.2017. CRISPR-Cas9 and CRISPR-Cpf1 mediated targeting of a stomatal developmental gene EPFL9 in rice. Plant Cell Rep, 36(5): 745-757. |
| [85] | Yu Q, Jalaludin A, Han H P, Chen M, Sammons R D, Powles S B.2015. Evolution of a double amino acid substitution in the 5-enolpyruvylshikimate-3-phosphate synthase in Eleusine indica conferring high-level glyphosate resistance. Plant Physiol, 167(4): 1440-1447. |
| [86] | Zhang J S, Zhang H, Botella J R, Zhu J K.2018. Generation of new glutinous rice by CRISPR/Cas9-targeted mutagenesis of the Waxy gene in elite rice varieties. J Integr Plant Biol, 60(5): 369-375. |
| [87] | Zhang Y, Liang Z, Zong Y, Wang Y P, Liu J X, Chen K L, Qiu J L, Gao C X.2016. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA.Nat Commun, 7: 12617. |
| [88] | Zhao C Z, Zheng X G, Qu W B, Li G L, Li X Y, Miao Y L, Han X S, Liu X D, Li Z H, Ma Y L, Shao Q Z, Li H W, Sun F, Xie S S, Zhao S H.2017. CRISPR-offinder: A CRISPR guide RNA design and off-target searching tool for user-defined protospacer adjacent motif.Int J Biol Sci, 13(12): 1470-1478. |
| [89] | Zhao T, Lin C Y, Shen Z C.2011. Development of transgenic glyphosate- resistant rice with G6 gene encoding 5-enolpyruvylshikimate- 3-phosphate synthase. Agric Sci China, 10(9): 1307-1312. |
| [90] | Zhou H, He M, Li J, Chen L, Huang Z F, Zheng S Y, Zhu L Y, Ni E D, Jiang D G, Zhao B R, Zhuang C X.2016. Development of commercial thermo-sensitive genic male sterile rice accelerates hybrid rice breeding using the CRISPR/Cas9-mediated TMS5 editing system. Sci Rep, 6: 37395. |
| [91] | Zhou J H, Peng Z, Long J Y, Sosso D, Liu B, Eom J S, Huang S, Liu S Z, Vera Cruz C, Frommer W B, White F F, Yang B.2015. Gene targeting by the TAL effector PthXo2 reveals cryptic resistance gene for bacterial blight of rice.Plant J, 82(4): 632-643. |
| [92] | Zhou J P, Xin X H, He Y, Chen H Q, Li Q, Tang X, Zhong Z H, Deng K J, Zheng X L, Akher S A, Cai G Z, Qi Y P, Zhang Y.2018. Multiplex QTL editing of grain-related genes improves yield in elite rice varieties.Plant Cell Rep, 38(4): 475-485. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIU Tingting, ZOU Jinpeng, YANG Xi, WANG Kejian, RAO Yuchun, WANG Chun. Development and Application of Prime Editing in Plants [J]. Rice Science, 2023, 30(6): 3-. |
| [7] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [8] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [9] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [10] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [11] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [12] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [13] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [14] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| [15] | Lu Xuedan, Li Fan, Xiao Yunhua, Wang Feng, Zhang Guilian, Deng Huabing, Tang Wenbang. Grain Shape Genes: Shaping the Future of Rice Breeding [J]. Rice Science, 2023, 30(5): 379-404. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||