
Rice Science ›› 2021, Vol. 28 ›› Issue (4): 368-378.DOI: 10.1016/j.rsci.2021.05.007
• Research Papers • Previous Articles Next Articles
Fuad Anshori Muhammad1,2, Sapta Purwoko Bambang3( ), Saraswati Dewi Iswari4, Wahyuning Ardie Sintho3, Bayuardi Suwarno Willy3
), Saraswati Dewi Iswari4, Wahyuning Ardie Sintho3, Bayuardi Suwarno Willy3
Received:2020-05-21
Accepted:2021-01-04
Online:2021-07-28
Published:2021-07-28
Fuad Anshori Muhammad, Sapta Purwoko Bambang, Saraswati Dewi Iswari, Wahyuning Ardie Sintho, Bayuardi Suwarno Willy. A New Approach to Select Doubled Haploid Rice Lines under Salinity Stress Using Indirect Selection Index[J]. Rice Science, 2021, 28(4): 368-378.
Add to citation manager EndNote|Ris|BibTeX
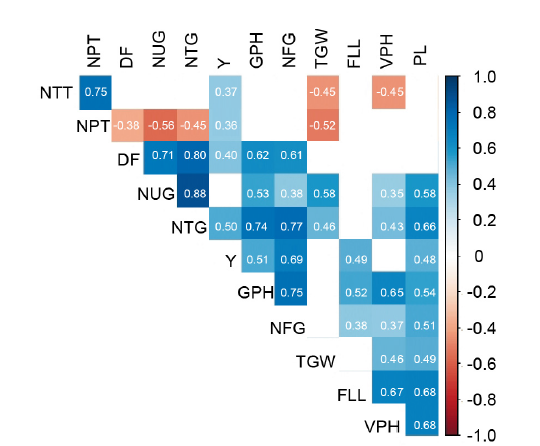
Fig. 1. Pearson correlations among all observed characters of double hybrid rice lines. NTT, Number of total tillers; NPT, Number of productive tillers; DF, Days to flowering; NUG, Number of unfilled grains per hill; NTG, Number of total grains per hill; Y, Yield; GPH, Generative plant height; NFG, Number of filled grains per panicle; TGW, 1000-grain weight; FLL, Flag leaf length; VPH, Vegetative plant height; PL, Panicle length.Significance was focused on the character Y. Color box (|r| > 0.34) indicates significant correlation at P < 0.01; Blank box, Not significant.
| Character | NTT | NPT | GPH | DF | FLL | PL | NFG | NTG |
|---|---|---|---|---|---|---|---|---|
| NTT | 0.02 | 0.39 | 0.01 | 0.00 | 0.00 | -0.07 | 0.00 | 0.01 |
| NPT | 0.02 | 0.52 | 0.03 | -0.14 | 0.02 | -0.06 | -0.06 | 0.02 |
| GPH | 0.00 | -0.12 | -0.13 | 0.23 | 0.06 | 0.16 | 0.36 | -0.04 |
| DF | 0.00 | -0.20 | -0.08 | 0.37 | 0.00 | 0.06 | 0.29 | -0.04 |
| FLL | 0.00 | 0.09 | -0.07 | -0.01 | 0.11 | 0.20 | 0.18 | -0.02 |
| PL | -0.01 | -0.10 | -0.07 | 0.08 | 0.07 | 0.29 | 0.24 | -0.03 |
| NFG | 0.00 | -0.06 | -0.10 | 0.22 | 0.04 | 0.15 | 0.47 | -0.04 |
| NTG | 0.00 | -0.24 | -0.09 | 0.29 | 0.03 | 0.19 | 0.36 | -0.05 |
Table S1. Path analysis of several characters of double haploid rice lines to the yield.
| Character | NTT | NPT | GPH | DF | FLL | PL | NFG | NTG |
|---|---|---|---|---|---|---|---|---|
| NTT | 0.02 | 0.39 | 0.01 | 0.00 | 0.00 | -0.07 | 0.00 | 0.01 |
| NPT | 0.02 | 0.52 | 0.03 | -0.14 | 0.02 | -0.06 | -0.06 | 0.02 |
| GPH | 0.00 | -0.12 | -0.13 | 0.23 | 0.06 | 0.16 | 0.36 | -0.04 |
| DF | 0.00 | -0.20 | -0.08 | 0.37 | 0.00 | 0.06 | 0.29 | -0.04 |
| FLL | 0.00 | 0.09 | -0.07 | -0.01 | 0.11 | 0.20 | 0.18 | -0.02 |
| PL | -0.01 | -0.10 | -0.07 | 0.08 | 0.07 | 0.29 | 0.24 | -0.03 |
| NFG | 0.00 | -0.06 | -0.10 | 0.22 | 0.04 | 0.15 | 0.47 | -0.04 |
| NTG | 0.00 | -0.24 | -0.09 | 0.29 | 0.03 | 0.19 | 0.36 | -0.05 |
| Predictors | Estimate | Standard Error | T-value | Probability (>|t|) |
|---|---|---|---|---|
| Intercept | -531.13 | 99.96 | -5.313 | 1.93e-06*** |
| Number of productive tillers | 32.64 | 4.26 | 7.659 | 2.82e-10*** |
| Number of filled grain | 3.09 | 0.87 | 3.553 | 0.000781*** |
| Number of total grain | 1.86 | 0.49 | 3.803 | 0.000355*** |
Table S2. Stepwise regression results of the yield of DH rice lines.
| Predictors | Estimate | Standard Error | T-value | Probability (>|t|) |
|---|---|---|---|---|
| Intercept | -531.13 | 99.96 | -5.313 | 1.93e-06*** |
| Number of productive tillers | 32.64 | 4.26 | 7.659 | 2.82e-10*** |
| Number of filled grain | 3.09 | 0.87 | 3.553 | 0.000781*** |
| Number of total grain | 1.86 | 0.49 | 3.803 | 0.000355*** |
| Parameter | PC1 | PC2 | PC3 | PC4 | PC5 | PC6 | PC7 | PC8 | PC9 | PC10 |
|---|---|---|---|---|---|---|---|---|---|---|
| VPH | 0.275 | -0.031 | 0.481 | -0.207 | 0.196 | 0.288 | -0.007 | -0.403 | 0.302 | -0.200 |
| NTT | -0.151 | -0.404 | -0.290 | 0.330 | 0.107 | 0.479 | 0.501 | 0.161 | 0.208 | -0.233 |
| NPT | -0.224 | -0.433 | 0.026 | 0.316 | 0.003 | -0.031 | -0.175 | -0.434 | -0.581 | 0.142 |
| GPH | 0.328 | -0.198 | -0.019 | -0.322 | 0.259 | 0.458 | 0.025 | -0.156 | -0.258 | 0.295 |
| DF | 0.280 | -0.026 | -0.509 | -0.125 | 0.127 | 0.075 | -0.220 | 0.240 | 0.077 | 0.473 |
| FLL | 0.190 | -0.329 | 0.422 | 0.168 | 0.314 | -0.069 | -0.291 | 0.647 | -0.145 | -0.147 |
| PL | 0.322 | -0.118 | 0.290 | 0.255 | -0.055 | -0.367 | 0.565 | -0.026 | 0.111 | 0.514 |
| TGW | 0.248 | 0.248 | 0.190 | 0.228 | -0.668 | 0.483 | -0.033 | 0.188 | -0.247 | 0.053 |
| NFG | 0.276 | -0.310 | -0.145 | -0.386 | -0.287 | -0.242 | 0.204 | 0.030 | -0.243 | -0.338 |
| NUG | 0.355 | 0.181 | -0.195 | 0.262 | 0.150 | -0.073 | -0.014 | -0.106 | -0.053 | -0.230 |
| NTG | 0.388 | -0.030 | -0.209 | -0.014 | -0.040 | -0.174 | 0.093 | -0.060 | -0.166 | -0.328 |
| Yield | 0.180 | -0.465 | -0.084 | 0.171 | -0.376 | -0.065 | -0.446 | -0.181 | 0.521 | 0.026 |
| SD | 2.44 | 1.68 | 1.32 | 0.98 | 0.75 | 0.61 | 0.44 | 0.39 | 0.31 | 0.29 |
| Proportion cumulative | 0.46 | 0.67 | 0.81 | 0.88 | 0.93 | 0.95 | 0.97 | 0.98 | 0.99 | 1.00 |
| Eigenvalue | 5.94 | 2.83 | 1.75 | 0.96 | 0.56 | 0.37 | 0.19 | 0.15 | 0.10 | 0.09 |
Table 1 Ten principal components derived from 12 characters of doubled haploid rice lines.
| Parameter | PC1 | PC2 | PC3 | PC4 | PC5 | PC6 | PC7 | PC8 | PC9 | PC10 |
|---|---|---|---|---|---|---|---|---|---|---|
| VPH | 0.275 | -0.031 | 0.481 | -0.207 | 0.196 | 0.288 | -0.007 | -0.403 | 0.302 | -0.200 |
| NTT | -0.151 | -0.404 | -0.290 | 0.330 | 0.107 | 0.479 | 0.501 | 0.161 | 0.208 | -0.233 |
| NPT | -0.224 | -0.433 | 0.026 | 0.316 | 0.003 | -0.031 | -0.175 | -0.434 | -0.581 | 0.142 |
| GPH | 0.328 | -0.198 | -0.019 | -0.322 | 0.259 | 0.458 | 0.025 | -0.156 | -0.258 | 0.295 |
| DF | 0.280 | -0.026 | -0.509 | -0.125 | 0.127 | 0.075 | -0.220 | 0.240 | 0.077 | 0.473 |
| FLL | 0.190 | -0.329 | 0.422 | 0.168 | 0.314 | -0.069 | -0.291 | 0.647 | -0.145 | -0.147 |
| PL | 0.322 | -0.118 | 0.290 | 0.255 | -0.055 | -0.367 | 0.565 | -0.026 | 0.111 | 0.514 |
| TGW | 0.248 | 0.248 | 0.190 | 0.228 | -0.668 | 0.483 | -0.033 | 0.188 | -0.247 | 0.053 |
| NFG | 0.276 | -0.310 | -0.145 | -0.386 | -0.287 | -0.242 | 0.204 | 0.030 | -0.243 | -0.338 |
| NUG | 0.355 | 0.181 | -0.195 | 0.262 | 0.150 | -0.073 | -0.014 | -0.106 | -0.053 | -0.230 |
| NTG | 0.388 | -0.030 | -0.209 | -0.014 | -0.040 | -0.174 | 0.093 | -0.060 | -0.166 | -0.328 |
| Yield | 0.180 | -0.465 | -0.084 | 0.171 | -0.376 | -0.065 | -0.446 | -0.181 | 0.521 | 0.026 |
| SD | 2.44 | 1.68 | 1.32 | 0.98 | 0.75 | 0.61 | 0.44 | 0.39 | 0.31 | 0.29 |
| Proportion cumulative | 0.46 | 0.67 | 0.81 | 0.88 | 0.93 | 0.95 | 0.97 | 0.98 | 0.99 | 1.00 |
| Eigenvalue | 5.94 | 2.83 | 1.75 | 0.96 | 0.56 | 0.37 | 0.19 | 0.15 | 0.10 | 0.09 |
| Direct Selection | Selection Indices | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Rank | Genotype | Y (ton ha-1) | Z yield | Rank | Genotype | NPT | NFG | Y (ton ha-1) | Z Index | |
| 1 | F47 | 6.00 | 1.87 | 1 | F51 | 19.9 | 130.7 | 6.00 | 1.73 | |
| 2 | F51 | 6.00 | 1.87 | 2 | F41 | 20.9 | 121.2 | 5.97 | 1.72 | |
| 3 | F41 | 5.97 | 1.83 | 3 | F40 | 21.2 | 123.8 | 5.52 | 1.56 | |
| 4 | F43 | 5.91 | 1.76 | 4 | F47 | 20.0 | 116.4 | 6.00 | 1.45 | |
| 5 | F44 | 5.67 | 1.44 | 5 | F44 | 19.2 | 130.2 | 5.67 | 1.37 | |
| 6 | F49 | 5.65 | 1.42 | 6 | F43 | 19.2 | 122.2 | 5.91 | 1.35 | |
| 7 | F46 | 5.59 | 1.34 | 7 | F46 | 20.5 | 118.3 | 5.59 | 1.35 | |
| 8 | F40 | 5.52 | 1.25 | 8 | F38 | 20.0 | 126.0 | 5.37 | 1.27 | |
| 9 | F22 | 5.49 | 1.21 | 9 | F22 | 19.7 | 123.3 | 5.49 | 1.23 | |
| 10 | F38 | 5.37 | 1.05 | 10 | F32 | 20.1 | 125.2 | 5.26 | 1.21 | |
| 11 | F37 | 5.35 | 1.03 | 11 | F39 | 19.7 | 129.7 | 5.24 | 1.20 | |
| 12 | F36 | 5.31 | 0.98 | 12 | F49 | 20.4 | 110.4 | 5.65 | 1.20 | |
| 13 | F19 | 5.30 | 0.96 | 13 | F37 | 20.9 | 108.5 | 5.35 | 1.09 | |
| 14 | F42 | 5.29 | 0.95 | 14 | F42 | 20.3 | 115.6 | 5.29 | 1.07 | |
| 15 | F32 | 5.26 | 0.91 | 15 | F36 | 18.4 | 133.5 | 5.31 | 1.06 | |
| 16 | F39 | 5.24 | 0.88 | 16 | F34 | 18.3 | 123.6 | 4.96 | 0.62 | |
| 17 | F16 | 5.14 | 0.75 | 17 | F45 | 19.2 | 115.4 | 4.88 | 0.59 | |
| 18 | F20 | 5.08 | 0.68 | 18 | F50 | 19.0 | 106.8 | 5.00 | 0.45 | |
| 19 | F21 | 5.05 | 0.64 | 19 | F16 | 14.0 | 152.6 | 5.14 | 0.43 | |
| 20 | F50 | 5.00 | 0.57 | 20 | F9 | 18.6 | 106.7 | 4.88 | 0.29 | |
| 21 | F34 | 4.96 | 0.52 | 21 | F52 | 18.1 | 111.9 | 4.71 | 0.19 | |
| 22 | F15 | 4.94 | 0.49 | 22 | F19 | 15.1 | 124.0 | 5.30 | 0.17 | |
| 23 | F45 | 4.88 | 0.42 | 23 | F25 | 19.0 | 108.0 | 4.50 | 0.17 | |
| 24 | F9 | 4.88 | 0.42 | 24 | F48 | 18.6 | 106.4 | 4.63 | 0.13 | |
| 25 | F52 | 4.71 | 0.19 | 25 | Inpari 34 Salin A | 17.0 | 126.4 | 4.48 | 0.12 | |
| 26 | F48 | 4.63 | 0.09 | 26 | F15 | 13.7 | 133.9 | 4.94 | -0.13 | |
| 27 | F14 | 4.59 | 0.04 | 27 | F20 | 15.3 | 113.0 | 5.08 | -0.14 | |
| 28 | F25 | 4.50 | -0.08 | 28 | F35 | 18.5 | 110.8 | 4.06 | -0.14 | |
| 29 | F13 | 4.48 | -0.10 | 29 | F21 | 13.1 | 134.4 | 5.05 | -0.18 | |
| 30 | Inpari 34 Salin A | 4.48 | -0.10 | 30 | F56 | 16.6 | 127.9 | 4.01 | -0.22 | |
| 32 | Inpari 29 | 4.40 | -0.21 | 33 | Inpari 29 | 15.3 | 128.8 | 4.40 | -0.23 | |
| 37 | Ciherang | 4.26 | -0.39 | 36 | Inpara 5 | 21.0 | 88.7 | 3.72 | -0.29 | |
| 52 | Inpara 5 | 3.72 | -1.09 | 37 | Ciherang | 17.7 | 102.1 | 4.26 | -0.37 | |
Table S3. Selected genotypes of DH rice lines resulting from direct and indirect selection.
| Direct Selection | Selection Indices | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Rank | Genotype | Y (ton ha-1) | Z yield | Rank | Genotype | NPT | NFG | Y (ton ha-1) | Z Index | |
| 1 | F47 | 6.00 | 1.87 | 1 | F51 | 19.9 | 130.7 | 6.00 | 1.73 | |
| 2 | F51 | 6.00 | 1.87 | 2 | F41 | 20.9 | 121.2 | 5.97 | 1.72 | |
| 3 | F41 | 5.97 | 1.83 | 3 | F40 | 21.2 | 123.8 | 5.52 | 1.56 | |
| 4 | F43 | 5.91 | 1.76 | 4 | F47 | 20.0 | 116.4 | 6.00 | 1.45 | |
| 5 | F44 | 5.67 | 1.44 | 5 | F44 | 19.2 | 130.2 | 5.67 | 1.37 | |
| 6 | F49 | 5.65 | 1.42 | 6 | F43 | 19.2 | 122.2 | 5.91 | 1.35 | |
| 7 | F46 | 5.59 | 1.34 | 7 | F46 | 20.5 | 118.3 | 5.59 | 1.35 | |
| 8 | F40 | 5.52 | 1.25 | 8 | F38 | 20.0 | 126.0 | 5.37 | 1.27 | |
| 9 | F22 | 5.49 | 1.21 | 9 | F22 | 19.7 | 123.3 | 5.49 | 1.23 | |
| 10 | F38 | 5.37 | 1.05 | 10 | F32 | 20.1 | 125.2 | 5.26 | 1.21 | |
| 11 | F37 | 5.35 | 1.03 | 11 | F39 | 19.7 | 129.7 | 5.24 | 1.20 | |
| 12 | F36 | 5.31 | 0.98 | 12 | F49 | 20.4 | 110.4 | 5.65 | 1.20 | |
| 13 | F19 | 5.30 | 0.96 | 13 | F37 | 20.9 | 108.5 | 5.35 | 1.09 | |
| 14 | F42 | 5.29 | 0.95 | 14 | F42 | 20.3 | 115.6 | 5.29 | 1.07 | |
| 15 | F32 | 5.26 | 0.91 | 15 | F36 | 18.4 | 133.5 | 5.31 | 1.06 | |
| 16 | F39 | 5.24 | 0.88 | 16 | F34 | 18.3 | 123.6 | 4.96 | 0.62 | |
| 17 | F16 | 5.14 | 0.75 | 17 | F45 | 19.2 | 115.4 | 4.88 | 0.59 | |
| 18 | F20 | 5.08 | 0.68 | 18 | F50 | 19.0 | 106.8 | 5.00 | 0.45 | |
| 19 | F21 | 5.05 | 0.64 | 19 | F16 | 14.0 | 152.6 | 5.14 | 0.43 | |
| 20 | F50 | 5.00 | 0.57 | 20 | F9 | 18.6 | 106.7 | 4.88 | 0.29 | |
| 21 | F34 | 4.96 | 0.52 | 21 | F52 | 18.1 | 111.9 | 4.71 | 0.19 | |
| 22 | F15 | 4.94 | 0.49 | 22 | F19 | 15.1 | 124.0 | 5.30 | 0.17 | |
| 23 | F45 | 4.88 | 0.42 | 23 | F25 | 19.0 | 108.0 | 4.50 | 0.17 | |
| 24 | F9 | 4.88 | 0.42 | 24 | F48 | 18.6 | 106.4 | 4.63 | 0.13 | |
| 25 | F52 | 4.71 | 0.19 | 25 | Inpari 34 Salin A | 17.0 | 126.4 | 4.48 | 0.12 | |
| 26 | F48 | 4.63 | 0.09 | 26 | F15 | 13.7 | 133.9 | 4.94 | -0.13 | |
| 27 | F14 | 4.59 | 0.04 | 27 | F20 | 15.3 | 113.0 | 5.08 | -0.14 | |
| 28 | F25 | 4.50 | -0.08 | 28 | F35 | 18.5 | 110.8 | 4.06 | -0.14 | |
| 29 | F13 | 4.48 | -0.10 | 29 | F21 | 13.1 | 134.4 | 5.05 | -0.18 | |
| 30 | Inpari 34 Salin A | 4.48 | -0.10 | 30 | F56 | 16.6 | 127.9 | 4.01 | -0.22 | |
| 32 | Inpari 29 | 4.40 | -0.21 | 33 | Inpari 29 | 15.3 | 128.8 | 4.40 | -0.23 | |
| 37 | Ciherang | 4.26 | -0.39 | 36 | Inpara 5 | 21.0 | 88.7 | 3.72 | -0.29 | |
| 52 | Inpara 5 | 3.72 | -1.09 | 37 | Ciherang | 17.7 | 102.1 | 4.26 | -0.37 | |
| Characters | MS Environment (E) | MS Genotype (G) | MS (G × E) | MS Error | CV (%) |
|---|---|---|---|---|---|
| Number of leaves | 219.882ns | 12.132** | 2.567ns | 2.624 | 15.96 |
| Number of tillers | 1.100ns | 0.124** | 0.043** | 0.030 | 13.37tr |
| Shoot height (cm) | 62,076.467** | 252.683** | 50.897** | 19.065 | 8.07 |
| Root length (cm) | 1,172.778ns | 26.123** | 8.863ns | 8.148 | 14.80tr |
| Shoot fresh weight (g) | 51.268** | 0.228** | 0.049** | 0.029 | 11.26tr |
| Root fresh weight (g) | 11.600** | 0.071** | 0.034ns | 0.025 | 14.68tr |
| Shoot dry weight (g) | 5.070* | 0.031** | 0.006ns | 0.007 | 13.05tr |
| Root dry weight (g) | 1.183* | 0.006** | 0.003ns | 0.002 | 13.67tr |
| Total fresh weight (g) | 60.969** | 0.282** | 0.070* | 0.046 | 11.50tr |
| Total dry weight (g) | 6.193* | 0.036** | 0.008ns | 0.008 | 12.58tr |
Table S4. ANOVA mean squares and coefficient of variation calculated from the combined analysis of normal and stress environments.
| Characters | MS Environment (E) | MS Genotype (G) | MS (G × E) | MS Error | CV (%) |
|---|---|---|---|---|---|
| Number of leaves | 219.882ns | 12.132** | 2.567ns | 2.624 | 15.96 |
| Number of tillers | 1.100ns | 0.124** | 0.043** | 0.030 | 13.37tr |
| Shoot height (cm) | 62,076.467** | 252.683** | 50.897** | 19.065 | 8.07 |
| Root length (cm) | 1,172.778ns | 26.123** | 8.863ns | 8.148 | 14.80tr |
| Shoot fresh weight (g) | 51.268** | 0.228** | 0.049** | 0.029 | 11.26tr |
| Root fresh weight (g) | 11.600** | 0.071** | 0.034ns | 0.025 | 14.68tr |
| Shoot dry weight (g) | 5.070* | 0.031** | 0.006ns | 0.007 | 13.05tr |
| Root dry weight (g) | 1.183* | 0.006** | 0.003ns | 0.002 | 13.67tr |
| Total fresh weight (g) | 60.969** | 0.282** | 0.070* | 0.046 | 11.50tr |
| Total dry weight (g) | 6.193* | 0.036** | 0.008ns | 0.008 | 12.58tr |
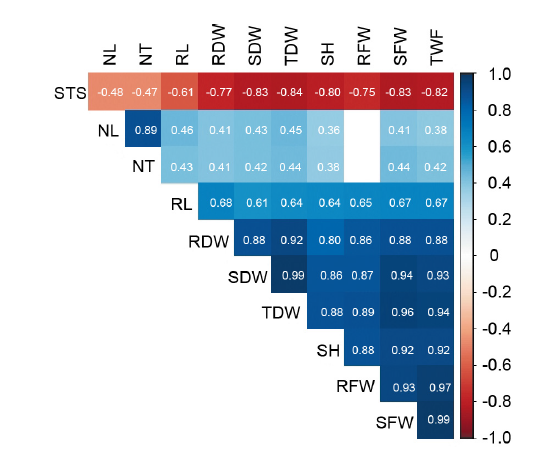
Fig. 2. Spearman correlations among all observed characters of DH rice lines under salinity stress. NT, Number of tillers; NL, Number of leaves; SH, Shoot height; RL, Root length; SFW, Shoot fresh weight; RFW, Root fresh weight; TFW, Total fresh weight; SDW, Shoot dry weight; RDW, Root dry weight; TDW, Total dry weight; STS, Salinity tolerance score.Color box (r > 0.35) indicates significant correlation at P < 0.01; Blank box, Not significant.
| Formula | SSI | Effectiveness |
|---|---|---|
| 1 | -1.7633 SH120 + 0.5314 SFW120 | 87.10 |
| 2 | 0.7998 RDSH + 0.8031 RDSFW | 93.55 |
| 3 | 1.2931 RDSH ‒ 0.1504 SFW120 | 83.87 |
| 4 | -0.5976 SH120 + 1.0274 RDSFW | 91.94 |
Table 2 SSI formula based on discriminant analysis.
| Formula | SSI | Effectiveness |
|---|---|---|
| 1 | -1.7633 SH120 + 0.5314 SFW120 | 87.10 |
| 2 | 0.7998 RDSH + 0.8031 RDSFW | 93.55 |
| 3 | 1.2931 RDSH ‒ 0.1504 SFW120 | 83.87 |
| 4 | -0.5976 SH120 + 1.0274 RDSFW | 91.94 |
| R | Formula 1 | G | GD | Formula 2 | G | GD | Formula 3 | G | GD | Formula 4 | G | GD |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | -5.49 | 61 | 1 | -3.38 | 28 | 1 | -3.51 | 61 | 1 | -3.88 | 61 | 1 |
| 2 | -1.56 | 11 | 1 | -2.78 | 11 | 1 | -2.46 | 11 | 1 | -3.43 | 28 | 1 |
| 3 | -1.33 | 36 | 1 | -2.65 | 23 | 1 | -2.42 | 36 | 1 | -2.82 | 23 | 1 |
| 4 | -1.27 | 54 | 1 | -2.47 | 61 | 1 | -2.08 | 28 | 1 | -2.45 | 11 | 1 |
| 5 | -1.01 | 28 | 1 | -2.20 | 36 | 1 | -1.85 | 47 | 1 | -1.88 | 26 | 1 |
| 6 | -0.93 | 13 | 1 | -1.98 | 47 | 1 | -1.81 | 46 | 1 | -1.76 | 41 | 1 |
| 7 | -0.87 | 26 | 1 | -1.91 | 41 | 1 | -1.61 | 26 | 1 | -1.57 | 47 | 1 |
| 8 | -0.86 | 37 | 1 | -1.90 | 26 | 1 | -1.56 | 41 | 1 | -1.54 | 36 | 1 |
| 9 | -0.86 | 10 | 1 | -1.69 | 46 | 1 | -1.28 | 23 | 1 | -1.52 | 30 | 1 |
| 10 | -0.83 | 41 | 1 | -1.57 | 30 | 1 | -1.22 | 30 | 1 | -1.35 | 10 | 1 |
| 11 | -0.82 | 46 | 1 | -1.36 | 52 | 1 | -1.19 | 54 | 1 | -1.28 | 52 | 1 |
| 12 | -0.76 | 30 | 1 | -1.33 | 10 | 1 | -1.15 | 43 | 1 | -1.25 | 37 | 1 |
| 13 | -0.76 | 8 | 1 | -1.31 | 37 | 1 | -1.15 | 48 | 1 | -1.16 | 46 | 1 |
| 14 | -0.71 | 47 | 1 | -1.15 | 59 | 1 | -1.11 | 37 | 1 | -1.12 | 13 | 1 |
| 15 | -0.71 | 33 | 1 | -0.96 | 25 | 1 | -1.05 | 10 | 1 | -1.02 | 54 | 1 |
| 16 | -0.70 | 3 | 1 | -0.93 | 13 | 1 | -0.99 | 25 | 1 | -0.85 | 31 | 1 |
| 17 | -0.70 | 12 | 1 | -0.91 | 48 | 1 | -0.97 | 59 | 1 | -0.80 | 24 | 1 |
| 18 | -0.64 | 25 | 1 | -0.91 | 24 | 1 | -0.91 | 22 | 1 | -0.79 | 22 | 1 |
| 19 | -0.63 | 22 | 1 | -0.88 | 22 | 1 | -0.81 | 29 | 1 | -0.79 | 14 | 1 |
| 20 | -0.61 | 49 | 1 | -0.87 | 54 | 1 | -0.75 | 50 | 1 | -0.74 | 33 | 1 |
| 21 | -0.56 | 20 | 1 | -0.87 | 43 | 1 | -0.71 | 45 | 1 | -0.70 | 25 | 1 |
| 22 | -0.51 | 45 | 1 | -0.80 | 31 | 1 | -0.66 | 24 | 1 | -0.58 | 48 | 1 |
| 23 | -0.45 | 43 | 1 | -0.66 | 33 | 1 | -0.62 | 13 | 1 | -0.54 | 49 | 1 |
| 24 | -0.42 | 42 | 1 | -0.66 | 29 | 1 | -0.58 | 52 | 1 | -0.53 | 20 | 1 |
| 25 | -0.34 | 48 | 1 | -0.48 | 49 | 1 | -0.47 | 33 | 1 | -0.49 | 59 | 1 |
| 26 | -0.32 | 5 | 1 | -0.40 | 27 | 1 | -0.37 | 49 | 1 | -0.36 | 27 | 1 |
| 27 | -0.28 | 23 | 1 | -0.32 | 38 | 1 | -0.34 | 42 | 1 | -0.33 | 43 | 1 |
| 28 | -0.27 | 50 | 1 | -0.30 | 50 | 1 | -0.30 | 38 | 1 | -0.27 | 3 | 1 |
| 29 | -0.26 | 29 | 1 | -0.30 | 39 | 1 | -0.26 | 3 | 1 | -0.26 | 39 | 1 |
| 30 | -0.26 | 24 | 1 | -0.16 | 14 | 1 | -0.20 | 31 | 1 | -0.17 | 29 | 1 |
| 31 | -0.11 | 38 | 1 | -0.14 | 3 | 1 | -0.20 | 27 | 1 | -0.16 | 12 | 1 |
| 32 | -0.11 | 18 | 1 | -0.11 | 20 | 1 | -0.13 | 40 | 1 | -0.07 | 38 | 1 |
| 33 | -0.07 | 27 | 1 | -0.10 | 32 | 1 | -0.11 | 32 | 1 | -0.01 | 50 | 1 |
| 34 | -0.06 | 16 | 2* | 0.00 | 42 | 1 | -0.11 | 39 | 1 | 0.03 | 32 | 1 |
| 35 | -0.01 | 40 | 1 | 0.01 | 45 | 1 | -0.09 | 53 | 1 | 0.09 | 21 | 1 |
| 36 | 0.01 | 4 | 1 | 0.13 | 40 | 1 | 0.04 | 60 | 2 | 0.23 | 8 | 1 |
| 37 | 0.05 | 19 | 1 | 0.26 | 12 | 1 | 0.13 | 8 | 1 | 0.24 | 40 | 1 |
| 38 | 0.06 | 39 | 1 | 0.44 | 8 | 1 | 0.16 | 20 | 1 | 0.27 | 42 | 1 |
| 39 | 0.07 | 9 | 2* | 0.53 | 53 | 1 | 0.32 | 5 | 1 | 0.31 | 19 | 1 |
| 40 | 0.08 | 2 | 2* | 0.61 | 5 | 1 | 0.45 | 12 | 1 | 0.43 | 5 | 1 |
| 41 | 0.08 | 31 | 1* | 0.78 | 60 | 2* | 0.70 | 14 | 1 | 0.52 | 18 | 1 |
| 42 | 0.11 | 7 | 2* | 0.80 | 19 | 1 | 0.72 | 2 | 2 | 0.53 | 45 | 1 |
| 43 | 0.20 | 52 | 1* | 0.83 | 51 | 1 | 0.77 | 1 | 2 | 0.60 | 51 | 1 |
| 44 | 0.24 | 32 | 1* | 0.89 | 21 | 1 | 0.77 | 35 | 2 | 0.80 | 9 | 2 |
| 45 | 0.29 | 21 | 1* | 1.01 | 18 | 1 | 0.91 | 9 | 2* | 0.81 | 7 | 2 |
| 46 | 0.29 | 14 | 1* | 1.04 | 9 | 2 | 0.98 | 19 | 1* | 0.94 | 16 | 2 |
| 47 | 0.30 | 59 | 1* | 1.06 | 35 | 2 | 1.01 | 51 | 1* | 1.07 | 44 | 2 |
| 48 | 0.34 | 6 | 2* | 1.12 | 7 | 2 | 1.04 | 7 | 2* | 1.10 | 2 | 2 |
| 49 | 0.50 | 35 | 2* | 1.23 | 2 | 2 | 1.04 | 18 | 1* | 1.10 | 4 | 1* |
| 50 | 0.60 | 51 | 1* | 1.35 | 16 | 2 | 1.04 | 16 | 2* | 1.11 | 35 | 2* |
| 51 | 0.90 | 62 | 2* | 1.40 | 4 | 1* | 1.11 | 4 | 1* | 1.29 | 53 | 1* |
| 52 | 1.00 | 53 | 1* | 1.52 | 1 | 2 | 1.42 | 58 | 2* | 1.59 | 15 | 2 |
| 53 | 1.10 | 60 | 2 | 1.56 | 44 | 2 | 1.42 | 6 | 2* | 1.64 | 60 | 2 |
| 54 | 1.22 | 1 | 2 | 1.92 | 58 | 2 | 1.49 | 21 | 1* | 1.99 | 1 | 2 |
| 55 | 1.25 | 34 | 2 | 2.07 | 56 | 2 | 1.78 | 62 | 2* | 2.15 | 55 | 2 |
| 56 | 1.33 | 17 | 2 | 2.23 | 6 | 2 | 1.87 | 34 | 2 | 2.20 | 6 | 2 |
| 57 | 1.52 | 44 | 2 | 2.30 | 55 | 2 | 1.90 | 56 | 2 | 2.22 | 62 | 2 |
| 58 | 2.37 | 15 | 2 | 2.36 | 15 | 2 | 2.00 | 44 | 2 | 2.39 | 34 | 2 |
| 59 | 2.40 | 58 | 2 | 2.41 | 62 | 2 | 2.64 | 57 | 2 | 2.43 | 56 | 2 |
| 60 | 2.65 | 56 | 2 | 2.44 | 34 | 2 | 3.05 | 17 | 2 | 2.61 | 58 | 2 |
| 61 | 3.22 | 57 | 2 | 2.80 | 57 | 2 | 3.13 | 15 | 2 | 2.70 | 17 | 2 |
| 62 | 3.88 | 55 | 2 | 3.37 | 17 | 2 | 3.18 | 55 | 2 | 3.04 | 57 | 2 |
Table S5. Discriminant grouping of DH rice lines and the check varieties under hydroponic salinity screening.
| R | Formula 1 | G | GD | Formula 2 | G | GD | Formula 3 | G | GD | Formula 4 | G | GD |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | -5.49 | 61 | 1 | -3.38 | 28 | 1 | -3.51 | 61 | 1 | -3.88 | 61 | 1 |
| 2 | -1.56 | 11 | 1 | -2.78 | 11 | 1 | -2.46 | 11 | 1 | -3.43 | 28 | 1 |
| 3 | -1.33 | 36 | 1 | -2.65 | 23 | 1 | -2.42 | 36 | 1 | -2.82 | 23 | 1 |
| 4 | -1.27 | 54 | 1 | -2.47 | 61 | 1 | -2.08 | 28 | 1 | -2.45 | 11 | 1 |
| 5 | -1.01 | 28 | 1 | -2.20 | 36 | 1 | -1.85 | 47 | 1 | -1.88 | 26 | 1 |
| 6 | -0.93 | 13 | 1 | -1.98 | 47 | 1 | -1.81 | 46 | 1 | -1.76 | 41 | 1 |
| 7 | -0.87 | 26 | 1 | -1.91 | 41 | 1 | -1.61 | 26 | 1 | -1.57 | 47 | 1 |
| 8 | -0.86 | 37 | 1 | -1.90 | 26 | 1 | -1.56 | 41 | 1 | -1.54 | 36 | 1 |
| 9 | -0.86 | 10 | 1 | -1.69 | 46 | 1 | -1.28 | 23 | 1 | -1.52 | 30 | 1 |
| 10 | -0.83 | 41 | 1 | -1.57 | 30 | 1 | -1.22 | 30 | 1 | -1.35 | 10 | 1 |
| 11 | -0.82 | 46 | 1 | -1.36 | 52 | 1 | -1.19 | 54 | 1 | -1.28 | 52 | 1 |
| 12 | -0.76 | 30 | 1 | -1.33 | 10 | 1 | -1.15 | 43 | 1 | -1.25 | 37 | 1 |
| 13 | -0.76 | 8 | 1 | -1.31 | 37 | 1 | -1.15 | 48 | 1 | -1.16 | 46 | 1 |
| 14 | -0.71 | 47 | 1 | -1.15 | 59 | 1 | -1.11 | 37 | 1 | -1.12 | 13 | 1 |
| 15 | -0.71 | 33 | 1 | -0.96 | 25 | 1 | -1.05 | 10 | 1 | -1.02 | 54 | 1 |
| 16 | -0.70 | 3 | 1 | -0.93 | 13 | 1 | -0.99 | 25 | 1 | -0.85 | 31 | 1 |
| 17 | -0.70 | 12 | 1 | -0.91 | 48 | 1 | -0.97 | 59 | 1 | -0.80 | 24 | 1 |
| 18 | -0.64 | 25 | 1 | -0.91 | 24 | 1 | -0.91 | 22 | 1 | -0.79 | 22 | 1 |
| 19 | -0.63 | 22 | 1 | -0.88 | 22 | 1 | -0.81 | 29 | 1 | -0.79 | 14 | 1 |
| 20 | -0.61 | 49 | 1 | -0.87 | 54 | 1 | -0.75 | 50 | 1 | -0.74 | 33 | 1 |
| 21 | -0.56 | 20 | 1 | -0.87 | 43 | 1 | -0.71 | 45 | 1 | -0.70 | 25 | 1 |
| 22 | -0.51 | 45 | 1 | -0.80 | 31 | 1 | -0.66 | 24 | 1 | -0.58 | 48 | 1 |
| 23 | -0.45 | 43 | 1 | -0.66 | 33 | 1 | -0.62 | 13 | 1 | -0.54 | 49 | 1 |
| 24 | -0.42 | 42 | 1 | -0.66 | 29 | 1 | -0.58 | 52 | 1 | -0.53 | 20 | 1 |
| 25 | -0.34 | 48 | 1 | -0.48 | 49 | 1 | -0.47 | 33 | 1 | -0.49 | 59 | 1 |
| 26 | -0.32 | 5 | 1 | -0.40 | 27 | 1 | -0.37 | 49 | 1 | -0.36 | 27 | 1 |
| 27 | -0.28 | 23 | 1 | -0.32 | 38 | 1 | -0.34 | 42 | 1 | -0.33 | 43 | 1 |
| 28 | -0.27 | 50 | 1 | -0.30 | 50 | 1 | -0.30 | 38 | 1 | -0.27 | 3 | 1 |
| 29 | -0.26 | 29 | 1 | -0.30 | 39 | 1 | -0.26 | 3 | 1 | -0.26 | 39 | 1 |
| 30 | -0.26 | 24 | 1 | -0.16 | 14 | 1 | -0.20 | 31 | 1 | -0.17 | 29 | 1 |
| 31 | -0.11 | 38 | 1 | -0.14 | 3 | 1 | -0.20 | 27 | 1 | -0.16 | 12 | 1 |
| 32 | -0.11 | 18 | 1 | -0.11 | 20 | 1 | -0.13 | 40 | 1 | -0.07 | 38 | 1 |
| 33 | -0.07 | 27 | 1 | -0.10 | 32 | 1 | -0.11 | 32 | 1 | -0.01 | 50 | 1 |
| 34 | -0.06 | 16 | 2* | 0.00 | 42 | 1 | -0.11 | 39 | 1 | 0.03 | 32 | 1 |
| 35 | -0.01 | 40 | 1 | 0.01 | 45 | 1 | -0.09 | 53 | 1 | 0.09 | 21 | 1 |
| 36 | 0.01 | 4 | 1 | 0.13 | 40 | 1 | 0.04 | 60 | 2 | 0.23 | 8 | 1 |
| 37 | 0.05 | 19 | 1 | 0.26 | 12 | 1 | 0.13 | 8 | 1 | 0.24 | 40 | 1 |
| 38 | 0.06 | 39 | 1 | 0.44 | 8 | 1 | 0.16 | 20 | 1 | 0.27 | 42 | 1 |
| 39 | 0.07 | 9 | 2* | 0.53 | 53 | 1 | 0.32 | 5 | 1 | 0.31 | 19 | 1 |
| 40 | 0.08 | 2 | 2* | 0.61 | 5 | 1 | 0.45 | 12 | 1 | 0.43 | 5 | 1 |
| 41 | 0.08 | 31 | 1* | 0.78 | 60 | 2* | 0.70 | 14 | 1 | 0.52 | 18 | 1 |
| 42 | 0.11 | 7 | 2* | 0.80 | 19 | 1 | 0.72 | 2 | 2 | 0.53 | 45 | 1 |
| 43 | 0.20 | 52 | 1* | 0.83 | 51 | 1 | 0.77 | 1 | 2 | 0.60 | 51 | 1 |
| 44 | 0.24 | 32 | 1* | 0.89 | 21 | 1 | 0.77 | 35 | 2 | 0.80 | 9 | 2 |
| 45 | 0.29 | 21 | 1* | 1.01 | 18 | 1 | 0.91 | 9 | 2* | 0.81 | 7 | 2 |
| 46 | 0.29 | 14 | 1* | 1.04 | 9 | 2 | 0.98 | 19 | 1* | 0.94 | 16 | 2 |
| 47 | 0.30 | 59 | 1* | 1.06 | 35 | 2 | 1.01 | 51 | 1* | 1.07 | 44 | 2 |
| 48 | 0.34 | 6 | 2* | 1.12 | 7 | 2 | 1.04 | 7 | 2* | 1.10 | 2 | 2 |
| 49 | 0.50 | 35 | 2* | 1.23 | 2 | 2 | 1.04 | 18 | 1* | 1.10 | 4 | 1* |
| 50 | 0.60 | 51 | 1* | 1.35 | 16 | 2 | 1.04 | 16 | 2* | 1.11 | 35 | 2* |
| 51 | 0.90 | 62 | 2* | 1.40 | 4 | 1* | 1.11 | 4 | 1* | 1.29 | 53 | 1* |
| 52 | 1.00 | 53 | 1* | 1.52 | 1 | 2 | 1.42 | 58 | 2* | 1.59 | 15 | 2 |
| 53 | 1.10 | 60 | 2 | 1.56 | 44 | 2 | 1.42 | 6 | 2* | 1.64 | 60 | 2 |
| 54 | 1.22 | 1 | 2 | 1.92 | 58 | 2 | 1.49 | 21 | 1* | 1.99 | 1 | 2 |
| 55 | 1.25 | 34 | 2 | 2.07 | 56 | 2 | 1.78 | 62 | 2* | 2.15 | 55 | 2 |
| 56 | 1.33 | 17 | 2 | 2.23 | 6 | 2 | 1.87 | 34 | 2 | 2.20 | 6 | 2 |
| 57 | 1.52 | 44 | 2 | 2.30 | 55 | 2 | 1.90 | 56 | 2 | 2.22 | 62 | 2 |
| 58 | 2.37 | 15 | 2 | 2.36 | 15 | 2 | 2.00 | 44 | 2 | 2.39 | 34 | 2 |
| 59 | 2.40 | 58 | 2 | 2.41 | 62 | 2 | 2.64 | 57 | 2 | 2.43 | 56 | 2 |
| 60 | 2.65 | 56 | 2 | 2.44 | 34 | 2 | 3.05 | 17 | 2 | 2.61 | 58 | 2 |
| 61 | 3.22 | 57 | 2 | 2.80 | 57 | 2 | 3.13 | 15 | 2 | 2.70 | 17 | 2 |
| 62 | 3.88 | 55 | 2 | 3.37 | 17 | 2 | 3.18 | 55 | 2 | 3.04 | 57 | 2 |
| G | RDSH | RDSFW | SSI | zSTS | SaTI | G | RDSH | RDSFW | SSI | zSTS | SaTI | G | RDSH | RDSFW | SSI | zSTS | SaTI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F1 | 0.43 | 0.77 | 1.52 | 0.95 | 1.24 | F22 | 0.33 | 0.57 | -0.88 | -0.84 | -0.86 | F43 | 0.31 | 0.60 | -0.87 | -0.92 | -0.90 |
| F2 | 0.43 | 0.73 | 1.23 | 1.35 | 1.29 | F23 | 0.32 | 0.37 | -2.65 | -1.09 | -1.87 | F44 | 0.51 | 0.67 | 1.56 | 1.22 | 1.39 |
| F3 | 0.37 | 0.62 | -0.14 | 0.14 | 0.00 | F24 | 0.35 | 0.55 | -0.91 | -0.88 | -0.90 | F45 | 0.34 | 0.69 | 0.01 | 0.14 | 0.07 |
| F4 | 0.45 | 0.72 | 1.40 | 0.14 | 0.77 | F25 | 0.33 | 0.58 | -0.96 | -0.23 | -0.60 | F46 | 0.28 | 0.54 | -1.69 | -0.88 | -1.29 |
| F5 | 0.41 | 0.68 | 0.61 | 0.06 | 0.33 | F26 | 0.30 | 0.49 | -1.90 | -0.92 | -1.41 | F47 | 0.28 | 0.50 | -1.98 | -1.09 | -1.53 |
| F6 | 0.46 | 0.81 | 2.23 | 1.14 | 1.68 | F27 | 0.37 | 0.59 | -0.40 | 0.14 | -0.13 | F48 | 0.32 | 0.59 | -0.91 | -0.88 | -0.90 |
| F7 | 0.45 | 0.69 | 1.12 | 1.14 | 1.13 | F28 | 0.27 | 0.33 | -3.38 | -1.09 | -2.24 | F49 | 0.36 | 0.59 | -0.48 | 0.14 | -0.17 |
| F8 | 0.39 | 0.67 | 0.44 | -0.86 | -0.21 | F29 | 0.33 | 0.61 | -0.66 | -0.88 | -0.77 | F50 | 0.34 | 0.64 | -0.30 | -1.09 | -0.70 |
| F9 | 0.44 | 0.69 | 1.04 | 0.51 | 0.77 | F30 | 0.32 | 0.51 | -1.57 | -0.88 | -1.23 | F51 | 0.44 | 0.65 | 0.83 | 0.14 | 0.49 |
| F10 | 0.33 | 0.53 | -1.33 | -0.92 | -1.13 | F31 | 0.37 | 0.53 | -0.80 | 0.14 | -0.33 | F52 | 0.35 | 0.49 | -1.36 | -0.27 | -0.81 |
| F11 | 0.24 | 0.45 | -2.78 | -0.84 | -1.81 | F32 | 0.38 | 0.62 | -0.10 | -0.84 | -0.47 | F53 | 0.37 | 0.71 | 0.53 | 0.14 | 0.33 |
| F12 | 0.41 | 0.62 | 0.26 | -0.23 | 0.02 | F33 | 0.36 | 0.57 | -0.66 | 0.14 | -0.26 | F54 | 0.33 | 0.59 | -0.87 | -0.23 | -0.55 |
| F13 | 0.35 | 0.55 | -0.93 | -0.84 | -0.89 | F34 | 0.49 | 0.80 | 2.44 | 0.40 | 1.42 | F55 | 0.57 | 0.68 | 2.30 | 2.59 | 2.44 |
| F14 | 0.43 | 0.54 | -0.16 | -0.84 | -0.50 | F35 | 0.43 | 0.71 | 1.06 | 1.10 | 1.08 | F56 | 0.49 | 0.75 | 2.07 | 2.22 | 2.15 |
| F15 | 0.57 | 0.69 | 2.36 | 1.61 | 1.99 | F36 | 0.24 | 0.52 | -2.20 | -1.09 | -1.64 | Inpara 5 | 0.53 | 0.79 | 2.80 | 2.18 | 2.49 |
| F16 | 0.45 | 0.72 | 1.35 | 0.38 | 0.87 | F37 | 0.32 | 0.54 | -1.31 | -0.88 | -1.10 | Ciherang | 0.46 | 0.77 | 1.92 | 2.12 | 2.02 |
| F17 | 0.56 | 0.83 | 3.37 | 0.95 | 2.16 | F38 | 0.36 | 0.61 | -0.32 | -0.86 | -0.59 | Inpari 29 | 0.32 | 0.56 | -1.15 | -0.23 | -0.69 |
| F18 | 0.45 | 0.67 | 1.01 | 0.06 | 0.53 | F39 | 0.38 | 0.59 | -0.30 | -0.88 | -0.59 | I34SA | 0.38 | 0.73 | 0.78 | 0.38 | 0.58 |
| F19 | 0.45 | 0.65 | 0.80 | 0.06 | 0.43 | F40 | 0.38 | 0.65 | 0.13 | -0.82 | -0.35 | IR29 | 0.49 | 0.80 | 2.41 | 2.59 | 2.50 |
| F20 | 0.40 | 0.59 | -0.11 | -0.23 | -0.17 | F41 | 0.30 | 0.49 | -1.91 | -0.88 | -1.40 | Pokkali | 0.22 | 0.52 | -2.47 | -1.09 | -1.78 |
| F21 | 0.48 | 0.62 | 0.89 | 0.14 | 0.51 | F42 | 0.36 | 0.66 | 0 | 0.14 | 0.07 |
Table 3 Salinity tolerance index and selection of doubled haploid rice lines tolerant to salinity stress.
| G | RDSH | RDSFW | SSI | zSTS | SaTI | G | RDSH | RDSFW | SSI | zSTS | SaTI | G | RDSH | RDSFW | SSI | zSTS | SaTI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F1 | 0.43 | 0.77 | 1.52 | 0.95 | 1.24 | F22 | 0.33 | 0.57 | -0.88 | -0.84 | -0.86 | F43 | 0.31 | 0.60 | -0.87 | -0.92 | -0.90 |
| F2 | 0.43 | 0.73 | 1.23 | 1.35 | 1.29 | F23 | 0.32 | 0.37 | -2.65 | -1.09 | -1.87 | F44 | 0.51 | 0.67 | 1.56 | 1.22 | 1.39 |
| F3 | 0.37 | 0.62 | -0.14 | 0.14 | 0.00 | F24 | 0.35 | 0.55 | -0.91 | -0.88 | -0.90 | F45 | 0.34 | 0.69 | 0.01 | 0.14 | 0.07 |
| F4 | 0.45 | 0.72 | 1.40 | 0.14 | 0.77 | F25 | 0.33 | 0.58 | -0.96 | -0.23 | -0.60 | F46 | 0.28 | 0.54 | -1.69 | -0.88 | -1.29 |
| F5 | 0.41 | 0.68 | 0.61 | 0.06 | 0.33 | F26 | 0.30 | 0.49 | -1.90 | -0.92 | -1.41 | F47 | 0.28 | 0.50 | -1.98 | -1.09 | -1.53 |
| F6 | 0.46 | 0.81 | 2.23 | 1.14 | 1.68 | F27 | 0.37 | 0.59 | -0.40 | 0.14 | -0.13 | F48 | 0.32 | 0.59 | -0.91 | -0.88 | -0.90 |
| F7 | 0.45 | 0.69 | 1.12 | 1.14 | 1.13 | F28 | 0.27 | 0.33 | -3.38 | -1.09 | -2.24 | F49 | 0.36 | 0.59 | -0.48 | 0.14 | -0.17 |
| F8 | 0.39 | 0.67 | 0.44 | -0.86 | -0.21 | F29 | 0.33 | 0.61 | -0.66 | -0.88 | -0.77 | F50 | 0.34 | 0.64 | -0.30 | -1.09 | -0.70 |
| F9 | 0.44 | 0.69 | 1.04 | 0.51 | 0.77 | F30 | 0.32 | 0.51 | -1.57 | -0.88 | -1.23 | F51 | 0.44 | 0.65 | 0.83 | 0.14 | 0.49 |
| F10 | 0.33 | 0.53 | -1.33 | -0.92 | -1.13 | F31 | 0.37 | 0.53 | -0.80 | 0.14 | -0.33 | F52 | 0.35 | 0.49 | -1.36 | -0.27 | -0.81 |
| F11 | 0.24 | 0.45 | -2.78 | -0.84 | -1.81 | F32 | 0.38 | 0.62 | -0.10 | -0.84 | -0.47 | F53 | 0.37 | 0.71 | 0.53 | 0.14 | 0.33 |
| F12 | 0.41 | 0.62 | 0.26 | -0.23 | 0.02 | F33 | 0.36 | 0.57 | -0.66 | 0.14 | -0.26 | F54 | 0.33 | 0.59 | -0.87 | -0.23 | -0.55 |
| F13 | 0.35 | 0.55 | -0.93 | -0.84 | -0.89 | F34 | 0.49 | 0.80 | 2.44 | 0.40 | 1.42 | F55 | 0.57 | 0.68 | 2.30 | 2.59 | 2.44 |
| F14 | 0.43 | 0.54 | -0.16 | -0.84 | -0.50 | F35 | 0.43 | 0.71 | 1.06 | 1.10 | 1.08 | F56 | 0.49 | 0.75 | 2.07 | 2.22 | 2.15 |
| F15 | 0.57 | 0.69 | 2.36 | 1.61 | 1.99 | F36 | 0.24 | 0.52 | -2.20 | -1.09 | -1.64 | Inpara 5 | 0.53 | 0.79 | 2.80 | 2.18 | 2.49 |
| F16 | 0.45 | 0.72 | 1.35 | 0.38 | 0.87 | F37 | 0.32 | 0.54 | -1.31 | -0.88 | -1.10 | Ciherang | 0.46 | 0.77 | 1.92 | 2.12 | 2.02 |
| F17 | 0.56 | 0.83 | 3.37 | 0.95 | 2.16 | F38 | 0.36 | 0.61 | -0.32 | -0.86 | -0.59 | Inpari 29 | 0.32 | 0.56 | -1.15 | -0.23 | -0.69 |
| F18 | 0.45 | 0.67 | 1.01 | 0.06 | 0.53 | F39 | 0.38 | 0.59 | -0.30 | -0.88 | -0.59 | I34SA | 0.38 | 0.73 | 0.78 | 0.38 | 0.58 |
| F19 | 0.45 | 0.65 | 0.80 | 0.06 | 0.43 | F40 | 0.38 | 0.65 | 0.13 | -0.82 | -0.35 | IR29 | 0.49 | 0.80 | 2.41 | 2.59 | 2.50 |
| F20 | 0.40 | 0.59 | -0.11 | -0.23 | -0.17 | F41 | 0.30 | 0.49 | -1.91 | -0.88 | -1.40 | Pokkali | 0.22 | 0.52 | -2.47 | -1.09 | -1.78 |
| F21 | 0.48 | 0.62 | 0.89 | 0.14 | 0.51 | F42 | 0.36 | 0.66 | 0 | 0.14 | 0.07 |
| G | SaTI | GAI | IASI | G | SaTI | GAI | IASI | G | SaTI | GAI | IASI | G | SaTI | GAI | IASI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F1 | 1.24 | -1.45 | -2.69 | F16 | 0.87 | 0.43 | -0.43 | F31 | -0.33 | -0.96 | -0.63 | F46 | -1.29 | 1.35 | 2.64 |
| F2 | 1.29 | -1.70 | -2.99 | F17 | 2.16 | -0.28 | -2.44 | F32 | -0.47 | 1.21 | 1.68 | F47 | -1.53 | 1.45 | 2.99 |
| F3 | 0 | -0.26 | -0.26 | F18 | 0.53 | -0.73 | -1.27 | F33 | -0.26 | -0.78 | -0.52 | F48 | -0.90 | 0.13 | 1.03 |
| F4 | 0.77 | -0.48 | -1.25 | F19 | 0.43 | 0.17 | -0.26 | F34 | 1.42 | 0.62 | -0.80 | F49 | -0.17 | 1.20 | 1.37 |
| F5 | 0.33 | -0.23 | -0.56 | F20 | -0.17 | -0.14 | 0.03 | F35 | 1.08 | -0.14 | -1.22 | F50 | -0.70 | 0.45 | 1.14 |
| F6 | 1.68 | -0.95 | -2.63 | F21 | 0.51 | -0.18 | -0.69 | F36 | -1.64 | 1.06 | 2.70 | F51 | 0.49 | 1.73 | 1.24 |
| F7 | 1.13 | -0.86 | -1.99 | F22 | -0.86 | 1.23 | 2.09 | F37 | -1.10 | 1.09 | 2.19 | F52 | -0.81 | 0.19 | 1.00 |
| F8 | -0.21 | -0.63 | -0.42 | F23 | -1.87 | -1.01 | 0.86 | F38 | -0.59 | 1.27 | 1.86 | F53 | 0.33 | -1.37 | -1.71 |
| F9 | 0.77 | 0.29 | -0.49 | F24 | -0.90 | -0.44 | 0.46 | F39 | -0.59 | 1.20 | 1.80 | F54 | -0.55 | -0.86 | -0.31 |
| F10 | -1.13 | -0.51 | 0.62 | F25 | -0.60 | 0.17 | 0.76 | F40 | -0.35 | 1.56 | 1.91 | F55 | 2.44 | -0.23 | -2.67 |
| F11 | -1.81 | -0.73 | 1.08 | F26 | -1.41 | -0.59 | 0.82 | F41 | -1.40 | 1.72 | 3.12 | F56 | 2.15 | -0.22 | -2.36 |
| F12 | 0.02 | -0.91 | -0.93 | F27 | -0.13 | -0.75 | -0.62 | F42 | 0.07 | 1.07 | 1.00 | Ciherang | 2.49 | -0.37 | -2.86 |
| F13 | -0.89 | -0.95 | -0.06 | F28 | -2.24 | -1.11 | 1.13 | F43 | -0.90 | 1.35 | 2.25 | Inpara 5 | 2.02 | -0.29 | -2.31 |
| F14 | -0.50 | -0.42 | 0.08 | F29 | -0.77 | -1.27 | -0.50 | F44 | 1.39 | 1.37 | -0.02 | Inpari 29 | -0.69 | -0.23 | 0.46 |
| F15 | 1.99 | -0.13 | -2.12 | F30 | -1.23 | -0.86 | 0.36 | F45 | 0.07 | 0.59 | 0.52 | I34SA | 0.58 | 0.12 | -0.46 |
Table 4 Indirect adaptability selection index (IASI) of doubled haploid rice lines adaptive to salinity stress.
| G | SaTI | GAI | IASI | G | SaTI | GAI | IASI | G | SaTI | GAI | IASI | G | SaTI | GAI | IASI |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| F1 | 1.24 | -1.45 | -2.69 | F16 | 0.87 | 0.43 | -0.43 | F31 | -0.33 | -0.96 | -0.63 | F46 | -1.29 | 1.35 | 2.64 |
| F2 | 1.29 | -1.70 | -2.99 | F17 | 2.16 | -0.28 | -2.44 | F32 | -0.47 | 1.21 | 1.68 | F47 | -1.53 | 1.45 | 2.99 |
| F3 | 0 | -0.26 | -0.26 | F18 | 0.53 | -0.73 | -1.27 | F33 | -0.26 | -0.78 | -0.52 | F48 | -0.90 | 0.13 | 1.03 |
| F4 | 0.77 | -0.48 | -1.25 | F19 | 0.43 | 0.17 | -0.26 | F34 | 1.42 | 0.62 | -0.80 | F49 | -0.17 | 1.20 | 1.37 |
| F5 | 0.33 | -0.23 | -0.56 | F20 | -0.17 | -0.14 | 0.03 | F35 | 1.08 | -0.14 | -1.22 | F50 | -0.70 | 0.45 | 1.14 |
| F6 | 1.68 | -0.95 | -2.63 | F21 | 0.51 | -0.18 | -0.69 | F36 | -1.64 | 1.06 | 2.70 | F51 | 0.49 | 1.73 | 1.24 |
| F7 | 1.13 | -0.86 | -1.99 | F22 | -0.86 | 1.23 | 2.09 | F37 | -1.10 | 1.09 | 2.19 | F52 | -0.81 | 0.19 | 1.00 |
| F8 | -0.21 | -0.63 | -0.42 | F23 | -1.87 | -1.01 | 0.86 | F38 | -0.59 | 1.27 | 1.86 | F53 | 0.33 | -1.37 | -1.71 |
| F9 | 0.77 | 0.29 | -0.49 | F24 | -0.90 | -0.44 | 0.46 | F39 | -0.59 | 1.20 | 1.80 | F54 | -0.55 | -0.86 | -0.31 |
| F10 | -1.13 | -0.51 | 0.62 | F25 | -0.60 | 0.17 | 0.76 | F40 | -0.35 | 1.56 | 1.91 | F55 | 2.44 | -0.23 | -2.67 |
| F11 | -1.81 | -0.73 | 1.08 | F26 | -1.41 | -0.59 | 0.82 | F41 | -1.40 | 1.72 | 3.12 | F56 | 2.15 | -0.22 | -2.36 |
| F12 | 0.02 | -0.91 | -0.93 | F27 | -0.13 | -0.75 | -0.62 | F42 | 0.07 | 1.07 | 1.00 | Ciherang | 2.49 | -0.37 | -2.86 |
| F13 | -0.89 | -0.95 | -0.06 | F28 | -2.24 | -1.11 | 1.13 | F43 | -0.90 | 1.35 | 2.25 | Inpara 5 | 2.02 | -0.29 | -2.31 |
| F14 | -0.50 | -0.42 | 0.08 | F29 | -0.77 | -1.27 | -0.50 | F44 | 1.39 | 1.37 | -0.02 | Inpari 29 | -0.69 | -0.23 | 0.46 |
| F15 | 1.99 | -0.13 | -2.12 | F30 | -1.23 | -0.86 | 0.36 | F45 | 0.07 | 0.59 | 0.52 | I34SA | 0.58 | 0.12 | -0.46 |
| SaTI | GAI | IASI | SSI | zSTS | Sukra yield | Bogor yield | AIa | BS | |
|---|---|---|---|---|---|---|---|---|---|
| SI | 1.000 | ||||||||
| GAI | 0.226 | 1.000 | |||||||
| AI | -0.576 | 0.667 | 1.000 | ||||||
| SSI | 0.955 | 0.336 | -0.450 | 1.000 | |||||
| zSTS | 0.871 | -0.005 | -0.670 | 0.687 | 1.000 | ||||
| Sukra yield | -0.430* | 0.435* | 0.694** | -0.346 | -0.482** | 1.000 | |||
| Bogor yield | 0.180 | 0.955 | 0.664 | 0.293 | -0.047 | 0.472** | 1.000 | ||
| AIa | -0.354 | 0.792 | 0.935 | -0.155 | -0.615 | 0.638** | 0.781 | 1.000 | |
| BS | -0.410 | 0.721 | 0.918 | -0.210 | -0.661 | 0.657** | 0.780 | 0.972 | 1.000 |
Table S6. Pearson correlation of validation test.
| SaTI | GAI | IASI | SSI | zSTS | Sukra yield | Bogor yield | AIa | BS | |
|---|---|---|---|---|---|---|---|---|---|
| SI | 1.000 | ||||||||
| GAI | 0.226 | 1.000 | |||||||
| AI | -0.576 | 0.667 | 1.000 | ||||||
| SSI | 0.955 | 0.336 | -0.450 | 1.000 | |||||
| zSTS | 0.871 | -0.005 | -0.670 | 0.687 | 1.000 | ||||
| Sukra yield | -0.430* | 0.435* | 0.694** | -0.346 | -0.482** | 1.000 | |||
| Bogor yield | 0.180 | 0.955 | 0.664 | 0.293 | -0.047 | 0.472** | 1.000 | ||
| AIa | -0.354 | 0.792 | 0.935 | -0.155 | -0.615 | 0.638** | 0.781 | 1.000 | |
| BS | -0.410 | 0.721 | 0.918 | -0.210 | -0.661 | 0.657** | 0.780 | 0.972 | 1.000 |
| Location | Cation content (cmol/kg) | ECe (dS/m) | ACC | pH | Climate | |||
|---|---|---|---|---|---|---|---|---|
| Ca | Mg | Na | T (ºC) | P (mm) | ||||
| Sawah Baru | 5.24 | 2.25 | 0.40 | 3.90 | 22.47 | 5.09 | 26.0 | 249.78 |
| Sukra | 18.64 | 15.34 | 5.66 | 12.07 | 41.81 | 4.96 | 27.4 | 144.93 |
Table S7. Description of soil and climate parameters in Sawah Baru and Sukra.
| Location | Cation content (cmol/kg) | ECe (dS/m) | ACC | pH | Climate | |||
|---|---|---|---|---|---|---|---|---|
| Ca | Mg | Na | T (ºC) | P (mm) | ||||
| Sawah Baru | 5.24 | 2.25 | 0.40 | 3.90 | 22.47 | 5.09 | 26.0 | 249.78 |
| Sukra | 18.64 | 15.34 | 5.66 | 12.07 | 41.81 | 4.96 | 27.4 | 144.93 |
| [1] | Acquaah G. 2007. Principles of Plant Genetics and Breeding. Oxford: Blackwell Publishing: 121‒145. |
| [2] | Akbar M R, Purwoko B S, Dewi I S, Suwarno W B, Sugiyanta.2019. Selection of doubled haploid lines of rainfed lowland rice in preliminary yield trial. Biodiversitas, 20(10): 2796-2801. |
| [3] | Akter S, Biswas B K, Azad A K, Hasanuzzaman M, Arifuzzaman M. 2010. Correlation and discriminant function analysis of some selected characters in fine rice (Oryza sativa L.) available in Bangladesh. Int J Sustain Crop Prod, 5(4): 30-35. |
| [4] | Akҫura M, Ҫeri S. 2011. Evaluation of drought tolerance indices for selection of Turkish oat (Avena sativa L.) landraces under various environmental conditions. Žemdirbystė, 98(2): 157-166. |
| [5] | Ali M N, Yeasmin L, Gantait S, Goswami R, Chakraborty S. 2014. Screening of rice landraces for salinity tolerance at seedling stage through morphological and molecular markers. Physiol Mol Biol Plants, 20(4): 411-423. |
| [6] | Al-Naggar A M M, Sabry S R S, Atta M M M, El-Aleem O M A. 2015. Effects of salinity on performance, heritability, selection gain and correlations in wheat (Triticum aestivum L.) doubled haploids. Sci Agric, 10(2): 70-83. |
| [7] | Alsabah R, Purwoko B S, Dewi I S, Wahyu Y. 2019. Selection index for selecting promising doubled haploid lines of black rice. SABRAO J Breed Genet, 51(4): 279-294. |
| [8] | Al-Sayed H M, Fateh H S, Fares W M, Attaya A S. 2012. Multivariate analysis of sugar yield factors in sugar cane. Am- Eurasian J Sustain Agric, 6(1): 44-50. |
| [9] | Anshori M F, Purwoko B S, Dewi I S, Ardie S W, Suwarno W B, Safitri H. 2018. Determination of selection criteria for screening of rice genotypes for salinity tolerance. SABRAO J Breed Genet, 50(3): 279-294. |
| [10] | Anshori M F, Purwoko B S, Dewi I S, Ardie S W, Suwarno W B. 2019. Selection index based on multivariate analysis for selecting doubled-haploid rice lines in lowland saline prone area. SABRAO J Breed Genet, 51(2): 161-174. |
| [11] | Anshori M F, Purwoko B S, Dewi I S, Ardie S W, Suwarno W B. 2020. Cluster heatmap for detection of good tolerance trait on doubled-haploid rice lines under hydroponic salinity screening. IOP Conf Ser Earth Environ Sci, 484: 012001. |
| [12] | Anshori M F. 2018. Characterization and selection of doubled haploid rice (Oryza sativa L.) lines adaptive to salinity stress. [Master Thesis]. Bogor, Indonesian: IPB University. (in Indonesian) |
| [13] | Asadi A, Zebarjadi A, Abdollahi M R, Seguí-Simarro J M. 2018. Assessment of different anther culture approaches to produce doubled haploids in cucumber (Cucumis sativus L.). Euphytica, 214(11): 216. |
| [14] | Azimi M H, Jozghasemi S, Barba-Gonzalez R. 2018. Multivariate analysis of morphological characteristics in Iris germanica hybrids. Euphytica, 214(9): 161. |
| [15] | Dallastra A, Unêda-Trevisoli S H, Ferraudo A S, Di Mauro A O D. 2014. Multivariate approach in the selection of superior soybean progeny which carry the RR gene. Rev Ciênc Agron, 45(3): 588-597. |
| [16] | De Leon T B, Linscombe S, Gregorio G, Subudhi P K. 2015. Genetic variation in Southern USA rice genotypes for seedling salinity tolerance. Front Plant Sci, 6: 374. |
| [17] | Dewi I S, Purwoko B S. 2012. Kultur antera untuk percepatan perakitan varietas padi di Indonesia. J AgroBiogen, 8(2): 78-88. (in Indonesian with English abstract). |
| [18] | Egdane J A, Vispo N A, Mohammadi R, Amas J, Katimbang M L, Platten J D, Ismail A, Gregorio G B. 2003. Phenotyping Protocols for Salinity and Other Problem Soils. Los Banos, the Philippines: International Rice Research Institute: 7‒8. |
| [19] | Fellahi Z E A, Hannachi A, Bouzerzour H. 2018. Analysis of direct and indirect selection and indices in bread wheat (Triticum aestivum L.) segregating progeny. Int J Agron, 2018: 8312857. |
| [20] | Gerona M E B, Deocampo M P, Egdane J A, Ismail A M, Dionisio- Sese M L. 2019. Physiological responses of contrasting rice genotypes to salt stress at reproductive stage. Rice Sci, 26(4): 207-219. |
| [21] | Ghosh B, Ali Md N, Saikat G. 2016. Response of rice under salinity stress: A review update. J Rice Res, 4(2): 1000167. |
| [22] | Godshalk E B, Timothy D H. 1988. Factor and principal component analyses as alternatives to index selection. Theor Appl Genet, 76(3): 352-360. |
| [23] | Hasan R, Akand M, Alam N, Bashar A, Huque A K M M. 2016. Genetic association analysis and selection indices for yield attributing traits in available chilli (Capsicum annum L.) genotypes. Mol Plant Breed, 7(19): 1-9. |
| [24] | Hidayatullah A, Purwoko B S, Dewi I S, Suwarno W B. 2018. Agronomic performance and yield of doubled haploid rice lines in advanced yield trial. SABRAO J Breed Genet, 50(3): 242-253. |
| [25] | Ilin A, Raiko T. 2010. Practical approaches to principal component analysis in the presence of missing values. J Mach Learn Res, 11: 1957‒2000. |
| [26] | Islam M R, Kayess M O, Hasanuzzaman M, Rahman M W, Uddin M J, Zaman M R. 2017. Selection index for genetic improvement of wheat (Triticum aestivum L.). J Chem Biol Phys Sci, 7(1): 1-8. |
| [27] | Ismail A M, Platten J D, Miro B. 2013. Physiological bases of tolerance of abiotic stresses in rice and mechanisms of adaptation. Oryza, 50(2): 91-99. |
| [28] | Janmohammadi M, Movahedi Z, Sabaghnia N. 2014. Multivariate statistical analysis of some traits of bread wheat for breeding under rainfed conditions. J Agric Sci, 59(1): 1-14. |
| [29] | Jolliffe I T. 2002. Principal Component Analysis. 2nd Edition. New York: Springer-Verlag: 167‒169. |
| [30] | Kose A, Onder O, Bilir O, Kosar F. 2018. Application of multivariate statistical analysis for breeding strategies of spring safflower (Carthamus tinctorius L.). Turk J Field Crops, 23(1): 12-19. |
| [31] | Kumar N, Paul S. 2016. Selection criteria of linseed genotypes for seed yield traits through correlation, path coefficient and principal component analysis. J Anim Plant Sci, 26(6): 1688-1695. |
| [32] | Lorencetti C, de Carvalho F I F, de Oliveira A C, Valério I P, Hartwig I, Benin G, Schmidt D A M. 2006. Applicability of phenotypic and canonic correlations and path coefficients in the selection of oat genotypes. Sci Agric, 3(1): 11-19. |
| [33] | Manjunatha G A, Kumar M S, Jayashree M. 2017. Character association and path analysis in rice (Oryza sativa L.) genotypes evaluated under organic management. J Pharmacogn Phytochem, 6(6): 1053-1058. |
| [34] | Mattjik A A, Sumertajaya I M. 2011. Multivariate Analysis Using SAS. Bogor, Indonesia: Statistika F-MIPA IPB: 119-128/223‒237/334‒335. (in Indonesian) |
| [35] | Mohamadi S F, Bagheri N, Kiani G, Jelodar N B. 2017. Evaluation of different rice genotypes in response to salinity stress. Biol Forum, 9(1): 174-182. |
| [36] | Ojulong H F, Labuschagne M T, Herselman L, Fregene M. 2010. Yield traits as selection indices in seedling populations of cassava. Crop Breed Appl Biotechnol, 10(3): 191-196. |
| [37] | Olivoto T, de Souza V Q, Nardino M, Carvalho I R, Ferrari M, de Pelegrin A J, Szareski V J, Schmidt D. 2017. Multicollinearity in path analysis: A simple method to reduce its effects. Agron J, 109(1): 131-142. |
| [38] | Peternelli L A, Moreira É F A, Nascimento M, Cruz C D. 2017. Artificial neural networks and linear discriminant analysis in early selection among sugarcane families. Crop Breed Appl Biotechnol, 17(4): 299-305. |
| [39] | Rajamani S, Sreekanth M, Naik V S, Ratnam M. 2016. Selection indices for yield attributing characters improvement in pigeon pea (Cajanus cajan L. Millspugh). Int J Life Sci Scienti Res, 2(2): 127-129. |
| [40] | Rawlings J O, Pantula S G, Dickey D A. 1998. Applied Regression Analysis: A Research Tool. 2nd Edition. New York, USA: Springer- Verlag: 213‒214. |
| [41] | Rezaei Z, Khadivi A, ValizadehKaji B, Abbasifar A. 2018. The selection of superior walnut (Juglans regia L.) genotypes as revealed by morphological characterization. Euphytica, 214(4): 69. |
| [42] | Sabouri H, Rabiei B, Fazlalipour M. 2008. Use of selection indices based on multivariate analysis for improving grain yield in rice. Rice Sci, 15(4): 303-310. |
| [43] | Sadeghi S M. 2011. Heritability, phenotypic correlation, and path coefficient studies for some agronomic characters in landrace rice varieties. World Appl Sci J, 13(5): 1229-1233. |
| [44] | Saed-Moucheshi A, Pessarakli M, Heidari B. 2013. Comparing relationships among yield and its related traits in mycorrhizal and nonmycorrhizal inoculated wheat cultivars under different water regimes using multivariate statistics. Int J Agron, 14: 682781. |
| [45] | Safitri H, Purwoko B S, Dewi I S, Ardie S W. 2016. Anther culture to obtain rice lines tolerant to salinity. J Agron Indonesia, 44(3): 221-227. |
| [46] | Safitri H. 2016. Development of salinity tolerant rice through anther culture. [PhD Thesis]. Bogor, Indonesia: IPB University. (in Indonesian) |
| [47] | Singh R K, Chaudhary B D. 2007. Biometrical Methods in Quantitative Genetic Analysis. New Delhi, India: Kalyani Publisher: 69‒78. |
| [48] | Spearman C. 2010. The proof and measurement of association between two things. Int J Epidemiol, 39(5): 1137-1150. |
| [49] | Sultana N, Ikeda T, Itoh R. 1999. Effect of NaCl salinity on photosynthesis and dry matter accumulation in developing rice grains. Environ Exper Bot, 42(3): 211-220. |
| [50] | Suwarno, Lubis E, Hairmansis A, Santoso.2009. Development of a package of 20 varieties for blast management on upland rice. In: Wang G L, Valent B. Advances in Genetics, Genomics and Control Rice Blast Disease. Dordrecht, Netherlands: Springer: 347‒357. |
| [51] | Türkan I, Demiral T. 2009. Recent developments in understanding salinity tolerance. Environ Exp Bot, 67(1): 2-9. |
| [52] | Vaisi H, Golpavar A R. 2013. Determination of the best indirect selection criteria to improve grain yield and seed weight in oat (Avena sativa L.) genotypes. Int J Farm Alli Sci, 2(19): 747-750. |
| [53] | Varthini N V, Sudhakar D, Raveendran M, Rajeswari S, Manonmani S, Tannidi S, Aravindhan P B, Ponniah G, Gunasekaran K, Robin S. 2017. Rice diversity panel evaluated for agro-morphological diversity by multivariate analysis. Int J Curr Microbiol App Sci, 6(11): 3887-3901. |
| [54] | Yamamoto A, Sawada H, Shim I S, Usui K, Fujihara S. 2011. Effect of salt on physiological response and leaves polyamine content in NERICA rice seedling. Plant Soil Environ, 57(12): 571-576. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||