
Rice Science ›› 2017, Vol. 24 ›› Issue (4): 32-40.DOI: 10.1016/j.rsci.2016.11.002
• Orginal Article • Next Articles
S. I. de Silva W.1,2, M. N. Perera M.1, L. N. S. Perera K.2, M. Wickramasuriya A.1, A. U. Jayasekera G.1( )
)
Received:2016-09-10
Accepted:2016-11-28
Online:2017-07-10
Published:2017-04-28
S. I. de Silva W., M. N. Perera M., L. N. S. Perera K., M. Wickramasuriya A., A. U. Jayasekera G.. In silico Analysis of osr40c1 Promoter Sequence Isolated from Indica Variety Pokkali[J]. Rice Science, 2017, 24(4): 32-40.
Add to citation manager EndNote|Ris|BibTeX
| Species | Gene ID | Gene name | Chromosome | Coding sequence | Gene description | |
|---|---|---|---|---|---|---|
| Start | End | |||||
| O. sativa ssp. indica | OSINDICA_03G20090 | OsIFCC006351 | 3 | 13335722 | 13337061 | No description available |
| OSINDICA_07G40440 | OsIFCC024829 | 7 | 27091888 | 27093301 | ||
| OSINDICA_07G40470 | OsIFCC024830 | 7 | 27114027 | 27114723 | ||
| O. sativa ssp. japonica | OS03G21040 | LOC_OS03G21040.2 | 3 | 11957978 | 11959491 | Stress responsive protein, putative, expressed |
| OS07G48490 | LOC_OS07G48490.2 | 7 | 29004296 | 29005809 | ||
| OS07G48500 | LOC_OS07G48500.1 | 7 | 29008973 | 29009669 | ||
Table 1 Descriptions of osr40c gene family members in rice.
| Species | Gene ID | Gene name | Chromosome | Coding sequence | Gene description | |
|---|---|---|---|---|---|---|
| Start | End | |||||
| O. sativa ssp. indica | OSINDICA_03G20090 | OsIFCC006351 | 3 | 13335722 | 13337061 | No description available |
| OSINDICA_07G40440 | OsIFCC024829 | 7 | 27091888 | 27093301 | ||
| OSINDICA_07G40470 | OsIFCC024830 | 7 | 27114027 | 27114723 | ||
| O. sativa ssp. japonica | OS03G21040 | LOC_OS03G21040.2 | 3 | 11957978 | 11959491 | Stress responsive protein, putative, expressed |
| OS07G48490 | LOC_OS07G48490.2 | 7 | 29004296 | 29005809 | ||
| OS07G48500 | LOC_OS07G48500.1 | 7 | 29008973 | 29009669 | ||
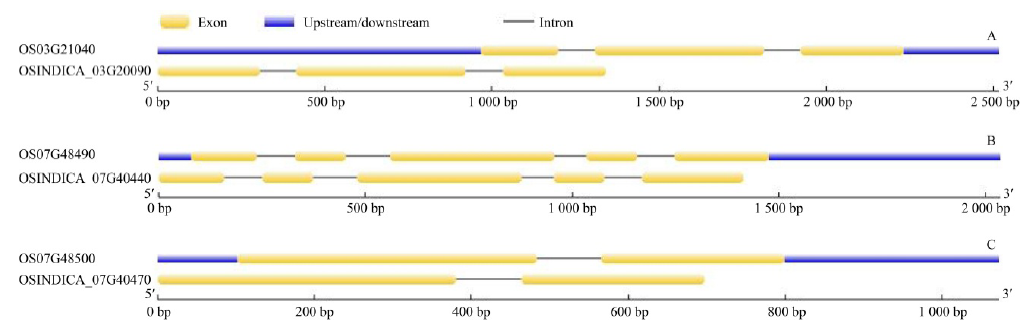
Fig. 1. Exon-intron structures of putative osr40c family members in rice. A, Gene structures of putative osr40c1 in indica (OSINDICA_03G20090) and japonica (OS03G21040); B, Gene structures of putative osr40g2 in indica (OSINDICA_07G40440) and japonica (OS07G48490); C, Gene structures of putative osr40g3 in indica (OSINDICA_07G40470) and japonica (OS07G48500).
| Stress | Changed folda | Gene expression omnibus accession number |
|---|---|---|
| Drought stress | 18.85 | GSE24048 |
| 6.44 | GSE25176 | |
| 5.64 | GSE24048 | |
| 4.4 | GSE25176 | |
| 3.91 | GSE26280 | |
| Cold stress | 7.74 | GSE37940 |
| 5.1 | GSE37940 | |
| 4.54 | GSE37940 | |
| 3.42 | GSE38023 | |
| Heat stress | 5.5 | GSE14275 |
| Photoperiod | 2.84 | GSE28124 |
| Biotic stress | 4.49 | GSE36272 |
Table 2 Expression of osr40c1 in rice plants in response to stress conditions from the Genevestigator database.
| Stress | Changed folda | Gene expression omnibus accession number |
|---|---|---|
| Drought stress | 18.85 | GSE24048 |
| 6.44 | GSE25176 | |
| 5.64 | GSE24048 | |
| 4.4 | GSE25176 | |
| 3.91 | GSE26280 | |
| Cold stress | 7.74 | GSE37940 |
| 5.1 | GSE37940 | |
| 4.54 | GSE37940 | |
| 3.42 | GSE38023 | |
| Heat stress | 5.5 | GSE14275 |
| Photoperiod | 2.84 | GSE28124 |
| Biotic stress | 4.49 | GSE36272 |
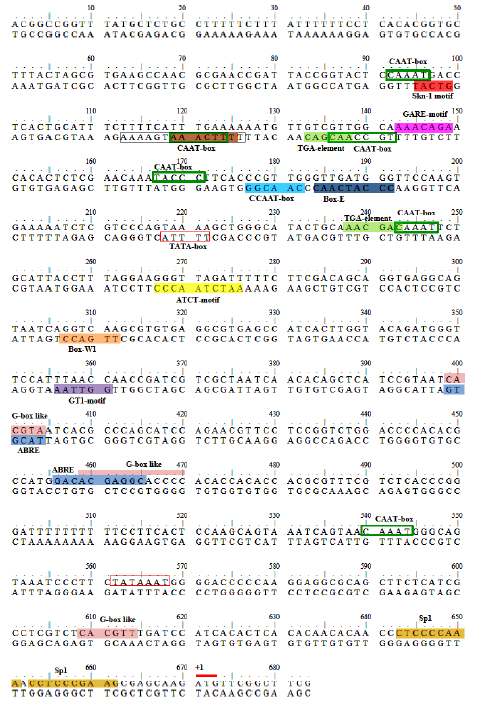

Fig. 2. Nucleotide sequence of Pokkali osr40c1 promoter along with putative cis-acting regulatory elements present in sequence. Putative translation initiation codon, ATG, is underlined in red and the first nucleotide of this codon is designated as ‘+1’. Putative core promoter elements, TATA and CAAT-boxes, are labelled and highlighted within red and green boxes, respectively. Highlighted brown colored sequence indicates the Box I motif and the sequence within the black box indicates the Gap-box motif. ABRE, ABA-responsive element; GARE, Gibberellin-responsive element.
| Type | Element | Motif function | Abundance of element |
|---|---|---|---|
| ABA-responsive element | ABRE | cis-acting regulatory element involved in the ABA responsiveness | 3 |
| Hormone-responsive element | GARE-motif | Gibberellin-responsive element | 1 |
| TGA-element | Auxin-responsive element | 2 | |
| Light-responsive element | ATCT-motif | Part of a conserved DNA module involved in light responsiveness | 1 |
| Box I | Light responsive element | 1 | |
| G-box like motif | cis-acting regulatory element involved in light responsiveness | 3 | |
| GT1-motif | Light responsive element | 1 | |
| Gap-box | Part of a light responsive element | 1 | |
| Sp1 | Light responsive element | 2 | |
| Stress-responsive element | CCAAT-box | MYBHv1 binding site | 1 |
| Fungal elicitor-responsive element | Box-W1 | Fungal elicitor responsive element | 1 |
| Box E | cis-acting regulatory element for induction upon fungal elicitation | 1 | |
| Tissue-specific element | Skn-1 motif | cis-acting regulatory element required for endosperm expression | 1 |
| Core promoter element | CAAT-box | Common cis-acting regulatory element in promoter and enhancer regions | 6 |
| TATA-box | Core promoter element around -30 of transcription start | 3 |
Table 3 Description of putative cis-acting regulatory elements in Pokkali osr40c1 promoter region from the PlantCARE database.
| Type | Element | Motif function | Abundance of element |
|---|---|---|---|
| ABA-responsive element | ABRE | cis-acting regulatory element involved in the ABA responsiveness | 3 |
| Hormone-responsive element | GARE-motif | Gibberellin-responsive element | 1 |
| TGA-element | Auxin-responsive element | 2 | |
| Light-responsive element | ATCT-motif | Part of a conserved DNA module involved in light responsiveness | 1 |
| Box I | Light responsive element | 1 | |
| G-box like motif | cis-acting regulatory element involved in light responsiveness | 3 | |
| GT1-motif | Light responsive element | 1 | |
| Gap-box | Part of a light responsive element | 1 | |
| Sp1 | Light responsive element | 2 | |
| Stress-responsive element | CCAAT-box | MYBHv1 binding site | 1 |
| Fungal elicitor-responsive element | Box-W1 | Fungal elicitor responsive element | 1 |
| Box E | cis-acting regulatory element for induction upon fungal elicitation | 1 | |
| Tissue-specific element | Skn-1 motif | cis-acting regulatory element required for endosperm expression | 1 |
| Core promoter element | CAAT-box | Common cis-acting regulatory element in promoter and enhancer regions | 6 |
| TATA-box | Core promoter element around -30 of transcription start | 3 |
| 1 | Ansari M U R, Shaheen T, Bukhari S, Husnain T.2015. Genetic improvement of rice for biotic and abiotic stress tolerance.Turk J Bot, 39(6): 911-919. |
| 2 | Borghi L.2010. Inducible gene expression systems for plants. In: Hennig L, Köhler C. Plant Developmental Biology. Totowa: Humana Press: 65-75. |
| 3 | Chen L, Jiang B J, Wu C X, Sun S, Hou W S, Han T F.2015. The characterization of GmTIP, a root-specific gene from soybean, and the expression analysis of its promoter.Plant Cell Tiss Organ Cult, 121(2): 259-274. |
| 4 | Doyle J J, Doyle J L.1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue.Phytochem Bull, 19: 11-15. |
| 5 | Furtado A, Henry R J.2005. The wheat Em promoter drives reporter gene expression in embryo and aleurone tissue of transgenic barley and rice.Plant Biotechnol J, 3(4): 421-434. |
| 6 | Hanahan D.1983. Studies on transformation of Escherichia coli with plasmids.J Mol Biol, 166(4): 557-580. |
| 7 | Hernandez-Garcia C M, Finer J J.2014. Identification and validation of promoters and cis-acting regulatory elements.Plant Sci, 217/218: 109-119. |
| 8 | Hsieh T H, Lee J T, Charng Y Y, Chan M T.2002. Tomato plants ectopically expressing Arabidopsis CBF1 show enhanced resistance to water deficit stress.Plant Physiol, 130(2): 618-626. |
| 9 | Huda K M K, Banu M S A, Pathi K M, Tuteja N.2013. Reproductive organ and vascular specific promoter of the rice plasma membrane Ca2+ ATPase mediates environmental stress responses in plants.PLoS One, 8: e57803. |
| 10 | International Rice Genome Sequencing Project.2005. The map-based sequence of the rice genome.Nature, 436: 793-800. |
| 11 | Jeon J S, Chung Y Y, Lee S, Yi G H, Oh B G, An G.1999. Isolation and characterization of an anther-specific gene, RA8, from rice (Oryza sativa L.).Plant Mol Biol, 39(1): 35-44. |
| 12 | Jiang C, Iu B, Singh J.1996. Requirement of a CCGAC cis-acting element for cold induction of the BN115 gene from winter Brassica napus.Plant Mol Biol, 30(3): 679-684. |
| 13 | Kasuga M, Liu Q, Miura S, Yamaguchi-Shinozaki K, Shinozaki K.1999. Improving plant drought, salt, and freezing tolerance by gene transfer of a single stress-inducible transcription factor.Nat Biotechnol, 17(3): 287-291. |
| 14 | Kasuga M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K.2004. A combination of the Arabidopsis DREB1A gene and stress-inducible rd29A promoter improved drought- and low- temperature stress tolerance in tobacco by gene transfer.Plant Cell Physiol, 45(3): 346-350. |
| 15 | Kawahara Y, de la Bastide M, Hamilton J P, Kanamori H, McCombie W R, Ouyang S, Schwartz D C, Tanaka T, Wu J Z, Zhou S G, Childs K L, Davidson R M, Lin H N, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee S S, Kim J, Numa H, Itoh T, Buell C R, Matsumoto T.2013. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data.Rice, 6: 1-10. |
| 16 | Khush G S.2001. Green revolution: The way forward.Nat Rev Genet, 2: 815-822. |
| 17 | Lan T, Gao J, Zeng Q Y.2013. Genome-wide analysis of the LEA (late embryogenesis abundant) protein gene family in Populus trichocarpa.Tree Genet Genom, 9(1): 253-264. |
| 18 | Langridge P, Fleury D.2011. Making the most of ‘omics’ for crop breeding.Trends Biotechnol, 29(1): 33-40. |
| 19 | Lata C, Yadav A, Prasad M.2011. Role of plant transcription factors in abiotic stress tolerance. In: Shanker A K, Venkateswarlu B. Abiotic Stress Response in Plants: Physiological, Biochemical and Genetic Perspectives. Croatia: InTech: 269-296. |
| 20 | Lescot M, Dehais P, Thijs G, Marchal K, Moreau Y, van de Peer Y, Rouze P, Rombauts S.2002. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences.Nucl Acids Res, 30(1): 325-327. |
| 21 | Mittler R.2006. Abiotic stress, the field environment and stress combination.Trends Plant Sci, 11(1): 15-19. |
| 22 | Moons A, Gielen J, vandekerckhove J, van der Straeten D, Gheysen G, van Montagu M.1997. An abscisic-acid- and salt-stress- responsive rice cDNA from a novel plant gene family.Planta, 202(4): 443-454. |
| 23 | Nakagawa H, Ohmiya K, Hattori T.1996. A rice bZIP protein, designated OSBZ8, is rapidly induced by abscisic acid.Plant J, 9(2): 217-227. |
| 24 | Nantel A, Quatrano R S.1996. Characterization of three rice basic/leucine zipper factors, including two inhibitors of EmBP-1 DNA binding activity.J Biol Chem, 271: 31296-31305. |
| 25 | Pan Y L, Ma X, Liang H W, Zhao Q, Zhu D Y, Yu J J.2015. Spatial and temporal activity of the foxtail millet (Setaria italica) seed-specific promoter pF128.Planta, 241(1): 57-67. |
| 26 | Shah S H, Jan S A, Ahmad N, Khan S U, Kumar T, Iqbal A, Nasir F, Noman M, Ali U A.2015. Use of different promoters in transgenic plant development: Current challenges and future perspectives.Am Eur J Agric Environ Sci, 15(4): 664-675. |
| 27 | Sundheep R, Arul L, Nagarajan P, Balasubramanian P.2008. Sprome: A database on promoters of abiotic stress inducible genes in rice.Int J Integr Biol, 2: 153-156. |
| 28 | Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K.2000. Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions.Proc Natl Acad Sci USA, 97: 11632-11637. |
| 29 | Wani S H, Sah S K.2014. Biotechnology and abiotic stress tolerance in rice.J Rice Res, 2: e105. |
| 30 | Yamaguchi-Shinozaki K, Shinozaki K.1994. A novel cis-acting element in an Arabidopsis gene is involved in responsiveness to drought, low-temperature, or high-salt stress.Plant Cell, 6(2): 251-264. |
| 31 | Yamaguchi-Shinozaki K, Shinozaki K.2006. Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses.Annu Rev Plant Biol, 57: 781-803. |
| 32 | Youm J W, Jeon J H, Choi D, Yi S Y, Joung H, Kim H S.2008. Ectopic expression of pepper CaPF1 in potato enhances multiple stresses tolerance and delays initiation of in vitro tuberization.Planta, 228(4): 701-708. |
| 33 | Yu J, Hu S N, Wang J, Wong G K, Li S G, Liu B, Deng Y J, Dai L, Zhou Y, Zhang X Q, Cao M L, Liu J, Sun J D, Tang J B, Chen Y J, Huang X B, Lin W, Ye C, Tong W, Cong L J, Geng J N, Han Y J, Li L, Li W, Hu G Q, Huang X G, Li W J, Li J, Liu Z W, Li L, Liu J P, Qi Q H, Liu J S, Li L, Li T, Wang X G, Lu H, Wu T T, Zhu M, Ni P X, Han H, Dong W, Ren X Y, Feng X L, Cui P, Li X R, Wang H, Xu X, Zhai W X, Xu Z, Zhang J S, He S J, Zhang J G, Xu J C, Zhang K L, Zheng X W, Dong J H, Zeng W Y, Tao L, Ye J, Tan J, Ren X D, Chen X W, He J, Liu D F, Tian W, Tian C G, Xia H A, Bao Q Y, Li G, Gao H, Cao T, Wang J, Zhao W M, Li P, Chen W, Wang X D, Zhang Y, Hu J F, Wang J, Liu S, Yang J, Zhang G Y, Xiong Y Q, Li Z J, Mao L, Zhou C S, Zhu Z, Chen R S, Hao B L, Zheng W M, Chen S Y, Guo W, Li G J, Liu S Q, Tao M, Wang J, Zhu L H, Yuan L P, Yang H M.2002. A draft sequence of the rice genome (Oryza sativa L. ssp. indica).Science, 296: 79-92. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||