
Rice Science ›› 2017, Vol. 24 ›› Issue (6): 336-348.DOI: 10.1016/j.rsci.2017.01.003
• Orginal Article • Previous Articles Next Articles
Chandra Roy Subhas1( ), Bhasker Reddy Lachagari Vijaya2
), Bhasker Reddy Lachagari Vijaya2
Received:2016-10-06
Accepted:2017-01-13
Online:2017-11-28
Published:2017-08-30
Chandra Roy Subhas, Bhasker Reddy Lachagari Vijaya. Assessment of SNP and InDel Variations Among Rice Lines of Tulaipanji x Ranjit[J]. Rice Science, 2017, 24(6): 336-348.
Add to citation manager EndNote|Ris|BibTeX
| Sample | Grain length (mm) | Grain width (mm) | 1000-grain weight (g) | Awn length (mm) |
|---|---|---|---|---|
| Tulaipanji | 7.73 | 1.88 | 15.72 | 21.01 |
| Ranjit | 8.07 | 2.39 | 18.62 | 0 |
| Progeny-awn | 7.85 | 1.98 | 15.45 | 14.77 |
| Progeny-awnless | 8.46 | 2.25 | 17.52 | 0 |
Table 1 Morphological characteristic of four rice samples.
| Sample | Grain length (mm) | Grain width (mm) | 1000-grain weight (g) | Awn length (mm) |
|---|---|---|---|---|
| Tulaipanji | 7.73 | 1.88 | 15.72 | 21.01 |
| Ranjit | 8.07 | 2.39 | 18.62 | 0 |
| Progeny-awn | 7.85 | 1.98 | 15.45 | 14.77 |
| Progeny-awnless | 8.46 | 2.25 | 17.52 | 0 |
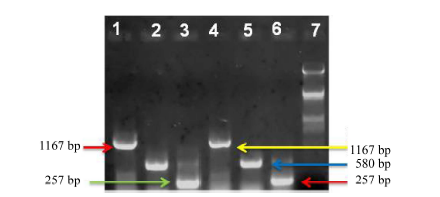
Fig. 2. Gene specific PCR amplified product fractioned on 1% agarose gel electrophoresis in Tulaipanji and Kalonunia (for check) landraces. Allele specific primer was used for SNP genotyping in respect to BAD2 gene and OsDREB transcription factor gene (abiotic stress responsive). Lanes 1 and 4, OsDREB amplified product of 1 167 bp; Lanes 2 and 5, 580 bp products as a positive control amplified by both external primers (ESP and EAP) of BAD2; Lanes 3 and 6, 257 bp amplified by the internal fragrant antisense primer (IFAP) and the external sense primer (ESP); Lane 7, Lambda DNA Hind III & EcoR I digested marker.
| Sample | Number of raw reads (bp) | Number of bases (bp) | GC content (%) | Percentage of data (%) | Raw read length | Number of preprocessed reads (bp) | |||
|---|---|---|---|---|---|---|---|---|---|
| Tulaipanji | 4 108 370 | 410 830 000 | 40.7 | 92.24 | 100×2 | 4 055 808 | |||
| Ranjit | 2 890 990 | 289 090 000 | 41.2 | 92.21 | 100×2 | 2 835 872 | |||
| Progeny-awn | 3 099 014 | 309 900 000 | 42.8 | 91.66 | 100×2 | 3 046 924 | |||
| Progeny-awnless | 3 131 956 | 313 190 000 | 43.1 | 91.91 | 100×2 | 3 091 894 | |||
| Sample | Percentage of preprocessed reads (%) | Number of reads aligned (bp) | Percentage of reads aligned (%) | Number of uniquely aligned reads (bp) | Percentage of uniquely aligned reads (%) | ||||
| Tulaipanji | 98.72 | 3 564 468 | 87.89 | 3 450 006 | 96.78 | ||||
| Ranjit | 98.09 | 2 417 235 | 85.24 | 2 326 571 | 96.24 | ||||
| Progeny-awn | 98.31 | 2 681 183 | 88.00 | 2 596 873 | 96.85 | ||||
| Progeny-awnless | 98.72 | 2 697 221 | 87.24 | 2 587 535 | 95.93 | ||||
Table 2 Summary and alignment statistics of next generation sequencing reads mapped of four rice lines with reference genome Nipponbare.
| Sample | Number of raw reads (bp) | Number of bases (bp) | GC content (%) | Percentage of data (%) | Raw read length | Number of preprocessed reads (bp) | |||
|---|---|---|---|---|---|---|---|---|---|
| Tulaipanji | 4 108 370 | 410 830 000 | 40.7 | 92.24 | 100×2 | 4 055 808 | |||
| Ranjit | 2 890 990 | 289 090 000 | 41.2 | 92.21 | 100×2 | 2 835 872 | |||
| Progeny-awn | 3 099 014 | 309 900 000 | 42.8 | 91.66 | 100×2 | 3 046 924 | |||
| Progeny-awnless | 3 131 956 | 313 190 000 | 43.1 | 91.91 | 100×2 | 3 091 894 | |||
| Sample | Percentage of preprocessed reads (%) | Number of reads aligned (bp) | Percentage of reads aligned (%) | Number of uniquely aligned reads (bp) | Percentage of uniquely aligned reads (%) | ||||
| Tulaipanji | 98.72 | 3 564 468 | 87.89 | 3 450 006 | 96.78 | ||||
| Ranjit | 98.09 | 2 417 235 | 85.24 | 2 326 571 | 96.24 | ||||
| Progeny-awn | 98.31 | 2 681 183 | 88.00 | 2 596 873 | 96.85 | ||||
| Progeny-awnless | 98.72 | 2 697 221 | 87.24 | 2 587 535 | 95.93 | ||||
| Sample | At read depth 2 | At read depth 5 | At read depth 10 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| No. of SNPs | No. of InDels | Total | No. of SNPs | No. of InDels | Total | No. of SNPs | No. of InDels | Total | |
| Tulaipanji | 12 972 | 925 | 13 897 | 10 218 | 701 | 10 919 | 7 476 | 489 | 7 965 |
| Ranjit | 18 042 | 1 703 | 19 745 | 13 510 | 1 300 | 14 810 | 9 966 | 950 | 10 916 |
| Progeny-awn | 31 303 | 2 383 | 33 686 | 25 760 | 1 988 | 27 748 | 19 559 | 1 514 | 21 073 |
| Progeny-awnless | 26 097 | 2 262 | 28 359 | 21 028 | 862 | 22 890 | 15 809 | 1 374 | 17 183 |
Table 3 Sample-wise polymorphic variant summary for four rice lines in different read depths.
| Sample | At read depth 2 | At read depth 5 | At read depth 10 | ||||||
|---|---|---|---|---|---|---|---|---|---|
| No. of SNPs | No. of InDels | Total | No. of SNPs | No. of InDels | Total | No. of SNPs | No. of InDels | Total | |
| Tulaipanji | 12 972 | 925 | 13 897 | 10 218 | 701 | 10 919 | 7 476 | 489 | 7 965 |
| Ranjit | 18 042 | 1 703 | 19 745 | 13 510 | 1 300 | 14 810 | 9 966 | 950 | 10 916 |
| Progeny-awn | 31 303 | 2 383 | 33 686 | 25 760 | 1 988 | 27 748 | 19 559 | 1 514 | 21 073 |
| Progeny-awnless | 26 097 | 2 262 | 28 359 | 21 028 | 862 | 22 890 | 15 809 | 1 374 | 17 183 |
| Sample | Nipponbare | Tulaipanji | Ranjit | Progeny-awnless |
|---|---|---|---|---|
| Tulaipanji (20.18%) | 3 698 | |||
| Ranjit (2.46%) | 10 855 | 10 013 | ||
| Progeny-awnless (2.24%) | 15 273 | 12 639 | 5 122 | |
| Progeny-awn (83.88%) | 1 662 | 341 | 1 499 | 951 |
Table 4 Common polymorphic homozygous markers at read depth 10.
| Sample | Nipponbare | Tulaipanji | Ranjit | Progeny-awnless |
|---|---|---|---|---|
| Tulaipanji (20.18%) | 3 698 | |||
| Ranjit (2.46%) | 10 855 | 10 013 | ||
| Progeny-awnless (2.24%) | 15 273 | 12 639 | 5 122 | |
| Progeny-awn (83.88%) | 1 662 | 341 | 1 499 | 951 |
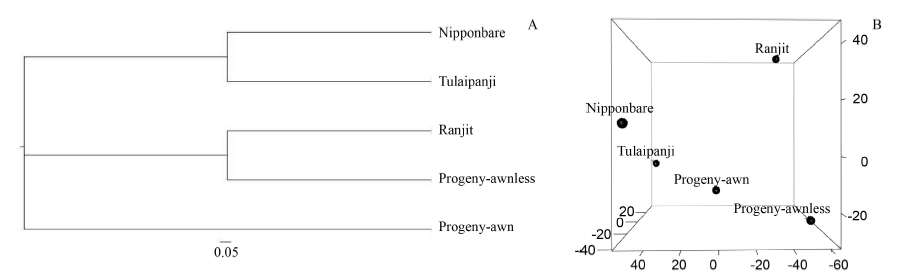

Fig. 3. Neighbour-joining tree (A) and principle component analysis (B) of Nipponbare and four rice lines based on similarity matrix of genotyping data.
| Sample | Allele | Heterozygous allele | Total missing allele | Total polymorphic allele | Recovery (%) | |||
|---|---|---|---|---|---|---|---|---|
| Tulaipanji | Ranjit | Tulaipanji | Ranjit | |||||
| Nipponbare | 7 500 | 2 476 | 0 | 37 | 9 939 | 75.46 | 24.54 | |
| Tulaipanji | 10 013 | 0 | 0 | 0 | 10 013 | 100.00 | 0.00 | |
| Ranjit | 0 | 10 013 | 0 | 0 | 10 013 | 0.00 | 100.00 | |
| Progeny-awnless | 1 006 | 8 629 | 47 | 331 | 9 304 | 10.81 | 89.19 | |
| Progeny-awn | 1 212 | 158 | 8 583 | 60 | 1 310 | 92.52 | 7.48 | |
Table 5 Genomic introgression based on polymorphic allelic from genotyping-by-sequencing.
| Sample | Allele | Heterozygous allele | Total missing allele | Total polymorphic allele | Recovery (%) | |||
|---|---|---|---|---|---|---|---|---|
| Tulaipanji | Ranjit | Tulaipanji | Ranjit | |||||
| Nipponbare | 7 500 | 2 476 | 0 | 37 | 9 939 | 75.46 | 24.54 | |
| Tulaipanji | 10 013 | 0 | 0 | 0 | 10 013 | 100.00 | 0.00 | |
| Ranjit | 0 | 10 013 | 0 | 0 | 10 013 | 0.00 | 100.00 | |
| Progeny-awnless | 1 006 | 8 629 | 47 | 331 | 9 304 | 10.81 | 89.19 | |
| Progeny-awn | 1 212 | 158 | 8 583 | 60 | 1 310 | 92.52 | 7.48 | |
| Class | At read depth 10 |
|---|---|
| Intergenic | 16 490 |
| Inside gene | 7 812 |
| Exonic | 3 377 |
| Exonic-CDS | 2 352 |
| Silent | 929 |
| Missense | 1 325 |
| Nonsense | 52 |
| Startloss | 1 |
| Stoploss | 12 |
| Frameshift-InDel | 22 |
| Inframe | 11 |
| Exonic-5′-UTR | 215 |
| Exonic-3′-UTR | 810 |
| Intronic | 4 435 |
| Intronic-3SPLICE_SITE | 109 |
| Intronic-5SPLICE_SITE | 92 |
| Intronic-others | 4 234 |
Table 6 Genome annotation and functional variation analysis.
| Class | At read depth 10 |
|---|---|
| Intergenic | 16 490 |
| Inside gene | 7 812 |
| Exonic | 3 377 |
| Exonic-CDS | 2 352 |
| Silent | 929 |
| Missense | 1 325 |
| Nonsense | 52 |
| Startloss | 1 |
| Stoploss | 12 |
| Frameshift-InDel | 22 |
| Inframe | 11 |
| Exonic-5′-UTR | 215 |
| Exonic-3′-UTR | 810 |
| Intronic | 4 435 |
| Intronic-3SPLICE_SITE | 109 |
| Intronic-5SPLICE_SITE | 92 |
| Intronic-others | 4 234 |
| [1] | Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R.2012. Genome sequencing reveals agronomically important loci in rice using MutMap.Nat Biotechnol, 30(2): 174-178. |
| [2] | Ahn S N, Bollisch C N, Tanksley S D.1992. RFLP tagging of a gene for aroma in rice.Theor Appl Genet, 84: 825-828. |
| [3] | Akhtar M, Jaiswal A, Taj G, Jaiswal J P, Qureshi M I, Singh N K.2012. DREB1/CBF transcription factors: Their structure, function and role in abiotic stress tolerance in plants.J Genet, 91(3): 385-395. |
| [4] | Arbelaez J D, Moreno L T, Singh N, Tung C W, Maron L G, Ospina Y, Martinez C P, Grenier C, Lorieux M, McCouch S.2015. Development and GBS-genotyping of introgression lines (ILs) using two wild species of rice, O. meridionalis and O. rufipogon, in a common recurrent parent, O. sativa cv. Curinga. Mol Breeding, 35: 81. |
| [5] | Asins M J.2002. Present and future of quantitative trait locus analysis in plant breeding.Plant Breeding, 121(4): 281-291. |
| [6] | Borevitz J O, Nordborg M.2003. The impact of genomics on the study of natural variation in Arabidopsis.Plant Physiol, 132(2): 718-725. |
| [7] | Bouman B, Barker R, Humphreys E, Tuong T P, Atlin G, Bennett J, Dawe D, Dittert K, Dobermann A, Facon T, Fujimoto N, Gupta R, Haefele S, Hosen Y, Ismail A, Johnson D, Johnson S, Khan S, Shan L, Masih I, Matsuno Y, Pandey S, Peng S, Muthukumarisami T, Wassman R.2007. Rice: Feeding the billions. In: Water for Food, Water for Life: A Comprehensive Assessment of Water Management in Agriculture. Colombo: International Water Management Institute: 515-549. |
| [8] | Bradbury L M, Fitgerald T L, Henry R J, Jin Q S, Waters D L E.2005. The gene for fragrance in rice.Plant Biotechnol J, 3(3): 363-370. |
| [9] | Bradbury P J, Zhang Z W, Kroon D E, Casstevens T M, Ramdoss Y, Buckler E S.2007. TASSEL: Software for association mapping of complex traits in diverse samples.Bioinformatics, 23: 2633-2635. |
| [10] | Brooks S A, Yan W G, Jackson A K, Deren C W.2008. A natural mutation in rc reverts white-rice-pericarp to red and results in a new, dominant, wild-type allele: Rc-g.Theor Appl Genet, 117(4): 575-580. |
| [11] | Buttery R G, Ling L C, Juliano B O, Turnbaugh J G.1983. Cooked rice aroma and 2-acetyl-1-pyroline.J Agric Food Chem, 31: 823-826. |
| [12] | Chin J H, Gamuyao R, Dalid C, Bustamam M, Prasetiyono J, Moeljopawiro S, Wissuwa M, Heuer S.2011. Developing rice with high yield under phosphorus deficiency: Pup1 sequence to application.Plant Physiol, 156(3): 1202-1216. |
| [13] | Collard B C Y, Mackill D J.2008. Marker-assisted selection: An approach for precision plant breeding in the twenty-first century.Phil Trans R Soc Lond B: Biol Sci, 363: 557-572. |
| [14] | Deschamps S, Llaca V, May G D.2012. Genotyping-by sequencing in plants.Biology, 1(3): 460-483. |
| [15] | Dubouzet J G, Sakuma Y, Ito Y, Kasuga M, Dubouzet E G, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K.2003. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold- responsive gene expression.Plant J, 33(4): 751-763. |
| [16] | Duitama J, Silva A, Sanabria Y, Cruz D F, Quintero C, Ballen C, Lorieux M, Scheffler B, Farmer A, Torres E, Oard J, Tohme J.2015. Whole genome sequencing of elite rice cultivars as a comprehensive information resource for marker assisted selection.PLoS One, 10: e0124617. |
| [17] | Elshire R J, Glaubitz J C, Sun Q, Poland J A, Kawamoto K, Buckler E S, Mitchell S E.2011. A robust, simple genotyping- by-sequencing (GBS) approach for high diversity species.PLoS One, 6(5): e19379. |
| [18] | Endelman J B, Jannink J L.2012. Shrinkage estimation of the realized relationship matrix.G3: Genes Genom Genet, 2(11): 1405-1413. |
| [19] | Feltus F A, Wan J, Schulze S R, Estill J C, Jiang N, Paterson A H.2004. An SNP resource for rice genetics and breeding based on subspecies indica and japonica genome alignments.Genom Res, 14: 1812-1819. |
| [20] | Furukawa T, Maekawa M, Oki T, Suda I, Iida S, Shimada H, Takamure I, Kadowaki K.2007. The Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp.Plant J, 49(1): 91-102. |
| [21] | Gao L Z, Zhang C H, Li D Y, Pan D J, Jia J Z, Dong Y S.2006. Genetic diversity within Oryza rufipogon germplasms preserved in Chinese field gene banks of wild rice as revealed by microsatellite markers.Biodiv Conserv, 15: 4059-4077. |
| [22] | Gao Q, Yue G D, Li W Q, Wang J Y, Xu J H, Yin Y.2012. Recent progress using high-throughput sequencing technologies in plant molecular breeding.J Integr Plant Biol, 54(4): 215-227. |
| [23] | Gill B S, Friebe B R, White F F.2011. Alien introgression represents a rich source of genes for crop improvement.Proc Natl Acad Sci USA, 108: 7657-7658. |
| [24] | Godfray H C J, Beddington J R, Crute I R, Haddad L, Lawrence D, Muir J F, Pretty J, Robinson S, Thomas S M, Toulmin C.2010. Food security: The challenge of feeding 9 billion people.Science, 327: 812-818. |
| [25] | Guo L B, Gao Z Y, Qian Q.2014. Application of resequencing to rice genomics, functional genomics and evolutionary analysis.Rice, 7: 4-7. |
| [26] | Han B, Huang X H.2013. Sequencing-based genome-wide association study in rice. Curr Opin Plant Biol, 16(2): 133-138. |
| [27] | Harushima Y, Nakagahra M, Yano M, Sasaki T, Kurata N.2001. A genome-wide survey of reproductive barriers in an intraspecific hybrid.Genetics, 159(2): 883-892. |
| [28] | Harushima Y, Nakagahra M, Yano M, Sasaki T, Kurata N.2002. Diverse variation of reproductive barriers in three intraspecific rice crosses.Genetics, 160(1): 313-322. |
| [29] | Heslot N, Rutkoski J, Poland J, Jannink J L, Sorrells M E.2013. Impact of marker ascertainment bias on genomic selection accuracy and estimates of genetic diversity.PLoS One, 8: e74612. |
| [30] | Huang R Y, Jiang L R, Zheng J S, Wang T S, Wang H C, Huang Y M, Hong Z L.2013. Genetic bases of rice grain shape: So many genes, so little known. Trends Plant Sci, 18(4): 218-226. |
| [31] | Huang X H, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan J P, Fan D L, Weng Q J, Huang T, Dong G J, Sang T, Han B.2009. High-throughput genotyping by whole-genome resequencing.Genom Res, 19(6): 1068-1076. |
| [32] | Huang X H, Wei X H, Sang T, Zhao Q, Feng Q, Zhao Y, Li C Y, Zhu C R, Lu T T, Zhang Z W, Li M, Fan D L, Guo Y L, Wang A, Wang L, Deng L W, Li W J, Lu Y Q, Weng Q J, Liu K, Huang T, Zhou T Y, Jing Y F, Li W, Lin Z, Buckler E S, Qian Q, Zhang Q F, Li J Y, Han B.2010. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet, 42: 961-967. |
| [33] | Huang X H, Lu T T, Han B.2013. Resequencing rice genomes: An emerging new era of rice genomics.Trends Genet, 29(4): 225-232. |
| [34] | Imai I, Kimball J A, Conway B, Yeater K M, McCouch S R, McClung A.2013. Validation of yield-enhancing quantitative trait loci from a low-yielding wild ancestor of rice.Mol Breeding, 32(1): 101-120. |
| [35] | IRGSP.2005. The map-based sequence of the rice genome.Nature, 7: 793-800. |
| [36] | Izawa T, Konishi S, Shomura A, Yano M.2009. DNA changes tell us about rice domestication.Curr Opin Plant Biol, 12(2): 185-192. |
| [37] | Jeong I S, Yoon U H, Lee G S, Ji H S, Lee H J, Han C D, Hahn J H, An G H, Kim T H.2013. SNP-based analysis of genetic diversity in anther-derived rice by whole genome sequencing.Rice, 7: 6. |
| [38] | Jin Q S, Waters D, Cordeiro G M, Henry R J, Reineke R F.2003. A single nucleotide polymorphism (SNP) marker linked to the fragrance gene in rice (Oryza sativa L.).Plant Sci, 165(2): 359-364. |
| [39] | Kawahara Y, de la Bastide M, Hamilton J P, Kanamori H, McCombie W R, Ouyang S, Schwartz D C, Tanaka T, Wu J Z, Zhou S G, Childs K L, Davidson R M, Lin H N, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee S S, Kim J, Numa H, Itoh T, Buell C R, Matsumoto T.2013. Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data.Rice, 6: 4-10. |
| [40] | Kilian B, Graner A.2012. NGS technologies for analyzing germplasm diversity in genebanks.Brief Funct Genom, 11(1): 38-50. |
| [41] | Kim R B, Jang S M, Chu S H, Bordiya Y, Akter M B, Lee J, Chin J H, Koh H J.2014. Analysis of segregation distortion and its relationship to hybrid barriers in rice.Rice, 7: 3. |
| [42] | Kimura M. 1983. The Neutral Theory of Molecular Evolution. Cambridge, UK Cambridge University Press. |
| [43] | Kovach M J, McCouch S R.2008. Leveraging natural diversity: Back through the bottleneck.Curr Opin Plant Biol, 11(2): 193-200. |
| [44] | Langmead B, Salzberg S L.2012. Fast gapped-read alignment with Bowtie 2.Nat Methods, 9: 357-359. |
| [45] | Li Z K, Zhang F.2013. Rice breeding in the post-genomics era: From concept to practice.Curr Opin Plant Biol, 16(2): 261-269. |
| [46] | Lipka A E, Tian F, Wang Q S, Peiffer J, Li M, Bradbury P J, Gore M A, Buckler E S, Zhang Z W.2012. GAPIT: Genome association and prediction integrated tool.Bioinformatics, 28: 2397-2399. |
| [47] | Little R R, Hilder G B, Dawson E H.1958. Differential effect of dilute alkali on 25 varieties of milled white rice.Cereal Chem, 35(2): 111-126. |
| [48] | Liu G J, Bernhardt J L, Jia M H, Wamishe Y A, Jia Y L.2008. Molecular characterization of the recombinant inbred line population derived from a japonica/indica rice cross.Euphytica, 159: 73-82. |
| [49] | Lorieux M, Tohme J, McCouch S R, Brondani C, Gridley H, Martinez C P, Diago M.2004. Exploring natural genetic variation: Developing genomic resources and introgression lines for four AA genome rice relatives. A proposal to the Generation Challenge Program Standard Grant: 1-43. |
| [50] | Matsushita S, Iseki T, Fukuta Y, Araki E, Kobayashi S, Osaki M, Yamagishi M.2003. Characterization of segregation distortion on chromosome 3 induced in wide hybridization between indica and japonica type rice varieties.Euphytica, 134(1): 27-32. |
| [51] | McCouch S, Baute G J, Bradeen J, Bramel P, Bretting P K, Buckler E, Burke J M, Charest D, Cloutier S, Cole G, Dempewolf H, Dingkuhn M, Feuillet C, Gepts P, Grattapaglia D, Guarino L, Jackson S, Knapp S, Langridge P, Lawton-Rauh A, Lijua Q, Lusty C, Michael T, Myles S, Naito K, Nelson R L, Pontarollo R, Richards C M, Rieseberg L, Ross-Ibarra J, Rounsley S, Hamilton R S, Schurr U, Stein N, Tomooka N, van der Knaap E, van Tassel D, Toll J, Valls J, Varshney R K, Ward J, Waugh R, Wenzl P, Zamir D.2013. Agriculture: Feeding the future.Nature, 499: 23-24. |
| [52] | McCouch S R, Zhao K, Wright M, Tung C W, Ebana K, Thomson M, Reynolds A, Wang D, DeClerck G, Ali M L, McClung A, Eizenga G, Bustamante C.2010. Development of genome-wide SNP assays for rice.Breeding Sci, 60: 524-535. |
| [53] | McCouch S R, McNally K L, Wang W, Hamilton R S.2012. Genomics of gene banks: A case study in rice.Am J Bot, 99(2): 407-423. |
| [54] | Mir R R, Varshney R K. 2012. Future prospects of molecular markers in plants. In: Henry R J. Molecular Markers in Plants. Oxford, UK Backwell Publishing. |
| [55] | Mishra B.2002. Varietal improvement for rice production in India. In: Nguyen V N. Genetic Diversity in Rice Production, Case Studies from Brazil, India and Nigeria. Rome, Italy: FAO: 37-91. |
| [56] | Nei M. 1987. Molecular Evolutionary Genetics. New York, USA: Columbia University Press. |
| [57] | Periyannan S, Moore J, Ayliffe M, Bansal U, Wang X J, Huang L, Deal K, Luo M C, Kong X Y, Bariana H, Mago R, McIntosh R, Dodds P, Dvorak J, Lagudah E.2013. The gene Sr33, an ortholog of barley Mla genes, encodes resistance to wheat stem rust race Ug99.Science, 341: 786-788. |
| [58] | Peterson B K, Weber J N, Kay E H, Fisher H S, Hoekstra H E.2012. Double digest RADseq: An inexpensive method for de novo SNP discovery and genotyping in model and non-model species.PLoS One, 7(5): e37135. |
| [59] | Poland J A, Rife T W.2012. Genotyping-by-sequencing for plant breeding and genetics.Plant Genome, 5(3): 92-102. |
| [60] | Rafalski A.2002. Applications of single nucleotide polymorphisms in crop genetics.Curr Opin Plant Biol, 5(2): 94-100. |
| [61] | Rowe H C, Renaut S, Guggisberg A.2011. RAD in the realm of next-generation sequencing technologies.Mol Ecol, 20(1): 3499-3502. |
| [62] | Saintenac C, Jiang D, Wang S, Akhunov E.2013. Sequence-based mapping of the polyploid wheat genome.G3: Genes Genom Genet, 3: 1105-1114. |
| [63] | Santure A W, Stapley J, Ball A D, Birkhead T R, Burke T, Slate J.2010. On the use of large marker panels to estimate inbreeding and relatedness: Empirical and simulation studies of a pedigreed Zebra finch population typed at 771 SNPs.Mol Ecol, 19(7): 1439-1451. |
| [64] | Sha X Y. 2013. Rice artificial hybridization for genetic analysis. In: Yang Y N. Rice Protocols, Methods in Molecular Biology. New York, USA Humana Press: 1-12. |
| [65] | Shekhar T C, Goyal A.2014. Antioxidant activity by DPPH radical scavenging method of Ageratum conyzoides linn. leaves.Am J Ethnomed, 1: 244-249. |
| [66] | Shirley B W.1998. Flavonoids in seeds and grains: Physiological function, agronomic importance and the genetics of biosynthesis.Seed Sci Res, 8(4): 415-422. |
| [67] | Sleper D A, Poehlman J M. 2007. Breeding rice. In: Breeding Field Crops. 5th edn. Iowa: Blackwell Publishing: 239-257. |
| [68] | Song W Y, Wang G L, Chen L L, Kim H S, Pi L Y, Holsten T, Gardner J, Wang B, Zhai W X, Zhu L H, Fauquet C, Ronald P.1995. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21.Science, 270: 1804-1806. |
| [69] | Sood B C, Siddiq E A.1978. A rapid technique for scent determination in rice.Ind J Genet Plant Breeding, 38(2): 268-275. |
| [70] | Spindel J, Wright M, Chen C, Cobb J, Gage J, Harrington S, Lorieux M, Ahmadi N, McCouch S.2013. Bridging the genotyping gap: Using genotyping by sequencing (GBS) to add high-density SNP markers and new value to traditional bi-parental mapping and breeding populations.Theor Appl Genet, 126(11): 2699-2716. |
| [71] | Spindel J, Begum H, Akdemir D, Virk P, Collard B, Redoña E, Atlin G, Jannink J L, McCouch S R.2015. Genomic selection and association mapping in rice (Oryza sativa): Effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines.PLoS Genet, 11(2): e1004982. |
| [72] | Subudhi P K, Sasaki T, Khush G S.2006. Rice. In: Genome Mapping and Molecular Breeding in Plants. Berlin, Heidelberg, Germany: Springer: 1-78. |
| [73] | Sun C Q, Wang X K, Li Z C, Yoshimura A, Iwata N.2001. Comparison of the genetic diversity of common wild rice (Oryza rufipogon Griff.) and cultivated rice (O. sativa L.) using RFLP markers.Theor Appl Genet, 102(1): 157-162. |
| [74] | Sweeney M T, Thomson M J, Cho Y G, Park Y J, Williamson S H, Bustamante C D, McCouch S R.2007. Global dissemination of a single mutation conferring white pericarp in rice.PLoS Genet, 3(8): e133. |
| [75] | Syvanen A C.2001. Accessing genetic variation: Genotyping single nucleotide polymorphisms.Nat Rev Genet, 2: 930-942. |
| [76] | Tajima F.1989. Statistical methods to test for nucleotide mutation hypothesis by DNA polymorphism.Genetics, 123: 585-595. |
| [77] | Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S, Innan H, Cano L M, Kamoun S, Terauchi R.2013. QTL-seq: Rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations.Plant J, 74(1): 174-183. |
| [78] | Tang W J, Wu T T, Ye J, Sun J, Jiang Y, Yu J, Tang J P, Chen G M, Wang C M, Wan J M.2016. SNP-based analysis of genetic diversity reveals important alleles associated with seed size in rice.BMC Plant Biol, 16: 93. |
| [79] | Tanksley S D, McCouch S R.1997. Seed banks and molecular maps, unlocking genetic potential from the wild.Science, 277: 1063-1066. |
| [80] | Thomson M J.2014. High-throughput SNP genotyping to accelerate crop improvement.Plant Breeding Biotechnol, 2(3): 195-212. |
| [81] | VaRaden P M.2008. Efficient methods to compute genomic predictions.J Dairy Sci, 91: 4414-4423. |
| [82] | Wang C M, Zhu C S, Zhai H Q, Wan J M.2005. Mapping segregation distortion loci and quantitative trait loci for spikelet sterility in rice (Oryza sativa L.).Genet Res, 86(2): 97-106. |
| [83] | Wang G L, MacKill D J, Bonman J M, McCouch S R, Champoux M C, Nelson R J.1994. RFLP mapping of genes conferring complete and partial resistance to blast in a durable resistant rice cultivar.Genetics, 136(4): 1421-1434. |
| [84] | Wang L, Wang A, Huang X H, Zhao Q, Dong G J, Qian Q, Sang T, Han B.2011. Mapping 49 quantitative trait loci at high resolution through sequencing-based genotyping of rice recombinant inbred lines.Theor Appl Genet, 122(2): 327-340. |
| [85] | Wang S H, Tan X L, Tan Y L, Zhang Z L, Wen J C, Kou S Y.2009. Segregation distortion detected in six rice F2 populations generated from reciprocal hybrids at three altitudes.Genet Res, 91(5): 345-353. |
| [86] | Wang Z Y, Second G, Tanksley S D.1992. Polymorphism and phylogenetic relationships among species in the genus Oryzae as determined by analysis of nuclear RFLPs.Theor Appl Genet, 83(5): 565-581. |
| [87] | Wu D H, Wu H P, Wang C S, Tseng H Y, Hwu K K.2013. Genome-wide InDel marker system for application in rice breeding and mapping studies.Euphytica, 192(1): 131-143. |
| [88] | Wu Y P, Ko P Y, Lee W C, Wei F J, Kuo S C, Ho S W, Hour A L, Hsing Y I, Lin Y R.2010. Comparative analyses of linkage maps and segregation distortion of two F2 populations derived from japonica crossed with indica rice.Hereditas, 147(5): 225-236. |
| [89] | Xiao J H, Li J M, Grsndillo S, Ahn S, Yuan L P, McCouch S R, Tanksley S D.1996. Genes from wild rice improved yield.Nature, 384: 223-224. |
| [90] | Xie X B, Jin F X, Song M H, Suh J P, Hwang H G, Kim Y G, McCouch S R, Ahn S N.2008. Fine mapping of a yield-enhancing QTL cluster associated with transgressive variation in an Oryza sativa × O. rufipogon cross.Theor Appl Genet, 116(5): 613-622. |
| [91] | Xu X, Liu X, Ge S, Jensen J D, Hu F, Li X, Dong Y, Gutenkunst R N, Fang L, Huang L, Li J X, He W M, Zhang G J, Zheng X M, Zhang F M, Li Y R, Yu C, Kristiansen K, Zhang X Q, Wang J, Wright M, McCouch S, Nielsen R, Wang J, Wang W.2012. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes.Nat Biotechnol, 30: 105-111. |
| [92] | Xu Y, Zhu L, Xiao J, Huang N, McCouch S R.1997. Chromosomal regions associated with segregation distortion of molecular markers in F2, backcross, doubled haploid, and recombinant inbred populations in rice (Oryza sativa L).Mol Gen Genet, 253(5): 535-545. |
| [93] | Xu Y B.2010. Molecular Plant Breeding. UK: CABI: 148-150. |
| [94] | Yamagishi M, Takeuchi Y, Tanakka I, Kono I, Murai K, Yano M.2010. Segregation distortion in F2 and double haploid populations of temperate japonica rice.J Genet, 89(2): 237-241. |
| [95] | Yonemaru J, Choi S H, Sakai H, Ando T, Shomura A, Yano M, Fukuoka S.2015. Genome-wide indel markers shared by diverse Asian rice cultivars compared to Japanese rice cultivar “Koshihikari”.Breeding Sci, 65(3): 249-256. |
| [96] | Yu J, Hu S N, Wang J, Wong G K S, Li S G, Liu B, Deng Y J, Dai L, Zhou Y, Zhang X Q, Cao M L, Liu J, Sun J D, Tang J B, Chen Y J, Huang X B, Lin W, Ye C, Tong W, Cong L J, Geng J N, Han Y J, Li L, Li W, Hu G Q, Huang X G, Li W J, Li J, Liu Z W, Li L, Liu J P, Qi Q H, Liu J S, Li L, Li T, Wang X G, Lu H, Wu T T, Zhu M, Ni P X, Han H, Dong W, Ren X Y, Feng X L, Cui P, Li X R, Wang H, Xu X, Zhai W X, Xu Z, Zhang J S, He S J, Zhang J G, Xu J C, Zhang K L, Zheng X W, Dong J H, Zeng W Y, Tao L, Ye J, Tan J, Ren X D, Chen X W, He J, Liu D F, Tian W, Tian C G, Xia H G, Bao Q Y, Li G, Gao H, Cao T, Wang J, Zhao W M, Li P, Chen W, Wang X D, Zhang Y, Hu J F, Wang J, Liu S, Yang J, Zhang G Y, Xiong Y Q, Li Z J, Mao L, Zhou C S, Zhu Z, Chen R S, Hao B L, Zheng W M, Chen S Y, Guo W, Li G J, Liu S Q, Tao M, Wang J, Zhu L H, Yuan L P, Yang H M.2002. A draft sequence of the rice genome (Oryza sativa L. ssp. indica).Science, 296: 79-92. |
| [97] | Zamir D.2001. Improving plant breeding with exotic genetic libraries.Nat Rev Genet, 2: 983-989. |
| [98] | Zhang Q, Lin S C, Zhao B Y, Wang C L, Yang W C, Zhou Y L, Li D Y, Chen C B, Zhu L H.1998. Identification and tagging of a new gene for resistance to bacterial blight (Xanthomonas oryzae pv. oryzae) from O. rufipogon.Rice Genet Newsl, 15: 138-142. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||