
Rice Science ›› 2022, Vol. 29 ›› Issue (4): 375-384.DOI: 10.1016/j.rsci.2021.11.007
• Research Paper • Previous Articles Next Articles
Suhas Gorakh Karkute1, Vishesh Kumar1, Mohd Tasleem1, Dwijesh Chandra Mishra2, Krishna Kumar Chaturvedi2, Anil Rai2, Amitha Mithra Sevanthi1, Kishor Gaikwad1, Tilak Raj Sharma3, Amolkumar U. Solanke1( )
)
Received:2021-08-11
Accepted:2021-11-12
Online:2022-07-28
Published:2021-12-31
Contact:
Amolkumar U. Solanke
Suhas Gorakh Karkute, Vishesh Kumar, Mohd Tasleem, Dwijesh Chandra Mishra, Krishna Kumar Chaturvedi, Anil Rai, Amitha Mithra Sevanthi, Kishor Gaikwad, Tilak Raj Sharma, Amolkumar U. Solanke. Genome-Wide Analysis of von Willebrand Factor A Gene Family in Rice for Its Role in Imparting Biotic Stress Resistance with Emphasis on Rice Blast Disease[J]. Rice Science, 2022, 29(4): 375-384.
Add to citation manager EndNote|Ris|BibTeX
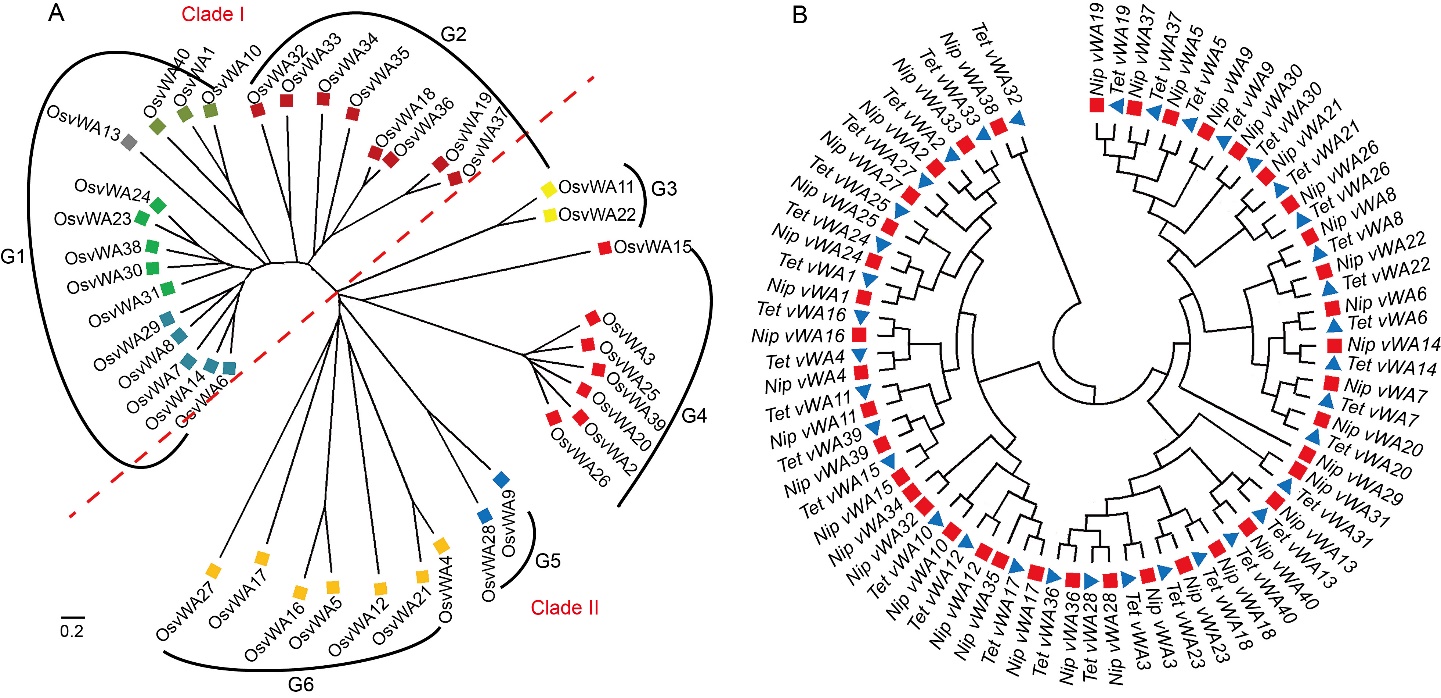
Fig. 2. Evolutionary relationship of vWA proteins (A) and promoter sequence of vWA genes (B) in rice. Different colored squares in A represent proteins in different subclades. Blue triangle and red square in B represent promoters from blast resistant variety Tetep (Tet) and susceptible variety Nipponbare (Nip), respectively. The evolutionary history was inferred using the neighbor-joining method. The evolutionary distances were computed using the Poisson correction method.
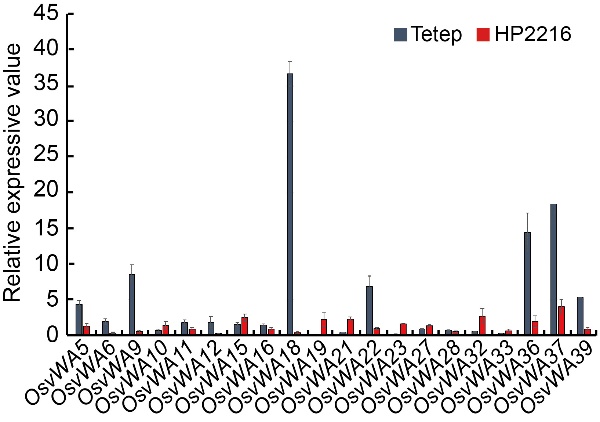
Fig. 3. Relative expression analysis of OsvWA genes in rice panicle tissues of Tetep and HP2216 genotypes inoculated with M. oryzae at 48 h post inoculation using qRT-PCR.
| [1] |
Balint-Kurti P. 2019. The plant hypersensitive response: Concepts, control and consequences. Mol Plant Pathol, 20(8): 1163-1178.
PMID |
| [2] |
Bharati K P, Prashanth U R. 2011. Von Willebrand disease: An overview. Indian J Pharm Sci, 73(1): 7-16.
PMID |
| [3] | Chen X L, Jia Y L, Wu B M. 2019. Evaluation of rice responses to the blast fungus Magnaporthe oryzae at different growth stages. Plant Dis, 103(1): 132-136. |
| [4] | Edgar R C. 2004. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res, 32(5): 1792-1797. |
| [5] | Fang N Y, Wei X R, Shen L T, Yu Y, Li M Y, Yin C F, He W W, Guan C H, Chen H, Zhang H S, Bao Y M. 2019. Fine mapping of a panicle blast resistance gene Pb-bd1 in japonica landrace Bodao and its application in rice breeding. Rice, 12(1): 18. |
| [6] |
Freemont P S. 2000. Ubiquitination: RING for destruction? Curr Biol, 10(2): R84-R87.
PMID |
| [7] |
Hirose N, Makita N, Kojima M, Kamada-Nobusada T, Sakakibara H. 2007. Overexpression of a type-A response regulator alters rice morphology and cytokinin metabolism. Plant Cell Physiol, 48(3): 523-539.
PMID |
| [8] |
Horton P, Park K J, Obayashi T, Fujita N, Harada H, Adams-Collier C J, Nakai K. 2007. WoLF PSORT: Protein localization predictor. Nucleic Acids Res, 35: W585-W587.
PMID |
| [9] | Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P. 2008. Genevestigator v3: A reference expression database for the meta-analysis of transcriptomes. Adv Bioinformatics, 2008: 420747. |
| [10] | Kazan K. 2017. The multitalented MEDIATOR25. Front Plant Sci, 8: 999. |
| [11] |
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones S J, Marra M A. 2009. Circos: An information aesthetic for comparative genomics. Genome Res, 19(9): 1639-1645.
PMID |
| [12] | Kumar S, Stecher G, Tamura K. 2016. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol, 33(7): 1870-1874. |
| [13] | Kumar V, Jain P, Venkadesan S, Karkute S G, Bhati J, Abdin M Z, Sevanthi A M, Mishra D C, Chaturvedi K K, Rai A, Sharma T R, Solanke A U. 2021. Understanding rice: Magnaporthe oryzae interaction in resistant and susceptible cultivars of rice under panicle blast infection using a time-course transcriptome analysis. Genes, 12(2): 301. |
| [14] | Li W T, Chern M, Yin J J, Wang J, Chen X W. 2019. Recent advances in broad-spectrum resistance to the rice blast disease. Curr Opin Plant Biol, 50: 114-120. |
| [15] | Liu J X, Jambunathan N, McNellis T W. 2005. Transgenic expression of the von Willebrand A domain of the BONZAI 1/ COPINE 1 protein triggers a lesion-mimic phenotype in Arabidopsis. Planta, 221(1): 85-94. |
| [16] | Liu Z, Wang T Z, Wang L, Zhao H, Yue E K, Yan Y, Irshad F, Zhou L, Duan M H, Xu J H. 2020. RTRIP: A comprehensive profile of transposon insertion polymorphisms in rice. Plant Biotechnol J, 18(12): 2379-2381. |
| [17] |
Livak K J, Schmittgen T D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods, 25(4): 402-408.
PMID |
| [18] | Polo S, Sigismund S, Faretta M, Guidi M, Capua M R, Bossi G, Chen H, de Camilli P, Di Fiore P P. 2002. A single motif responsible for ubiquitin recognition and monoubiquitination in endocytic proteins. Nature, 416: 451-455. |
| [19] | Rawat N, Naga N C, Meenakshi S R, Nair S, Bentur J S. 2012. A novel mechanism of gall midge resistance in the rice variety Kavya revealed by microarray analysis. Funct Integr Genomics, 12(2): 249-264. |
| [20] | Reininger A J. 2008. Function of von Willebrand factor in haemostasis and thrombosis. Haemophilia, 14: 11-26. |
| [21] | Sandhu M, Sureshkumar V, Prakash C, Dixit R, Solanke A U, Sharma T R, Mohapatra T, Sv A M. 2017. RiceMetaSys for salt and drought stress responsive genes in rice: A web interface for crop improvement. BMC Bioinformatics, 18(1): 432. |
| [22] | Sureshkumar V, Dutta B, Kumar V, Prakash G, Mishra D C, Chaturvedi K K, Rai A, Sevanthi A M, Solanke A U. 2019. RiceMetaSysB: A database of blast and bacterial blight responsive genes in rice and its utilization in identifying key blast-resistant WRKY genes. Database, 2019: baz015. |
| [23] | Swarbrick P J, Huang K, Liu G, Slate J, Press M C, Scholes J D. 2008. Global patterns of gene expression in rice cultivars undergoing a susceptible or resistant interaction with the parasitic plant Striga hermonthica. New Phytol, 179(2): 515-529. |
| [24] | Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller L A, Rhee S Y, Stitt M. 2004. MAPMAN: A user‐driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J, 37(6): 914-939. |
| [25] | Tuckwell D. 1999. Evolution of von Willebrand factor A (VWA) domains. Biochem Soc Trans, 27(6): 835-840. |
| [26] |
Voorrips R E. 2002. MapChart: Software for the graphical presentation of linkage maps and QTLs. J Hered, 93(1): 77-78.
PMID |
| [27] | Wang L, Zhao L N, Zhang X H, Zhang Q J, Jia Y X, Wang G, Li S M, Tian D C, Li W H, Yang S H. 2019. Large-scale identification and functional analysis of NLR genes in blast resistance in the Tetep rice genome sequence. Proc Natl Acad Sci USA, 116(37): 18479-18487. |
| [28] |
Whittaker C A, Hynes R O. 2002. Distribution and evolution of von Willebrand/integrin A domains: Widely dispersed domains with roles in cell adhesion and elsewhere. Mol Biol Cell, 13(10): 3369-3387.
PMID |
| [29] | Yang S H, Yang H J, Grisafi P, Sanchatjate S, Fink G R, Sun Q, Hua J. 2006. The BON/CPN gene family represses cell death and promotes cell growth in Arabidopsis. Plant J, 45(2): 166-179. |
| [30] | Yin X, Zou B H, Hong X X, Gao M J, Yang W B, Zhong X B, He Y, Kuai P, Lou Y G, Huang J R, Hua J, He Z H. 2018. Rice copine genes OsBON1 and OsBON3 function as suppressors of broad-spectrum disease resistance. Plant Biotechnol J, 16(8): 1476-1487. |
| [31] | Yu J, Wang J, Lin W, Li S G, Li H, Zhou J, Ni P X, Dong W, Hu S N, Zeng C Q, Zhang J G, Zhang Y, Li R Q, Xu Z Y, Li S T, Li X R, Zheng H K, Cong L J, Lin L, Yin J N, Geng J N, Li G Y, Shi J P, Liu J, Lv H, Li J, Wang J, Deng Y J, Ran L H, Shi X L, Wang X Y, Wu Q F, Li C F, Ren X Y, Wang J Q, Wang X L, Li D W, Liu D Y, Zhang X W, Ji Z D, Zhao W M, Sun Y Q, Zhang Z P, Bao J Y, Han Y J, Dong L L, Ji J, Chen P, Wu S M, Liu J S, Xiao Y, Bu D B, Tan J L, Yang L, Ye C, Zhang J F, Xu J Y, Zhou Y, Yu Y P, Zhang B, Zhuang S L, Wei H B, Liu B, Lei M, Yu H, Li Y Z, Xu H, Wei S L, He X M, Fang L J, Zhang Z J, Zhang Y Z, Huang X G, Su Z X, Tong W, Li J H, Tong Z Z, Li S L, Ye J, Wang L S, Fang L, Lei T T, Chen C, Chen H, Xu Z, Li H H, Huang H Y, Zhang F, Xu H Y, Li N, Zhao C F, Li S T, Dong L J, Huang Y Q, Li L, Xi Y, Qi Q H, Li W J, Zhang B, Hu W, Zhang Y L, Tian X J, Jiao Y Z, Liang X H, Jin J, Gao L, Zheng W M, Hao B L, Liu S Q, Wang W, Yuan L P, Cao M L, McDermott J, Samudrala R, Wang J, Wong G K S, Yang H M. 2005. The genomes of Oryza sativa: A history of duplications. PLoS Biol, 3(2): e38. |
| [1] | Li Shuyan, Yan Qiling, Wang Jieyu, Jiang Huidan, Li Zuren, Peng Qiong. Root Endophyte Shift and Key Genera Discovery in Rice under Barnyardgrass Stress [J]. Rice Science, 2023, 30(2): 160-170. |
| [2] | Qiu Diyang, Hu Rui, Li Ji, Li Ying, Ding Jierong, Xia Kuaifei, Zhong Xuhua, Fang Zhongming, Zhang Mingyong. Peptide Transporter OsNPF8.1 Contributes to Sustainable Growth under Salt and Drought Stresses, and Grain Yield under Nitrogen Deficiency in Rice [J]. Rice Science, 2023, 30(2): 113-126. |
| [3] | M. Iqbal R. Khan, Sarika Kumari, Faroza Nazir, Risheek Rahul Khanna, Ravi Gupta, Himanshu Chhillar. Defensive Role of Plant Hormones in Advancing Abiotic Stress-Resistant Rice Plants [J]. Rice Science, 2023, 30(1): 15-35. |
| [4] | Ruili Li, Jiaoling Wang, Lei Xu, Meihao Sun, Keke Yi, Hongyu Zhao. Functional Analysis of Phosphate Transporter OsPHT4 Family Members in Rice [J]. Rice Science, 2020, 27(6): 493-503. |
| [5] | Matías Romero Fernando, Gatica-Arias Andrés. CRISPR/Cas9: Development and Application in Rice Breeding [J]. Rice Science, 2019, 26(5): 265-281. |
| [6] | Chandran Jisha Kolothodi, Thomas Puthur Jos. Seed Priming with Beta-Amino Butyric Acid Improves Abiotic Stress Tolerance in Rice Seedlings [J]. Rice Science, 2016, 23(5): 242-254. |
| [7] | Challagulla Vineela, Bhattarai Surya, J. Midmore David. In-vitro vs in-vivo Inoculation: Screening for Resistance of Australian Rice Genotypes Against Blast Fungus [J]. Rice Science, 2015, 22(3): 132-137. |
| [8] | GAO Qing-song, ZHANG Dan, XU Liang, XU Chen-wu. Systematic Identification of Rice ABC1 Gene Family and Its Response to Abiotic Stress [J]. RICE SCIENCE, 2011, 18(3): 167-177. |
| [9] | Shigeto MORITA, Shinya NAKATANI, Tomokazu KOSHIBA, Takehiro MASUMURA, Yasunari OGIHARA, Kunisuke TANAKA. Differential Expression of Two Cytosolic Ascorbate Peroxidase and Two Superoxide Dismutase Genes in Response to Abiotic Stress in Rice [J]. RICE SCIENCE, 2011, 18(3): 157-166. |
| [10] | CHEN Xi-feng, GU Zhi-min, LIU Feng, MA Bo-jun, ZHANG Hong-sheng. Molecular Analysis of Rice CIPKs Involved in Both Biotic and Abiotic Stress Responses [J]. RICE SCIENCE, 2011, 18(1): 1-9 . |
| [11] | LIANG Wei-hong, BI Jia-jia, PENG Wei-feng, ZHANG Fan, SHI Hong-hao, LI Li. Cloning and Expression Analysis of a Mitogen-Activated Protein Kinase Gene OsMPK14 in Rice [J]. RICE SCIENCE, 2010, 17(4): 269-275 . |
| [12] | MA Hao-li, ZHOU Han-lin, ZHANG Huai-yu, ZHAO Jie. Cloning and Expression Analysis of an AP2/ERF Gene and Its Responses to Phytohormones and Abiotic Stresses in Rice [J]. RICE SCIENCE, 2010, 17(1): 1-9 . |
| [13] | MA Bo-jun, GU Zhi-min, TANG Hai-juan, CHEN Xi-feng, LIU Feng, ZHANG Hong-sheng . Preliminary Study on Function of Calcineurin B-Like Protein Gene OsCBL8 in Rice [J]. RICE SCIENCE, 2010, 17(1): 10-18 . |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||