
Rice Science ›› 2021, Vol. 28 ›› Issue (1): 58-68.DOI: 10.1016/j.rsci.2020.11.007
• Research Papers • Previous Articles Next Articles
Xun Xu1,2, Song Ge1,2, Fumin Zhang1,2( )
)
Received:2020-04-07
Accepted:2020-06-20
Online:2021-01-28
Published:2021-01-28
Xun Xu, Song Ge, Fumin Zhang. Genetic and Geographic Patterns of Duplicate DPL Genes Causing Genetic Incompatibility Within Rice: Implications for Multiple Domestication Events in Rice[J]. Rice Science, 2021, 28(1): 58-68.
Add to citation manager EndNote|Ris|BibTeX
| Species/Population/ Group | Number | Frequency (%) | S | π | h | Hd | Tajima’s D | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DPL1- | DPL2- | DPL1- | DPL2- | DPL1 | DPL2 | DPL1 | DPL2 | DPL1 | DPL2 | DPL1 | DPL2 | DPL1 | DPL2 | |||||||
| O. nivara | 91 | 0 | 37.0 | 0.0 | 41 | 12 | 0.00336 | 0.00090 | 23 | 9 | 0.819 | 0.267 | -1.73831 | -1.85460* | ||||||
| O. rufipogon | 16 | 18 | 5.0 | 4.9 | 79 | 34 | 0.00421 | 0.00290 | 53 | 25 | 0.914 | 0.429 | -2.19350** | -2.11384** | ||||||
| O. sativa | 15 | 49 | 10.2 | 30.4 | 11 | 6 | 0.00118 | 0.00363 | 7 | 4 | 0.223 | 0.571 | -1.21896 | 1.04607 | ||||||
| nIND | 18 | 0 | 45.0 | 0.0 | 12 | 5 | 0.00210 | 0.00140 | 6 | 5 | 0.638 | 0.410 | -1.18432 | -1.17921 | ||||||
| rIND | 2 | 0 | 11.8 | 0.0 | 17 | 8 | 0.00422 | 0.00292 | 9 | 7 | 0.955 | 0.584 | -1.31101 | -1.22445 | ||||||
| nTHA | 5 | 0 | 62.5 | 0.0 | 6 | 3 | 0.00370 | 0.00105 | 3 | 3 | 0.762 | 0.295 | 1.59761 | -1.65231 | ||||||
| rTHA | 2 | 0 | 13.3 | 0.0 | 12 | 3 | 0.00345 | 0.00110 | 5 | 3 | 0.813 | 0.362 | -1.08464 | -1.34917 | ||||||
| nKHM | 29 | 0 | 93.5 | 0.0 | 11 | 1 | 0.00202 | 0.00014 | 5 | 2 | 0.743 | 0.125 | -1.19393 | -1.14244 | ||||||
| rKHM | 6 | 0 | 22.2 | 0.0 | 6 | 0 | 0.00182 | 0.00000 | 5 | 4 | 0.667 | 0.200 | -0.48253 | NA | ||||||
| nMMR | 0 | 0 | 0.0 | 0.0 | 0 | 3 | 0.00000 | 0.00050 | 1 | 2 | 0.000 | 0.063 | NA | -1.56135 | ||||||
| rMMR | 0 | 0 | 0.0 | 0.0 | 22 | 10 | 0.00429 | 0.00361 | 9 | 1 | 0.714 | 0.000 | -1.52558 | -1.00632 | ||||||
| nNEP | 0 | 0 | 0.0 | 0.0 | 4 | 1 | 0.00083 | 0.00028 | 3 | 3 | 0.530 | 0.170 | -0.76705 | -0.77374 | ||||||
| rNEP | 2 | 0 | 7.1 | 0.0 | 5 | 3 | 0.00123 | 0.00071 | 3 | 7 | 0.574 | 0.696 | -0.51862 | -1.37016 | ||||||
| nLAO1 | 20 | 0 | 100.0 | 0.0 | 13 | 1 | 0.00341 | 0.00015 | 4 | 2 | 0.591 | 0.067 | 1.061730 | -1.14700 | ||||||
| rLAO1 | 0 | 0 | 0.0 | 0.0 | 12 | 13 | 0.00278 | 0.00240 | 8 | 5 | 0.837 | 0.261 | -1.04354 | -2.24274** | ||||||
| nLAO2 | 19 | 0 | 90.5 | 0.0 | 8 | 0 | 0.00142 | 0.00000 | 4 | 1 | 0.348 | 0.000 | -1.54473 | NA | ||||||
| rLAO2 | 1 | 0 | 3.4 | 0.0 | 13 | 2 | 0.00297 | 0.00102 | 7 | 2 | 0.817 | 0.228 | -0.78196 | -0.24788 | ||||||
| nLKA04 | 0 | 0 | 0.0 | 0.0 | 1 | 0 | 0.00023 | 0.00000 | 2 | 1 | 0.200 | 0.000 | -1.11173 | NA | ||||||
| nLKA05 | 0 | 0 | 0.0 | 0.0 | 3 | 0 | 0.00187 | 0.00000 | 2 | 1 | 0.533 | 0.000 | 1.83053* | NA | ||||||
| nLKA07 | 0 | 0 | 0.0 | 0.0 | 2 | 0 | 0.00047 | 0.00000 | 3 | 1 | 0.378 | 0.000 | -1.40085 | NA | ||||||
| nLKA08 | 0 | 0 | 0.0 | 0.0 | 1 | 0 | 0.00026 | 0.00000 | 2 | 1 | 0.222 | 0.000 | -1.88230 | NA | ||||||
| nLKA09 | 0 | 0 | 0.0 | 0.0 | 0 | 0 | 0.00000 | 0.00000 | 1 | 1 | 0.000 | 0.000 | NA | NA | ||||||
| nLKA10 | 0 | 0 | 0.0 | 0.0 | 0 | 3 | 0.00000 | 0.00198 | 1 | 3 | 0.000 | 0.600 | NA | -0.65748 | ||||||
| nLKA11 | 0 | 0 | 0.0 | 0.0 | 3 | 0 | 0.00098 | 0.00000 | 3 | 1 | 0.417 | 0.000 | -0.93613 | NA | ||||||
| nLKA12 | 0 | 0 | 0.0 | 0.0 | 4 | 2 | 0.00201 | 0.00091 | 2 | 1 | 0.429 | 0.000 | 0.48523 | -1.40085 | ||||||
| rLKA01 | 0 | 0 | 0.0 | 0.0 | 3 | 1 | 0.00195 | 0.00076 | 4 | 2 | 1.000 | 0.333 | 0.16766 | -0.93302 | ||||||
| rLKA06 | 0 | 0 | 0.0 | 0.0 | 1 | 0 | 0.00033 | 0.00000 | 1 | 1 | 0.000 | 0.000 | -1.00623 | NA | ||||||
| rLKA13 | 0 | 0 | 0.0 | 0.0 | 0 | 1 | 0.00000 | 0.00046 | 1 | 1 | 0.000 | 0.000 | NA | -1.11173 | ||||||
| rGDGZ | 0 | 10 | 0.0 | 47.6 | 8 | 4 | 0.00152 | 0.00354 | 7 | 3 | 0.569 | 0.648 | -1.48037 | 1.12924 | ||||||
| rGXBH | 0 | 0 | 0.0 | 0.0 | 6 | 2 | 0.00285 | 0.00198 | 2 | 2 | 1.000 | 0.400 | -0.49605 | -0.05002 | ||||||
| rGXHZ | 0 | 8 | 0.0 | 34.8 | 8 | 4 | 0.00213 | 0.00236 | 6 | 4 | 0.748 | 0.679 | -0.60490 | -0.12320 | ||||||
| rGXTD | 0 | 0 | 0.0 | 0.0 | 18 | 2 | 0.00440 | 0.00230 | 11 | 2 | 0.912 | 0.500 | -1.13215 | 1.81115 | ||||||
| rHNCL | 0 | 0 | 0.0 | 0.0 | 7 | 2 | 0.00235 | 0.00249 | 4 | 3 | 0.810 | 0.607 | -0.85010 | 1.82766 | ||||||
| rHNJY | 0 | 0 | 0.0 | 0.0 | 11 | 3 | 0.00407 | 0.00294 | 3 | 3 | 0.600 | 0.679 | -0.38658 | 0.77501 | ||||||
| rHNDZ | 0 | 0 | 0.0 | 0.0 | 8 | 4 | 0.00306 | 0.00370 | 2 | 3 | 0.476 | 0.733 | -0.51253 | 0.56555 | ||||||
| rHNWC | 0 | 0 | 0.0 | 0.0 | 11 | 5 | 0.00332 | 0.00332 | 7 | 4 | 0.944 | 0.750 | -0.23157 | -0.56682 | ||||||
| rJXDX | 0 | 0 | 0.0 | 0.0 | 8 | 2 | 0.00331 | 0.00213 | 6 | 2 | 0.952 | 0.533 | -0.25553 | 1.03299 | ||||||
| rYNJH | 0 | 0 | 0.0 | 0.0 | 5 | 1 | 0.00251 | 0.00046 | 5 | 2 | 0.806 | 0.200 | 0.28702 | -1.11173 | ||||||
| rIDN | 0 | 0 | 0.0 | 0.0 | 5 | 4 | 0.00304 | 0.00365 | 4 | 3 | 1.000 | 0.700 | 0.56199 | -1.09380 | ||||||
| rPNG | 3 | 0 | 60.0 | 0.0 | 5 | 2 | 0.00281 | 0.00178 | 3 | 2 | 0.700 | 0.400 | 0.00000 | 0.19590 | ||||||
| sARO | 0 | 1 | 0.0 | 25.0 | 0 | 4 | 0.00000 | 0.00456 | 1 | 2 | 0.000 | 0.500 | NA | -0.78012 | ||||||
| sAUS | 12 | 1 | 70.6 | 4.8 | 7 | 3 | 0.00286 | 0.00068 | 4 | 2 | 0.596 | 0.100 | 0.61839 | -1.72331 | ||||||
| sINDI | 2 | 5 | 3.0 | 7.5 | 5 | 5 | 0.00035 | 0.00135 | 2 | 3 | 0.060 | 0.252 | -1.64155 | -0.99668 | ||||||
| sRAY | 0 | 2 | 0.0 | 66.7 | 1 | 3 | 0.00078 | 0.00456 | 2 | 2 | 0.667 | 0.667 | NA | NA | ||||||
| sTEJ | 0 | 36 | 0.0 | 87.8 | 1 | 4 | 0.00007 | 0.00172 | 2 | 3 | 0.056 | 0.292 | -1.13321 | -0.46012 | ||||||
| sTRJ | 1 | 4 | 4.8 | 16.0 | 7 | 5 | 0.00078 | 0.00417 | 3 | 4 | 0.186 | 0.684 | -2.13123* | 1.11236 | ||||||
Table 1 Nucleotide polymorphism and results of neutral test in O. rufipogon, O. nivara and O. sativa.
| Species/Population/ Group | Number | Frequency (%) | S | π | h | Hd | Tajima’s D | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| DPL1- | DPL2- | DPL1- | DPL2- | DPL1 | DPL2 | DPL1 | DPL2 | DPL1 | DPL2 | DPL1 | DPL2 | DPL1 | DPL2 | |||||||
| O. nivara | 91 | 0 | 37.0 | 0.0 | 41 | 12 | 0.00336 | 0.00090 | 23 | 9 | 0.819 | 0.267 | -1.73831 | -1.85460* | ||||||
| O. rufipogon | 16 | 18 | 5.0 | 4.9 | 79 | 34 | 0.00421 | 0.00290 | 53 | 25 | 0.914 | 0.429 | -2.19350** | -2.11384** | ||||||
| O. sativa | 15 | 49 | 10.2 | 30.4 | 11 | 6 | 0.00118 | 0.00363 | 7 | 4 | 0.223 | 0.571 | -1.21896 | 1.04607 | ||||||
| nIND | 18 | 0 | 45.0 | 0.0 | 12 | 5 | 0.00210 | 0.00140 | 6 | 5 | 0.638 | 0.410 | -1.18432 | -1.17921 | ||||||
| rIND | 2 | 0 | 11.8 | 0.0 | 17 | 8 | 0.00422 | 0.00292 | 9 | 7 | 0.955 | 0.584 | -1.31101 | -1.22445 | ||||||
| nTHA | 5 | 0 | 62.5 | 0.0 | 6 | 3 | 0.00370 | 0.00105 | 3 | 3 | 0.762 | 0.295 | 1.59761 | -1.65231 | ||||||
| rTHA | 2 | 0 | 13.3 | 0.0 | 12 | 3 | 0.00345 | 0.00110 | 5 | 3 | 0.813 | 0.362 | -1.08464 | -1.34917 | ||||||
| nKHM | 29 | 0 | 93.5 | 0.0 | 11 | 1 | 0.00202 | 0.00014 | 5 | 2 | 0.743 | 0.125 | -1.19393 | -1.14244 | ||||||
| rKHM | 6 | 0 | 22.2 | 0.0 | 6 | 0 | 0.00182 | 0.00000 | 5 | 4 | 0.667 | 0.200 | -0.48253 | NA | ||||||
| nMMR | 0 | 0 | 0.0 | 0.0 | 0 | 3 | 0.00000 | 0.00050 | 1 | 2 | 0.000 | 0.063 | NA | -1.56135 | ||||||
| rMMR | 0 | 0 | 0.0 | 0.0 | 22 | 10 | 0.00429 | 0.00361 | 9 | 1 | 0.714 | 0.000 | -1.52558 | -1.00632 | ||||||
| nNEP | 0 | 0 | 0.0 | 0.0 | 4 | 1 | 0.00083 | 0.00028 | 3 | 3 | 0.530 | 0.170 | -0.76705 | -0.77374 | ||||||
| rNEP | 2 | 0 | 7.1 | 0.0 | 5 | 3 | 0.00123 | 0.00071 | 3 | 7 | 0.574 | 0.696 | -0.51862 | -1.37016 | ||||||
| nLAO1 | 20 | 0 | 100.0 | 0.0 | 13 | 1 | 0.00341 | 0.00015 | 4 | 2 | 0.591 | 0.067 | 1.061730 | -1.14700 | ||||||
| rLAO1 | 0 | 0 | 0.0 | 0.0 | 12 | 13 | 0.00278 | 0.00240 | 8 | 5 | 0.837 | 0.261 | -1.04354 | -2.24274** | ||||||
| nLAO2 | 19 | 0 | 90.5 | 0.0 | 8 | 0 | 0.00142 | 0.00000 | 4 | 1 | 0.348 | 0.000 | -1.54473 | NA | ||||||
| rLAO2 | 1 | 0 | 3.4 | 0.0 | 13 | 2 | 0.00297 | 0.00102 | 7 | 2 | 0.817 | 0.228 | -0.78196 | -0.24788 | ||||||
| nLKA04 | 0 | 0 | 0.0 | 0.0 | 1 | 0 | 0.00023 | 0.00000 | 2 | 1 | 0.200 | 0.000 | -1.11173 | NA | ||||||
| nLKA05 | 0 | 0 | 0.0 | 0.0 | 3 | 0 | 0.00187 | 0.00000 | 2 | 1 | 0.533 | 0.000 | 1.83053* | NA | ||||||
| nLKA07 | 0 | 0 | 0.0 | 0.0 | 2 | 0 | 0.00047 | 0.00000 | 3 | 1 | 0.378 | 0.000 | -1.40085 | NA | ||||||
| nLKA08 | 0 | 0 | 0.0 | 0.0 | 1 | 0 | 0.00026 | 0.00000 | 2 | 1 | 0.222 | 0.000 | -1.88230 | NA | ||||||
| nLKA09 | 0 | 0 | 0.0 | 0.0 | 0 | 0 | 0.00000 | 0.00000 | 1 | 1 | 0.000 | 0.000 | NA | NA | ||||||
| nLKA10 | 0 | 0 | 0.0 | 0.0 | 0 | 3 | 0.00000 | 0.00198 | 1 | 3 | 0.000 | 0.600 | NA | -0.65748 | ||||||
| nLKA11 | 0 | 0 | 0.0 | 0.0 | 3 | 0 | 0.00098 | 0.00000 | 3 | 1 | 0.417 | 0.000 | -0.93613 | NA | ||||||
| nLKA12 | 0 | 0 | 0.0 | 0.0 | 4 | 2 | 0.00201 | 0.00091 | 2 | 1 | 0.429 | 0.000 | 0.48523 | -1.40085 | ||||||
| rLKA01 | 0 | 0 | 0.0 | 0.0 | 3 | 1 | 0.00195 | 0.00076 | 4 | 2 | 1.000 | 0.333 | 0.16766 | -0.93302 | ||||||
| rLKA06 | 0 | 0 | 0.0 | 0.0 | 1 | 0 | 0.00033 | 0.00000 | 1 | 1 | 0.000 | 0.000 | -1.00623 | NA | ||||||
| rLKA13 | 0 | 0 | 0.0 | 0.0 | 0 | 1 | 0.00000 | 0.00046 | 1 | 1 | 0.000 | 0.000 | NA | -1.11173 | ||||||
| rGDGZ | 0 | 10 | 0.0 | 47.6 | 8 | 4 | 0.00152 | 0.00354 | 7 | 3 | 0.569 | 0.648 | -1.48037 | 1.12924 | ||||||
| rGXBH | 0 | 0 | 0.0 | 0.0 | 6 | 2 | 0.00285 | 0.00198 | 2 | 2 | 1.000 | 0.400 | -0.49605 | -0.05002 | ||||||
| rGXHZ | 0 | 8 | 0.0 | 34.8 | 8 | 4 | 0.00213 | 0.00236 | 6 | 4 | 0.748 | 0.679 | -0.60490 | -0.12320 | ||||||
| rGXTD | 0 | 0 | 0.0 | 0.0 | 18 | 2 | 0.00440 | 0.00230 | 11 | 2 | 0.912 | 0.500 | -1.13215 | 1.81115 | ||||||
| rHNCL | 0 | 0 | 0.0 | 0.0 | 7 | 2 | 0.00235 | 0.00249 | 4 | 3 | 0.810 | 0.607 | -0.85010 | 1.82766 | ||||||
| rHNJY | 0 | 0 | 0.0 | 0.0 | 11 | 3 | 0.00407 | 0.00294 | 3 | 3 | 0.600 | 0.679 | -0.38658 | 0.77501 | ||||||
| rHNDZ | 0 | 0 | 0.0 | 0.0 | 8 | 4 | 0.00306 | 0.00370 | 2 | 3 | 0.476 | 0.733 | -0.51253 | 0.56555 | ||||||
| rHNWC | 0 | 0 | 0.0 | 0.0 | 11 | 5 | 0.00332 | 0.00332 | 7 | 4 | 0.944 | 0.750 | -0.23157 | -0.56682 | ||||||
| rJXDX | 0 | 0 | 0.0 | 0.0 | 8 | 2 | 0.00331 | 0.00213 | 6 | 2 | 0.952 | 0.533 | -0.25553 | 1.03299 | ||||||
| rYNJH | 0 | 0 | 0.0 | 0.0 | 5 | 1 | 0.00251 | 0.00046 | 5 | 2 | 0.806 | 0.200 | 0.28702 | -1.11173 | ||||||
| rIDN | 0 | 0 | 0.0 | 0.0 | 5 | 4 | 0.00304 | 0.00365 | 4 | 3 | 1.000 | 0.700 | 0.56199 | -1.09380 | ||||||
| rPNG | 3 | 0 | 60.0 | 0.0 | 5 | 2 | 0.00281 | 0.00178 | 3 | 2 | 0.700 | 0.400 | 0.00000 | 0.19590 | ||||||
| sARO | 0 | 1 | 0.0 | 25.0 | 0 | 4 | 0.00000 | 0.00456 | 1 | 2 | 0.000 | 0.500 | NA | -0.78012 | ||||||
| sAUS | 12 | 1 | 70.6 | 4.8 | 7 | 3 | 0.00286 | 0.00068 | 4 | 2 | 0.596 | 0.100 | 0.61839 | -1.72331 | ||||||
| sINDI | 2 | 5 | 3.0 | 7.5 | 5 | 5 | 0.00035 | 0.00135 | 2 | 3 | 0.060 | 0.252 | -1.64155 | -0.99668 | ||||||
| sRAY | 0 | 2 | 0.0 | 66.7 | 1 | 3 | 0.00078 | 0.00456 | 2 | 2 | 0.667 | 0.667 | NA | NA | ||||||
| sTEJ | 0 | 36 | 0.0 | 87.8 | 1 | 4 | 0.00007 | 0.00172 | 2 | 3 | 0.056 | 0.292 | -1.13321 | -0.46012 | ||||||
| sTRJ | 1 | 4 | 4.8 | 16.0 | 7 | 5 | 0.00078 | 0.00417 | 3 | 4 | 0.186 | 0.684 | -2.13123* | 1.11236 | ||||||
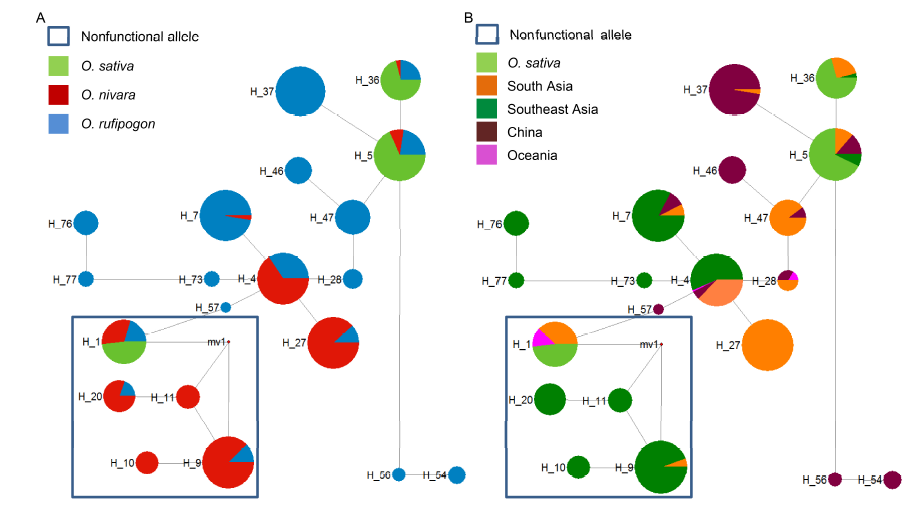
Fig. 1. Median-joining haplotype networks of DPL1.Sections of each circle represent the proportion species (A) or locations (B) present in each haplotype. The size of each circle corresponds to the number of sequences of each haplotype. The lengths of the gray lines correspond to the number of mutations. Nonfunctional haplotypes are indicated by hollow squares.
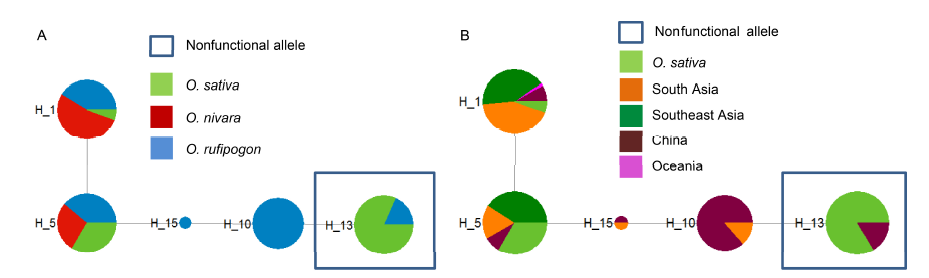
Fig. 2. Median-joining haplotype networks of DPL2.Sections of each circle represent the proportion species (A) or locations (B) present in each haplotype. The size of each circle corresponds to the number of sequences of each haplotype. The lengths of the gray lines correspond to the number of mutations. Nonfunctional haplotypes are indicated by hollow squares.
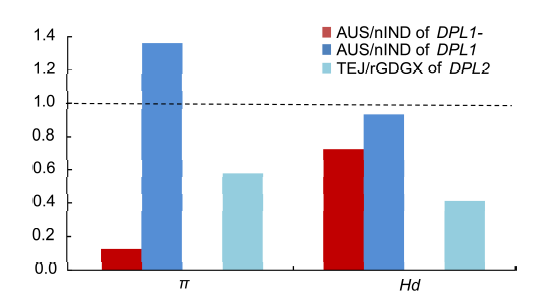
Fig. 3. Ratios of nucleotide diversity (π) and haplotype diversity (Hd) between groups of O. sativa and ancestral populations.AUS and TEJ indicate aus and temperate japonica rice, respectively. nIND and rGDGX indicate the ancestral populations: O. nivara from India and the two O. rufipogon populations carrying DPL2- from South China. The ratio of DPL2- nucleotide diversity between temperate japonica and the ancestral populations is represented by a blank shape, since DPL2- of temperate japonica has no diversity.
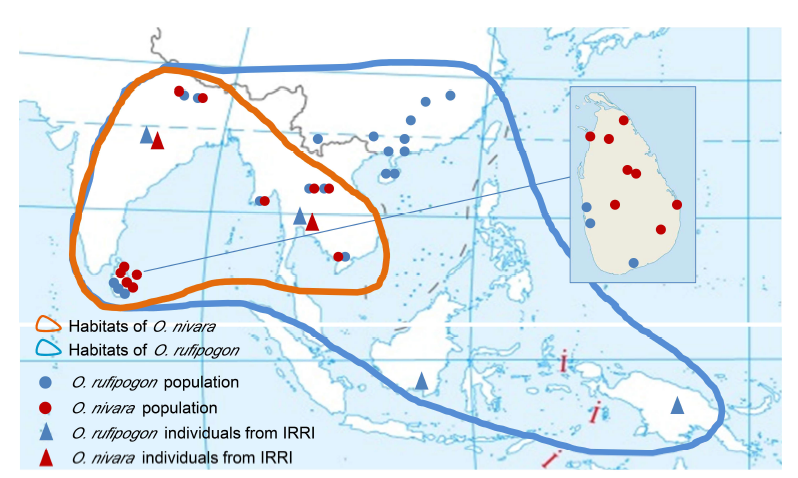
Fig. 4. Distribution of wild rice samplings, as well as functional and nonfunctional alleles of DPLs.Populations of O. nivara and O. rufipogon are indicated with spots. Triangles indicate individual samples from the International Rice Research Institute (IRRI). Red and blue lines indicate O. nivara and O. rufipogon, respectively. Circles indicate the frequencies of functional and nonfunctional alleles.
| [1] | Bandelt H J, Forster P, Rohl A. 1999. Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol, 16(1): 37-48 |
| [2] | Bank C, Burger R, Hermisson J. 2012. The limits to parapatric speciation: Dobzhansky-Muller incompatibilities in a Continent- Island Model. Genetics, 191(3): 845-345 |
| [3] | Cai Z, Zhou L, Ren N N, Xu X, Liu R, Huang L, Zheng X M, Meng Q L, Du Y S, Wang M X, Geng M F, Chen W L, Jing C Y, Zou X H, Guo J, Chen C B, Zeng H Z, Liang Y T, Wei X H, Guo Y L, Zhou H F, Zhang F M, Ge S. 2019. Parallel speciation of wild rice associated with habitat shifts. Mol Biol Evol, 36(5): 875-889 |
| [4] | Chen J J, Ding J H, Ouyang Y D, Du H Y, Yang J Y, Cheng K, Zhao J, Qiu S Q, Zhang X L, Yao J L, Liu K D, Wang L, Xu C G, Li X H, Xue Y B, Xia M, Ji Q, Lu J F, Xu M L, Zhang Q F. 2008. A triallelic system of S5 is a major regulator of the reproductive barrier and compatibility of indica-japonica hybrids in rice. Proc Natl Acad Sci USA, 105(32): 11436-11441 |
| [5] | Civan P, Craig H, Cox C J, Brown T A. 2015. Three geographically separate domestications of Asian rice. Nat Plants, 1(11): 1-5 |
| [6] | Corbett-Detig R B, Zhou J, Clark A G, Hartl D L, Ayroles J F. 2013. Genetic incompatibilities are widespread within species. Nature, 504: 135-137 |
| [7] | Craig S M, Reagon M, Resnick L E, Caicedo A L. 2014. Allele distributions at hybrid incompatibility loci facilitate the potential for gene flow between cultivated and weedy rice in the US. PLoS One, 9(1): e86647. |
| [8] | Cutter A D. 2012. The polymorphic prelude to Bateson-Dobzhansky- Muller incompatibilities. Trends Ecol Evol, 27(4): 209-218 |
| [9] | Du H, Ouyang Y, Zhang C, Zhang Q. 2011. Complex evolution of S5, a major reproductive barrier regulator, in the cultivated rice Oryza sativa and its wild relatives. New Phytol, 191(1): 275-287 |
| [10] | Garris A J, Tai T H, Coburn J, Kresovich S, McCouch S. 2005. Genetic structure and diversity in Oryza sativa L. Genetics, 169(3): 1631-1638 |
| [11] | Gavrilets S. 1997. Hybrid zones with Dobzhansky-type epistatic selection. Evolution, 51(4): 1027-1035 |
| [12] | Hall T A. 1999. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser, 41: 95-98 |
| [13] | Huang X H, Kurata N, Wei X H, Wang Z X, Wang A, Zhao Q, Zhao Y, Liu K Y, Lu H Y, Li W J, Guo Y L, Lu Y Q, Zhou C C, Fan D L, Weng Q J, Zhu C R, Huang T, Zhang L, Wang Y C, Feng L, Furuumi H, Kubo T, Miyabayashi T, Yuan X P, Xu Q, Dong G J, Zhan Q L, Li C Y, Fujiyama A, Toyoda A, Lu T, Feng Q, Qian Q, Li J Y, Han B. 2012. A map of rice genome variation reveals the origin of cultivated rice. Nature, 490: 497-501 |
| [14] | Khush G S. 1997. Origin, dispersal, cultivation and variation of rice. Plant Mol Biol, 35(1): 25-34 |
| [15] | Kovach M J, Sweeney M T, McCouch S R. 2007. New insights into the history of rice domestication. Trends Genet, 23(11): 578-587 |
| [16] | Li G W, Li X T, Wang Y, Mi J M, Xing F, Zhang D H, Dong Q Y, Li X H, Xiao J H, Zhang Q F, Ouyang Y D. 2017. Three representative inter and intra-subspecific crosses reveal the genetic architecture of reproductive isolation in rice. Plant J, 92(3): 349-362 |
| [17] | Librado P, Rozas J. 2009. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25(11): 1451-1452 |
| [18] | Lindtke D, Buerkle C A.2015. The genetic architecture of hybrid incompatibilities and their effect on barriers to introgression in secondary contact. Evolution, 69(8): 1987‒2004. |
| [19] | Liu R, Zheng X M, Zhou L, Zhou H F, Ge S. 2015. Population genetic structure of Oryza rufipogon and Oryza nivara: Implications for the origin of O. nivara. Mol Ecol, 24(20): 5211-5228 |
| [20] | Londo J P, Chiang Y C, Hung K H, Chiang T Y, Schaal B A. 2006. Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa. Proc Natl Acad Sci USA, 103(25): 9578-9583 |
| [21] | Long Y M, Zhao L F, Niu B X, Su J, Wu H, Chen Y L, Zhang Q Y, Guo J X, Zhuang C X, Mei M T, Xia J X, Wang L, Wu H B, Liu Y G. 2008. Hybrid male sterility in rice controlled by interaction between divergent alleles of two adjacent genes. Proc Natl Acad Sci USA, 105(48): 18871-18876 |
| [22] | Mizuta Y, Harushima Y, Kurata N. 2010. Rice pollen hybrid incompatibility caused by reciprocal gene loss of duplicated genes. Proc Natl Acad Sci USA, 107(47): 20417-20422 |
| [23] | Morishima H, Sano Y, Oka H.1992. Evolutionary Studies in Cultivated Rice and Its Wild Relatives. Oxford: Oxford University Press. |
| [24] | Oka H I. 1988. Origin of Cultivated Rice. Amsterdam, Holland: Elsevier. |
| [25] | Ouyang Y D, Liu Y G, Zhang Q F. 2010. Hybrid sterility in plant: Stories from rice. Curr Opin Plant Biol, 13(2): 186-192 |
| [26] | Ouyang Y D, Zhang Q F. 2013. Understanding reproductive isolation based on the rice model. Annu Rev Plant Biol, 64(1): 111-135 |
| [27] | Ouyang Y D, Zhang Q F. 2018. The molecular and evolutionary basis of reproductive isolation in plants. J Genet Genom, 45(11): 613-620 |
| [28] | Polzin T, Daneshmand S V. 2003. On Steiner trees and minimum spanning trees in hypergraphs. Oper Res Lett, 31(1): 12-20 |
| [29] | Rieseberg L H, Blackman B K. 2010. Speciation genes in plants. Ann Bot, 106(3): 439-455 |
| [30] | Sang T, Ge S. 2007. The puzzle of rice domestication. J Integr Plant Biol, 49(6): 760-768 |
| [31] | Seidel H S, Rockman M V, Kruglyak L. 2008. Widespread genetic incompatibility in C. elegans maintained by balancing selection. Science, 319: 589-594 |
| [32] | Shen R X, Wang L, Liu X P, Wu J, Jin W W, Zhao X C, Xie X R, Zhu Q L, Tang H W, Li Q, Chen L T, Liu Y G. 2017. Genomic structural variation-mediated allelic suppression causes hybrid male sterility in rice. Nat Commun, 8: 1310. |
| [33] | Shuker D M, Underwood K, King T M, Butlin R K. 2005. Patterns of male sterility in a grasshopper hybrid zone imply accumulation of hybrid incompatibilities without selection. Proc R Soc Lond Ser B-Biol Sci, 272: 2491-2497 |
| [34] | Sicard A, Kappel C, Josephs E B, Lee Y W, Marona C, Stinchcombe J R, Wright S I, Lenhard M. 2015. Divergent sorting of a balanced ancestral polymorphism underlies the establishment of gene-flow barriers inCapsella. Nat Commun, 6: 7960. |
| [35] | Sweeney M, McCouch S. 2007. The complex history of the domestication of rice. Ann Bot, 100(5): 951-957 |
| [36] | Tajima F. 1989. Statistical-method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics, 123(3): 585-595 |
| [37] | Thompson J D, Gibson T J, Plewniak F, Jeanmougin F, Higgins D G. 1997. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl Acids Res, 25(24): 4876-4882 |
| [38] | Vaughan D A, Lu B R, Tomooka N. 2008. The evolving story of rice evolution. Plant Sci, 174(4): 394-408 |
| [39] | Wang C H, Zheng X M, Xu Q, Yuan X P, Huang L, Zhou H F, Wei X H, Ge S. 2014. Genetic diversity and classification of Oryza sativa with emphasis on Chinese rice germplasm. Heredity, 112: 489-496 |
| [40] | Woerner A E, Cox M P, Hammer M F. 2007. Recombination- filtered genomic datasets by information maximization. Bioinformatics, 23(14): 1851-1853 |
| [41] | Xie Y Y, Shen R X, Chen L T, Liu Y G. 2019. Molecular mechanisms of hybrid sterility in rice. Sci China Life Sci, 62(6): 737-743 |
| [42] | Yamagata Y, Yamamoto E, Aya K, Win K T, Doi K, Sobrizal , Ito T, Kanamori H, Wu J, Matsumoto T, Matsuoka M, Ashikari M, Yoshimura A. 2010. Mitochondrial gene in the nuclear genome induces reproductive barrier in rice. Proc Natl Acad Sci USA, 107(4): 1494‒1499. |
| [43] | Yang C C, Kawahara Y, Mizuno H, Wu J Z, Matsumoto T, Itoh T. 2012. Independent domestication of Asian rice followed by gene flow from japonica to indica. Mol Biol Evol, 29(5): 1471-1479 |
| [44] | Yang J Y, Zhao X B, Cheng K, Du H Y, Ouyang Y D, Chen J J, Qiu S Q, Huang J Y, Jiang Y H, Jiang L W, Ding J H, Wang J, Xu C G, Li X H, Zhang Q F. 2012. A killer-protector system regulates both hybrid sterility and segregation distortion in rice. Science, 337: 1336-1340 |
| [45] | Zheng X M, Ge S. 2010. Ecological divergence in the presence of gene flow in two closely related Oryza species (Oryza rufipogon and O. nivara). Mol Ecol, 19(12): 2439-2454 |
| [46] | Zuellig M P, Sweigart A L. 2018. A two-locus hybrid incompatibility is widespread, polymorphic, and active in natural populations of Mimulus. Evolution, 72(11): 2394-2405 |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||