
Rice Science ›› 2018, Vol. 25 ›› Issue (4): 208-217.DOI: 10.1016/j.rsci.2018.06.004
• Orginal Article • Previous Articles Next Articles
Zhiguo E1, Tingting Li1, Chen Chen2, Lei Wang1( )
)
Received:2017-03-23
Accepted:2018-05-03
Online:2018-06-20
Published:2018-04-10
Zhiguo E, Tingting Li, Chen Chen, Lei Wang. Genome-Wide Survey and Expression Analysis of P1B-ATPases in Rice, Maize and Sorghum[J]. Rice Science, 2018, 25(4): 208-217.
Add to citation manager EndNote|Ris|BibTeX
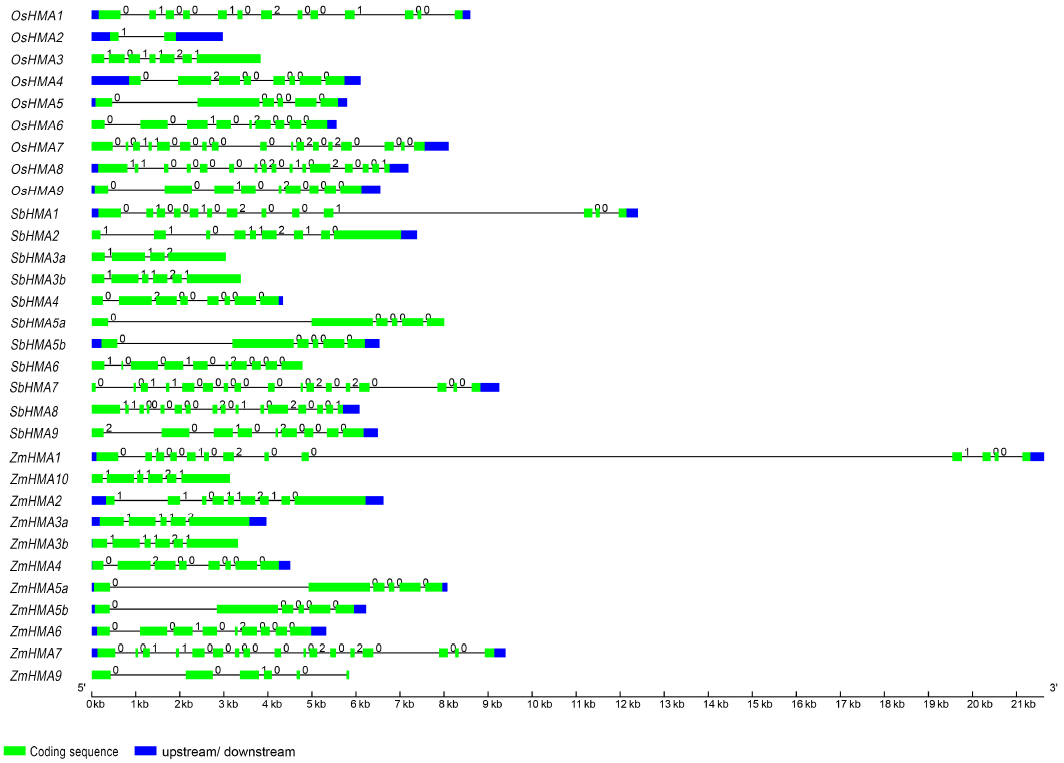
Fig. 1. Intron-exon structures of heavy metal P1B-ATPase (HMA) genes in rice, maize and sorghum.Names for the genes are on the left. The numbers 0, 1 and 2 indicate the phase of introns.

Fig. 2. Multiple sequence alignment of heavy metal P1B-ATPase (HMA) domains in rice, maize and sorghum.Identical and conservative amino acid residues are shaded in blue and light blue, respectively. The pink column below the alignment indicates the similarity of conserved amino acid residues of HMAs. The conserved motif is bordered (red rectangles), namely GMxCxxCE motif.
| Species | Chr | Gene name | Locus name a | No. of introns | Amino acid (aa) | TM b | Subcellular localizationc | Location (bp) | Strand |
|---|---|---|---|---|---|---|---|---|---|
| Oryza sativa (rice) | |||||||||
| 6 | OsHMA1 | LOC_Os06g47550 | 12 | 822 | 6 | Chloroplast | 28 797 545-28 789 015 | Reverse | |
| 6 | OsHMA2 | LOC_Os06g48720 | 8 | 1 067 | 6 | PM | 29 480 905-29 477 949 | Reverse | |
| 7 | OsHMA3 | LOC_Os07g12900 | 6 | 1 005 | 5 | Tonoplast | 7 409 553-7 405 745 | Reverse | |
| 2 | OsHMA4 | LOC_Os02g10290 | 8 | 979 | 8 | PM | 5 404 703-5 410 764 | Forward | |
| 4 | OsHMA5 | LOC_Os04g46940 | 5 | 1 003 | 8 | PM | 27 829 775-27 835 535 | Forward | |
| 2 | OsHMA6 | LOC_Os02g07630 | 8 | 1 013 | 7 | PM | 3 955 899-3 950 378 | Reverse | |
| 8 | OsHMA7 | LOC_Os08g37950 | 16 | 960 | 7 | Chloroplast | 24 031 272-24 039 316 | Forward | |
| 3 | OsHMA8 | LOC_Os03g08070 | 15 | 886 | 5 | Chloroplast | 4 126 933-4 119 945 | Reverse | |
| 6 | OsHMA9 | LOC_Os06g45500 | 8 | 1 004 | 7 | PM | 27 523 604-27 517 100 | Reverse | |
| Zea mays (maize) | |||||||||
| 5 | ZmHMA1 | GRMZM2G067853 | 12 | 823 | 6 | Chloroplast | 52 117 376-52 139 003 | Forward | |
| 5 | ZmHMA2 | GRMZM2G099191 | 8 | 1 099 | 4 | PM | 56 423 032-56 429 658 | Reverse | |
| 2 | ZmHMA3a | GRMZM2G175576 | 4 | 993 | 6 | Tonoplast | 158 406 991-158 410 957 | Reverse | |
| 2 | ZmHMA3b | GRMZM2G455491 | 5 | 927 | 5 | Tonoplast | 158 386 097-158 389 415 | Reverse | |
| 5 | ZmHMA4 | GRMZM2G029951 | 7 | 974 | 8 | PM | 121 407 958-121 412 467 | Reverse | |
| 2 | ZmHMA5a | GRMZM2G143512 | 5 | 999 | 8 | PM | 18 705 084-18 713 162 | Reverse | |
| 2 | ZmHMA5b | GRMZM2G144083 | 5 | 1 001 | 8 | PM | 18 620 426-18 626 656 | Reverse | |
| 4 | ZmHMA6 | GRMZM2G010152 | 8 | 998 | 7 | PM | 235 734 155-235 739 482 | Forward | |
| 4 | ZmHMA7 | GRMZM2G315931 | 16 | 928 | 7 | Chloroplast | 197 051 975-197 061 374 | Forward | |
| 9 | ZmHMA9 | GRMZM2G404702 | 5 | 597 | 5 | PM | 109 888 623-109 894 466 | Reverse | |
| 7 | ZmHMA10 | AC205008.4_FG002 | 5 | 883 | 5 | Tonoplast | 21 633 432-21 636 570 | Forward | |
| Sorghum bicolor (sorghum) | |||||||||
| 10 | SbHMA1 | Sb10g028080 | 12 | 828 | 6 | Chloroplast | 57 972 772-57 985 175 | Reverse | |
| 10 | SbHMA2 | Sb10g028920 | 8 | 1069 | 6 | PM | 58 715 561-58 722 945 | Reverse | |
| 2 | SbHMA3a | Sb02g006940 | 3 | 895 | 6 | Tonoplast | 8 908 023-8 911 069 | Forward | |
| 2 | SbHMA3b | Sb02g006950 | 5 | 933 | 8 | Tonoplast | 8 913 910-8 917 295 | Forward | |
| 4 | SbHMA4 | Sb04g006600 | 7 | 974 | 8 | PM | 6 611 424-6 615 768 | Forward | |
| 6 | SbHMA5a | Sb06g024900 | 5 | 1 002 | 8 | PM | 53 929 840-53 937 843 | Forward | |
| 6 | SbHMA5b | Sb06g024910 | 5 | 998 | 8 | PM | 53 941 501-53 948 034 | Forward | |
| 4 | SbHMA6 | Sb04g004820 | 9 | 1 011 | 8 | PM | 4 611 722-4 616 506 | Reverse | |
| 7 | SbHMA7 | Sb07g029010 | 16 | 817 | 6 | Chloroplast | 64 072 258-64 081 513 | Reverse | |
| 1 | SbHMA8 | Sb01g045340 | 15 | 877 | 6 | Chloroplast | 68 452 000-68 458 070 | Forward | |
| 10 | SbHMA9 | Sb10g026600 | 8 | 996 | 8 | PM | 56 024 169-56 030 667 | Reverse | |
| Chr, Chromosome; TM, Transmembrane structure; PM, Plasma membrane. a Locus identity number of HMAs assigned by Gramene; b Prediction of transmembrane helices in proteins using TMHMM Server v.2.0 (http://www.cbs.dtu.dk/services/TMHMM/); c Subcellular location prediction using TargetP 1.1 Server (http://www.cbs.dtu.dk/services/TargetP/). | |||||||||
Table 1 General information of 31 heavy metal P1B-ATPase (HMA) genes.
| Species | Chr | Gene name | Locus name a | No. of introns | Amino acid (aa) | TM b | Subcellular localizationc | Location (bp) | Strand |
|---|---|---|---|---|---|---|---|---|---|
| Oryza sativa (rice) | |||||||||
| 6 | OsHMA1 | LOC_Os06g47550 | 12 | 822 | 6 | Chloroplast | 28 797 545-28 789 015 | Reverse | |
| 6 | OsHMA2 | LOC_Os06g48720 | 8 | 1 067 | 6 | PM | 29 480 905-29 477 949 | Reverse | |
| 7 | OsHMA3 | LOC_Os07g12900 | 6 | 1 005 | 5 | Tonoplast | 7 409 553-7 405 745 | Reverse | |
| 2 | OsHMA4 | LOC_Os02g10290 | 8 | 979 | 8 | PM | 5 404 703-5 410 764 | Forward | |
| 4 | OsHMA5 | LOC_Os04g46940 | 5 | 1 003 | 8 | PM | 27 829 775-27 835 535 | Forward | |
| 2 | OsHMA6 | LOC_Os02g07630 | 8 | 1 013 | 7 | PM | 3 955 899-3 950 378 | Reverse | |
| 8 | OsHMA7 | LOC_Os08g37950 | 16 | 960 | 7 | Chloroplast | 24 031 272-24 039 316 | Forward | |
| 3 | OsHMA8 | LOC_Os03g08070 | 15 | 886 | 5 | Chloroplast | 4 126 933-4 119 945 | Reverse | |
| 6 | OsHMA9 | LOC_Os06g45500 | 8 | 1 004 | 7 | PM | 27 523 604-27 517 100 | Reverse | |
| Zea mays (maize) | |||||||||
| 5 | ZmHMA1 | GRMZM2G067853 | 12 | 823 | 6 | Chloroplast | 52 117 376-52 139 003 | Forward | |
| 5 | ZmHMA2 | GRMZM2G099191 | 8 | 1 099 | 4 | PM | 56 423 032-56 429 658 | Reverse | |
| 2 | ZmHMA3a | GRMZM2G175576 | 4 | 993 | 6 | Tonoplast | 158 406 991-158 410 957 | Reverse | |
| 2 | ZmHMA3b | GRMZM2G455491 | 5 | 927 | 5 | Tonoplast | 158 386 097-158 389 415 | Reverse | |
| 5 | ZmHMA4 | GRMZM2G029951 | 7 | 974 | 8 | PM | 121 407 958-121 412 467 | Reverse | |
| 2 | ZmHMA5a | GRMZM2G143512 | 5 | 999 | 8 | PM | 18 705 084-18 713 162 | Reverse | |
| 2 | ZmHMA5b | GRMZM2G144083 | 5 | 1 001 | 8 | PM | 18 620 426-18 626 656 | Reverse | |
| 4 | ZmHMA6 | GRMZM2G010152 | 8 | 998 | 7 | PM | 235 734 155-235 739 482 | Forward | |
| 4 | ZmHMA7 | GRMZM2G315931 | 16 | 928 | 7 | Chloroplast | 197 051 975-197 061 374 | Forward | |
| 9 | ZmHMA9 | GRMZM2G404702 | 5 | 597 | 5 | PM | 109 888 623-109 894 466 | Reverse | |
| 7 | ZmHMA10 | AC205008.4_FG002 | 5 | 883 | 5 | Tonoplast | 21 633 432-21 636 570 | Forward | |
| Sorghum bicolor (sorghum) | |||||||||
| 10 | SbHMA1 | Sb10g028080 | 12 | 828 | 6 | Chloroplast | 57 972 772-57 985 175 | Reverse | |
| 10 | SbHMA2 | Sb10g028920 | 8 | 1069 | 6 | PM | 58 715 561-58 722 945 | Reverse | |
| 2 | SbHMA3a | Sb02g006940 | 3 | 895 | 6 | Tonoplast | 8 908 023-8 911 069 | Forward | |
| 2 | SbHMA3b | Sb02g006950 | 5 | 933 | 8 | Tonoplast | 8 913 910-8 917 295 | Forward | |
| 4 | SbHMA4 | Sb04g006600 | 7 | 974 | 8 | PM | 6 611 424-6 615 768 | Forward | |
| 6 | SbHMA5a | Sb06g024900 | 5 | 1 002 | 8 | PM | 53 929 840-53 937 843 | Forward | |
| 6 | SbHMA5b | Sb06g024910 | 5 | 998 | 8 | PM | 53 941 501-53 948 034 | Forward | |
| 4 | SbHMA6 | Sb04g004820 | 9 | 1 011 | 8 | PM | 4 611 722-4 616 506 | Reverse | |
| 7 | SbHMA7 | Sb07g029010 | 16 | 817 | 6 | Chloroplast | 64 072 258-64 081 513 | Reverse | |
| 1 | SbHMA8 | Sb01g045340 | 15 | 877 | 6 | Chloroplast | 68 452 000-68 458 070 | Forward | |
| 10 | SbHMA9 | Sb10g026600 | 8 | 996 | 8 | PM | 56 024 169-56 030 667 | Reverse | |
| Chr, Chromosome; TM, Transmembrane structure; PM, Plasma membrane. a Locus identity number of HMAs assigned by Gramene; b Prediction of transmembrane helices in proteins using TMHMM Server v.2.0 (http://www.cbs.dtu.dk/services/TMHMM/); c Subcellular location prediction using TargetP 1.1 Server (http://www.cbs.dtu.dk/services/TargetP/). | |||||||||

Fig. 3. Phylogenetic relationship and functional domains of protein sequences of heavy metal P1B-ATPases (HMAs) in rice, maize and sorghum.A, Phylogenetic tree of 31 HMAs using maximum-likelihood methods. Scale bar represents 0.2 amino acid substitution per site. The proteins on the tree can be divided into two distinct subfamilies. The number in the line represents the goodness of fit (%). B, Functional domains of protein sequences of HMAs. Blue, green and red rectangles represent HMA domains, E1-E2_ATPase domains and hydrolase domains, respectively.
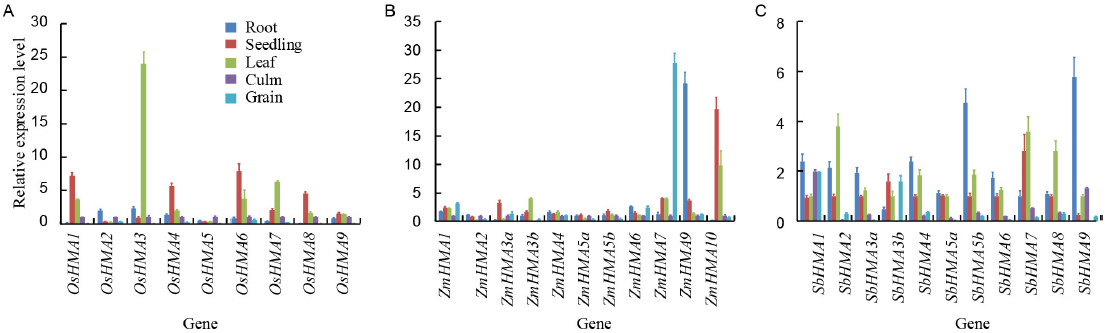
Fig. 4. Quantitative real-time PCR analysis of tissue-specific expression of heavy metal P1B-ATPase (HMA) genes in rice (A), maize (B) and sorghum (C). Relative mRNA expression levels of individual genes normalized to reference genes are shown, indicating their preferential expression in roots, seedlings, leafs, stems and grains. Error bars indicate standard deviations of three independent biological replicates.
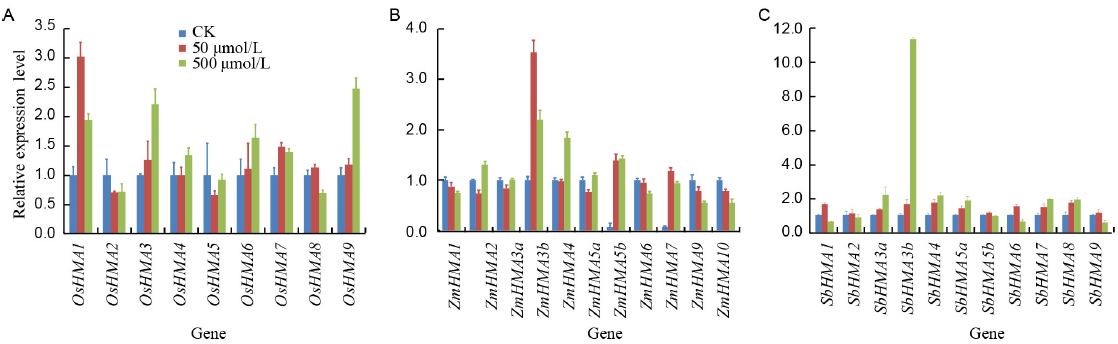
Fig. 5. Quantitative real-time PCR analysis of heavy metal P1B-ATPase (HMA) genes after Cu treatment in seedling in rice (A), maize (B) and sorghum (C).Relative mRNA expression levels of individual genes normalized to reference genes are shown, indicating their induced expression in seedlings treated with 50 μmol/L and 500 μmol/L Cu at 14 d after sowing. Error bars indicate standard deviations of three independent biological replicates.

Fig. 6. Quantitative real-time PCR analysis of heavy metal P1B-ATPase (HMA) genes after Cd treatment in seedling in rice (A), maize (B) and sorghum (C).Relative mRNA expression levels of individual genes normalized to reference genes are shown, indicating their induced expression in seedlings treated with 50 μmol/L and 500 μmol/L Cd at 14 d after sowing. Error bars indicate standard deviations of three independent biological replicates.
| [1] | Axelsen K B, Palmgren M G.2001. Inventory of the superfamily of P-type ion pumps inArabidopsis. Plant Physiol, 126(2): 696-706. |
| [2] | Deng F L, Yamaji N, Xia J X, Ma J F.2013. A member of the heavy metal P-type ATPase OsHMA5 is involved in xylem loading of copper in rice.Plant Physiol, 163(3): 1353-1362. |
| [3] | E Z G, Zhang Y P, Li T T, Wang L, Zhao H M.2015. Characterization of the ubiquitin-conjugating enzyme gene family in rice and evaluation of expression profiles under abiotic stresses and hormone treatments.PLoS One, 10: e0122621. |
| [4] | Huang X Y, Chao D Y, Gao J P, Zhu M Z, Shi M, Lin H X.2009. A previously unknown zinc finger protein, DST, regulates drought and salt tolerance in rice via stomatal aperture control.Genes Dev, 23: 1805-1817. |
| [5] | Huang X Y, Deng F L, Yamaji N, Pinson S R M, Fujii-Kashino M, Danku J, Douglas A, Guerinot M L, Salt D E, Ma J F.2016. A heavy metal P-type ATPase OsHMA4 prevents copper accumulation in rice grain.Nat Commun, 7: 12138. |
| [6] | Hussain D, Haydon M J, Wang Y, Wong E, Sherson S M, Young J, Camakaris J, Harper J F, Cobbett C S.2004. P-type ATPase heavy metal transporters with roles in essential zinc homeostasis inArabidopsis. Plant Cell, 16(5): 1327-1339. |
| [7] | Kim Y Y, Choi H, Segami S, Cho H T, Martinoia E, Maeshima M, Lee Y.2009. AtHMA1 contributes to the detoxification of excess Zn(II) inArabidopsis. Plant J, 58(5): 737-753. |
| [8] | Lee S, Kim Y Y, Lee Y, An G.2007. Rice P1B-type heavy-metal ATPase, OsHMA9, is a metal efflux protein.Plant Physiol, 145(3): 831-842. |
| [9] | Mikkelsen M D, Pedas P, Schiller M, Vincze E, Mills R F, Borg S, Møller A, Schjoerring J K, Williams L E, Baekgaard L, Holm P B, Palmgren M G.2012. Barley HvHMA1 is a heavy metal pump involved in mobilizing organellar Zn and Cu and plays a role in metal loading into grains.PLoS One, 7(11): e49027. |
| [10] | Mills R F, Krijger G C, Baccarini P J, Hall J L, Williams L E.2003. Functional expression of AtHMA4, a P1B-type ATPase of the Zn/Co/Cd/Pb subclass.Plant J, 35(2): 164-176. |
| [11] | Mills R F, Peaston K A, Runions J, Williams L E.2012. HvHMA2, a P1B-ATPase from barley, is highly conserved among cereals and functions in Zn and Cd transport.PLoS One, 7: e42640. |
| [12] | Miyadate H, Adachi S, Hiraizumi A, Tezuka K, Nakazawa N, Kawamoto T, Katou K, Kodama I, Sakurai K, Takahashi H, Satoh-Nagasawa N, Watanabe A, Fujimura T, Akagi H.2011. OsHMA3, a P1B-type of ATPase affects root-to-shoot cadmium translocation in rice by mediating efflux into vacuoles.New Phytol, 189(1): 190-199. |
| [13] | Penaud S, Fernandez A, Boudebbouze S, Ehrlich S D, Maguin E, van de Guchte M.2006. Induction of heavy-metal-transporting CPX-type ATPases during acid adaptation inLactobacillus bulgaricus. Appl Environ Microbiol, 72: 7445-7454. |
| [14] | Peng X X, Bai N N, Wang H H.2017. Isolation and expression profiles of cadmium stress-responsive rice WRKY15 transcription factor gene.Chin J Rice Sci, 32(2): 103-110. (in Chinese with English abstract) |
| [15] | Rascioa N, Navari-Izzob F.2011. Heavy metal hyperaccumulating plants: How and why do they do it? And what makes them so interesting?Plant Sci, 180(2): 169-181. |
| [16] | Satoh-Nagasawa N, Mori M, Nakazawa N, Kawamoto T, Nagato Y, Sakurai K, Takahashi H, Watanabe A, Akagi H.2012. Mutations in rice (Oryza sativa) heavy metal ATPase 2(OsHMA2) restrict the translocation of zinc and cadmium. Plant Cell Physiol, 53(1): 213-224. |
| [17] | Seigneurin-Berny D, Gravot A, Auroy P, Mazard C, Kraut A, Finazzi G, Grunwald D, Rappaport F, Vavasseur A, Joyard J, Richaud P, Rolland N.2005. HMA1, a new Cu-ATPase of the chloroplast envelope, is essential for growth under adverse light conditions.J Biol Chem, 281(5): 2882-2892. |
| [18] | Takahashi R, Bashir K, Ishimaru Y, Nishizawa N K, Nakanishi H.2012a. The role of heavy-metal ATPases, HMAs, in zinc and cadmium transport in rice.Plant Signal Behav, 7(12): 1605-1607. |
| [19] | Takahashi R, Ishimaru Y, Shimo H, Ogo Y, Senoura T, Nishizawa N K, Nakanishi H.2012b. TheOsHMA2 transporter is involved in root-to-shoot translocation of Zn and Cd in rice. Plant Cell Environ, 35(11): 1948-1957. |
| [20] | Tan J J, Wang J W, Chai T Y, Zhang Y X, Feng S S, Li Y, Zhao H J, Liu H M, Chai X P.2013. Functional analyses of TaHMA2, a P1B-type ATPase in wheat.Plant Biotechnol J, 11: 420-431. |
| [21] | Williams L E, Pittman J K, Hall J L.2000. Emerging mechanisms for heavy metal transport in plants.Biochim Biophys Acta, 1465(1/2): 104-126. |
| [22] | Williams L E, Mills R F.2005. P1B-ATPases: An ancient family of transition metal pumps with diverse functions in plants.Trends Plant Sci, 10: 491-502. |
| [23] | Xie D L, Cui J H, Chang J H.2013. Cloning and expression analysis ofSbDREB2 gene from Sorghum bicolor. Acta Agron Sin, 39: 1352-1359. (in Chinese with English abstract) |
| [24] | Xu F Y, Zhang M X, Zeng H Q, Zhu Y Y.2016. Involvement of plasma member H+-ATPase in uptake of nitrogen, phosphorus and potassium by rice root.Chin J Rice Sci, 30(1): 106-110. (in Chinese with English abstract) |
| [25] | Yamaji N, Xia J X, Mitani-Ueno N, Yokosho K, Ma J F.2013. Preferential delivery of zinc to developing tissues in rice is mediated by P-type heavy metal ATPase OsHMA2.Plant Physiol, 162(2): 927-939. |
| [26] | Zuo W L, Chao Q, Zhang N, Ye J R, Tan G Q, Li B L, Xing Y X, Zhang B Q, Liu H J, Fengler K A, Zhao J, Zhao X R, Chen Y S, Lai J S, Yan J B, Xu M L.2015. A maize wall-associated kinase confers quantitative resistance to head smut.Nat Genet, 47: 151-157. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||