
Rice Science ›› 2017, Vol. 24 ›› Issue (6): 360-364.DOI: 10.1016/j.rsci.2017.10.002
• Orginal Article • Previous Articles
Naga Bheema Lingeswara Reddy Inja, Kim Sung-Mi, Kim Beom-Ki, Yoon In-Sun, Kwon Taek-Ryoun( )
)
Online:2017-11-28
Published:2017-08-30
Naga Bheema Lingeswara Reddy Inja, Kim Sung-Mi, Kim Beom-Ki, Yoon In-Sun, Kwon Taek-Ryoun. Identification of Rice Accessions Associated with K+/Na+ Ratio and Salt Tolerance Based on Physiological and Molecular Responses[J]. Rice Science, 2017, 24(6): 360-364.
Add to citation manager EndNote|Ris|BibTeX
| Accession | Origin | Type | Sub-population |
|---|---|---|---|
| IT001158 | India | Introduced cultivar | Indica |
| IT246674 | Bangladesh | Introduced cultivar | Not available |
| IT260533 | India | Not available | Indica |
| IT291341 | Iran | Introduced cultivar | Indica |
| IT168699 | South Korea | Weedy | Japonica |
| IT175218 | South Korea | Weedy | Japonica |
| IT219993 | China | Not available | Not available |
| IT227934 | South Korea | Landrace | Japonica |
| Dongjin | South Korea | Breeding line | Japonica |
Table 1 List of accessions used in this study.
| Accession | Origin | Type | Sub-population |
|---|---|---|---|
| IT001158 | India | Introduced cultivar | Indica |
| IT246674 | Bangladesh | Introduced cultivar | Not available |
| IT260533 | India | Not available | Indica |
| IT291341 | Iran | Introduced cultivar | Indica |
| IT168699 | South Korea | Weedy | Japonica |
| IT175218 | South Korea | Weedy | Japonica |
| IT219993 | China | Not available | Not available |
| IT227934 | South Korea | Landrace | Japonica |
| Dongjin | South Korea | Breeding line | Japonica |
| Name | Forward primer (5′-3′) | Reverse primer (5′-3′) | AT (oC) | Polymorphic | Reference |
|---|---|---|---|---|---|
| RM5 | TGCAACTTCTAGCTGCTCGA | GCATCCGATCTTGATGGG | 56 | Monomorphic | Koyama et al, 2001 |
| RM261 | CTACTTCTCCCCTTGTGTCG | TGTACCATCGCCAAATCTCC | 56 | Monomorphic | Koyama et al, 2001 |
| RM136 | GAGAGCTCAGCTGCTGCCTCTAGC | GAGGAGCGCCACGGTGTACGCC | 64 | Monomorphic | Yao et al, 2005 |
| RM13 | TCCAACATGGCAAGAGAGAG | GGTGGCATTCGATTCCAG | 56 | Monomorphic | Yao et al, 2005 |
| RM345 | ATTGGTAGCTCAATGCAAGC | GTGCAACAACCCCACATG | 55 | Polymorphic | Yao et al, 2005 |
| RM176 | CGGCTCCCGCTACGACGTCTCC | AGCGATGCGCTGGAAGAGGTGC | 64 | Monomorphic | Yao et al, 2005 |
| RM262 | CATTCCGTCTCGGCTCAACT | CAGAGCAAGGTGGCTTGC | 58 | Monomorphic | Yao et al, 2005 |
| RM318 | GTACGGAAAACATGGTAGGAAG | TCGAGGGAAGGATCTGGTC | 55 | Polymorphic | Yao et al, 2005 |
| RM104 | GGAAGAGGAGAGAAAGATGTGTGTCG | TCAACAGACACACCGCCACCGC | 64 | Not reproducible | Yao et al, 2005 |
| RM336 | CTTACAGAGAAACGGCATCG | GCTGGTTTGTTTCAGGTTCG | 55 | Not reproducible | Bhowmik et al, 2009 |
| RM7075 | TATGGACTGGAGCAAACCTC | GGCACAGCACCAATGTCTC | 55 | Polymorphic | Bhowmik et al, 2009 |
| RM253 | TCCTTCAAGAGTGCAAAACC | GCATTGTCATGTCGAAGCC | 55 | Polymorphic | Bhowmik et al, 2009 |
| RM8053 | AGACATTGCCGATGATAGG | AAGTACCCCACCGAATAGAG | 55 | Polymorphic | Senguttuvel et al, 2010 |
| RM9 | GGTGCCATTGTCGTCCTC | ACGGCCCTCATCACCTTC | 58 | Not reproducible | Senguttuvel et al, 2010 |
| RM212 | CCACTTTCAGCTACTACCAG | CACCCATTTGTCTCTCATTATG | 55 | Monomorphic | Senguttuvel et al, 2010 |
| RM23 | CATTGGAGTGGAGGCTGG | GTCAGGCTTCTGCCATTCTC | 55 | Monomorphic | Senguttuvel et al, 2010 |
| RM493 | TAGCTCCAACAGGATCGACC | GTACGTAAACGCGGAAGGTG | 58 | Monomorphic | Senguttuvel et al, 2010 |
Table 2 Description of simple sequence repeat (SSR) markers used in this study.
| Name | Forward primer (5′-3′) | Reverse primer (5′-3′) | AT (oC) | Polymorphic | Reference |
|---|---|---|---|---|---|
| RM5 | TGCAACTTCTAGCTGCTCGA | GCATCCGATCTTGATGGG | 56 | Monomorphic | Koyama et al, 2001 |
| RM261 | CTACTTCTCCCCTTGTGTCG | TGTACCATCGCCAAATCTCC | 56 | Monomorphic | Koyama et al, 2001 |
| RM136 | GAGAGCTCAGCTGCTGCCTCTAGC | GAGGAGCGCCACGGTGTACGCC | 64 | Monomorphic | Yao et al, 2005 |
| RM13 | TCCAACATGGCAAGAGAGAG | GGTGGCATTCGATTCCAG | 56 | Monomorphic | Yao et al, 2005 |
| RM345 | ATTGGTAGCTCAATGCAAGC | GTGCAACAACCCCACATG | 55 | Polymorphic | Yao et al, 2005 |
| RM176 | CGGCTCCCGCTACGACGTCTCC | AGCGATGCGCTGGAAGAGGTGC | 64 | Monomorphic | Yao et al, 2005 |
| RM262 | CATTCCGTCTCGGCTCAACT | CAGAGCAAGGTGGCTTGC | 58 | Monomorphic | Yao et al, 2005 |
| RM318 | GTACGGAAAACATGGTAGGAAG | TCGAGGGAAGGATCTGGTC | 55 | Polymorphic | Yao et al, 2005 |
| RM104 | GGAAGAGGAGAGAAAGATGTGTGTCG | TCAACAGACACACCGCCACCGC | 64 | Not reproducible | Yao et al, 2005 |
| RM336 | CTTACAGAGAAACGGCATCG | GCTGGTTTGTTTCAGGTTCG | 55 | Not reproducible | Bhowmik et al, 2009 |
| RM7075 | TATGGACTGGAGCAAACCTC | GGCACAGCACCAATGTCTC | 55 | Polymorphic | Bhowmik et al, 2009 |
| RM253 | TCCTTCAAGAGTGCAAAACC | GCATTGTCATGTCGAAGCC | 55 | Polymorphic | Bhowmik et al, 2009 |
| RM8053 | AGACATTGCCGATGATAGG | AAGTACCCCACCGAATAGAG | 55 | Polymorphic | Senguttuvel et al, 2010 |
| RM9 | GGTGCCATTGTCGTCCTC | ACGGCCCTCATCACCTTC | 58 | Not reproducible | Senguttuvel et al, 2010 |
| RM212 | CCACTTTCAGCTACTACCAG | CACCCATTTGTCTCTCATTATG | 55 | Monomorphic | Senguttuvel et al, 2010 |
| RM23 | CATTGGAGTGGAGGCTGG | GTCAGGCTTCTGCCATTCTC | 55 | Monomorphic | Senguttuvel et al, 2010 |
| RM493 | TAGCTCCAACAGGATCGACC | GTACGTAAACGCGGAAGGTG | 58 | Monomorphic | Senguttuvel et al, 2010 |
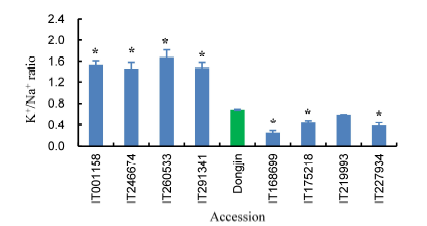
Fig. 1. K+/Na+ ratio in response to 50 mmol/L NaCl.* means significant difference at the 0.05 level in comparison to Dongjin. Data are means ± SE (n = 3).
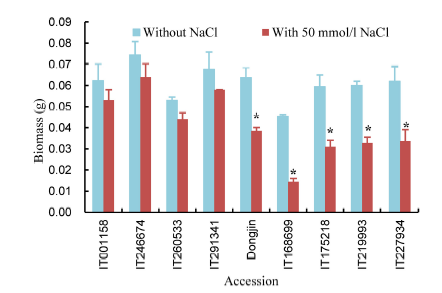
Fig. 2. Biomass changes in response to 50 mmol/L NaCl.* means significant difference at the 0.05 level between the two treatment. Data are means ± SE (n = 3).
| Variable | K+/Na+ ratio | Na+ concentration | K+ concentration |
|---|---|---|---|
| Biomass | 0.805* | -0.245 | 0.815* |
| K+/Na+ ratio | -0.552 | 0.758* | |
| Na+ concentration | -0.068 |
Table 3 Pearson correlation matrix among traits.
| Variable | K+/Na+ ratio | Na+ concentration | K+ concentration |
|---|---|---|---|
| Biomass | 0.805* | -0.245 | 0.815* |
| K+/Na+ ratio | -0.552 | 0.758* | |
| Na+ concentration | -0.068 |
| [1] | Bhowmik S K, Titov S, Islam M M, Siddika A, Sultana S, Haque M D S.2009. Phenotypic and genotypic screening of rice genotypes at seedling stage for salt tolerance.Afric J Biotechnol, 8(23): 6490-6494. |
| [2] | Blumwald E.2000. Sodium transport and salt tolerance in plants.Curr Opin Cell Biol, 12(4): 431-434. |
| [3] | Bonilla P, Dvorak J, Mackill D, Deal K, Gregorio G.2002. RFLP and SSL mapping of salinity tolerance genes in chromosome 1 of rice (Oryza sativa L.) using recombinant inbred lines.Phil J Agric Sci, 85: 68-76. |
| [4] | Flowers T J.2004. Improving crop salt tolerance.J Exp Bot, 55: 307-319. |
| [5] | Gill K S, Singh O S.1995. Studies on the growth, ionic composition and yield characters associated with salt tolerance in rice.J Potash Res, 11: 166-175. |
| [6] | Gregorio G B, Senadhira D.1993. Genetic analysis of salinity tolerance in rice (Oryza sativa L.).Theor Appl Genet, 86: 333-338. |
| [7] | Gregorio G B.1997. Tagging Salinity Tolerance Genes in Rice Using Amplified Fragment Length Polymorphism (AFLP). Los Baños, the Philippines: University of the Philippines: 118. |
| [8] | Gregorio G B, Senadhira D, Mendoza R D.1997. Screening Rice for Salinity Tolerance. Los Baños, the Philippines: International Rice Research Institute: 1-30. |
| [9] | Gregorio G B, Senadhira D, Mendoza R D, Manigbas N L, Roxas J P, Guerta C Q.2002. Progress in breeding for salinity tolerance and associated abiotic stresses in rice.Field Crops Res, 76(2): 91-101. |
| [10] | IRRI Rice Knowledge Bank. Knowledgebank.irri.org. Retrieved on February 20, 2017. |
| [11] | Kanawapee N, Sanitchon J, Lontom W, Threerakulpisut P.2012. Evaluation of salt tolerance at the seedling stage in rice genotypes by growth performance, ion accumulation, proline and chlorophyll content.Plant Soil, 358: 235-249. |
| [12] | Khush G S.2005. What it will take to feed 5.0 billion rice consumers in 2030.Plant Mol Biol, 59: 1-6. |
| [13] | Koyama M L, Levesley A, Koebner R M D, Flowers T J, Yeo A R.2001. Quantitative trait loci for component physiological traits determining salt tolerance in rice.Plant Physiol, 125: 406-422. |
| [14] | Moradi F, Ismail A M.2007. Responses of photosynthesis, chlorophyll fluorescence and ROS scavenging systems to salt stress during seedling and reproductive stages in rice.Ann Bot, 9(6): 1161-1173. |
| [15] | Munns R, James R A.2003. Screening methods for salinity tolerance: A case study with tetraploid wheat.Plant Soil, 253(1): 201-218. |
| [16] | Munns R, Tester M.2008. Mechanisms of salinity tolerance.Annu Rev Plant Biol, 59: 651-681. |
| [17] | Oliveira E J, Pádua J G, Zucchi M I, Vencovsky R, Viera M L C.2006. Origin, evolution and genome distribution of microsatellites.Genet Mol Biol, 29(2): 294-307. |
| [18] | Pandey U K, Srivastava R D L.1991. Leaf potassium as an index of salt tolerance in paddy.Natl Acad Sci Lett, 14(4): 161-164. |
| [19] | Pires I S, Negrão S, Oliveira M M, Purugganan M D.2015. Comprehensive phenotypic analysis of rice (Oryza sativa) response to salinity stress.Physiol Plant, 155(1): 43-54. |
| [20] | Pushparajan N, Krishnasamy V, Babu R C, Kannanbabu J R.2011. Association mapping of salinity tolerance in rice using molecular markers.Int J Biores Stress Manag, 2(3): 307-312. |
| [21] | Reddy M A, Francies R M, Rasool S N, Reddy V R P.2014. Breeding for tolerance to stress triggered by salinity in rice.Int J Appl Biol Pharm Technol, 5(1): 166-176. |
| [22] | Sabouri H, Biabani A.2009. Toward the mapping of agronomic characters on a rice genetic map: Quantitative trait loci analysis under saline condition.Biotechnology, 8(1): 144-149. |
| [23] | Sakina A, Ahmed I, Shahzad A, Iqbal M, Asif M.2016. Genetic variation for salinity tolerance in Pakistani rice (Oryza sativa L.) germplasm.J Agron Crop Sci, 202(1): 25-36. |
| [24] | Saqib M, Akhtar J, Qureshi R H, Aslam M, Nawaz S.1999. Effect of salinity and hypoxia on growth and ionic composition of different genotypes of wheat.Pak J Soil Sci, 17: 1-8. |
| [25] | Senguttuvel P, Raveendran M, Vijayalakshmi C, Thiyagarajan K, Bapu J R K, Viraktamath B C.2010. Molecular mechanism of salt tolerance for genetic diversity analysed in association with Na+/K+ ratio through SSR markers in rice (Oryza sativa L.).Int J Agric Res, 5(9): 708-719. |
| [26] | Shabala S, Cuin T A.2008. Potassium transport and plant salt tolerance.Physiol Planta, 133(4): 651-669. |
| [27] | Soubir T, Salil K B, Mirza M I, Ayesha S, Sharmin S, Shahidul L.2009. Phenotypic and genotypic screening of rice genotypes at seedling stage for salt tolerance.UDO Agric Magaz, 9(4): 770-775. |
| [28] | Thomson M J, Ismail A M, McCouch S R, Mackill M J.2010. Marker assisted breeding. In: Pareek A, Sopory S K, Bohnert H J, Govindjee. Abiotic Stress Adaptation in Plants, Physiological, Molecular and Genomic Foundation. New York: Springer: 451-469. |
| [29] | Waziri A, Kumar P, Purty R S.2016. Saltol QTL and their role in salinity tolerance in rice.Aust J Biotechnol Bioeng, 3(3): 1067-1072. |
| [30] | Yao M Z, Wang J F, Chen H Y, Zhai H Q, Zhang H S.2005. Inheritance and QTL mapping of salt tolerance in rice.Rice Sci, 12(1): 25-32. |
| [31] | Yoshida S, Forno D A, Cock J H, Gomez K A.1976. Laboratory Manual for Physiological Studies of Rice. 3rd edn. Los Baños, the Philippines: International Rice Research Institute: 83. |
| [32] | Zeng L H, Shannon M C, Lesch S M.2001. Timing of salinity stress affects rice growth and yield components.Agric Water Manag, 48(3): 191-206. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||