
Rice Science ›› 2017, Vol. 24 ›› Issue (6): 299-321.DOI: 10.1016/j.rsci.2017.08.001
• Orginal Article • Next Articles
Srivastava Deepti1, Shamim Md2, Kumar Mahesh2, Mishra Anurag3, Pandey Pramila4, Kumar Deepak4, Yadav Prashant5, Harrish Siddiqui Mohammed1, Narayan Singh Kapildeo4( )
)
Received:2017-05-21
Accepted:2017-08-08
Online:2017-11-28
Published:2017-08-30
Srivastava Deepti, Shamim Md, Kumar Mahesh, Mishra Anurag, Pandey Pramila, Kumar Deepak, Yadav Prashant, Harrish Siddiqui Mohammed, Narayan Singh Kapildeo. Current Status of Conventional and Molecular Interventions for Blast Resistance in Rice[J]. Rice Science, 2017, 24(6): 299-321.
Add to citation manager EndNote|Ris|BibTeX
| Fungicide | Nature | Mode of action | Active ingredient per hm2 | Reference |
|---|---|---|---|---|
| Probenazole | Systemic | Activates plant defence response | 1000-2400 mL | Filippi and Prabhu, 1997 |
| Tricyclazole | Systemic | Melanin biosynthesis inhibitor-R (polyhydroxyl-naphthalene reductase) | 225-300 mL | Anwar and Bhat, 2005 |
| Azoxystrobin | Systemic | QOI complex 3 inhibitor | 170 mL | Neelakanth et al, 2017 |
| Isoprothiolane | Systemic | Phospholipid biosynthesis (methyltransferase) and/or choline biosynthesis inhibitor | 300 mL | Sachin and Rana, 2011 |
| Propiconazole | Systemic | Sterol biosynthesis (14-demethylase) inhibitor | 125 mL | Prasanna Kumar and Veerabhadraswamy, 2014 |
| Benomyl | Systemic | Benomyl binds to microtubules, interfering with cell functions, such as meiosis and intracellular transportation | 120 mg | Kapoor and Singh, 1982 |
| Edifenphos | Systemic | Inhibitor of the biosynthesis of phosphatidylcholine | 250-300 mg | Anon, 1992; Filippi and Prabhu, 1997 |
| Iprobenfos | Systemic | Inhibitor of the biosynthesis of phosphatidylcholine | 300-500 mL | Gohel and Chauhan, 2015 |
| Pyroquilon | Systemic | Inhibitor of melanin biosynthesis inhibitors (MBIs) | 500 mL | Tsuda et al, 1998 |
| Diclocymet | Systemic | Inhibitor of melanin biosynthesis inhibitors (MBIs) | 300 mL | Kurahash, 2001 |
| Carpropamid | Root-systemic | Inhibits the enzyme scytalone dehydratase essential for the synthesis of the melanin biosynthesis and also induce resistance in plants | 139 mL | Motoyama et al, 1999 |
| Fenoxanil | Systemic | Inhibitor of melanin biosynthesis inhibitors (MBIs) | 120-150 mg | Nishimura and Hino, 2002 |
| Metominostrobin | Systemic | Inhibits respiratory electron transfer | 100 mg | - |
| Zineb | Contact | When being applied, it is converted to an isothiocyanate, and inactivates the sulphahydral groups enzymes of fungi | 1500 mg | - |
| Hexaconazole | Systemic | Inhibits ergosterol biosynthesis (steroid dimethylation inhibitor) | 50 mL | Prasanna Kumar et al, 2011 |
| Carbendszim 12% + Mancozeb 63% | Contact and systemic | Carbendazim works by inhibiting spindle formation at the mitosis stage and mancozeb affects the nervous system | 750 mg | Venkata Rao and Muralidharan, 1983 |
| Eprobenfos | Systemic | Eprobenfos block the synthesis of phospholipid and alters the membrane structure of fungus by increasing the permeability with consequent loss of vital cellular component | 240 mL | - |
| Kresoxim methyl | Contact and systemic (local) | Acts by binding to Qo site, blocking electron transfer and respiration of the fungi | 250 mg | Prasanna Kumar et al, 2011 |
| Tebuconazole | Systemic | Dimethylase inhibitor: interferes in process of building the structure of fungal cell wall | 187.5 mL | Ghazanfar et al, 2009 |
Table 1 Most common fungicides used against rice blast disease.
| Fungicide | Nature | Mode of action | Active ingredient per hm2 | Reference |
|---|---|---|---|---|
| Probenazole | Systemic | Activates plant defence response | 1000-2400 mL | Filippi and Prabhu, 1997 |
| Tricyclazole | Systemic | Melanin biosynthesis inhibitor-R (polyhydroxyl-naphthalene reductase) | 225-300 mL | Anwar and Bhat, 2005 |
| Azoxystrobin | Systemic | QOI complex 3 inhibitor | 170 mL | Neelakanth et al, 2017 |
| Isoprothiolane | Systemic | Phospholipid biosynthesis (methyltransferase) and/or choline biosynthesis inhibitor | 300 mL | Sachin and Rana, 2011 |
| Propiconazole | Systemic | Sterol biosynthesis (14-demethylase) inhibitor | 125 mL | Prasanna Kumar and Veerabhadraswamy, 2014 |
| Benomyl | Systemic | Benomyl binds to microtubules, interfering with cell functions, such as meiosis and intracellular transportation | 120 mg | Kapoor and Singh, 1982 |
| Edifenphos | Systemic | Inhibitor of the biosynthesis of phosphatidylcholine | 250-300 mg | Anon, 1992; Filippi and Prabhu, 1997 |
| Iprobenfos | Systemic | Inhibitor of the biosynthesis of phosphatidylcholine | 300-500 mL | Gohel and Chauhan, 2015 |
| Pyroquilon | Systemic | Inhibitor of melanin biosynthesis inhibitors (MBIs) | 500 mL | Tsuda et al, 1998 |
| Diclocymet | Systemic | Inhibitor of melanin biosynthesis inhibitors (MBIs) | 300 mL | Kurahash, 2001 |
| Carpropamid | Root-systemic | Inhibits the enzyme scytalone dehydratase essential for the synthesis of the melanin biosynthesis and also induce resistance in plants | 139 mL | Motoyama et al, 1999 |
| Fenoxanil | Systemic | Inhibitor of melanin biosynthesis inhibitors (MBIs) | 120-150 mg | Nishimura and Hino, 2002 |
| Metominostrobin | Systemic | Inhibits respiratory electron transfer | 100 mg | - |
| Zineb | Contact | When being applied, it is converted to an isothiocyanate, and inactivates the sulphahydral groups enzymes of fungi | 1500 mg | - |
| Hexaconazole | Systemic | Inhibits ergosterol biosynthesis (steroid dimethylation inhibitor) | 50 mL | Prasanna Kumar et al, 2011 |
| Carbendszim 12% + Mancozeb 63% | Contact and systemic | Carbendazim works by inhibiting spindle formation at the mitosis stage and mancozeb affects the nervous system | 750 mg | Venkata Rao and Muralidharan, 1983 |
| Eprobenfos | Systemic | Eprobenfos block the synthesis of phospholipid and alters the membrane structure of fungus by increasing the permeability with consequent loss of vital cellular component | 240 mL | - |
| Kresoxim methyl | Contact and systemic (local) | Acts by binding to Qo site, blocking electron transfer and respiration of the fungi | 250 mg | Prasanna Kumar et al, 2011 |
| Tebuconazole | Systemic | Dimethylase inhibitor: interferes in process of building the structure of fungal cell wall | 187.5 mL | Ghazanfar et al, 2009 |
| Chr | Gene | Tightly linked marker | Map position (cM) | Donor rice | Resistance type | Reference | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Type | Name | ||||||||||||||||||
| 1 | Pit | SNP | t311, t256, t8042 | 16.1 | Tjahaja | Complete | Hayashi et al, 2006 | ||||||||||||
| Pi27(t) | SSR | RM151, RN259 | 28.4-38.8 | Q14 | Complete | Sallaud et al, 2006 | |||||||||||||
| Pitp(t) | SSR | RM246 | 114.1 | Tetep | Partial | Barman et al, 2004 | |||||||||||||
| Pi35(t) | SSR | RM1216, RM1003 | 132.0-136.6 | Hokkai 188 | Partial | Nguyen et al, 2006 | |||||||||||||
| Pi37 | SSR | RM302, RM212, FPSM1, FPSM2, FPSM4 | 136.1 | St.No.1 | Complete | Lin et al, 2007 | |||||||||||||
| STS | S15628, FSTS1, FSTS2, FSTS3, FSTS4 | ||||||||||||||||||
| Pi64 | SSR | RM11715, RM11787 | - | Yangmaogu | Complete | Ma et al, 2015 | |||||||||||||
| Pish | RFLP | - | 148.7-154.8 | Shin 2 | Complete | Zhu et al, 2004 | |||||||||||||
| 2 | Pid1(t) | SSR | RM262 | 87.5-89.9 | Digu | Complete | Chen et al, 2004 | ||||||||||||
| Pig(t) | SSR | RM166, RM208 | 142.0-154.1 | Guangchangzhan | Complete | Shi et al, 2012 | |||||||||||||
| Pitq5 | RFLP | RG520, RZ446B, RZ446A, RG654, RG256 | 150.5-157.9 | Teqing | Complete | Zhou et al, 2004 | |||||||||||||
| Piy1(t) | SSR | RM3248, RM20 | 153.2-154.1 | Yanxian 1 | - | Fukuta et al, 2004 | |||||||||||||
| Piy2(t) | SSR | RM3248, RM20 | 153.2-154.1 | Yanxian 1 | - | Fukuta et al, 2004 | |||||||||||||
| Pib | SNP | b213, b28, b2, b3989, Pibdom | 154.1 | Tohoku, IL9, Koshiihikari | Complete | Hayashi et al, 2006 | |||||||||||||
| Pi14(t) | Isozyme | Amp-1 | 32.6 | Maowang | Complete | Tabien et al, 2000 | |||||||||||||
| Pi16(t) | Isozyme | Amp-1 | 34.3 | AUS373, Maowamg | Complete | Pan et al, 1996 | |||||||||||||
| Pi-Da(t) | SSR | RM5529, RM211 | 12.6 | Dacca 6 | - | Lei et al, 2005; Shi et al, 2012 | |||||||||||||
| 3 | Pi66(t) | SSR | RM487, RM16, RM55, RM168 | AS201 | Partial | Liang et al, 2016 | |||||||||||||
| 4 | Pi21 | STS | P702D03-#79 | 58.6 | Owarihatamochi | Partial | Pan et al, 1998 | ||||||||||||
| Pikur1 | Isozyme | - | 86.0 | Kuroka | - | Fukuoka et al, 2007 | |||||||||||||
| Pi39(t) | SSR | RM3843, RM5473 | 107.4-108.2 | Chubu 111 (Haonaihuan) | - | Shinoda et al, 1971 | |||||||||||||
| Pi(t) | - | - | 12.2 | P167(1) | - | Causse et al, 1994 | |||||||||||||
| Pi5(t) | RFLP | RG788, RG498 | 12.0 | RIL29 (Morobere kan) | Complete | Wang et al, 1994 | |||||||||||||
| 5 | Pi26(t) | RFLP | RG313 | 22.5-24.7 | Azucena | - | Ahn et al, 1996; Nguyen et al, 2006 | ||||||||||||
| Pi23(t) | - | - | 59.3-99.5 | Sweon 3655 | - | Terashima et al, 2008 | |||||||||||||
| Pi10 | InDel | OPF62700 | 88.5-102.8 | Tongil | Complete | Wu et al, 2005 | |||||||||||||
| 6 | Pi22(t) | RFLP | - | 38.7-41.9 | Sweon 365 | - | Terashima et al, 2008 | ||||||||||||
| Pi26 | RFLP | K17, K2123 | 22.5-63.2 | Gumei 2 | Complete | Ahn et al, 1996 | |||||||||||||
| Pi27(t) | RFLP | Est-2 | 51.9 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Pi40(t) | SSR | RM3330, RM527 | 54.1-61.6 | IR65482-4-136-2-2 | - | ||||||||||||||
| CAPS | S2539 | ||||||||||||||||||
| Piz-5 | STS | BS2-Pi9, NBS4-Pi9 | 58.7 | Tadukan | Complete | Deng et al, 2006 | |||||||||||||
| Piz | InDel | z4794 | 58.7 | Zenith | Complete | Hayashi et al, 2006 | |||||||||||||
| SNP | z60510, z5765, z56592, z565962 | Wang et al, 2012 | |||||||||||||||||
| Piz-t | InDel | z4794 | 58.7 | Torde 1 | Complete | Deng et al, 2006 | |||||||||||||
| Pi9 | - | - | 58.7 | 75-1-127 (101141) | Complete | Qu et al, 2006; Koide et al, 2013 | |||||||||||||
| Pi25(t) | RFLP | A7-RG456 | 63.2-64.6 | Gumei 2 | - | Ahn et al, 1996 | |||||||||||||
| Pid2 | CAPS | CAPS1, CAPS8 | 65.8 | Digu | Complete | Chen et al, 2006 | |||||||||||||
| Pigm(t) | CAPS | C26348 | 65.8 | Gumei 4 | - | Naqvi and Chattoo, 1996 | |||||||||||||
| Pitq1 | RFLP | RZ682, C236, RG653, RZ508 | 103.0-124.4 | Teqing | Complete | Zhou et al, 2004 | |||||||||||||
| Pi8 | Isozyme | Amp-3, pgi-2, Amp-3 | 98.0 | Kasalath | Complete | Tabien et al, 2000 | |||||||||||||
| Pi13(t) | Isozyme | Amp-3 | 74.6-78.2 | Mawong | Complete | Pan et al, 1996 | |||||||||||||
| Pi13 | RFLP | R2123, R538 | 68.1 | Kasalath | - | Fjellstrom et al, 2006; Hayasaka et al, 1995 | |||||||||||||
| Pi2(t) | RFLP | RG64 | 2.8 | Cultivar 5173 | Complete | Yu et al, 1991; Zhou et al, 2006 | |||||||||||||
| Pi2-2 | SSR | AP5659-3, RM19817 | 58.7 | Jefferson | Ballini et al, 2008 | ||||||||||||||
| Pi50(t) | InDel | GDAP51, GDAP16 | 46.8 | - | Jiang et al, 2012 | ||||||||||||||
| Pi40(t) | SSR, CAPS | RM3330, RM527 S2539 | 54.1-61.6 | CO39, IR50 | - | Jeung et al, 2007 | |||||||||||||
| Pi59(t) | SSR | RM19835 | Haoru × US-2 | - | Zhou et al, 2006 | ||||||||||||||
| 7 | Pi17(t) | - | - | 94.0-104.0 | Kasalath | Complete | Zhu et al, 2004 | ||||||||||||
| 8 | Pi36 | SSR | RM5647 | 21.6-25.2 | Q61 | - | Zhu et al, 1993 | ||||||||||||
| CAPS | CRG2, CRG3, CRG4 | ||||||||||||||||||
| Pi33 | SSR | RM72, RM44 | 45.4 | IR64, Bala | Complete | Pan et al, 1995 | |||||||||||||
| Chr | Gene | Tightly linked marker | Map position (cM) | Donor rice | Resistance type | Reference | |||||||||||||
| Type | Name | ||||||||||||||||||
| 8 | Pizh/Pi11(t) | RFLP | RZ617 | 53.2-84.8 | Zhaiyeqing | Complete | Berruyer et al, 2003 | ||||||||||||
| Pi29(t) | RFLP | RZ617, RGA-IR86 | 69.0 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| PiGD-1(t) | RFLP | - | 11.3 | Sanhuangzan 2 | He et al, 2012 | ||||||||||||||
| Pi55(t) | SSR, STS | H2, H66 | 100.6 | Yuejingsimiao 2, Sanhuangzan 2 | Liu et al, 2005; He et al, 2012 | ||||||||||||||
| 9 | Pii2(t) | - | - | Ishikari Shiroke | - | Pan et al, 2003 | |||||||||||||
| Pi5(t) | CAPS | 94A20r, 76B14f, 40N23r | 31.3-33.0 | RIL125, RIL249, RIL260 (Moroberekan) | Complete | Mackill and Bonman, 1992 | |||||||||||||
| SNP | JJ817 | Kinoshita and Kiyosawa, 1997 | |||||||||||||||||
| Pi3(t) | - | - | 31.3-33.0 | Pai-Kan-Tao | Complete | Liu et al, 2004 | |||||||||||||
| Pi15 | RAPD | BAPi15486, BAPi15782, BAPil5844 | 31.3-34.9 | GA25 | Complete | ||||||||||||||
| Pi56(t) | SSR | RM24022 | 31.3 | SHZ-2, BC-10 × TXZ-13 | Complete | Jeon et al, 2003 | |||||||||||||
| Pihk2 | SSR InDel | RM24048, RM24065, RM7390, RM3912, RM24019 ID3, ILP-7, ID-5, ID-9, ILP-19 | Heikezijing | Complete | He at al, 2017 | ||||||||||||||
| 10 | Pi28(t) | RFLP | RZ500 | 114.7 | Azucena | - | Nguyen et al, 2006 | ||||||||||||
| PiGD-2(t) | RFLP | R16 | 3.9 | Sanhuangzhan 2 | - | He et al, 2012 | |||||||||||||
| 11 | Pia | CAPS | Yca72 | 36 | Aichi Asahi | Complete | Zenbayashi et al, 2002 | ||||||||||||
| PiCO39(t) | CAPS | RGA8, RZ141, RGACO39 | 49.1 | CO39 | Complete | Kwon et al, 2008 | |||||||||||||
| Pilm2 | RFLP | L457B, G2132b, RZ536, RG1109 | 56.2-117.9 | Lemont | Complete | Zhou et al, 2004 | |||||||||||||
| Pi30(t) | RFLP | OpZ11-f, RGA-IR14 | 59.4-60.4 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Pi7(t) | - | - | 71.4-84.3 | RIL29 (Moroberekan) | Complete | Miyamoto et al, 2001 | |||||||||||||
| Pi34 | RFLP | C1172, E2021 | 79.1-91.4 | Chubu 32 | Partial | Yunoki et al, 1970 | |||||||||||||
| Pi38 | SSR | RM206, RM21 | 79.1-88.7 | Tadukan | - | Goto, 1970 | |||||||||||||
| Pif | - | - | - | St No.1 | Partial | Gowda et al, 2006 | |||||||||||||
| Pb1 | - | - | 85.7-91.4 | Modan | Partial | Fuentes et al, 2008 | |||||||||||||
| Pi44(t) | AFLP | AF348, AF349 | 91.4-117.9 | RIL29 (Moroberekan) | Complete | Chen et al, 1999; Chauhan et al, 2002 | |||||||||||||
| Pikh/Pi54 | SSR | RM206, RM144, RM224, RM1233 | 101.9 | Tetep, Taipei 309 | Complete | Fjellstrom et al, 2006; Hayasaka et al, 1995; Wu et al, 2013 | |||||||||||||
| Pi1 | SNP | CRG11-7, K28 | 112.1-117.9 | C101LAC (Lac23) | Prasad et al, 2009 | ||||||||||||||
| Pi7(t) | RFLP | RG103A, RG16 | 71.4-84.3 | RIL29 (Moroberekan) | Complete | Wang et al, 1994 | |||||||||||||
| Pikm | InDel SSR | k6861, k2167 RM254, RM144 | 115.1-117.0 | Tsuyake | Complete | Sun et al, 2013; Li et al, 2007 | |||||||||||||
| SNP | k641, k6441, k473, k7237 | ||||||||||||||||||
| Pi18(t) | RFLP | RZ536 | 117.9 | Suweon 365 | Complete | Li et al, 2007; Ahn et al, 2000 | |||||||||||||
| Pik | InDel | k6816, k2167 | 119.9-120.3 | Kusabue | Complete | Hayashi et al, 2006 | |||||||||||||
| Pik-p | SNP | k641, k39575, k403, k3957 | 119.9-120.3 | HR22 | Complete | Hayashi et al, 2006 | |||||||||||||
| Pik-s | SSR | RM144, RM224, RM1233 | 115.1-117.3 | Shin 2 | Complete | Fjellstrom et al, 2006 | |||||||||||||
| Pik-g | - | - | - | GA20 | Complete | Tabien et al, 2000 | |||||||||||||
| Pise1 | - | - | - | Sensho | - | Izawall and Iwasakizl, 2000 | |||||||||||||
| Pi-hk1 | SSR | RM27248, RM27318 | - | Heikezijing | Complete | Liu et al, 2012 | |||||||||||||
| Pikur2 | - | - | - | Kuroka | - | Goto, 1970 | |||||||||||||
| Pi-1(t) | SSR | RM12331, RM224 | 112.1-117.9 | ILs C10LAC and C101A5 | - | Sharma et al, 2005 | |||||||||||||
| Pi47 | SSR | RM206, RM224 | XZ3150 × CO39 | Complete | Ahn et al, 2000 | ||||||||||||||
| Pi49 | STS | K10, K134 | 1.01-1.89 | CO39 | Chen et al, 1999 | ||||||||||||||
| Pi60(t) | InDel | K1-4, E12 | 93-11 | Complete | Huang et al, 2006 | ||||||||||||||
| Pi-jnw1 | SSR InDel | RM27150, RM27381 W26, W28, BS33, BS39, BS71 | Jiangnanwan | Complete | Wang et al, 2016 | ||||||||||||||
| Pi65(t) | SNP InDel | SNP-2, SNP-8 InDel-1 | - | Guangyu 129 | Complete | Zheng et al, 2016 | |||||||||||||
| 12 | Pi24 | RGA | RGA3 | 10.3 | Zhong 156 | Complete | Zhuang et al, 2002; Hua et al, 2012 | ||||||||||||
| Pi62(t) | RFLP | RG869 | 12.2-26.0 | Yashiromochi | - | Kumar et al, 2010 | |||||||||||||
| Pitq6 | RFLP | RG341a, RG869, L102, G1468a, RZ397, RZ257 | 29.2-47.5 | Teqing | Complete | Zhu et al, 2000 | |||||||||||||
| Chr | Gene | Tightly linked marker | Map position (cM) | Donor rice | Resistance type | Reference | |||||||||||||
| Type | Name | ||||||||||||||||||
| 12 | Pi6(t) | RFLP | RG869 | 32.6-63.2 | Apura | Complete | Yu et al, 1991 | ||||||||||||
| Pi12(t) | - | - | 42.8-53.0 | Hongjiaozhan | Complete | - | |||||||||||||
| Pi31(t) | RFLP | O10-800 | 44.3 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Pi32(t) | RFLP | AF6 | 47.5 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Ipi(t) | RFLP | RG241X | 47.6-58.3 | BS125 × WL02 | - | Causse et al, 1994 | |||||||||||||
| IPi3(t) | RFLP | RG241X | 47.6-58.3 | BS125 × WL02 | - | Causse et al, 1994 | |||||||||||||
| Pi157 | - | - | 49.5-62.2 | Moroberecan | - | - | |||||||||||||
| Pita | SNP RAPD | ta642, ta801, ta3, ta577, ta5, pita440, pita1042, pita403 SP4B9, SP9F3 | 50.4 | Taducan Yashiro-mochi, Tsuyuake | Complete | Hayashi et al, 1998 | |||||||||||||
| Pi39(t) | CAPS | 39M6, 39M7 | 50.4 | Q15 | Bryan et al, 2000 | ||||||||||||||
| Pi20(t) | SSR | RM1337, RM5364, RM7102 | 51.5-51.8 | IR24 | Complete | Liu et al, 2007 | |||||||||||||
| PiGD-3(t) | SSR | RM179 | 55.8 | Sanhuangzhan 2 | - | Liu et al, 2007 | |||||||||||||
| Pi42(t) | STS SSR | STS5 RRS44, RRS51, RRS60, RRS63, RRS6 | DHR9 | Kumar et al, 2010 | |||||||||||||||
| Pi4(t) | RFLP | RG869, RZ397 | 47.5 | Apura | Nguyen et al, 2006 | ||||||||||||||
| Pita-2 | SNP | ta642, ta801, ta3, ta577 | 50.4 | Shimokita | Complete | Hyashi et al, 1998 | |||||||||||||
| Pi19(t) | - | - | Aichi Asahi | Complete | Inukai and Nelson, 1994 | ||||||||||||||
| Pi21(t) | RFLP | - | 43.4-59.6 | Suweon 365 | Terashima et al, 2008 | ||||||||||||||
| Pi58(t) | SSR | RM27954, RM27933, RM3103 | Haoru × US-2 | Zhou et al, 2004 | |||||||||||||||
| Pi48 | SSR | RM5364, RM7102 | XZ3150 × CO39 | Complete | Ahn et al, 2000 | ||||||||||||||
| Pi61(t) | InDel | M2, S29 | 93-11 | Complete | Huang et al, 2011 | ||||||||||||||
Table 2 Blast resistance genes and tightly linked markers in rice for successful breeding.
| Chr | Gene | Tightly linked marker | Map position (cM) | Donor rice | Resistance type | Reference | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Type | Name | ||||||||||||||||||
| 1 | Pit | SNP | t311, t256, t8042 | 16.1 | Tjahaja | Complete | Hayashi et al, 2006 | ||||||||||||
| Pi27(t) | SSR | RM151, RN259 | 28.4-38.8 | Q14 | Complete | Sallaud et al, 2006 | |||||||||||||
| Pitp(t) | SSR | RM246 | 114.1 | Tetep | Partial | Barman et al, 2004 | |||||||||||||
| Pi35(t) | SSR | RM1216, RM1003 | 132.0-136.6 | Hokkai 188 | Partial | Nguyen et al, 2006 | |||||||||||||
| Pi37 | SSR | RM302, RM212, FPSM1, FPSM2, FPSM4 | 136.1 | St.No.1 | Complete | Lin et al, 2007 | |||||||||||||
| STS | S15628, FSTS1, FSTS2, FSTS3, FSTS4 | ||||||||||||||||||
| Pi64 | SSR | RM11715, RM11787 | - | Yangmaogu | Complete | Ma et al, 2015 | |||||||||||||
| Pish | RFLP | - | 148.7-154.8 | Shin 2 | Complete | Zhu et al, 2004 | |||||||||||||
| 2 | Pid1(t) | SSR | RM262 | 87.5-89.9 | Digu | Complete | Chen et al, 2004 | ||||||||||||
| Pig(t) | SSR | RM166, RM208 | 142.0-154.1 | Guangchangzhan | Complete | Shi et al, 2012 | |||||||||||||
| Pitq5 | RFLP | RG520, RZ446B, RZ446A, RG654, RG256 | 150.5-157.9 | Teqing | Complete | Zhou et al, 2004 | |||||||||||||
| Piy1(t) | SSR | RM3248, RM20 | 153.2-154.1 | Yanxian 1 | - | Fukuta et al, 2004 | |||||||||||||
| Piy2(t) | SSR | RM3248, RM20 | 153.2-154.1 | Yanxian 1 | - | Fukuta et al, 2004 | |||||||||||||
| Pib | SNP | b213, b28, b2, b3989, Pibdom | 154.1 | Tohoku, IL9, Koshiihikari | Complete | Hayashi et al, 2006 | |||||||||||||
| Pi14(t) | Isozyme | Amp-1 | 32.6 | Maowang | Complete | Tabien et al, 2000 | |||||||||||||
| Pi16(t) | Isozyme | Amp-1 | 34.3 | AUS373, Maowamg | Complete | Pan et al, 1996 | |||||||||||||
| Pi-Da(t) | SSR | RM5529, RM211 | 12.6 | Dacca 6 | - | Lei et al, 2005; Shi et al, 2012 | |||||||||||||
| 3 | Pi66(t) | SSR | RM487, RM16, RM55, RM168 | AS201 | Partial | Liang et al, 2016 | |||||||||||||
| 4 | Pi21 | STS | P702D03-#79 | 58.6 | Owarihatamochi | Partial | Pan et al, 1998 | ||||||||||||
| Pikur1 | Isozyme | - | 86.0 | Kuroka | - | Fukuoka et al, 2007 | |||||||||||||
| Pi39(t) | SSR | RM3843, RM5473 | 107.4-108.2 | Chubu 111 (Haonaihuan) | - | Shinoda et al, 1971 | |||||||||||||
| Pi(t) | - | - | 12.2 | P167(1) | - | Causse et al, 1994 | |||||||||||||
| Pi5(t) | RFLP | RG788, RG498 | 12.0 | RIL29 (Morobere kan) | Complete | Wang et al, 1994 | |||||||||||||
| 5 | Pi26(t) | RFLP | RG313 | 22.5-24.7 | Azucena | - | Ahn et al, 1996; Nguyen et al, 2006 | ||||||||||||
| Pi23(t) | - | - | 59.3-99.5 | Sweon 3655 | - | Terashima et al, 2008 | |||||||||||||
| Pi10 | InDel | OPF62700 | 88.5-102.8 | Tongil | Complete | Wu et al, 2005 | |||||||||||||
| 6 | Pi22(t) | RFLP | - | 38.7-41.9 | Sweon 365 | - | Terashima et al, 2008 | ||||||||||||
| Pi26 | RFLP | K17, K2123 | 22.5-63.2 | Gumei 2 | Complete | Ahn et al, 1996 | |||||||||||||
| Pi27(t) | RFLP | Est-2 | 51.9 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Pi40(t) | SSR | RM3330, RM527 | 54.1-61.6 | IR65482-4-136-2-2 | - | ||||||||||||||
| CAPS | S2539 | ||||||||||||||||||
| Piz-5 | STS | BS2-Pi9, NBS4-Pi9 | 58.7 | Tadukan | Complete | Deng et al, 2006 | |||||||||||||
| Piz | InDel | z4794 | 58.7 | Zenith | Complete | Hayashi et al, 2006 | |||||||||||||
| SNP | z60510, z5765, z56592, z565962 | Wang et al, 2012 | |||||||||||||||||
| Piz-t | InDel | z4794 | 58.7 | Torde 1 | Complete | Deng et al, 2006 | |||||||||||||
| Pi9 | - | - | 58.7 | 75-1-127 (101141) | Complete | Qu et al, 2006; Koide et al, 2013 | |||||||||||||
| Pi25(t) | RFLP | A7-RG456 | 63.2-64.6 | Gumei 2 | - | Ahn et al, 1996 | |||||||||||||
| Pid2 | CAPS | CAPS1, CAPS8 | 65.8 | Digu | Complete | Chen et al, 2006 | |||||||||||||
| Pigm(t) | CAPS | C26348 | 65.8 | Gumei 4 | - | Naqvi and Chattoo, 1996 | |||||||||||||
| Pitq1 | RFLP | RZ682, C236, RG653, RZ508 | 103.0-124.4 | Teqing | Complete | Zhou et al, 2004 | |||||||||||||
| Pi8 | Isozyme | Amp-3, pgi-2, Amp-3 | 98.0 | Kasalath | Complete | Tabien et al, 2000 | |||||||||||||
| Pi13(t) | Isozyme | Amp-3 | 74.6-78.2 | Mawong | Complete | Pan et al, 1996 | |||||||||||||
| Pi13 | RFLP | R2123, R538 | 68.1 | Kasalath | - | Fjellstrom et al, 2006; Hayasaka et al, 1995 | |||||||||||||
| Pi2(t) | RFLP | RG64 | 2.8 | Cultivar 5173 | Complete | Yu et al, 1991; Zhou et al, 2006 | |||||||||||||
| Pi2-2 | SSR | AP5659-3, RM19817 | 58.7 | Jefferson | Ballini et al, 2008 | ||||||||||||||
| Pi50(t) | InDel | GDAP51, GDAP16 | 46.8 | - | Jiang et al, 2012 | ||||||||||||||
| Pi40(t) | SSR, CAPS | RM3330, RM527 S2539 | 54.1-61.6 | CO39, IR50 | - | Jeung et al, 2007 | |||||||||||||
| Pi59(t) | SSR | RM19835 | Haoru × US-2 | - | Zhou et al, 2006 | ||||||||||||||
| 7 | Pi17(t) | - | - | 94.0-104.0 | Kasalath | Complete | Zhu et al, 2004 | ||||||||||||
| 8 | Pi36 | SSR | RM5647 | 21.6-25.2 | Q61 | - | Zhu et al, 1993 | ||||||||||||
| CAPS | CRG2, CRG3, CRG4 | ||||||||||||||||||
| Pi33 | SSR | RM72, RM44 | 45.4 | IR64, Bala | Complete | Pan et al, 1995 | |||||||||||||
| Chr | Gene | Tightly linked marker | Map position (cM) | Donor rice | Resistance type | Reference | |||||||||||||
| Type | Name | ||||||||||||||||||
| 8 | Pizh/Pi11(t) | RFLP | RZ617 | 53.2-84.8 | Zhaiyeqing | Complete | Berruyer et al, 2003 | ||||||||||||
| Pi29(t) | RFLP | RZ617, RGA-IR86 | 69.0 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| PiGD-1(t) | RFLP | - | 11.3 | Sanhuangzan 2 | He et al, 2012 | ||||||||||||||
| Pi55(t) | SSR, STS | H2, H66 | 100.6 | Yuejingsimiao 2, Sanhuangzan 2 | Liu et al, 2005; He et al, 2012 | ||||||||||||||
| 9 | Pii2(t) | - | - | Ishikari Shiroke | - | Pan et al, 2003 | |||||||||||||
| Pi5(t) | CAPS | 94A20r, 76B14f, 40N23r | 31.3-33.0 | RIL125, RIL249, RIL260 (Moroberekan) | Complete | Mackill and Bonman, 1992 | |||||||||||||
| SNP | JJ817 | Kinoshita and Kiyosawa, 1997 | |||||||||||||||||
| Pi3(t) | - | - | 31.3-33.0 | Pai-Kan-Tao | Complete | Liu et al, 2004 | |||||||||||||
| Pi15 | RAPD | BAPi15486, BAPi15782, BAPil5844 | 31.3-34.9 | GA25 | Complete | ||||||||||||||
| Pi56(t) | SSR | RM24022 | 31.3 | SHZ-2, BC-10 × TXZ-13 | Complete | Jeon et al, 2003 | |||||||||||||
| Pihk2 | SSR InDel | RM24048, RM24065, RM7390, RM3912, RM24019 ID3, ILP-7, ID-5, ID-9, ILP-19 | Heikezijing | Complete | He at al, 2017 | ||||||||||||||
| 10 | Pi28(t) | RFLP | RZ500 | 114.7 | Azucena | - | Nguyen et al, 2006 | ||||||||||||
| PiGD-2(t) | RFLP | R16 | 3.9 | Sanhuangzhan 2 | - | He et al, 2012 | |||||||||||||
| 11 | Pia | CAPS | Yca72 | 36 | Aichi Asahi | Complete | Zenbayashi et al, 2002 | ||||||||||||
| PiCO39(t) | CAPS | RGA8, RZ141, RGACO39 | 49.1 | CO39 | Complete | Kwon et al, 2008 | |||||||||||||
| Pilm2 | RFLP | L457B, G2132b, RZ536, RG1109 | 56.2-117.9 | Lemont | Complete | Zhou et al, 2004 | |||||||||||||
| Pi30(t) | RFLP | OpZ11-f, RGA-IR14 | 59.4-60.4 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Pi7(t) | - | - | 71.4-84.3 | RIL29 (Moroberekan) | Complete | Miyamoto et al, 2001 | |||||||||||||
| Pi34 | RFLP | C1172, E2021 | 79.1-91.4 | Chubu 32 | Partial | Yunoki et al, 1970 | |||||||||||||
| Pi38 | SSR | RM206, RM21 | 79.1-88.7 | Tadukan | - | Goto, 1970 | |||||||||||||
| Pif | - | - | - | St No.1 | Partial | Gowda et al, 2006 | |||||||||||||
| Pb1 | - | - | 85.7-91.4 | Modan | Partial | Fuentes et al, 2008 | |||||||||||||
| Pi44(t) | AFLP | AF348, AF349 | 91.4-117.9 | RIL29 (Moroberekan) | Complete | Chen et al, 1999; Chauhan et al, 2002 | |||||||||||||
| Pikh/Pi54 | SSR | RM206, RM144, RM224, RM1233 | 101.9 | Tetep, Taipei 309 | Complete | Fjellstrom et al, 2006; Hayasaka et al, 1995; Wu et al, 2013 | |||||||||||||
| Pi1 | SNP | CRG11-7, K28 | 112.1-117.9 | C101LAC (Lac23) | Prasad et al, 2009 | ||||||||||||||
| Pi7(t) | RFLP | RG103A, RG16 | 71.4-84.3 | RIL29 (Moroberekan) | Complete | Wang et al, 1994 | |||||||||||||
| Pikm | InDel SSR | k6861, k2167 RM254, RM144 | 115.1-117.0 | Tsuyake | Complete | Sun et al, 2013; Li et al, 2007 | |||||||||||||
| SNP | k641, k6441, k473, k7237 | ||||||||||||||||||
| Pi18(t) | RFLP | RZ536 | 117.9 | Suweon 365 | Complete | Li et al, 2007; Ahn et al, 2000 | |||||||||||||
| Pik | InDel | k6816, k2167 | 119.9-120.3 | Kusabue | Complete | Hayashi et al, 2006 | |||||||||||||
| Pik-p | SNP | k641, k39575, k403, k3957 | 119.9-120.3 | HR22 | Complete | Hayashi et al, 2006 | |||||||||||||
| Pik-s | SSR | RM144, RM224, RM1233 | 115.1-117.3 | Shin 2 | Complete | Fjellstrom et al, 2006 | |||||||||||||
| Pik-g | - | - | - | GA20 | Complete | Tabien et al, 2000 | |||||||||||||
| Pise1 | - | - | - | Sensho | - | Izawall and Iwasakizl, 2000 | |||||||||||||
| Pi-hk1 | SSR | RM27248, RM27318 | - | Heikezijing | Complete | Liu et al, 2012 | |||||||||||||
| Pikur2 | - | - | - | Kuroka | - | Goto, 1970 | |||||||||||||
| Pi-1(t) | SSR | RM12331, RM224 | 112.1-117.9 | ILs C10LAC and C101A5 | - | Sharma et al, 2005 | |||||||||||||
| Pi47 | SSR | RM206, RM224 | XZ3150 × CO39 | Complete | Ahn et al, 2000 | ||||||||||||||
| Pi49 | STS | K10, K134 | 1.01-1.89 | CO39 | Chen et al, 1999 | ||||||||||||||
| Pi60(t) | InDel | K1-4, E12 | 93-11 | Complete | Huang et al, 2006 | ||||||||||||||
| Pi-jnw1 | SSR InDel | RM27150, RM27381 W26, W28, BS33, BS39, BS71 | Jiangnanwan | Complete | Wang et al, 2016 | ||||||||||||||
| Pi65(t) | SNP InDel | SNP-2, SNP-8 InDel-1 | - | Guangyu 129 | Complete | Zheng et al, 2016 | |||||||||||||
| 12 | Pi24 | RGA | RGA3 | 10.3 | Zhong 156 | Complete | Zhuang et al, 2002; Hua et al, 2012 | ||||||||||||
| Pi62(t) | RFLP | RG869 | 12.2-26.0 | Yashiromochi | - | Kumar et al, 2010 | |||||||||||||
| Pitq6 | RFLP | RG341a, RG869, L102, G1468a, RZ397, RZ257 | 29.2-47.5 | Teqing | Complete | Zhu et al, 2000 | |||||||||||||
| Chr | Gene | Tightly linked marker | Map position (cM) | Donor rice | Resistance type | Reference | |||||||||||||
| Type | Name | ||||||||||||||||||
| 12 | Pi6(t) | RFLP | RG869 | 32.6-63.2 | Apura | Complete | Yu et al, 1991 | ||||||||||||
| Pi12(t) | - | - | 42.8-53.0 | Hongjiaozhan | Complete | - | |||||||||||||
| Pi31(t) | RFLP | O10-800 | 44.3 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Pi32(t) | RFLP | AF6 | 47.5 | IR64 | - | Nguyen et al, 2006 | |||||||||||||
| Ipi(t) | RFLP | RG241X | 47.6-58.3 | BS125 × WL02 | - | Causse et al, 1994 | |||||||||||||
| IPi3(t) | RFLP | RG241X | 47.6-58.3 | BS125 × WL02 | - | Causse et al, 1994 | |||||||||||||
| Pi157 | - | - | 49.5-62.2 | Moroberecan | - | - | |||||||||||||
| Pita | SNP RAPD | ta642, ta801, ta3, ta577, ta5, pita440, pita1042, pita403 SP4B9, SP9F3 | 50.4 | Taducan Yashiro-mochi, Tsuyuake | Complete | Hayashi et al, 1998 | |||||||||||||
| Pi39(t) | CAPS | 39M6, 39M7 | 50.4 | Q15 | Bryan et al, 2000 | ||||||||||||||
| Pi20(t) | SSR | RM1337, RM5364, RM7102 | 51.5-51.8 | IR24 | Complete | Liu et al, 2007 | |||||||||||||
| PiGD-3(t) | SSR | RM179 | 55.8 | Sanhuangzhan 2 | - | Liu et al, 2007 | |||||||||||||
| Pi42(t) | STS SSR | STS5 RRS44, RRS51, RRS60, RRS63, RRS6 | DHR9 | Kumar et al, 2010 | |||||||||||||||
| Pi4(t) | RFLP | RG869, RZ397 | 47.5 | Apura | Nguyen et al, 2006 | ||||||||||||||
| Pita-2 | SNP | ta642, ta801, ta3, ta577 | 50.4 | Shimokita | Complete | Hyashi et al, 1998 | |||||||||||||
| Pi19(t) | - | - | Aichi Asahi | Complete | Inukai and Nelson, 1994 | ||||||||||||||
| Pi21(t) | RFLP | - | 43.4-59.6 | Suweon 365 | Terashima et al, 2008 | ||||||||||||||
| Pi58(t) | SSR | RM27954, RM27933, RM3103 | Haoru × US-2 | Zhou et al, 2004 | |||||||||||||||
| Pi48 | SSR | RM5364, RM7102 | XZ3150 × CO39 | Complete | Ahn et al, 2000 | ||||||||||||||
| Pi61(t) | InDel | M2, S29 | 93-11 | Complete | Huang et al, 2011 | ||||||||||||||
| Trait | Gene/QTL | Marker type | Technique | Application | Reference |
|---|---|---|---|---|---|
| Blast resistance | Pi1, Piz-5, Pita | RFLP | MAS | Pyramiding of three near isogenic lines (C101LAC, C101A51 and C101PKT) for blast resistance into a single cultivar CO39, each carrying the major genes Pi1, Piz-5 and Pita, respectively | Hittalmani et al, 2000 |
| Blast resistance | Pi1 | SSR, ISSR | MABB | Applied for backcross breeding of variety | Liu et al, 2002 |
| Bacterial blight resistance and blast resistance | Xa21, Piz | SSR | MAS | Pyramiding of target traits | Narayanan et al, 2002 |
| Blast resistance | Pid1, Pib, Pita, Pi2 | SSR | MAS | Pid1, Pib and Pita genes were introduced into G46B, while Pi2 was introduced into Zhenshan 97B | Chen et al, 2004 |
| Blast resistance | Piz | SSR | MAS | Successfully used for selection of blast resistance in a wide array of rice germplasm | Fjellstrom et al, 2006 |
| Blast resistance and bacterial blight resistance and sheath blight resistance | Xa13, Xa21, Pi54, qSBR11 | SSR | MAS | Transfer genes conferring the resistances toward three different diseases in rice | Singh V K et al, 2012 |
| Blast resistance | Pita | Gene specific | MAS | Existence of Pita gene in 141 rice germplasms was determined, but the results were more articulated when Pita gene was introduced through advanced breeding lines | Wang et al, 2007 |
| Blast resistance | Pi1, Pi2, Pi33 | SSR | MABB | Introgressed into Jin23B | Chen et al, 2008 |
| Blast resistance and bacterial blight | Pi1, Pi2, Xa23 | SSR | MABB | Sucessfully applied for breeding variety Rongfeng B | Fu et al, 2012 |
| Blast resistance | Piz-5, Pi54 | SSR | MABB | Transfer blast resistance genes from donor lines (C101A51 and Tetep) into PRR78 to develop Pusa 1602 (PRR78 + Piz-5) and Pusa 1603(PRR78 + Pi54), respectively | Singh et al, 2013 |
| Blast resistance | Pi9 | Gene specific | MABB | Applied to introgress the cultivar Luhui 17 | Wen and Gao, 2011 |
| Blast resistance | Pi1, Piz | SSR | MABB | Pyramiding of Pi1 and Piz-5 genes into introduced PRR78 | Gouda et al, 2013 |
| Blast resistance | Pi39 | InDel | MABB | Introgressed into Chinese cultivar Q15 | Hua et al, 2015 |
| Blast resistance and bacterial blight | Pi2, Xa21, Xa33 | SSR | MABB | Introgressed into RPHR-1005 | Kumar et al, 2016 |
| Blast resistance | Pi40 | SSR | MABB | Introgressed into elite cultivars Turkish, Osmancik-97 and Halilbey | Beser et al, 2016 |
| Blast resistance | Pi1, Pi2 | SSR | MABB | Introgressed into Intan variety and BPT5204 | Hegde and Prashanthi, 2016 |
| Blast resistance | Pi46, Pita | SSR | MABB | Introgressed into Hang hui 179 (HH179) | Xiao et al, 2016 |
| Blast resistance | Pi2, Pi9 | SNP | MABB | Introgressed into R179 | Luo et al, 2016 |
| Blast resistance | Pizt, Pi2, Pigm, Pi40, Pi9, Piz | SSR | MAS | Introgressed into Yangdao 6 | Wu et al, 2016 |
| Blast resistance | Pi1, Pi2, Pi33 | SSR | MAS | Introgressed into Russian rice varieties | Usatov et al, 2016 |
| Blast resistance | Pi9, Pizt, Pi54 | SNP | MABB | Introgressed into japonica rice 07GY31 | Xiao et al, 2016 |
| Blast resistance | Pi-b, Pik-h | SSR | MAS | Introgressed into MR219 | Tanweer et al, 2015 |
Table 3 Successful examples of marker-assisted selection (MAS) and marker-assisted backcrossing breeding (MABB) in rice for blast resistance breeding.
| Trait | Gene/QTL | Marker type | Technique | Application | Reference |
|---|---|---|---|---|---|
| Blast resistance | Pi1, Piz-5, Pita | RFLP | MAS | Pyramiding of three near isogenic lines (C101LAC, C101A51 and C101PKT) for blast resistance into a single cultivar CO39, each carrying the major genes Pi1, Piz-5 and Pita, respectively | Hittalmani et al, 2000 |
| Blast resistance | Pi1 | SSR, ISSR | MABB | Applied for backcross breeding of variety | Liu et al, 2002 |
| Bacterial blight resistance and blast resistance | Xa21, Piz | SSR | MAS | Pyramiding of target traits | Narayanan et al, 2002 |
| Blast resistance | Pid1, Pib, Pita, Pi2 | SSR | MAS | Pid1, Pib and Pita genes were introduced into G46B, while Pi2 was introduced into Zhenshan 97B | Chen et al, 2004 |
| Blast resistance | Piz | SSR | MAS | Successfully used for selection of blast resistance in a wide array of rice germplasm | Fjellstrom et al, 2006 |
| Blast resistance and bacterial blight resistance and sheath blight resistance | Xa13, Xa21, Pi54, qSBR11 | SSR | MAS | Transfer genes conferring the resistances toward three different diseases in rice | Singh V K et al, 2012 |
| Blast resistance | Pita | Gene specific | MAS | Existence of Pita gene in 141 rice germplasms was determined, but the results were more articulated when Pita gene was introduced through advanced breeding lines | Wang et al, 2007 |
| Blast resistance | Pi1, Pi2, Pi33 | SSR | MABB | Introgressed into Jin23B | Chen et al, 2008 |
| Blast resistance and bacterial blight | Pi1, Pi2, Xa23 | SSR | MABB | Sucessfully applied for breeding variety Rongfeng B | Fu et al, 2012 |
| Blast resistance | Piz-5, Pi54 | SSR | MABB | Transfer blast resistance genes from donor lines (C101A51 and Tetep) into PRR78 to develop Pusa 1602 (PRR78 + Piz-5) and Pusa 1603(PRR78 + Pi54), respectively | Singh et al, 2013 |
| Blast resistance | Pi9 | Gene specific | MABB | Applied to introgress the cultivar Luhui 17 | Wen and Gao, 2011 |
| Blast resistance | Pi1, Piz | SSR | MABB | Pyramiding of Pi1 and Piz-5 genes into introduced PRR78 | Gouda et al, 2013 |
| Blast resistance | Pi39 | InDel | MABB | Introgressed into Chinese cultivar Q15 | Hua et al, 2015 |
| Blast resistance and bacterial blight | Pi2, Xa21, Xa33 | SSR | MABB | Introgressed into RPHR-1005 | Kumar et al, 2016 |
| Blast resistance | Pi40 | SSR | MABB | Introgressed into elite cultivars Turkish, Osmancik-97 and Halilbey | Beser et al, 2016 |
| Blast resistance | Pi1, Pi2 | SSR | MABB | Introgressed into Intan variety and BPT5204 | Hegde and Prashanthi, 2016 |
| Blast resistance | Pi46, Pita | SSR | MABB | Introgressed into Hang hui 179 (HH179) | Xiao et al, 2016 |
| Blast resistance | Pi2, Pi9 | SNP | MABB | Introgressed into R179 | Luo et al, 2016 |
| Blast resistance | Pizt, Pi2, Pigm, Pi40, Pi9, Piz | SSR | MAS | Introgressed into Yangdao 6 | Wu et al, 2016 |
| Blast resistance | Pi1, Pi2, Pi33 | SSR | MAS | Introgressed into Russian rice varieties | Usatov et al, 2016 |
| Blast resistance | Pi9, Pizt, Pi54 | SNP | MABB | Introgressed into japonica rice 07GY31 | Xiao et al, 2016 |
| Blast resistance | Pi-b, Pik-h | SSR | MAS | Introgressed into MR219 | Tanweer et al, 2015 |
| Trait | Parental line | Pyramided gene | Marker | Reference |
|---|---|---|---|---|
| Blast resistance | C101LAC, C101A51 | Pi1, Pi2, Pi33 | SSR | Chen et al, 2008 |
| Blast resistance | IR5, IR8, IR20, IR22, IR24, IR26, IR28, IR29, IR30, IR32, IR34, IR36, IR38, IR40, IR42, IR43, IR44, IR45, IR46, IR48, IR50 IR52, IR54, IR56, IR58, IR60, IR62, IR64, IR65, IR66, IR68, IR70, IR72, IR74 | Pib, Pita | SSR | Fujita et al, 2009 |
| Blast resistance | CO39 | Pish, Pib | SSR | Koide et al, 2010 |
| Blast resistance Blast resistance | IR64, JHN JHN × RD6 | Multiple resistance QTLs qBl1, qBl11 | SSR SSR | Sreewongchai et al, 2010 Wongsaprom et al, 2010 |
| Blast resistance | Rongfeng B | Pi1, Pi2, Xa23 | Fu et al, 2012 | |
| Blast resistance | C101LAC, C101A51 | Pi1, Pi2 | RG64, C481 | Mahdian and Shahsavari, 2013 |
| Blast and bacterial leaf blight resistance | RD6 × P0489, RD6 × JHN | Four QTLs for blast resistance and one gene (xa5) for bacterial leaf blight | SSR | Pinta et al, 2013 |
| Blast resistance Blast and bacterial blight resistance | C101A51, Tetep BL122 and CBB23 are donors while Rongfeng 3A is recurrent parent | Piz-5, Pi54 Pi1, Pi2, Xa23 | SSR SSR | Singh et al, 2013 Fu et al, 2012 |
| Blast resistance | Carnaroli, Baldo, Arborio | Piz, Pi5 | SSR | Urso et al, 2013 |
| Leaf blast resistance | Koshihikari | Pi21, Pi34, Pi35 | SSR | Yasuda et al, 2014 |
| Blast resistance | GZ63-4S | Pi2, Xa23 | SSR | Jiang et al, 2015b |
| Blast resistance Blast and bacterial blight resistance | Aichiasahi × Owarihatamochi 6G241 × GZ63S | pi21, Pi34, qBR4-2, qBR12-1 Pi9, Xa23 | SSR Gene-linked marker | Fukuoka et al, 2015; Ni et al, 2015 |
| Blast and bacterial blight resistance | Samba Mahsuri (possessing Xa21 and xa13) and NLR145 (possessing Pi54) | Pi54, Xa21, Xa13 | SSR | Arunakumari et al, 2016 |
| Blast resistance | RD6 × PO489, RD6 × JHN | Four QTLs located on chromosomes 1, 2, 11 and 12, respectively | SSR | Suwannual et al, 2017 |
| Blast, bacterial blight and brown planthopper resistance | CBB23, HN88, Zhongzu 14, Shuhui 162 | Xa23, Xa5, Pita, Pi1, Pi2, Bph3 | SSR | Ji et al, 2016 |
| Blast and bacterial blight resistance | RPBio Patho-1 and FBR1-15 are donors, and RPHR-1005 is recurrent | Xa21, Xa33, Pi2, Rf3, Rf4w | SSR | Kumar et al, 2016 |
| Blast, bacterial blight and submergence tolerance | WH21, IR64 and 75-1-127 are donors, and Chinese variety 9211 is recurrent | Pi9, Xa21, Xa27, Sub1A | STS, SSR | Luo et al, 2017 |
| Blast resistance | Monogenic near isogenic lines Pusa 1637- 18-7-6-20 and Pusa 1633-3-8-8-16-1 are donors and Pusa Basmati 1 is recurrent | Pi9, Pita | Gene-linked marker | Khanna et al, 2015 |
| Blast, bacterial blight, gall midge, submergence and salt tolerance | C101A51, WHD-18-75-1-127, Kavya, Abhaya, FR13A, Saltol, Improved Lalat | Pi2, Pi9, Gm1, Gm4, Sub1, Saltol, Xa21, xa13, xa5 | SSR, Gene-linked marker | Das and Rao, 2015 |
Table 4 Successful examples of gene pyramiding in important rice varieties against blast disease.
| Trait | Parental line | Pyramided gene | Marker | Reference |
|---|---|---|---|---|
| Blast resistance | C101LAC, C101A51 | Pi1, Pi2, Pi33 | SSR | Chen et al, 2008 |
| Blast resistance | IR5, IR8, IR20, IR22, IR24, IR26, IR28, IR29, IR30, IR32, IR34, IR36, IR38, IR40, IR42, IR43, IR44, IR45, IR46, IR48, IR50 IR52, IR54, IR56, IR58, IR60, IR62, IR64, IR65, IR66, IR68, IR70, IR72, IR74 | Pib, Pita | SSR | Fujita et al, 2009 |
| Blast resistance | CO39 | Pish, Pib | SSR | Koide et al, 2010 |
| Blast resistance Blast resistance | IR64, JHN JHN × RD6 | Multiple resistance QTLs qBl1, qBl11 | SSR SSR | Sreewongchai et al, 2010 Wongsaprom et al, 2010 |
| Blast resistance | Rongfeng B | Pi1, Pi2, Xa23 | Fu et al, 2012 | |
| Blast resistance | C101LAC, C101A51 | Pi1, Pi2 | RG64, C481 | Mahdian and Shahsavari, 2013 |
| Blast and bacterial leaf blight resistance | RD6 × P0489, RD6 × JHN | Four QTLs for blast resistance and one gene (xa5) for bacterial leaf blight | SSR | Pinta et al, 2013 |
| Blast resistance Blast and bacterial blight resistance | C101A51, Tetep BL122 and CBB23 are donors while Rongfeng 3A is recurrent parent | Piz-5, Pi54 Pi1, Pi2, Xa23 | SSR SSR | Singh et al, 2013 Fu et al, 2012 |
| Blast resistance | Carnaroli, Baldo, Arborio | Piz, Pi5 | SSR | Urso et al, 2013 |
| Leaf blast resistance | Koshihikari | Pi21, Pi34, Pi35 | SSR | Yasuda et al, 2014 |
| Blast resistance | GZ63-4S | Pi2, Xa23 | SSR | Jiang et al, 2015b |
| Blast resistance Blast and bacterial blight resistance | Aichiasahi × Owarihatamochi 6G241 × GZ63S | pi21, Pi34, qBR4-2, qBR12-1 Pi9, Xa23 | SSR Gene-linked marker | Fukuoka et al, 2015; Ni et al, 2015 |
| Blast and bacterial blight resistance | Samba Mahsuri (possessing Xa21 and xa13) and NLR145 (possessing Pi54) | Pi54, Xa21, Xa13 | SSR | Arunakumari et al, 2016 |
| Blast resistance | RD6 × PO489, RD6 × JHN | Four QTLs located on chromosomes 1, 2, 11 and 12, respectively | SSR | Suwannual et al, 2017 |
| Blast, bacterial blight and brown planthopper resistance | CBB23, HN88, Zhongzu 14, Shuhui 162 | Xa23, Xa5, Pita, Pi1, Pi2, Bph3 | SSR | Ji et al, 2016 |
| Blast and bacterial blight resistance | RPBio Patho-1 and FBR1-15 are donors, and RPHR-1005 is recurrent | Xa21, Xa33, Pi2, Rf3, Rf4w | SSR | Kumar et al, 2016 |
| Blast, bacterial blight and submergence tolerance | WH21, IR64 and 75-1-127 are donors, and Chinese variety 9211 is recurrent | Pi9, Xa21, Xa27, Sub1A | STS, SSR | Luo et al, 2017 |
| Blast resistance | Monogenic near isogenic lines Pusa 1637- 18-7-6-20 and Pusa 1633-3-8-8-16-1 are donors and Pusa Basmati 1 is recurrent | Pi9, Pita | Gene-linked marker | Khanna et al, 2015 |
| Blast, bacterial blight, gall midge, submergence and salt tolerance | C101A51, WHD-18-75-1-127, Kavya, Abhaya, FR13A, Saltol, Improved Lalat | Pi2, Pi9, Gm1, Gm4, Sub1, Saltol, Xa21, xa13, xa5 | SSR, Gene-linked marker | Das and Rao, 2015 |
| Mapping population | Parent | Total No. of QTLs | Marker | Reference |
|---|---|---|---|---|
| RIL | CT9993-5-10-1-m × KDML105 (F8); Zhenshan 97 × Minghui 63 (RILs); Moroberekan × Co39 (F7); Lemont × Teqing (F8); Lemont × Teqing (F14); Bala × Azucena (F6); Zhong 156 × Gumei 2 (F8); OryzicaLlanos 5 × Fanny (F5 and F6); SHZ-2 × Lijiangxintuanheigu (LTH) (RILs); KDML105 × JHN (F6); Suweon 365 × Chucheong (RILs) | 186 | RFLP, SSR, RAPD, Isozymes, AFLP, DR gene markers | Cho et al, 2008 |
| Double haploid | IR64 × Azucena; IR64 × Azucena; ZYQ8 × JX17 | 146 | RFLP, RAPD, Isozymes | Xu et al, 2004 |
| SSSL | Developed by HXJ74 as recipient and 24 accessions as donors | 11 | SSR | Zhang et al, 2012 |
| Backcross population | WayRarem × OryzicaLlanos 5 (IRGC117017); MR219 × O. rufipogon IRGC105491; SHZ-2 × TXZ-13; Oryza rufipogon × IR64 | 45 | SSR, SNP | Rahim et al, 2012 |
| F2, F3 and F4 | Nipponbare × Owarihatamochi (F4); Kahei × Koshihikari (F2:3); Tainung 69 × Koshihikari (F2); URN12 × Koshihikari (F2); Norin 29 × Chubu 32 (F3); PongsuSeribu 2 × Mahsuri (F2:3); TAM × KHZ (F2:3); Junambyeo × O. minuta introgression line IR71033-121-15 (F2:3); Danghang-Shali × Hokkai 188 (F2:3) | 60 | RFLP, SSR, STS | Ashkani et al, 2013a, b |
| RIL | 162 RILs derived from Heikezijing × Suyunuo | 2 | SSR | Fang et al, 2016 |
| RIL | 122 RILs derived from Dagad Deshi × Danteshwari | 5 | SSR | Mandal et al, 2017 |
| F3 | Nekken 2 × Hokuriku 193 | 3 | SNP | Nagaoka et al, 2017 |
Table 5 Examples of QTL mapping for resistance to blast disease in rice.
| Mapping population | Parent | Total No. of QTLs | Marker | Reference |
|---|---|---|---|---|
| RIL | CT9993-5-10-1-m × KDML105 (F8); Zhenshan 97 × Minghui 63 (RILs); Moroberekan × Co39 (F7); Lemont × Teqing (F8); Lemont × Teqing (F14); Bala × Azucena (F6); Zhong 156 × Gumei 2 (F8); OryzicaLlanos 5 × Fanny (F5 and F6); SHZ-2 × Lijiangxintuanheigu (LTH) (RILs); KDML105 × JHN (F6); Suweon 365 × Chucheong (RILs) | 186 | RFLP, SSR, RAPD, Isozymes, AFLP, DR gene markers | Cho et al, 2008 |
| Double haploid | IR64 × Azucena; IR64 × Azucena; ZYQ8 × JX17 | 146 | RFLP, RAPD, Isozymes | Xu et al, 2004 |
| SSSL | Developed by HXJ74 as recipient and 24 accessions as donors | 11 | SSR | Zhang et al, 2012 |
| Backcross population | WayRarem × OryzicaLlanos 5 (IRGC117017); MR219 × O. rufipogon IRGC105491; SHZ-2 × TXZ-13; Oryza rufipogon × IR64 | 45 | SSR, SNP | Rahim et al, 2012 |
| F2, F3 and F4 | Nipponbare × Owarihatamochi (F4); Kahei × Koshihikari (F2:3); Tainung 69 × Koshihikari (F2); URN12 × Koshihikari (F2); Norin 29 × Chubu 32 (F3); PongsuSeribu 2 × Mahsuri (F2:3); TAM × KHZ (F2:3); Junambyeo × O. minuta introgression line IR71033-121-15 (F2:3); Danghang-Shali × Hokkai 188 (F2:3) | 60 | RFLP, SSR, STS | Ashkani et al, 2013a, b |
| RIL | 162 RILs derived from Heikezijing × Suyunuo | 2 | SSR | Fang et al, 2016 |
| RIL | 122 RILs derived from Dagad Deshi × Danteshwari | 5 | SSR | Mandal et al, 2017 |
| F3 | Nekken 2 × Hokuriku 193 | 3 | SNP | Nagaoka et al, 2017 |
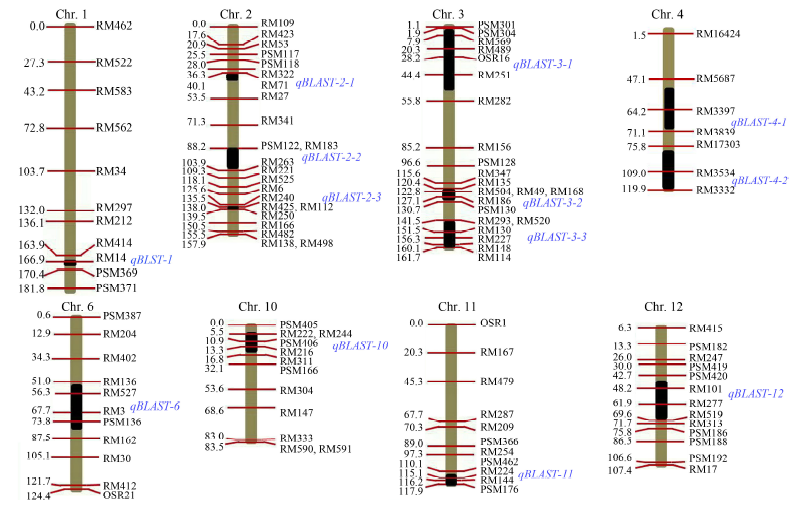
Fig. 1. Chromosomal location of 13 QTLs for blast resistance in rice. SSR markers are indicated on the right of the chromosomes. Genetic distance (cM) is shown on the left of the chromosomes. Black bars in each chromosome are the location intervals of QTLs for blast resistance with their names on the right (Zhang et al, 2012).
| R gene/locus | Chromosome | Rice germplasm | Reference |
|---|---|---|---|
| Pita | 12 | From wild rice species [O. rufipogon (Griff) and O. rufipogon (ETOR)] | Geng et al, 2008 |
| Pita | 12 | From O. rufipogon | Huang et al, 2008 |
| Pita | 12 | From cultivated (AA) and wild species and invasive weedy rice | Lee et al, 2011 |
| Pita | 12 | In 26 accessions, consisting of wild rice (O. rufipogon), cultivated rice (O. sativa) and related wild rice species (O. meridionalis and O. officinalis) | Yoshida and Miyashita, 2009 |
| Pita | 12 | From landraces and wild Oryza species | Ramkumar et al, 2010 |
| Pita | 12 | In Indian landraces of rice | Sharma et al, 2010 |
| Pita | 12 | From Indian landraces of rice collected from different ecogeographical regions | Thakur et al, 2013a |
| Pikh (Pi54) | 11 | From wild and cultivated species of rice | Rai et al, 2011 |
| Pikh (Pi54) | 11 | From the blast-resistant wild species of rice, O. rhizomatis | Das et al, 2012 |
| Pikh (Pi54) | 11 | From six cultivated rice lines and eight wild rice species | Kumari et al, 2013 |
| Pikh (Pi54) | 11 | In Indian landraces of rice | Sharma et al, 2010 |
| Pi-z(t) | 6 | In Indian landraces of rice | Sharma et al, 2010 |
| Pi-z(t) | 6 | In 529 landraces of rice collected at three different geographical locations of India | Thakur et al, 2013b |
| Pid3 | 6 | From 36 accessions of wild rice O. rufipogon | Xu et al, 2014 |
| Pid3-A4 | 6 | From wild rice A4 (O. rufipogon) | Lv et al, 2013 |
| Pi9 | 6 | In different rice species, five AA genome Oryza species including two cultivated rice species (O. sativa and O. glaberrima) and three wild rice species (O. nivara, O. rufipogon and O. barthii) | Liu et al, 2011 |
| Pi9 | 6 | From 338 rice landraces | Imam et al, 2016 |
| AC134922 | 11 | Rice lines from various sources | Wang D et al, 2014 |
| Pid3 | 6 | A set of 289 cultivated varieties including 140 indica and 149 japonica ones from China | Lv et al, 2017 |
Table 6 Example of allele mining for blast resistance in rice based on Ashikani et al (2015).
| R gene/locus | Chromosome | Rice germplasm | Reference |
|---|---|---|---|
| Pita | 12 | From wild rice species [O. rufipogon (Griff) and O. rufipogon (ETOR)] | Geng et al, 2008 |
| Pita | 12 | From O. rufipogon | Huang et al, 2008 |
| Pita | 12 | From cultivated (AA) and wild species and invasive weedy rice | Lee et al, 2011 |
| Pita | 12 | In 26 accessions, consisting of wild rice (O. rufipogon), cultivated rice (O. sativa) and related wild rice species (O. meridionalis and O. officinalis) | Yoshida and Miyashita, 2009 |
| Pita | 12 | From landraces and wild Oryza species | Ramkumar et al, 2010 |
| Pita | 12 | In Indian landraces of rice | Sharma et al, 2010 |
| Pita | 12 | From Indian landraces of rice collected from different ecogeographical regions | Thakur et al, 2013a |
| Pikh (Pi54) | 11 | From wild and cultivated species of rice | Rai et al, 2011 |
| Pikh (Pi54) | 11 | From the blast-resistant wild species of rice, O. rhizomatis | Das et al, 2012 |
| Pikh (Pi54) | 11 | From six cultivated rice lines and eight wild rice species | Kumari et al, 2013 |
| Pikh (Pi54) | 11 | In Indian landraces of rice | Sharma et al, 2010 |
| Pi-z(t) | 6 | In Indian landraces of rice | Sharma et al, 2010 |
| Pi-z(t) | 6 | In 529 landraces of rice collected at three different geographical locations of India | Thakur et al, 2013b |
| Pid3 | 6 | From 36 accessions of wild rice O. rufipogon | Xu et al, 2014 |
| Pid3-A4 | 6 | From wild rice A4 (O. rufipogon) | Lv et al, 2013 |
| Pi9 | 6 | In different rice species, five AA genome Oryza species including two cultivated rice species (O. sativa and O. glaberrima) and three wild rice species (O. nivara, O. rufipogon and O. barthii) | Liu et al, 2011 |
| Pi9 | 6 | From 338 rice landraces | Imam et al, 2016 |
| AC134922 | 11 | Rice lines from various sources | Wang D et al, 2014 |
| Pid3 | 6 | A set of 289 cultivated varieties including 140 indica and 149 japonica ones from China | Lv et al, 2017 |
| Transferred gene | Transformation method | Variety in which gene was transferred | Reference |
|---|---|---|---|
| Cht-2 or Cht-3 (class-I chitinase gene) | Agrobacterium-mediated | Japonica varieties Nipponbare and Koshihikari | Nishizawa et al,1999 |
| Rirlb (putative defense gene) | Biolistic | Japonica variety Taipei 309 | Schaffrath et al, 2000 |
| Barnase and Barstar | Biolistic | Japonica Zhonghua 8 and Taipei 309 | Mao et al, 2003 |
| Pid3 | Agrobacterium-mediated | Japonica variety Taipei 309 | Shang et al, 2009 |
| ACS2 (1-aminocyclopropane-1-carboxylic acid synthase, a key enzyme of ET biosynthesis) | Agrobacterium-mediated | Kitaake | Helliwell et al, 2013 |
| Maize regulatory genes C1 (coloured-1), R (red) and the structural gene C2 (coloured-2, encoding chalcone synthase) | Bioloistic | Taipei 309 | Gandikota et al, 2001 |
| cecropin A (cecropins, antimicrobial peptides) | Agrobacterium-mediated | Japonica rice Senia | Coca et al, 2006 |
| Aspergillus giganteus antifungal protein (AFP) | Agrobacterium-mediated | Japonica rice Senia | Coca et al, 2004; Moreno et al, 2005 |
| pinA and/or pinB (ouroindoline genes) | - | M202 | Krishnamurthy et al, 2001 |
| WRKY45 | Agrobacterium-mediated | Nipponbare | Shimono et al, 2007 |
| Thanatin (antimicrobial peptide) | Agrobacterium-mediated | Nipponbare | Imamura et al, 2010 |
| BjNPR1 (Brassica juncea non expressor of pathogenesis-related gene 1) | Agrobacterium-mediated | Indica varieties Chaitanya and Samba Mahsuri | Sadumpatia et al, 2013 |
| Pikh | Agrobacterium-mediated | MR219 | Azizi et al, 2016 |
Table 7 Examples of transgenic rice having blast resistance.
| Transferred gene | Transformation method | Variety in which gene was transferred | Reference |
|---|---|---|---|
| Cht-2 or Cht-3 (class-I chitinase gene) | Agrobacterium-mediated | Japonica varieties Nipponbare and Koshihikari | Nishizawa et al,1999 |
| Rirlb (putative defense gene) | Biolistic | Japonica variety Taipei 309 | Schaffrath et al, 2000 |
| Barnase and Barstar | Biolistic | Japonica Zhonghua 8 and Taipei 309 | Mao et al, 2003 |
| Pid3 | Agrobacterium-mediated | Japonica variety Taipei 309 | Shang et al, 2009 |
| ACS2 (1-aminocyclopropane-1-carboxylic acid synthase, a key enzyme of ET biosynthesis) | Agrobacterium-mediated | Kitaake | Helliwell et al, 2013 |
| Maize regulatory genes C1 (coloured-1), R (red) and the structural gene C2 (coloured-2, encoding chalcone synthase) | Bioloistic | Taipei 309 | Gandikota et al, 2001 |
| cecropin A (cecropins, antimicrobial peptides) | Agrobacterium-mediated | Japonica rice Senia | Coca et al, 2006 |
| Aspergillus giganteus antifungal protein (AFP) | Agrobacterium-mediated | Japonica rice Senia | Coca et al, 2004; Moreno et al, 2005 |
| pinA and/or pinB (ouroindoline genes) | - | M202 | Krishnamurthy et al, 2001 |
| WRKY45 | Agrobacterium-mediated | Nipponbare | Shimono et al, 2007 |
| Thanatin (antimicrobial peptide) | Agrobacterium-mediated | Nipponbare | Imamura et al, 2010 |
| BjNPR1 (Brassica juncea non expressor of pathogenesis-related gene 1) | Agrobacterium-mediated | Indica varieties Chaitanya and Samba Mahsuri | Sadumpatia et al, 2013 |
| Pikh | Agrobacterium-mediated | MR219 | Azizi et al, 2016 |
| [1] | Ahloowalia B S, Maluszynski M, Nichterlein K.2004. Global impact of mutation-derived varieties.Euphytica, 135(2): 187-204. |
| [2] | Ahn S N, Kim Y K, Han S S, Choi H C, Moon H P, McCouch S R.1996. Molecular mapping of a gene for resistance to a Korean isolate of rice blast.Rice Genet Newsl, 13: 74-76. |
| [3] | Ahn S N, Kim Y K, Hong H C, Han S S, Kwon S J, Choi H C, Moon H P, McCouch S R.2000. Molecular mapping of a new gene for resistance to rice blast (Pyricularia grisea Sacc.).Euphytica, 116: 17-22. |
| [4] | Allard R.1999. Principles of Plant Breeding. 2nd edn. New York: Willey. |
| [5] | Amante-Bordeos A, Sitch L A, Nelson R, Dalmacio R D, Oliva N P, Aswidinnoor H, Leung H.1992. Transfer of bacterial blight and blast resistance from the tetraploid wild rice Oryza minuta to cultivated rice Oryza sativa.Theor Appl Genet, 84: 345-354. |
| [6] | Anon.1992. Statistics of Agro-Chemicals in Japan. Tokyo, Japan: Japan Plant Protection Association. |
| [7] | Anupam A, Imam J, Quatadah S M, Siddaiah A, Das S P, Variar M, Mandal N P.2017. Genetic diversity analysis of rice germplasm in Tripura state of Northeast India using drought and blast linked markers.Rice Sci, 24(1): 10-20. |
| [8] | Anwar A, Bhat M S.2005. Efficacy of fungicides as seed treatment in the management of blast disease of rice in nursery bed.Agric Sci Digest, 25(4): 293-295. |
| [9] | Arunakumari K, Durgarani C V, Sattru V, Sarikonda K R, Chitoor P D R, Vutukuri B, Laha G S, Nelli A P K, Gatty S, Jamal M, Prasadbabu A, Hajira S, Sundaram R M.2016. Marker assisted pyramiding of genes conferring resistance against bacterial blight and blast diseases into Indian rice variety MTU1010.Rice Sci, 23(6): 306-316. |
| [10] | Asghar A, Rashid H, Ashraf M, Khan M H, Chaudhry Z.2007. Improvement of basmati rice against fungus infection through gene transfer technology.Pak J Bot, 39(4): 1277-1283. |
| [11] | Ashkani S, Rafii M Y, Rusli I, Sariah M, Abdullah S N A, Rahim H A, Latif M A.2012. SSRs for marker-assisted selection for blast resistance in rice (Oryza sativa L.).Plant Mol Biol Rep, 30(1): 79-86. |
| [12] | Ashkani S, Rafii M Y, Rahim H A, Latif M A.2013a. Genetic dissection of rice blast resistance by QTL mapping approach using an F3 population.Mol Biol Rep, 40(3): 2503-2515. |
| [13] | Ashkani S, Rafii M Y, Rahim H A, Latif M A.2013b. Mapping of the quantitative trait locus (QTL) conferring partial resistance to rice leaf blast disease.Biotechnol Lett, 35(5): 799-810. |
| [14] | Ashkani S, Rafil M Y, Shabanimofrad M, Miah G, Sahebi M, Azizi P, Tanweer F A, Akhtar M S, Nasehi A.2015. Molecular breeding strategy and challenges towards the improvement of blast disease resistance in rice crop.Front Plant Sci, 6: 1-14. |
| [15] | Ashkani S, Rafii M Y, Shabanimofrad M, Ghasemzadeh A, Ravanfar S A, Latif M A.2016. Molecular progress on the mapping and cloning of functional genes for blast disease in rice (Oryza sativa L.): Current status and future considerations.Crit Rev Biotechnol, 36: 353-367. |
| [16] | Azizi P, Rafii M Y, Abdullah S N A, Hanafi M M, Maziah M, Sahebi M, Ashkani S, Taheri S, Jahromi M F.2016. Over-expression of the Pikh gene with a CaMV 35S promoter leads to improved blast disease (Magnaporthe oryzae) tolerance in rice.Front Plant Sci, 7: 773. |
| [17] | Baldrich P, Campo S, Wu M T, Liu T T, Hsing Y I, Segundo B S.2015. MicroRNA-mediated regulation of gene expression in the response of rice plants to fungal elicitors.RNA Biol, 12: 847-863. |
| [18] | Ballini E, Morel J B, Droc G, Price A, Courtois B, Notteghem J L, Tharreau D.2008. A genome-wide meta analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol Plant Microb Interact, 21: 859-868. |
| [19] | Barman S R, Gowda M, Venu R C, Chattoo B B.2004. Identification of a major blast resistance gene in the rice cultivar Tetep.Plant Breeding, 123(3): 300-302. |
| [20] | Berruyer R, Adreit H, Milazzo J, Gaillard S, Berger A, Dioh W, Lebrun M H, Tharreau D.2003. Identification and fine mapping of Pi33, the rice resistance gene corresponding to the Magnaporthe grisea avirulence gene ACE1.Theor Appl Genet, 107(6): 1139-1147. |
| [21] | Beser N, Del Valle M M, Kim S M, Vinarao B R, Surek H, Jena K K.2016. Marker-assisted introgression of a broad-spectrum resistance gene, Pi40 improved blast resistance of two elite rice (Oryza sativa L.).Mol Plant Breeding, 7(33): 1-15. |
| [22] | Bevitori R, Ghini R.2014. Rice blast disease in climate change times.J Rice Res, 3: e111. |
| [23] | Bryan G T, Wu K S, Farrall L, Jia Y L, Hershey H P, McAdams S A, Faulk K N, Donaldson G K, Tarchini R, Valent B.2000. Single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell, 12: 2033-2046. |
| [24] | Bundo M, Coca M.2015. Enhancing blast disease resistance by overexpression of the calcium-dependent protein kinase OsCPK4 in rice.Plant Biotechnol J, 14(6): 1357-1367. |
| [25] | Calvero Jr S B, Coakley S M, Teng P S.1996. Development of empirical forecasting models for rice blast based on weather factors.Plant Pathol, 45(4): 667-678. |
| [26] | Causse M A, Fulton T M, Cho Y G, Ahn S N, Chunwongse J, Wu K S, Xiao J H, Yu Z H, Ronald P C, Harrington S E, Second G, McCouch S R, Tanksley S D.1994. Saturated molecular map of the rice genome based on an interspecific backcross population.Genetics, 138(4): 1251-1274. |
| [27] | Chen D H, Dela Vina M, Inukai T, Mackill D J, Ronald P C, Nelson R J.1999. Molecular mapping of the blast resistance gene, Pi44(t), in a line derived from a durably resistant rice cultivar.Theor Appl Genet, 98: 1046-1053. |
| [28] | Chen H Q, Chen Z X, Ni S, Zuo S M, Pan X B, Zhu X D.2008. Pyramiding three genes with resistance to blast by marker assisted selection to improve rice blast resistance of Jin23B.Chin J Rice Sci, 22: 23-27. (in Chinese with English abstract) |
| [29] | Chen X W, Li S G, Ma Y Q, Li H Y, Zhou K D, Zhu L H.2004. Marker-assisted selection and pyramiding for three blast resistance genes, Pi-d(t)1, Pi-b, Pi-ta2, in rice.Chin J Biotechnol, 20(5): 708-714. (in Chinese with English abstract) |
| [30] | Chen X W, Shang J J, Chen D X, Lei C L, Zou Y, Zhai W X, Liu G Z, Xu J C, Ling Z Z, Cao G, Ma B T, Wang Y P, Zhao X F, Li S G, Zhu L H.2006. A β-lectin receptor kinase gene conferring rice blast resistance.Plant J, 46(5): 794-804. |
| [31] | Chen X W, Ronald P C.2011. Innate immunity in rice.Trends Plant Sci, 16(8): 451-459. |
| [32] | Cho Y C, Kwon S W, Suh J P, Kim J J, Lee J H, Roh J H, Oh M K, Kim M K, Ahn S N, Koh H J.2008. QTLs identification and confirmation of field resistance to leaf blast in temperate japonica rice (Oryza sativa L.).J Crop Sci Biotechnol, 11: 269-276. |
| [33] | Choi W J, Park E W, Lee E J.1988. LEAF BLAST: A computer simulation model for leaf blast development on rice. Kor J Plant Pathol (Korea R), 4: 25-32. |
| [34] | Coca M, Bortolotti C, Rufat M, Pen G, Eritja R, Tharreau D, del Pozo A M, Messeguer J, Segundo B S.2004. Transgenic rice plants expressing the antifungal AFP protein from Aspergillus giganteus show enhanced resistance to the rice blast fungus Magnaporthe grisea.Plant Mol Biol, 54(2): 245-259. |
| [35] | Coca M, Peaas G, Gomez J, Campo S, Bortolotti C, Messeguer J, Segundo B S.2006. Enhanced resistance to the rice blast fungus Magnaporthe grisea conferred by expression of a cecropin A gene in transgenic rice.Planta, 233(3): 392-406. |
| [36] | Collard B C Y, Mackill D J.2008. Marker-assisted selection: An approach for precision plant breeding in the twenty-first century.Phil Trans R Soc B Biol Sci, 363: 557-572. |
| [37] | Costanzo S, Jia Y.2010. Sequence variation at the rice blast resistance gene Pi-km locus: Implications for the development of allele specific markers.Plant Sci, 178(6): 523-530. |
| [38] | Couch B C, Kohn L M.2002. A multilocus gene genealogy concordant with host preference indicates segregation of a new species, Magnaporthe oryzae, fromM. grisea. Mycologia, 94: 683-693. |
| [39] | Dangl J L, Horvath D M, Staskawicz B J.2013. Pivoting the plant immune system from dissection to deployment.Science, 341: 746-751. |
| [40] | Das A, Soubam D, Singh P K, Thakur S, Singh N K, Sharma T R.2012. A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct Integr Genom, 12(2): 215-228. |
| [41] | Das G, Rao G J N.2015. Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar.Front Plant Sci, 6: 1-18. |
| [42] | Dean R A, Talbot N J, Ebbole D J, Farman M L, Mitchell T K, Orbach M J, Thon M, Kulkarni R, Xu J R, Pan H, Read N D, Lee Y H, Carbone I, Brown D, Oh Y Y, Donofrio N, Jeong J S, Soanes D M, Djonovic S, Kolomiets E, Rehmeyer C, Li W, Harding M, Kim S, Lebrun M H, Bohnert H, Coughlan S, Butler J, Calvo S, Ma L J, Nicol R, Purcell S, Nusbaum C, Galagan E J, Birren B W.2005. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature, 434: 980-986. |
| [43] | Deng Y W, Zhu X D, Shen Y, He Z H.2006. Genetic characterization and fine mapping of the blast resistance locus Pigm(t) tightly linked to Pi2 and Pi9 in a broad-spectrum resistant Chinese variety.Theor Appl Genet, 113(4): 705-713. |
| [44] | Divya B, Robin S, Rabindran R, Senthil S, Raveendran M, Joel A J.2014. Marker assisted backcross breeding approach to improve blast resistance in Indian rice (Oryza sativa) variety ADT43.Euphytica, 200(1): 61-77. |
| [45] | Dong T G, Ho B T, Yoder-Himes D R, Mekalanos J J.2013. Identification of T6SS-dependent effector and immunity proteins by Tn-seq Vibrio cholera. inProc Natl Acad Sci USA, 110(7): 2623-2628. |
| [46] | Fang N Y, Wang R S, He W W, Yin C F, Guan C H, Chen H, Huang J, Wang J F, Bao Y M, Zhang H S.2016. QTL mapping of panicle blast resistance in japonica landrace Heikezieng and its application in rice breeding.Mol Breeding, 36: 171-179. |
| [47] | Fjellstrom R, Mc-Clung A M, Shank A R.2006. SSR markers closely linked to the Pi-z locus are useful for selection of blast resistance in abroad array of rice germplasm.Mol Breeding, 17(2): 149-157. |
| [48] | Filippi M C, Prabhu A S.1997. Integrated effect of host plant resistance and fungicidal seed treatment on rice blast control in Brazil.Plant Dis, 81(4): 351-355. |
| [49] | Fu C Y, Wu T, Liu W G, Wang F, Li J H, Zhu X Y, Huang H J, Liu Z R, Liao Y L, Zhu M S, Chen J W, Huang Y J.2012. Genetic improvement of resistance to blast and bacterial blight of the elite maintainer line Rongfeng B in hybrid rice (Oryza sativa L.) by using marker-assisted selection.Afr J Biotechnol, 11(67): 13104-13114. |
| [50] | Fujimski H.1979. Recurrent selection by using male sterility for rice improvement.Jpn Agric Res Quart, 13: 153-156. |
| [51] | Fujita D, Ebron L A, Kobayashi N, Fukuta Y.2009. DNA marker analysis of blast resistance genes Pib and Pita in IRRI-bred rice varieties comparison with gene estimation by a differential system. In: Wang G L, Valent B. Advances in Genetics, Genomics and Control of Rice Blast Disease. Berlin: Springer: 315-324. |
| [52] | Fuentes J L, Correa-Victoria F J, Escobar F, Prado G, Aricapa G, Duque M C, Tohme J.2008. Identification of microsatellite markers linked to the blast resistance gene Pi-1(t) in rice.Euphytica, 160: 295-304. |
| [53] | Fukuoka S, Okuno K.2001. QTL analysis and mapping of pi21, a recessive gene for field resistance to rice blast in Japanese upland rice.Theor Appl Genet, 103: 185-190. |
| [54] | Fukuoka S, Okuno K, Kawase M.2007. Rice blast disease gene Pi21, resistance gene pi21 and utilization thereof. Patent WO/2007/ 000880. |
| [55] | Fukuoka S, Saka N, Mizukami Y, Koga H, Yamanouchi U, Yoshioka Y, Hayashi N, Ebana K, Mizobuchi R, Yano M.2015. Gene pyramiding enhances durable blast disease resistance in rice.Sci Rep, 5: 7773. |
| [56] | Fukuta Y, Araki E, Yanoria M J T, Imbe T, Tsunematsu H, Kato H, Ebron L A, Mercado-Escueta D, Khush G S.2004. Development of differential varieties for blast resistance in IRRI-Japan collaborative research project. In: Kawasaki S. Rice Blast: Interaction with Rice and Control. Kluwer, Dordrecht: Springer: 229-233. |
| [57] | Gandikota M, de Kochko A, Chen L, Ithal N, Fauquet C, Reddy A R.2001. Development of transgenic rice plants expressing maize anthocyanin genes and increased blast resistance.Mol Breeding, 7(1): 73-83. |
| [58] | Geng X S, Yang M Z, Huang X Q, Cheng Z Q, Fun J, Sun T, Li J.2008. Cloning and analyzing of rice blast resistance gene Pi-ta+ allele from Jinghong erect type of common wild rice (Oryza rufipogon Griff) in Yunnan.Hereditas, 30(1): 109-114. |
| [59] | Ghazanfar M U, Wakil W, Sahi S T, Saleem-il-Yasin.2009. Influence of various fungicides on the management of rice blast disease.Mycopath, 7(1): 29-34. |
| [60] | Gohel N M, Chauhan H L.2015. Integrated management of leaf and neck blast disease of rice caused by Pyricularia oryzae.Afr J Agric Res, 10(19): 2038-2040. |
| [61] | Goto I.1970. Genetic studies on the resistance of rice plant to the blast fungus: I. Inheritance of resistance in crosses SenshoXH-79 and Imochi-shirazu XH-79.Ann Phytopathol Soc, 36: 304-312. |
| [62] | Goto I.1988. Genetic studies on resistance of rice plant to blast fungus: 7. Blast resistance genes of Kuroka.Ann Phytopathol Soc Jpn, 54: 460-465. |
| [63] | Gouda P K, Saikumar S, Varma C M K, Nagesh K, Thippeswamy S, Shenoy V, Ramesha M S, Shashidhar H E.2013. Marker-assisted breeding of Pi-1 and Piz-5 genes imparting resistance to rice blast in PRR78 restorer line of Pusa RH-10 Basmati rice hybrid.Plant Breeding, 132(1): 61-69. |
| [64] | Gowda M, Roy-Barman S, Chatoo B B.2006. Molecular mapping of a novel blast resistance gene Pi38 in rice using SSLP and AFLP markers.Plant Breeding, 125(6): 596-599. |
| [65] | Gowda M, Shirke M D, Mahesh H B, Chandarana P, Rajamani A, Chattoo B B.2015. Genome analysis of rice-blast fungus Magnaporthe oryzae field isolates from southern India.Genom Data, 5: 284-291. |
| [66] | Guimaraes E P, Correa-Victoria F.1997. Use of recurrent selection for develop resistance Pyricularria grisea Sacc. resistance in rice. In: Guimares E P. Advances in Rice Population Improvement. Cali, Colombia: CIAT: 165-175. |
| [67] | Guo L Y, Guo W, Zhao H W, Wang J G, Liu H L, Sun J, Zheng H L, Sha H J, Zou D T.2015. Association mapping and resistant alleles’ analysis for japonica rice blast resistance.Plant Breeding, 134(6): 646-652. |
| [68] | Hasan M M, Rafii M Y, Ismail M R, Mahmood M, Rahim H A, Alam M A, Ashkani S, Malek M A, Latif M A.2015. Marker-assisted backcrossing: A useful method for rice improvement.Biotechnol Biotechnol Equip, 29: 237-254. |
| [69] | Hayashi K, Yoshida H, Ashikawa I.2006. Development of PCR-based allele specific and InDel marker sets for nine rice blast resistance genes.Theor Appl Genet, 113(2): 251-260. |
| [70] | Hayashi N, Ando I, Imbe T.1998. Identification of a new resistance gene to a Chinese blast fungus isolate in the Japanese rice cultivar Aichi Asahi.Phytopathology, 88: 822-827. |
| [71] | He W W, Fang N Y, Wang R S, Wu Y Y, Zen G Y, Guan C H, Chen H, Huang J, Wang J F, Bao Y M, Zhang H S.2017. Fine mapping of a new race-specific blast resistance gene, Pi-hk2, in japonica Heikezijing from Taihu region of China.Phytopathology, 107: 84-91. |
| [72] | He X Y, Liu X Q, Wang L, Wang L, Lin F, Cheng Y S, Chen Z M, Liao Y P, Pan Q H.2012. Identification of the novel recessive gene pi55(t) conferring resistance to Magnaporthe oryzae.Sci China Life Sci, 55: 141-149. |
| [73] | Hegde S S, Prashanthi S K.2016. Identification of polymorphic markers and introgression of Pi1 and Pi2 genes for blast resistance in rice.J Farm Sci, 29: 327-331. |
| [74] | Helliwell E E, Wang Q, Yang Y.2013. Transgenic rice with inducible ethylene production exhibits broad spectrum disease resistance to the fungal pathogens Magnaporthe oryzae and Rhizoctonia solani. Plant Biot J, 11(1): 33-42. |
| [75] | Hittalmani S, Parco A, Mew T V, Zeigler R S, Huang N.2000. Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice.Theor Appl Genet, 100(7): 1121-1128. |
| [76] | Hua L X, Wu J Z, Chen C X, Wu W H, He X Y, Lin F, Wang L, Ashikawa I, Matsumoto T, Wang L, Pan Q H.2012. The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast.Theor Appl Genet, 125: 1047-1055. |
| [77] | Hua L X, Liang L Q, He X Y, Wang L, Zhang W S, Liu W, Liu X Q, Lin F.2015. Development of a marker specific for the rice blast resistance gene Pi39 in the Chinese cultivar Q15 and its use in genetic improvement.Biotech Biotechnol Equip, 29(3): 448-456. |
| [78] | Huang C L, Hwang S Y, Chiang Y C, Lin T P.2008. Molecular evolution of the Pi-ta gene resistant to rice blast in wild rice (Oryza rufipogon).Genetics, 179(3): 1527-1538. |
| [79] | Huang H M, Huang L, Feng G P, Wang S H, Wang Y, Liu J L, Jiang N, Yan W T, Xu L C, Sun P Y, Li Z Q, Pan S J, Liu X L, Xiao Y H, Liu E M, Dai L Y, Wang G L.2011. Molecular mapping of the new blast resistance genes Pi47 and Pi48 in the durably resistant local rice cultivar Xiangzi 3150.Phytopathology, 101: 620-626. |
| [80] | Hubert J, Mabagala R, Mamiro D.2015. Efficacy of selected plant extacts against Pyricularia grisea, causal agent of rice blast disease.Am J Plant Sci, 6: 602-611. |
| [81] | Imam J, Mandal N P, Variar M, Shukla P.2016. Allele mining and selective patterns of Pi9 gene in a set of rice landraces from India.Front Plant Sci, 10: 1846. |
| [82] | Imamura T, Yasuda M, Kusano H, Nakashita H, Ohno Y, Kamakura T, Taguchi S, Shimada H.2010. Acquired resistance to the rice blast in transgenic rice accumulating the antimicrobial peptide thanatin.Transg Res, 19(3): 415-424. |
| [83] | Inukai T, Nelson R.1994. Mapping for blast resistance gene H-3 derived from rice cultivar Pai-Kan-Tao. Rep Hokkaido Br Crop Sol See Jpn Soc Breeding, 35: 54-55. |
| [84] | Izawall T, Iwasakizl M.2000. Identification of a RFLP marker tightly linked to the panicle blast resistance gene, Pb1, in rice.Breeding Sci, 50: 183-188. |
| [85] | Jain P, Singh P K, Kapoor R, Khanna A, Solanke A U, Krishnan S G, Singh A K, Sharma V, Sharma T R.2017. Understanding host-pathogen interactions with expression profiling of NILs carrying rice-blast resistance Pi9 gene.Front Plant Sci, 8: 93. |
| [86] | Jena K K, Khush G S.2000. Exploitation of species in rice improvement opportunities, achievements and future challenges. In: Nanda J S. Rice Breeding and Genetic Research Priorities and Challenges. New Hampshiire USA: Enfield: Science Publication: 269-284. |
| [87] | Jeon J S, Chen D, Yi G H, Wang G L, Ronald P C.2003. Genetic and physical mapping of Pi5(t), a locus associated with broad-spectrum resistance to rice blast.Mol Genet Genom, 269: 280-289. |
| [88] | Jeung J U, Kim B R, Cho Y C, Han S S, Moon H P, Lee Y T, Jena K K.2007. A novel gene, Pi40(t), linked to the DNA markers derived from NBSLRR motifs confers broad spectrum of blast resistance in rice.Theor Appl Genet, 115(8): 1163-1177. |
| [89] | Ji Z J, Yang S D, Zeng Y X, Liang Y, Yang C D, Qian Q.2016. Pyramiding blast, bacterial blight and brown planthopper resistance genes in rice restorer lines.J Intl Agric, 15(7): 1432-1440. |
| [90] | Jiang J F, Mou T M, Yu H H, Zhou F S.2015a. Molecular breeding of thermo-sensitive genic male sterile (TGMS) lines of rice for blast resistance using Pi2 gene.Rice, 8: 11. |
| [91] | Jiang J F, Yang D B, Ali J, Mou T M.2015b. Molecular marker-assisted pyramiding of broad-spectrum disease resistance genes, Pi2 and Xa23, into GZ63-4S, an elite thermo-sensitive genic male-sterile line in rice.Mol Breeding, 35: 1-12. |
| [92] | Jiang N, Li Z Q, Wu J, Wang Y, Wu L Q, Wang S H, Wang D, Wen T, Liang Y, Sun P Y, Liu J L, Dai L Y, Wang Z L, Wang C, Luo M Z, Liu X L, Wan G L.2012. Molecular mapping of the Pi2/9 allelic gene Pi2-2 conferring broad-spectrum resistance to Magnaporthe oryzae in the rice cultivar Jefferson.Rice, 5: 29-36. |
| [93] | Kadotani N, Nakayashiki H, Tosa Y, Mayama S.2003. RNA silencing in the phytopathogenic fungus Magnaporthe oryza.Mol Plant Microb Int, 16(9): 769-776. |
| [94] | Kapoor A S, Singh B M.1982. Evaluation of some fungicides for the control of rice blast.Ind Phytopath, 53: 283. |
| [95] | Katiyar-Agarwal S, Jin H L.2010. Role of small RNAs in host-microbe interactions.Annu Rev Phytopathol, 48: 225-246. |
| [96] | Kaur S, Padmanabhan S Y, Rao M.1975. Induction of resistance to blast disease (Pyricularia oryzae) in the high yielding variety, Ratna (IRE 9 TKM 6). In: Proceedings of the IAEA research coordination. Geoling, Ames, Iowa: 141-145. |
| [97] | Khanna A, Sharma V, Ellur R K, Shikari A B, Gopala Krishnan S, Singh U D, Prakash G, Sharma T R, Rathour R, Variar M, Prashanthi S K, Nagarajan M, Vinod K K, Bhowmick P K, Singh N K, Prabhu K V, Singh B D, Singh A K.2015. Development and evaluation of near-isogenic lines for major blast resistance gene(s) in Basmati rice.Theor Appl Genet, 128(7): 1243-1259. |
| [98] | Kim C K, Kim C H.1993. The rice leaf blast simulation model EPIBLAST. In: Penning de Vries F, Teng P, Metselaar K. Systems Approaches for Agricultural Development. Dordrecht: Springer: 309-321. |
| [99] | Kim J A, Cho K, Singh R, Jung Y H, Jeong S H, Kim S H, Lee J E, Cho Y S, Agrawal G K, Rakwal R, Tamogami S, Kersten B, Jeon J S, An G, Jwa N S.2009. Rice OsACDR1 (Oryza sativa accelerated cell death and resistance 1) is a potential positive regulator of fungal disease resistance.Mol Cells, 28(5): 431-439. |
| [100] | Kinoshita T, Kiyosawa S.1997. Some considerations on linkage relationships between Pii and Piz in the blast resistance of rice.Rice Genet Newsl, 14: 57-59. |
| [101] | Koide Y, Kobayashi N, Xu D, Fukuta Y.2009. Resistance genes and selection DNA markers for blast disease in rice (Oryza sativa L.).Jpn Agric Res Q, 43(4): 255-280. |
| [102] | Koide Y, Kawasaki A, Telebancoâaaa-Yanoria M J, Hairmansis A, Nguyet N T M, Bigirimana J, Fujita D, Kobayashi N, Fukuta Y.2010. Development of pyramided lines with two resistance genes, Pish and Pib, for blast disease (Magnaporthe oryzae B. Couch) in rice (Oryza sativa L.).Plant Breeding, 129(6): 670-675. |
| [103] | Koide Y, Ebron L A, Kato H, Tsunematsu H, Telebanco-Yanoria M J, Kobayashi N, Yokoo M, Maruyama S, Imbe T, Fukuta Y.2011. A set of near-isogenic lines for blast resistance genes with an indica-type rainfed lowland elite rice (Oryza sativa L.) genetic background.Field Crops Res, 123(1): 19-27. |
| [104] | Koide Y, Telebanco-Yanoria M J, Fukuta Y, Kobayashi N.2013. Detection of novel blast resistance genes, Pi58(t) and Pi59(t), in a Myanmar rice landrace based on a standard differential system.Mol Breeding, 32: 241-252. |
| [105] | Kontargyri V T, Tsekouras G J, Gialketsi A A, Kontaxis P A.2008. Comparison between artificial neural network algorithms for the estimation of the flashover voltage on insulators. In: 9th WSEAS International Conference on Neural Networks. Sofia, Bulgaria. 2-5 May: 225-230. |
| [106] | Korinsak S, Sirithunya P, Meakwatanakarn P, Sarkarung S, Vanavichit A, Toojinda T.2011. Changing allele frequencies associated with specific resistance genes to leaf blast in backcross introgression lines of Khao Dawk Mali 105 developed from a conventional selection program.Field Crops Res, 122(1): 32-39. |
| [107] | Krishnamurthy K, Balconi C, Sherwood J E, Giroux M J.2001. Wheat puroindolines enhance fungal disease resistance in transgenic rice.Mol Plant Microb Int, 14: 1255-1260. |
| [108] | Kumar M K P, Gowda D K S, Moudgal R, Kumar N K, Gowda K T P, Vishwanath K.2013. Impact of fungicides on rice production in India. In: Nita M. Fungicides: Showcases of Integrated Plant Disease Management from Around the World. InTech: 77-98. |
| [109] | Kumar P, Pathania S, Katoch P, Sharma T R, Plaha P, Rathour R.2010. Genetic and physical mapping of blast resistance gene Pi42(t) on the short arm of rice chromosome 12.Mol Breeding, 25(2): 217-228. |
| [110] | Kumar V A, Balachiranjeevi C H, Naik S B, Rambabu R, Rekha G, Harika G, Hajira S K, Pranathi K, Vijay S, Anila M, Mahadevaswamy H K, Kousik M, Yugander A, Aruna J, Hari Prasad M S, Madhav M S, Laha G S, Balachandran S M, Prasad M S, Ravindra Babu V, Sundaram R M.2016. Marker-assisted improvement of the elite restorer line of rice, RPHR-1005 for resistance against bacterial blight and blast diseases.J Genet, 95(4): 895-903. |
| [111] | Kumari A, Das A, Devanna B N, Thakur S, Singh P K, Singh N K, Sharma T R.2013. Mining of rice blast resistance gene Pi54 shows effect of single nucleotide polymorphisms on phenotypic expression of the alleles.Eur J Plant Pathol, 137(1): 55-65. |
| [112] | Kurahashi Y2001. Melanin biosynthesis inhibitors (MBIs) for control of rice blast.Pestic outlook, 12: 32-35. |
| [113] | Kwon S W, Cho Y C, Kim Y G, Suh J P, Jeung J U, Roh J H, Lee S K, Jeon J S, Yang S J, Lee Y T.2008. Development of near isogenic japonica rice lines with enhanced resistance to Magnaporthe grisea.Mol Cell, 25: 407-416. |
| [114] | Lee S, Costanzo S, Jia Y, Olsen K M, Caicedo A L.2009. Evolutionary dynamics of the genomic region around the blast resistance gene Pi-ta in AA genome Oryza species.Genetics, 183(4): 1315-1325. |
| [115] | Lee S, Jia Y, Jia M, Gealy D R, Olsen K M, Caicedo A L.2011. Molecular evolution of the rice blast resistance gene Pi-ta in invasive weedy rice in the USA.PLoS One, 6: e26260. |
| [116] | Lei C, Huang D, Li W, Wang J L, Liu Z L, Wang Z L, Wang X T, Shi K, Cheng Z J, Zhang X, Ling Z, Wan J M.2005. Molecular mapping of a blast resistance gene in an indica rice cultivar Yanxian No. 1.J Rice Genet News, 22: 76-77. |
| [117] | Li L Y, Wang L, Jing J X, Li Z, Lin F, Huang L F, Pan Q H.2007. The Pikm gene, conferring stable resistance to isolates of Magnaporthe oryzae, was finely mapped in a crossover-cold region on rice chromosome 11.Mol Breeding, 20: 179-188. |
| [118] | Li T, Liu B, Spalding M H, Weeks D P, Yang B.2012. High-efficiency TALEN-based gene editing produces disease- resistant rice.Nat Biotechnol, 30: 390-392. |
| [119] | Li Y, Lu Y G, Shi Y, Wu L, Xu Y J, Huang F, Guo X Y, Zhang Y, Fan J, Zhao J Q, Zhang H Y, Xu P Z, Zhou J M, Wu X J, Wang P R, Wang W M.2014. Multiple rice microRNAs are involved in immunity against the blast fungus Magnaporthe oryzae.Plant Physiol, 164(2): 1077-1092. |
| [120] | Li Y, Zhao S L, Li J L, Hu X H, Wang H, Cao X L, Xu Y J, Zhao Z X, Xiao Z Y, Yang N, Fan J, Huang F, Wang W M.2017. Osa-miR169 negatively regulates rice immunity against the blast fungus Magnaporthe oryzae.Front Plant Sci, 8: 2. |
| [121] | Liang Z J, Wang L, Pan Q H.2016. A new recessive gene conferring resistance against rice blast.Rice, 9: 47-52. |
| [122] | Lin F, Liu Y, Wang L, Liu X Q, Pan Q H.2007a. A high-resolution map of the rice blast resistance gene Pi15 constructed by sequence ready markers.Plant Breeding, 126(3): 287-290. |
| [123] | Lin F, Chen S, Que Z Q, Wang L, Liu X Q, Pan Q H.2007b. The blast resistance gene Pi37 encodes an NBS-LRR protein and is a member of a resistance gene cluster on rice chromosome 1.Genetics, 177: 1871-1880. |
| [124] | Liu B, Zhang S, Zhu X, Yang Q, Wu S, Mei M, Mauleon R, Leach J, Mew T, Leung H.2004. Candidate defense genes as predictors of quantitative blast resistance in rice.Mol Plant Microb Interact, 17: 1146-1152. |
| [125] | Liu S P, Li X, Wang C Y, Li X H, He Y Q.2002. Improvement of resistance to rice blast in Zhenshan 97 by molecular marker- aided selection.Acta Bot Sin, 45: 1346-1350. |
| [126] | Liu X Q, Wang L, Chen S, Lin F, Pan Q H.2005. Genetic and physical mapping of Pi36(t), a novel rice blast resistance gene located on rice chromosome 8.Mol Genet Gen, 274(4): 394-401. |
| [127] | Liu X Q, Lin F, Wang L, Pan Q H.2007. The in silico map-based cloning of Pi36, a rice coiled-coil-nucleotide-binding site- leucine-rich repeat gene that confers race-specific resistance to blast fungus.Genetics, 176: 2541-2549. |
| [128] | Liu Y, Zhu X Y, Zhang S H, Bernardo M, Edwards J, Galbraith D W, Leach J, Zhang G S, Liu B, Leung H.2011. Dissecting quantitative resistance against blast disease using heterogeneous inbred family lines in rice.Theor Appl Genet, 122(2): 341-353. |
| [129] | Liu Y, Liu B, Zhu X Y, Yang J Y, Bordeos A, Wang G L, Leach J E, Leung H.2012. Fine-mapping and molecular marker development for Pi56(t), a NBS-LRR gene conferring broadspectrum resistance to Magnaporthe oryzae in rice.Theor Appl Genet, 126: 985-998. |
| [130] | Liu Z Y, Wu H F, Huang J F.2010. Application of neural networks to discriminate fungal infection levels in rice panicles using hyperspectral reflectance and principal components analysis.Comp Elect Agric, 72(2): 99-106. |
| [131] | Luo W L, Huang M, Guo T, Xiao W M, Wang J F, Yang G L, Liu Y Z, Wang H, Chen Z H, Zhuang C X.2016. Marker-assisted selection for rice blast resistance genes Pi2 and Pi9 through high-resolution melting of a gene-targeted amplicon.Plant Breeding, 136: 67-73. |
| [132] | Luo Y C, Ma T C, Zhang A F, Ong K H, Luo Z X, Li Z F, Yang J B, Yin Z C.2017. Marker assisted breeding of Chinese elite rice cultivar 9311 for disease resistance to rice blast and bacterial blight and tolerance to submergence.Mol Breeding, 37: 106-128. |
| [133] | Lv Q M, Xu X, Shang J J, Jiang G H, Pang Z Q, Zhou Z Z, Wang J, Liu Y, Li T, Li X B, Xu J C, Cheng Z K, Zhao X F, Li S G, Zhu L H.2013. Functional analysis of Pid3-A4, an ortholog of rice blast resistance gene Pid3 revealed by allele mining in common wild rice.Phytopathology, 103(6): 594-599. |
| [134] | Lv Q M, Huang Z Y, Xu X, Tang L, Liu H, Wang C C, Zhou Z Z, Xin Y Y, Xing J J, Peng Z R, Li X B, Zheng T Q, Zhu L H.2017. Allelic variation of the rice blast resistance gene Pid3 in cultivated rice worldwide.Sci Rep, 7: 10362-10375. |
| [135] | Ma J, Lei C L, Xu X T, Hao K, Wang J L, Cheng Z J, Ma X D, Ma J, Zhou K N, Zhang X, Guo X P, Wu F Q, Lin Q B, Wang C M, Zhai H Q, Wang H Y, Wan J M.2015. Pi-64, encoding a novel CC-NBS-LRR protein, confers resistance to leaf and neck blast in rice.Mol Plant Microb Inter, 28(5): 558-568. |
| [136] | Mackill D, Bonman J.1992. Inheritance of blast resistance in nearisogenic lines of rice.Phytopathology, 82: 746-749. |
| [137] | Mahdian S A, Shahsavari A.2013. Pyramiding of blast resistance genes Pi-1 and Pi-2 in tarom mahalli rice cultivar.Seed Plant Improv J, 29: 391-395. |
| [138] | Mahesh H B, Shirke M D, Singh S, Rajamani A, Hittalmani S, Wang G L, Gowda M.2016. Indica rice genome assembly, annotation and mining of blast disease resistance genes.BMC Genom, 17: 242. |
| [139] | Malzahn A, Lowder L, Qi Y P.2017. Plant genome editing with TALEN and CRISPR.Cell Biosci, 7: 21. |
| [140] | Mandal L, Verma S K, Kotasthane A, Verulkar S.2017. Identification of quantitative trait loci for leaf blast resistance of rice (Oryza sativa L.).Biotechnol J Inter, 19(2): 1-14. |
| [141] | Manibhushanrao K, Krishnan P.1991. Epidemiology of blast (EPIBLA): A simulation model and forecasting system for tropical rice in India. In: Sreenivasaprasad S, Johnson R. Rice Blast Modeling and Forecasting. Manila, the Philippines: IRRI: 31-38. |
| [142] | Mao S J, Gu H Y, Qu L J, Chen Z L.2003. Obtaining transgenic rice resistant to rice fungal blast disease by controlled cell death strategy.Chin Sci Bull, 48(16): 1753-1759. |
| [143] | McCouch S R, Wright M H, Tung C W, Maron L G, McNally K L, Fitzgerald M, Singh N, DeClerck G, Agosto-Perez F, Korniliev P, Greenberg A J, Naredo M E B, Mercado S M Q, Harrington S E, Shi Y, Branchini D A, Kuser-Falcao P R, Leung H, Ebana K, Yano M, Eizenga G, McClung A, Mezy J.2016. Open access resources for genome wide association mapping in rice.Nat Commun, 7: 10532. |
| [144] | McDonald B A, Linde C.2002. Pathogen population genetics, evolutionary potential, and durable resistance.Annu Rev Phytopathol, 40: 349-379. |
| [145] | Mgonja E M, Park C H, Kang H, Balimponya E G, Opiyo S, Bellizzi M, Mutiga S K, Rotich F, Ganeshan V D, Mabagala R, Smeller C, Correll J, Zhou B, Talbot N J, Mitchell T K, Wang G L.2017. Genotyping-by-sequencing-based genetic analysis of African rice cultivars and association mapping of blast resistance genes against Magnaporthe oryzae populations in Africa.Phytopathology, 107(9): 1039-1046. |
| [146] | Miah G, Rafii M Y, Ismail M R, Puteh A B, Rahim H A, Asfaliza R, Latif M A.2013. Blast resistance in rice: A review of conventional breeding to molecular approaches.Mol Biol Rep, 40: 2369-2388. |
| [147] | Miyamoto M, Yano M, Hirasawa H.2001. Mapping of quantitative trait loci conferring blast field resistance in the Japanese upland rice variety Kahei.Breeding Sci, 51(4): 257-261. |
| [148] | Moreno A B, Penas G, Rufat M, Bravo J M, Estopà M, Messeguer J, Segundo B S.2005. Pathogen-induced production of the antifungal AFP protein from Aspergillus giganteus confers resistance to the blast fungus Magnaporthe grisea in transgenic rice.Mol Plant Microb Inter, 18(9): 960-972. |
| [149] | Motoyama T, Nakasako M, Yamaguchi I, Lyr H, Russell P E, Dehne H W, Sisler H D.1999. Molecular action mechanism of a new melanin biosynthesis inhibitor. In: Modern Fungicides and Antifungal Compounds II. 12th International Reinhardsbrunn Symposium. 24th-29th May, 1998. Friedrichroda, Thuringia, Germany: 111-119. |
| [150] | Nagato Y, Yoshimura A.1998. Report of the committee on gene symbolization, nomenclature and linkage groups.Rice Genet News Lett, 15: 13-74. |
| [151] | Nagaoka I, Sasahara H, Tabuchi H, Shigemune A, Matsushita K, Maeda H, Goto A, Fukuoka S, Ando T, Miura K.2017. Quantitative trait loci analysis of blast resistance in Oryza sativa L. Hokuriku 193.Breeding Sci, 67: 159-164. |
| [152] | Nakata Y, Ueno M, Kihara J, Ichii M, Taketa S, Arase S.2008. Rice blast disease and susceptibility to pests in a silicon uptake-deficient mutant.Crop Prot, 27: 865-868. |
| [153] | Naqvi N I, Chattoo B B.1996. Development of a sequence characterized amplified region (SCAR) based indirect selection method for a dominant blast-resistance gene in rice.Genome, 39: 26-30. |
| [154] | Narayanan N N, Baisakh N, Cruz C M V, Gnanamanickam S S, Datta K, Datta S K.2002. Molecular breeding for the development of blast and bacterial blight resistance in rice cv. IR50.Crop Sci, 42(6): 2072-2079. |
| [155] | Neelakanth, Sidde Gowda D K, Chethana B S, Parasappa H H.2017.In vitro and in vivo evaluation of fungicides against Pyricularia oryzae causing blast of rice.Int J Pure App Biosci, 5(3): 259-263. |
| [156] | Nguyen T T T, Koizumi S, La T N, Zenbayashi K S, Ashizawa T, Yasuda N, Imazaki I, Miyasaka A.2006. Pi35(t), a new gene conferring partial resistance to leaf blast in the rice cultivar Hokkai 188.Theor Appl Genet, 113(4): 697-704. |
| [157] | Ni D H, Song F S, Ni J L, Zhang A F, Wang C L, Zhao K J, Yang Y C, Wei P C, Yang J B, Li L.2015. Marker-assisted selection of two-line hybrid rice for disease resistance to rice blast and bacterial blight.Field Crops Res, 184: 1-8. |
| [158] | Nishimura A, Hino I.2002. Fenoxanil (ACHI-BU (R)): A new systemic funicide for the control of rice blast.Agrochem Jpn, 81: 13-15. |
| [159] | Nishizawa Y, Nishio Z, Nakazono K, Soma M, Nakajima E, Ugaki M, Hibi T.1999. Enhanced resistance to blast (Magnaporthe grisea) in transgenic japonica rice by constitutive expression of rice chitinase.Theor Appl Genet, 99: 383-390. |
| [160] | Ohyanagi H, Tanaka T, Sakai H, Shigemoto Y, Yamaguchi K, Habara T, Fuji Y, Antonio B A, Nagamura Y, Imanishi T, Ikeo K, Itoh T, Gojobori T, Sasaki T.2006. The rice annotation project database (RAP-DB): Hub for Oryza sativa ssp. japonica genome information.Nucl Acids Res, 34: D741-D744. |
| [161] | Oliva R, Win J, Raffaele S, Boutemy L, Bozkurt T O, Chaparro-Garcia A, Segretin M E, Stam R, Schornack S, Cano L M, van Damme M, Huitema E, Thines M, Banfield M J, Kamoun S.2010. Recent developments in effector biology of filamentous plant pathogens.Cell Microb, 12(6): 705-715. |
| [162] | Ou S H.1985. Rice Diseases. 2nd edn. Kew, Surrey, England: Commonwealth Mycological Institute: 61-96. |
| [163] | Pan Q H, Wang L, Ikehashi H, Tanisaka T.1996. Identification of a new blast resistance gene in the indica rice cultivar Kasalath using Japanese differential cultivars and isozyme markers.Phytopath, 86: 1071-1075. |
| [164] | Pan Q H, Wang L, Ikehashi H, Yamagata H, Tanisaka T.1998. Identification of two new genes conferring resistance to rice blast in the Chinese native cultivar Maowangu.Plant Breeding, 117: 27-31. |
| [165] | Pan Q H, Hu Z D, Takatoshi T, Wang L.2003. Fine mapping of the blast resistance gene Pi15, linked to Pii, on rice chromosome 9.Acta Bot Sin, 45(7): 871-877. |
| [166] | Petit-Houdenot Y, Fudal I.2017. Complex interactions between fungal avirulence genes and their corresponding plant resistance genes and consequences for disease resistance management.Front Plant Sci, 8: 1072. |
| [167] | Pinta W, Toojinda T, Thummabenjapone P, Sanitchon J.2013. Pyramiding of blast and bacterial leaf blight resistance genes in to rice cultivar RD6 using marker assisted selection.Afr J Biotechnol, 12: 4432-4438. |
| [168] | Prasad M S, Kanthi B A, Balachandran S M, Seshumadhav M, Mohan K M, Viraktamath B C.2009. Molecular mapping of rice blast resistance gene Pi-1(t) in the elite indica variety Samba mahsuri. World J Microbiol Biotechnol, 25: 1765-1769. |
| [169] | Prasanna Kumar M K, Siddegowda D K, Atheekur Rehaman H M, Pandurange Gowda K T, Sudarshan G K.2011. New strobilurin group fungicide in rice disease management.Pestology, 35(9): 34-39. |
| [170] | Prasanna Kumar M K, Veerabhadraswamy A L.2014. Appraise a combination of fungicides against blast and sheath blight diseases of paddy (Oryza sativa L.).J Exp Biol Agric Sci, 2(1): 49-57. |
| [171] | Qu S H, Liu G F, Zhou B, Bellizzi M, Zeng L R, Dai L Y, Han B, Wang G L.2006. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics, 172: 1901-1914. |
| [172] | Raboin L M, Ballini E, Tharreau D, Ramanantsoanirina A, Frouin J, Courtois B, Ahmadi N.2016. Association mapping of resistance to rice blast in upland field conditions.Rice, 9: 59. |
| [173] | Rahim H A, Bhuiyan M A R, Lim L S, Sabu K K, Saad A, Azhar M, Wickneswari R.2012. Identification of quantitative trait loci for blast resistance in BC2F3 and BC2F5 advanced backcross families of rice.Genet Mol Res, 11(3): 3277-3289. |
| [174] | Rai A K, Kumar S P, Gupta S K, Gautam N, Singh N K, Sharma T R.2011. Functional complementation of rice blast resistance gene Pi-kh (Pi54) conferring resistance to diverse strains of Magnaporthe oryzae.J Plant Biochem Biotechnol, 20(1): 55-65. |
| [175] | Ramkumar G, Biswal A K, Mohan K M, Sakthivel K, Sivaranjani A K P, Neeraja C N, Ram T, Balachandran S M, Sundaram R M, Prasad M S, Viraktamath B C, Madhav M S.2010. Identifying novel alleles of rice blast resistance genes Pikh and Pita through allele mining.Int Rice Res Notes, 117: 185. |
| [176] | Ramsey F, Schafer D W.1993. The Statistical Sleuth: An Intermediate Course in Statistical Methods. Corvallis: Oregon State University: 177. |
| [177] | Rani R, Yadav P, Barbadikar K M, Baliyan N, Malhotra E V, Singh B K, Kumar A, Singh D.2016. CRISPR/Cas9: A promising way to exploit genetic variation in plants.Biotechnol Lett, 11(1): 1-8. |
| [178] | Sachin U, Rana S K.2011. Effect of fungicides on neck blast incidence and grain yield of rice in mid hills of Himachal Pradesh.Plant Diseas Res, 26(2): 196. |
| [179] | Sadek S, Al-Hamadi A, Michaelis B.2013. Toward real-world activity recognition: An SVM based system using fuzzy directional features.WSEAS Trans Inf Sci Appl, 10: 116-127. |
| [180] | Sadumpatia V, Kalamburb M, Vudema D R, Kirti P B, Khareedua V R.2013. Transgenic indica rice lines, expressing Brassica juncea nonexpressor of pathogenesis-related genes 1 (BjNPR1), exhibit enhanced resistance to major pathogens.J Biotechnol, 166(3): 114-121. |
| [181] | Sallaud C, Lorieux M, Roumen E, Tharreau D, Berruyer R, Svestasrani P, Garsmeur O, Ghesquiere A, Notteghem J L.2003. Identification of five new blast resistance genes in the highly blast-resistant rice variety IR64 using a QTL mapping strategy.Theor Appl Genet, 106(5): 794-803. |
| [182] | Sato H, Takeuchi Y, Hirabayashi H, Nemoto H, Hirayama M, Kato H, Imbe T, Ando I.2006. Mapping QTLs for field resistance to rice blast in the Japanese upland rice variety Norin 12.Breeding Sci, 56(4): 415-418. |
| [183] | Schaffrath U, Mauch F, Freydl E, Schweizer P, Dudler R.2000. Constitutive expression of the defense-related Rir1b gene in transgenic rice plants confers enhanced resistance to the rice blast fungus Magnaporthe grisea.Plant Mol Biol, 43(1): 59-66. |
| [184] | Selin C, de Kievit T R, Belmonte M F, Fernando E G D.2016. Elucidating the role of effectors in plant-fungal interactions: Progress and challenges.Front Microbiol, 7: 967. |
| [185] | Shan Q W, Zhang Y, Chen K L, Zhang K, Gao C X.2015. Creation of fragrant rice by targeted knockout of the OsBADH2 gene using TALEN technology.Plant Biotechnol J, 13(6): 791-800. |
| [186] | Shang J J, Tao Y, Chen X W, Zou Y, Lei C L, Wang J, Li X B, Zhao X F, Zhang M J, Lu Z K, Xu J C, Cheng Z K, Wan J M, Zhu L H.2009. Identification of a new rice blast resistance gene, Pid3, by genome wide comparison of paired nucleotide-binding site leucine-rich repeat genes and their pseudogene alleles between the two sequenced rice genomes.Genetics, 182(4): 1303-1311. |
| [187] | Sharma T R, Madhav M S, Singh B K, Shanker P, Jana T K, Dalal V, Pandit A, Singh A, Gaikwad K, Upreti H C, Singh N K.2005. High resolution mapping, cloning and molecular characterization of the Pi-kh gene of rice, which confers resistance to M. grisea. Mol Genet Genom, 274(6): 569-578. |
| [188] | Sharma T R, Rai A K, Gupta S K, Singh N K.2010. Broad-spectrum blast resistance gene Pi-kh cloned from rice line Tetep designated Pi54.J Plant Biochem Biotechnol, 191: 87-89. |
| [189] | Sharma T R, Rai A K, Gupta S K, Vijayan J, Devanna B N, Ray S.2012. Rice blast management through host-plant resistance: Retrospect and prospects.Agric Res, 1(1): 37-52. |
| [190] | Shi B, Zhang J, Zheng Y, Liu Y Q, Cruz C M, Zheng T Q, Zhao M F.2012. Identification of a new resistance gene Pi-Da(t) from Dacca6 against rice blast fungus (Magnaporthe oryzae) in Jin23B background.Mol Breeding, 30: 1089-1096. |
| [191] | Shimono M, Sugano S, Nakayama A, Jiang C J, Ono K, Toki S, Takatsuji H.2007. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance.Plant Cell, 19(6): 2064-2076. |
| [192] | Shinoda, Toriyama H, Yunoki K, Ezuka T, Sakurai Y.1971. Studies on the varietal resistance of rice to blast: 6. Linkage relationship of blast resistance genes.Bull Chugoku Agric Exp Stn Ser A, 20: 1-25. (in Japanese with English abstract) |
| [193] | Shu G Y.2009. Induced Plant Mutations in the Genomics Era. Rome: Food and Agriculture Organization of the United Nations: 425-427. |
| [194] | Singh A, Singh V K, Singh S P, Pandian R T P, Ellur R K, Singh D, Bhowmick P K, Gopala Krishnan S, Nagarajan M, Vinod K K, Singh U D, Prabhu K V, Sharma T R, Mohapatra T, Singh A K. 2012. Molecular breeding for the development of multiple disease resistance in Basmati rice.AoB Plants, 2012: pls029. |
| [195] | Singh V K, Singh A, Singh S P, Ellur R K, Choudhary V, Sarkel S, Singh D, Gopala Krishnan S, Nagarajan M, Vinod K K, Singh U D, Rathore R, Prasanthi S K, Agrawal P K, Bhatt J C, Mohapatra T, Prabhu K V, Singh A K.2012. Incorporation of blast resistance gene in elite Basmati rice restorer line PRR78, using marker assisted selection.Field Crops Res, 128: 8-16. |
| [196] | Singh V K, Singh A, Singh S P, Ellur R K, Singh D, Gopala Krishnan S, Bhowmick P K, Nagarajan M, Vinod K K, Singh U D, Rathore R, Prasanthi S K, Agrawal P K, Bhatt J C, Mohapatra T, Prabhu K V, Singh A K.2013. Marker-assisted simultaneous but step wise back cross breeding for pyramiding blast resistance genes Piz5 and Pi54 into an elite Basmati rice restorer line ‘PRR78’.Plant Breeding, 132: 486-495. |
| [197] | Siregar A F, Sipahutar I A, Husnai H, Wibowo H, Sato K, Wakatsuki T, Masunaga T.2016. Influence of water management and silica application on rice growth and productivity in central Java, Indonesia.J Agric Sci, 8: 86-96. |
| [198] | Skamnioti P, Gurr S J.2009. Against the grain, safeguarding rice from rice blast disease.Trends Biotechnol, 27(3): 141-150. |
| [199] | Song F M, Goodman R M.2001. Molecular biology of disease resistance in rice.Physiol Mol Plant Pathol, 59(1): 1-11. |
| [200] | Spence C, Alff E, Johnson C, Ramos C, Donofrio N, Sundaresan V, Bais H.2014. Natural rice rhizospheric microbes suppress rice blast infections.BMC Plant Biol, 14: 130. |
| [201] | Sreewongchai T, Toojinda T, Thanintorn N, Kosawang C, Vanavichit A, Tharreau D, Sirithumya P.2010. Development of elite indica rice lines with wide spectrum of resistance to Thai blast isolates by pyramiding multiple resistance QTLs.Plant Breeding, 129(2): 176-180. |
| [202] | Sun P Y, Liu J L, Wang Y, Jiang N, Wang S H, Dai Y S, Gao J, Li Z Q, Pan S J, Wang D, Li W, Liu X L, Xiao Y H, Liu E M, Wang G L, Dai L Y.2013. Molecular mapping of the blast resistance gene Pi49 in the durably resistant rice cultivar Mowanggu.Euphytica, 192: 45-54. |
| [203] | Sundaram R M, Vishnupriya M R, Laha G S, Rani N S, Rao P S, Balachandran S M, Reddy G A, Sarma N P, Sonti Dr R V.2009. Introduction of bacterial blight resistance into Triguna, a high yielding, mid-early duration rice variety.Biotechnol J, 4(3): 400-407. |
| [204] | Suwannual T, Chankaew S, Monkham T, Saksirirat W, Sanitchon J.2017. Pyramiding of four blast resistance QTLs into Thai rice cultivar RD6 through marker-assisted selection.Czech J Genet Plant Breeding, 53: 1-8. |
| [205] | Tabien R, Li Z, Paterson A, marchetti M A, Stansel J W, Pinson S R M, Park W D.2000. Mapping of four major rice blast resistance genes from ‘Lemont’ and ‘Teqing’ and evaluation of their combinatorial effect for field resistance.Theor Appl Genet, 101: 1215-1225. |
| [206] | Tanksley S D, Grandillo S, Fulton T M, Zamir D, Eshed Y, Petiard V, Lopez J, Beck-Bunn T.1996. Advanced backcross QTL analysis in a cross between an elite processing line of tomato and its wild relative L. pimpinellifolium.Theor Appl Genet, 92(2): 213-224. |
| [207] | Tanweer F A, Rafii M Y, Sijam K, Rahim H A, Ahmed F, Ashkani S, Latif M.2015. Introgression of blast resistance genes (putative Pi-b and Pi-kh) into elite rice cultivar MR219 through marker-assisted selection.Front Plant Sci, 6: 1-11. |
| [208] | Terashima T, Fukuoka S, Saka N, Kudo S.2008. Mapping of a blast field resistance gene Pi39(t) of elite rice strain Chubu 111.Plant Breeding, 127: 485-489. |
| [209] | Thakur S, Gupta Y K, Singh P K, Rathour R, Variar M, Prashanthi S K, Singh A K, Singh U D, Chand D, Rana J C.2013a. Molecular diversity in rice blast resistance gene Pi-ta makes it highly effective against dynamic population of Magnaporthe oryzae.Funct Int Genom, 13(3): 309-322. |
| [210] | Thakur S, Singh P K, Rathour R, Variar M, Prashanthi S K, Singh A K, Singh U D, Chand D, Singh N K, Sharma T R.2013b. Positive selection pressure on rice blast resistance allele Piz-t makes it divergent in Indian landraces.J Plant Int, 8: 34-44. |
| [211] | Tian D G, Chen Z J, Chen Z Q, Zhou Y C, Wang Z H, Wang F, Chen S B.2016. Allele-specific marker-based assessment revealed that the rice blast resistance genes Pi2 and Pi9 have not been widely deployed in Chinese indica rice cultivars.Rice, 9(1): 19. |
| [212] | Toojinda T, Tragoonrung S, Vanavichit A, Siangliw J L, Pa-In N, Jantaboon J, Siangliw M, Fukai S.2005. Molecular breeding for rainfed lowland rice in the Mekong region.Plant Prod Sci, 8(3): 330-333. |
| [213] | Tsuda M H, Takahi Y, Tanizawa K, Kunitomo A, Tsnjino Y, Tanaka H, Miura T, Karto P.1998. Rice blast control with a throw in packed formulation containing pyroquilon.J Pest Sci, 23: 230-234. |
| [214] | Urso S, Orasen G, Perrini R, Tacconi G, Delfanti S, Biselli C, Cattivelli L, Vale G.2013. Pyramiding of Pi resistance genes to increase blast resistance in Italian rice varieties using marker-assisted selection approaches. In: Proceedings of the 57th Italian Society of Agricultural Genetics Annual Congress. 16-19 September.Foggia, Itlay. |
| [215] | Usatov A V, Kostylev P V, Azarin K V, Markin K V, Markareno M S, Khachumov V A, Bibov M Y.2016. Introgression of rice blast resistance gene Pi1, Pi2 and Pi33 into Russian rice varieties by marker assisted selection.Ind J Gent Plant Breeding, 76(1): 18-23. |
| [216] | Venkata Rao G, Muralidharan K.1983. Fungicides and control of leaf blast in dry nursery.Ind Phytopath, 36: 355. |
| [217] | Wang D, Guo C J, Huang J, Yang S H, Tian D C, Zhang X H.2014. Allele mining of rice blast resistance genes at AC134922 locus.Biochem Biophy Res Comm, 446(4): 1085-1090. |
| [218] | Wang F J, Wang C L, Liu P Q, Lei C L, Hao W, Gao Y, Liu Y G, Zhao K J.2016. Enhanced rice blast resistance by CRISPR/Cas9- targeted mutagenesis of the ERF transcription factor gene OsERF922.PLoS One, 11: e0154027. |
| [219] | Wang G L, Mackill D J, Bonman J M, McCouch S R, Champoux M C, Nelson R J.1994. RFLP mapping of genes conferring complete and partial resistance to blast in a durably resistant rice cultivar.Genetics, 136: 1421-1434. |
| [220] | Wang X Y, Lee S H, Wang J C, Ma J B, Bianco T, Jia Y L.2014. Current advances on genetic resistance to rice blast disease. In: Yan W G, Bao J S. Rice: Germplasm, Genetics and Improvement. InTech. DOI: 10.5772/56824. |
| [221] | Wang Z, Jia Y, Rutger J N, Xia Y.2007. Rapid survey for presence of a blast resistance gene Pi-ta in rice cultivars using the dominant DNA markers derived from portions of the Pi-ta gene.Plant Breeding, 126(1): 36-42. |
| [222] | Wang Z Z, Han Q, Zi Q, Lv S, Qiu D W, Zeng H M.2017. Enhanced disease resistance and drought tolerance in transgenic rice plants overexpressing protein elicitors from Magnaporthe oryzae. PLoS One, 12(4): e0175734. |
| [223] | Wen S S, Gao B J.2011. Introgressing blast resistant gene Pi-9(t) into elite rice restorer Luhui 17 by marker assisted selection.Rice Genom Genet, 2: 31-36. |
| [224] | Wongsaprom C, Sirithunya P, Vanavichit A, Pantuwan G, Jongdee B, Sidhiwong N, Lanceras-Siangliw J, Toojinda T.2010. Two introgressed quantitative trait loci confer a broad-spectrum resistance to blast disease in the genetic background of the cultivar RD6, a Thai glutinous jasmine rice.Field Crops Res, 119: 245-251. |
| [225] | Wu J L, Sinha P K, Varivar M, Zheng K L, Leach J E, Courtois B, Leaung H.2004. Association between molecular markers and blast resistance in an advanced backcross population of rice.Theor Appl Genet, 108(6): 1024-1032. |
| [226] | Wu J L, Fan Y Y, Li D B, Zheng K L, Leung H, Zhuang J Y.2005. Genetic control of rice blast resistance in the durably resistant cultivar Gumei 2 against multiple isolates. Theor Appl Genet, 111(1): 50-56. |
| [227] | Wu Y Y, Bao Y M, Xie L J, Su Y Y, Chu R Z, He W W, Huang J, Wang J F, Zhang H S.2013. Fine mapping and identification of blast resistance gene Pi-hk1 in a broad-spectrum resistant japonica rice landrace.Phytopathology, 103: 1162-1168. |
| [228] | Wu Y Y, Yu L, Pan C H, Dai Z Y, Li Y H, Xiao N, Zhang X X, Ji H J, Huang N S, Zhao B H, Zhou C H, Liu G Q, Liu X J, Pan X B, Liang C Z, Li A H.2016. Development of near isogenic lines with different alleles of Piz locus and analysis of their breeding effect under Yangdao 6 background.Mol Breeding, 36: 1-12. |
| [229] | Xi Z Y, He F H, Zeng R Z, Zhang Z M, Ding X H, Li W T, Zhang G Q.2008. Development of a wide population of chromosome single-segment substitution lines in the genetic background of an elite cultivar of rice (Oryza sativa L.). Genome, 49(5): 476-484. |
| [230] | Xiao N, Wu Y Y, Pan C H, Yu L, Chen Y, Liu G Q, Li Y H, Zhang X X, Wang Z P, Dai Z Y, Liang C Z, Li A H.2017. Improving of rice blast resistances in japonica by pyramiding major R genes.Front Plant Sci, 7: 1918. |
| [231] | Xiao W M, Luo L X, Wang H, Guo T, Liu Y Z, Zhou J Y, Zhu X Y, Yang Q Y, Chen Z Q.2016. Pyramiding of Pi46 and Pita to improve blast resistance and to evaluate the resistance effect of the two R genes.J Integr Agric, 15: 2290-2298. |
| [232] | Xu J C, Wang J L, Ling Z Z, Zhu L H.2004. Analysis of rice blast resistance genes by QTL mapping.Chin Sci Bull, 49(4): 337-342. |
| [233] | Xu X, Lv Q M, Shang J J, Pang Z Q, Zhou Z Z, Wang J, Jiang G H, Tao Y, Xu Q, Li X B, Zhao X F, Li S G, Xu J C, Zhu L H.2014. Excavation of Pid3 orthologs with differential resistance spectra to Magnaporthe oryzae in rice resource.PLoS One, 9: e93275. |
| [234] | Yan L, Yan B Y, Peng Y L, Ji Z J, Zeng Y X, Wu H L, Yang C D.2017. Molecular screening of blast resistance genes in rice germplasms resistant to Magnaporthe oryzae.Rice Sci, 24(1): 41-47. |
| [235] | Yang Q Z, Saito K, Yang P W, Wang Q, Sunohara Y, Zheng F P, Ye C R, Li J R, Kato A.2001. Molecular mapping of a new blast resistance gene Pi25(t) possessed in a japonica rice cultivar, Oryza sativa L. cv Yunxi 2. In: Proceedings of the 1st rice blast congress in China. Kunming, China: 49-55. |
| [236] | Yasuda N, Mitsunaga T, Hayashi K, Koizumi S, Fujita Y.2014. Effects of pyramiding quantitative resistance genes Pi21, Pi34, and Pi35 on rice leaf blast disease.Plant Dis, 99(7): 904-909. |
| [237] | Yoshida K, Miyashita N T.2009. DNA polymorphism in the blast disease resistance gene Pita of the wild rice Oryza rufipogon and its related species.Genes Genet Syst, 84(2): 121-136. |
| [238] | Young N D.1996. QTL mapping and quantitative disease resistance in plants.Ann Rev Phytopathol, 34: 479-501. |
| [239] | Yu J M, Holland J B, McMullen M D, Buckler E S.2008. Genetic design and statistical power of nested association mapping in maize.Genetics, 178(1): 539-551. |
| [240] | Yu Z, Mackill D, Bonman J, Tanksley S.1991. Tagging genes for blast resistance in rice via linkage to RFLP markers.Theor Appl Genet, 81: 471-476. |
| [241] | Yunoki T, Ezuka A, Morinaka T, Sakurai Y.1970. Studies on the varietal resistance to rice blast: 4. Variation of field resistance due to fungus strains.Bull Chugoku Agric Exp Stn Ser A, 6: 21-41. (in Japanese with English abstract) |
| [242] | Zenbayashi K, Ashizawa T, Tani T, Koizumi S.2002. Mapping of the QTL (quantitative trait locus) conferring partial resistance to leaf blast in rice cultivar Chubu 32.Theor Appl Genet, 104: 547-552. |
| [243] | Zhang H J, Li G J, Li W, Song F M.2009. Transgenic strategies for improving rice disease resistance.Afr J Biotechnol, 8(9): 1750-1757. |
| [244] | Zhang J, Li X, Jiang G, Xu Y, He Y Q.2006. Pyramiding of Xa7 and Xa21 for the improvement of disease resistance to bacterial blight in hybrid rice.Plant Breeding, 125(6): 600-605. |
| [245] | Zhang M X, Xu J L, Luo R T, Shi D, Li Z K.2003. Genetic analysis and breeding use of blast resistance in a japonica rice mutant R917.Euphytica, 130: 71-76. |
| [246] | Zhang Y X, Yang J Y, Shan Z L, Chen S, Qiao W H, Zhu X Y, Xie Q J, Zhu H T, Zhang Z M, Zeng R Z, Ding X H, Zhang G Q.2012. Substitution mapping of QTLs for blast resistance with SSSLs in rice (Oryza sativa L.).Euphytica, 184(1): 141-150. |
| [247] | Zheng W J, Wang Y, Wang L L, Ma Z B, Zhao J M, Wang P, Zhang L X, Liu Z H, Lu X C.2016. Genetic mapping and molecular marker development of Pi65(t) a novel broad spectrum resistance gene to rice blast using next generation sequencing.Theor Appl Genet, 129(5): 1035-1044. |
| [248] | Zhou B, Qu S, Liu G, Dolan M, Sakai H, Lu G, Belleizi M, Wang G L.2006. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea.Mol Plant Microbe Interact, 19: 1216-1228. |
| [249] | Zhou J H, Wang J L, Xu J C, Lei C L, Ling Z Z.2004. Identification and mapping of a rice blast resistance gene Pi-g(t) in the cultivar Guangchangzhan.Plant Pathol, 53(2): 191-196. |
| [250] | Zhu L H, Chen Y, Xu Y B, Xu J C, Cai H W, Ling Z Z.1993. Construction of a molecular map of rice and gene mapping using a double haploid population of a cross between indica and japonica varieties.Rice Genet Newsl, 10: 132-134. |
| [251] | Zhu M L, Wang L, Pan Q H.2004. Identification and characterization of a new blast resistance gene located on rice chromosome 1 through linkage and differential analyses.Phytopathology, 94: 515-519. |
| [252] | Zhu Y Y, Chen H R, Fan J H, Wang Y Y, Li Y, Chen J B, Fan J X, Yang S S, Hu L P, Leung H, Mew T W, Teng P S, Wang Z H, Mund C C.2000. Genetic diversity and disease control in rice. Nature, 406: 718-722. |
| [253] | Zhuang J Y, Ma W B, Wu J L, Chai R Y, Lu J, Fan Y Y, Jin Z M, Leung H, Zheng K L.2002. Mapping of leaf and neck blast resistance genes with resistance gene analog, RAPD and RFLP in rice.Euphytica, 128: 363-370. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||