
Rice Science ›› 2019, Vol. 26 ›› Issue (3): 189-194.DOI: 10.1016/j.rsci.2018.08.007
• Short Communications • Previous Articles Next Articles
Puttaswamaiah Ranjitha Hurugalavadi1( ), Gowda Rame1, Nethra Nagarajappa1, Amruta Narayanappa1, Kumar Kandikattu Hemanth2
), Gowda Rame1, Nethra Nagarajappa1, Amruta Narayanappa1, Kumar Kandikattu Hemanth2
Received:2018-03-08
Accepted:2018-08-28
Online:2019-05-28
Published:2019-01-25
Puttaswamaiah Ranjitha Hurugalavadi, Gowda Rame, Nethra Nagarajappa, Amruta Narayanappa, Kumar Kandikattu Hemanth. Biochemical and Metabolomics on Rice Cultivars[J]. Rice Science, 2019, 26(3): 189-194.
Add to citation manager EndNote|Ris|BibTeX
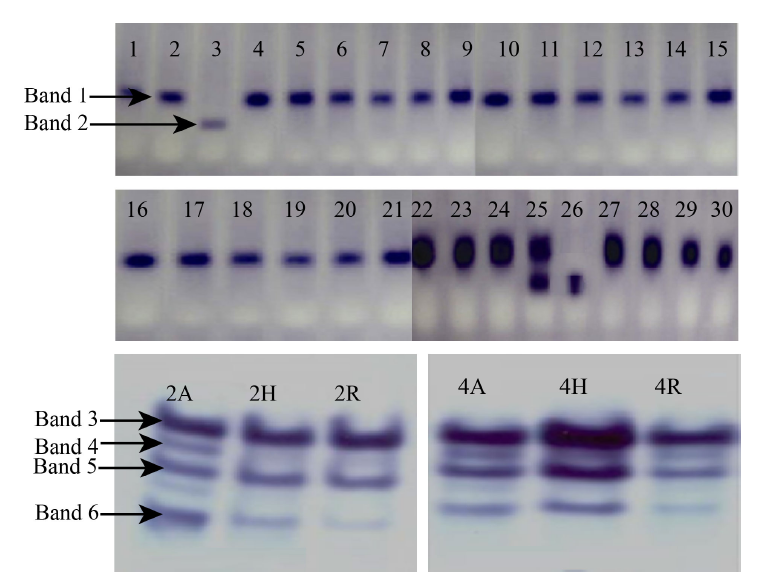
Fig. 1. Alcohol dehydrogenase activity of rice cultivars. 1, Jaya; 2, Mandya Sona 1; 3, CTH1; 4, BI33; 5, Ratna Choodi; 6, KMP200; 7, MAS946-1; 8, Rajamudi; 9, JGL1798; 10, KMO201; 11, Mandya Sona 2; 12, MAS26; 13, BR2655; 14, BPT5204; 15, KMP175; 16, Mandya Sona 3; 17, Thanu; 18, KMP128; 19, MTU1001; 20, Gangavathi Sona; 21, Rasi; 22, Raksha; 23, Tella Hamsa; 24, MTU1001; 25, IR30864; 26, IR64; 27, Jyothi; 28, CTH3; 29, KMP153; 30, Tunga; 2A, IR58025; 2H, KRH-2; 2R, KMR-3; 4A, CRMS32A; 4H, KRH-4; 4R, MSN36.

Supplemental Fig. 1 Malate dehydrogenase (A), catalase (B), peroxidase (C), esterase (D), and phenylalanine ammonia lyase (E) activities of thirty rice cultivars.
| CN | Compound Name | Quantification peak | Confirming peak | ||||||
|---|---|---|---|---|---|---|---|---|---|
| RT(min) | Precursor | Product | CE | Precursor | Product | CE | |||
| 1 | l-Alanine, N-(trimethylsilyl)-, trimethylsilyl ester | 6.37 | 99 | 69 | 30 | 99 | 71 | 10 | |
| 2 | Glycine, N-(trimethylsilyl)-, trimethylsilyl ester | 5.69 | 100.1 | 59.1 | 10 | 174.1 | 59.1 | 20 | |
| 3 | Phosphoric acid, bis(trimethylsilyl)monomethyl ester | 6.92 | 78.1 | 64 | 12 | 78.1 | 71.3 | 12 | |
| 4 | L-Valine, N-(trimethylsilyl)-, trimethylsilyl ester | 7.44 | 148.8 | 450 | 30 | 148.8 | 75 | 10 | |
| 5 | 2-(4-Methoxyphenyl)-2-(4-trimethoxysilyloxy)propane | 8.73 | 11.4 | 45.6 | 18 | 118.4 | 75.2 | 8 | |
| 6 | Butane, 2,3-bis(trimethylsiloxy)- | 10.06 | 116.1 | 43 | 28 | 116.1 | 45.1 | 18 | |
| 7 | 1,2-Bis(trimethylsiloxy)ethane | 10.48 | 102.1 | 45.1 | 20 | 102.1 | 58 | 30 | |
| 8 | Silanamine, N,N'-methanetetraylbis[1,1,1-trimethyl- | 11.55 | 133 | 115 | 10 | 163.1 | 133.1 | 10 | |
| 9 | Ethylbis(trimethylsilyl)amine | 12.32 | 144.1 | 43 | 32 | 144.1 | 58.1 | 32 | |
| 10 | N,N'-Bis(trimethylsilyl)trifluoroacetamidine | 13.36 | 299.1 | 151.1 | 10 | 300.5 | 74.1 | 10 | |
| 11 | Glycerol, tris(trimethylsilyl) ether | 13.43 | 116.9 | 59.1 | 10 | 116.9 | 89.1 | 10 | |
| 12 | Butanedioic acid, bis(trimethylsilyl) ester | 14.14 | 74.8 | 47 | 10 | 148.5 | 45.2 | 30 | |
| 13 | N,O,O-Tris(trimethylsilyl)-L-threonine | 14.94 | 205.5 | 205.3 | 10 | 218.2 | 45 | 30 | |
| 14 | Serine, N,O-bis(trimethylsilyl)-, trimethylsilyl ester | 15.37 | 100.6 | 86.1 | 10 | 117.1 | 43 | 30 | |
| 15 | Threitol, 1,2,3,4-tetrakis-O-(trimethylsilyl)-, D- | 16.57 | 229.1 | 47 | 32 | 229.1 | 131.1 | 12 | |
| 16 | Amine, N,N,N-tris((trimethylsilyloxy)ethyl)- | 17.04 | 189.7 | 147.1 | 10 | 233.1 | 73.1 | 10 | |
| 17 | Decanoic acid, trimethylsilyl ester | 17.23 | 100 | 59.1 | 10 | 115 | 59.1 | 20 | |
| 18 | Butanedioic acid, [(trimethylsilyl)oxy]-, bis (trimethylsilyl) ester | 17.35 | 205.3 | 117.1 | 8 | 205.3 | 147.1 | 8 | |
| 19 | 3-Trifluoromethylbenzylamine, N,N-dinonyl | 17.53 | 232.1 | 100.1 | 8 | 232.1 | 188.2 | 8 | |
| 20 | Arabinofuranose, 1,2,3,5-tetrakis-O-(trimethylsilyl)- | 17.73 | 171.1 | 73.1 | 22 | 314.1 | 73.1 | 22 | |
| 21 | Myo-Inositol, 1,2,3,4,5,6-hexakis-O-(trimethylsilyl)- | 19.12 | 128.1 | 73.1 | 10 | 246.1 | 73.1 | 20 | |
| 22 | Oleic acid, trimethylsilyl ester | 19.24 | 232.2 | 73.1 | 22 | 262.3 | 82.1 | 12 | |
| 23 | Myristic acid, 2,3-bis(trimethylsiloxy)propyl ester | 19.94 | 231.1 | 116.1 | 8 | 231.1 | 132.2 | 8 | |
| 24 | D-(+)-Trehalose, octakis(trimethylsilyl) ether | 20.84 | 204.8 | 117.1 | 10 | 204.8 | 147.2 | 10 | |
| 25 | D-Turanose, heptakis(trimethylsilyl)- Isomer 1 | 21.59 | 299.2 | 147.2 | 20 | 299.2 | 225.2 | 10 | |
| 26 | α-D-Galactopyranoside, methyl 2,3,4,6-tetrakis-O-(trimethylsilyl)- | 21.81 | 155.1 | 45 | 28 | 156 | 45.1 | 28 | |
| 27 | 1,2-Propanediol-1-phosphate, tris(trimethylsilyl)- isomer 1 | 22.41 | 437.4 | 191.2 | 10 | 437.4 | 257.2 | 10 | |
| 28 | 1,2-Propanediol-1-phosphate, tris(trimethylsilyl)- isomer 2 | 22.69 | 273.1 | 67.1 | 10 | 273.1 | 183.2 | 10 | |
| 29 | Silane, [[(3β,24R)-ergost-5-en-3-yl]oxy]trimethyl- | 23.3 | 285.2 | 95.2 | 12 | 285.2 | 131.1 | 12 | |
| 30 | β-Sitosterol trimethylsilyl ether | 22.86 | 518.5 | 188.3 | 12 | 518.5 | 428.3 | 12 | |
| 31 | D-Glucose, 4-O-[2,3,4,6-tetrakis-O-(trimethylsilyl) -β-D-galactopyranosyl]-2,3,5,6-tetrakis-O-(trimethylsilyl)- | 24.34 | 191.1 | 45.1 | 30 | 191.1 | 147.2 | 10 | |
| 32 | 2-O-Glycerol-α-d-galactopyranoside, hexa-TMS isomer 1 | 25.02 | 218.1 | 73.1 | 8 | 218.1 | 100.1 | 8 | |
| 33 | D-Turanose, heptakis(trimethylsilyl)- isomer 3 | 27.31 | 73.1 | 45.1 | 12 | 73.1 | 58 | 22 | |
| 34 | 9,19-Cyclolanostan-3-ol, 24-methylene-, (3β)- | 28.15 | 305.2 | 147.2 | 8 | 305.2 | 217.2 | 8 | |
| 35 | 2-Methyl-2(p-methoxy)mandelate, bis(trimethylsilyl)- | 30.27 | 202.1 | 45.1 | 32 | 202.1 | 58 | 32 | |
| 36 | α-D-Galactopyranoside, methyl 2,3,4,6-tetrakis-O-(trimethylsilyl)Isomer1 | 30.7 | 81 | 79.1 | 10 | 337.2 | 131.1 | 10 | |
| 37 | β-D-Xylopyranose,1,2,3,4-tetrakis-O-(trimethylsilyl)- | 30.82 | 339.5 | 75.1 | 20 | 339.5 | 131.1 | 10 | |
| 38 | β-D-Xylopyranose, 1,2,3,4-tetrakis-O-(trimethylsilyl) | 32.65 | 146.9 | 45 | 30 | 146.9 | 131.2 | 10 | |
| 39 | D-Turanose, heptakis(trimethylsilyl)- Isomer 2 | 32.78 | 85 | 43.9 | 12 | 98.1 | 55.1 | 12 | |
| 40 | 1,2-Propanediol-1-phosphate, tris(trimethylsilyl)- | 34.4 | 211.2 | 71.1 | 10 | 343.5 | 95.1 | 10 | |
| 41 | β-D-Xylopyranose, 1,2,3,4-tetrakis-O-(trimethylsilyl)- | 34.64 | 318.4 | 147.2 | 20 | 318.4 | 215.2 | 10 | |
| 42 | 9-Octadecenoic acid, 2-[(trimethylsilyl)oxy]-1-[[(trimethylsilyl)oxy]methyl]ethyl ester | 34.73 | 225.1 | 79.1 | 10 | 225.1 | 93.1 | 10 | |
| 43 | Sucrose, octakis(trimethylsilyl) ether | 35.27 | 117 | 47.1 | 20 | 117 | 75.1 | 10 | |
| 44 | Sucrose, octakis(trimethylsilyl) ether | 38 | 147.1 | 45.1 | 32 | 147.1 | 131.1 | 12 | |
| 45 | β-D-Galactopyranoside, methyl 2,3-bis-O-(trimethylsilyl)-, cyclic methylboronate | 39.09 | 352.7 | 169.2 | 10 | 362.7 | 170.2 | 10 | |
| 46 | Myo-Inositol, pentakis-O-(trimethylsilyl)-, bis(trimethylsilyl) phosphate | 40.81 | 361.4 | 169.1 | 10 | 361.4 | 243.3 | 10 | |
| 47 | 2-Pyrrolidinone, 1-(9-octadecenyl)- | 41 | 103 | 45.1 | 20 | 103 | 58.1 | 30 | |
| 48 | α-L-Galactopyranose, 6-deoxy-1,2,3,4-tetrakis-O-imethylsilyl)- | 41.54 | 399.3 | 81.1 | 10 | 399.3 | 95.5 | 10 | |
| 49 | 9,12-Octadecadienoic acid (Z,Z)-, trimethylsilyl ester | 42.68 | 298.9 | 147.1 | 20 | 298.9 | 225.2 | 10 | |
| 50 | Tryptophan, bis(trimethylsilyl)- | 42.81 | 373.4 | 167.1 | 12 | 373.4 | 211.2 | 12 | |
| 51 | Hexadecanoic acid, trimethylsilyl ester | 43 | 361.1 | 169.2 | 10 | 361.1 | 243.3 | 10 | |
| 52 | N,O,O'-Tris-(trimethylsilyl)tyrosine | 43.67 | 147.2 | 45.1 | 32 | 147.2 | 131.1 | 12 | |
| 53 | Mannose, 2,3,4,5,6-pentakis-O-(trimethylsilyl)-, D- | 44.71 | 204 | 73.1 | 10 | 204 | 189.2 | 10 | |
| 54 | β-Hydroxypyruvic acid, trimethylsilyl ether, trimethylsilyl ester | 44.81 | 147.3 | 131.1 | 10 | 129.3 | 45 | 20 | |
| 55 | Tetradecanoic acid, trimethylsilyl ester | 45.01 | 488.7 | 222.8 | 20 | 488.7 | 223.5 | 10 | |
| 56 | 1,2,3-Propanetricarboxylic acid, 2-[(trimethylsilyl)oxy]-, tris(trimethylsilyl) ester | 45.85 | 357.1 | 225.1 | 18 | 357.1 | 341.2 | 8 | |
| 57 | l-Glutamine, tris(trimethylsilyl) deriv. | 46.57 | 205.2 | 45.3 | 30 | 205.2 | 190.1 | 10 | |
| 58 | Phosphoric acid, bis(trimethylsilyl) 2,3-bis[(trimethylsilyl)oxy]propyl ester | 46.81 | 222 | 45.1 | 32 | 222 | 194.1 | 12 | |
| 59 | Arabinitol, pentakis-O-(trimethylsilyl)- | 48.44 | 211 | 115.1 | 30 | 211 | 133.1 | 10 | |
| 60 | L-Asparagine, N,N2-bis(trimethylsilyl)-, trimethylsilyl ester | 48.84 | 343.2 | 95.2 | 20 | 434.2 | 121.1 | 10 | |
| 61 | Glutamic acid, N-(trimethylsilyl)-, bis(trimethylsilyl) ester, L- | 50.16 | 357.3 | 95.1 | 20 | 357.3 | 107.1 | 20 | |
| 62 | L-Aspartic acid, N-(trimethylsilyl)-, bis(trimethylsilyl) ester | 51.24 | 362.5 | 169.2 | 12 | 362.5 | 170.2 | 12 | |
| 63 | Acetic acid, iodo-, trimethylsilyl ester | 52.13 | 147 | 105.1 | 10 | 147 | 119.1 | 10 | |
| 64 | α-D-Glucopyranoside, 1,3,4,6-tetrakis-O-(trimethylsilyl)-β-D-fructofuranosyl 2,3,4,6-tetrakis-O-(trimethylsilyl)- | 53.49 | 217.2 | 45.1 | 28 | 361.2 | 169.2 | 8 | |
| 65 | D-Glucose, 4-O-[2,3,4,6-tetrakis-O-(trimethylsilyl)-β-D-galactopyranosyl]-2,3,5,6-tetrakis-O-(trimethylsilyl)- | 54.95 | 217.1 | 45.1 | 32 | 217.1 | 143.1 | 12 | |
| 66 | α-D-Glucopyranoside, 1,3,4,6-tetrakis-O-(trimethylsilyl)-β-D-fructofuranosyl 2,3,4,6-tetrakis-O-(trimethylsilyl)- | 59.21 | 204.1 | 45.1 | 30 | 204.1 | 189.2 | 10 | |
Supplemental Table 1 Metabolites obtained from GC/MS SRM transitions method for 12 diverse rice cultivars selected from Phase-I discovery.
| CN | Compound Name | Quantification peak | Confirming peak | ||||||
|---|---|---|---|---|---|---|---|---|---|
| RT(min) | Precursor | Product | CE | Precursor | Product | CE | |||
| 1 | l-Alanine, N-(trimethylsilyl)-, trimethylsilyl ester | 6.37 | 99 | 69 | 30 | 99 | 71 | 10 | |
| 2 | Glycine, N-(trimethylsilyl)-, trimethylsilyl ester | 5.69 | 100.1 | 59.1 | 10 | 174.1 | 59.1 | 20 | |
| 3 | Phosphoric acid, bis(trimethylsilyl)monomethyl ester | 6.92 | 78.1 | 64 | 12 | 78.1 | 71.3 | 12 | |
| 4 | L-Valine, N-(trimethylsilyl)-, trimethylsilyl ester | 7.44 | 148.8 | 450 | 30 | 148.8 | 75 | 10 | |
| 5 | 2-(4-Methoxyphenyl)-2-(4-trimethoxysilyloxy)propane | 8.73 | 11.4 | 45.6 | 18 | 118.4 | 75.2 | 8 | |
| 6 | Butane, 2,3-bis(trimethylsiloxy)- | 10.06 | 116.1 | 43 | 28 | 116.1 | 45.1 | 18 | |
| 7 | 1,2-Bis(trimethylsiloxy)ethane | 10.48 | 102.1 | 45.1 | 20 | 102.1 | 58 | 30 | |
| 8 | Silanamine, N,N'-methanetetraylbis[1,1,1-trimethyl- | 11.55 | 133 | 115 | 10 | 163.1 | 133.1 | 10 | |
| 9 | Ethylbis(trimethylsilyl)amine | 12.32 | 144.1 | 43 | 32 | 144.1 | 58.1 | 32 | |
| 10 | N,N'-Bis(trimethylsilyl)trifluoroacetamidine | 13.36 | 299.1 | 151.1 | 10 | 300.5 | 74.1 | 10 | |
| 11 | Glycerol, tris(trimethylsilyl) ether | 13.43 | 116.9 | 59.1 | 10 | 116.9 | 89.1 | 10 | |
| 12 | Butanedioic acid, bis(trimethylsilyl) ester | 14.14 | 74.8 | 47 | 10 | 148.5 | 45.2 | 30 | |
| 13 | N,O,O-Tris(trimethylsilyl)-L-threonine | 14.94 | 205.5 | 205.3 | 10 | 218.2 | 45 | 30 | |
| 14 | Serine, N,O-bis(trimethylsilyl)-, trimethylsilyl ester | 15.37 | 100.6 | 86.1 | 10 | 117.1 | 43 | 30 | |
| 15 | Threitol, 1,2,3,4-tetrakis-O-(trimethylsilyl)-, D- | 16.57 | 229.1 | 47 | 32 | 229.1 | 131.1 | 12 | |
| 16 | Amine, N,N,N-tris((trimethylsilyloxy)ethyl)- | 17.04 | 189.7 | 147.1 | 10 | 233.1 | 73.1 | 10 | |
| 17 | Decanoic acid, trimethylsilyl ester | 17.23 | 100 | 59.1 | 10 | 115 | 59.1 | 20 | |
| 18 | Butanedioic acid, [(trimethylsilyl)oxy]-, bis (trimethylsilyl) ester | 17.35 | 205.3 | 117.1 | 8 | 205.3 | 147.1 | 8 | |
| 19 | 3-Trifluoromethylbenzylamine, N,N-dinonyl | 17.53 | 232.1 | 100.1 | 8 | 232.1 | 188.2 | 8 | |
| 20 | Arabinofuranose, 1,2,3,5-tetrakis-O-(trimethylsilyl)- | 17.73 | 171.1 | 73.1 | 22 | 314.1 | 73.1 | 22 | |
| 21 | Myo-Inositol, 1,2,3,4,5,6-hexakis-O-(trimethylsilyl)- | 19.12 | 128.1 | 73.1 | 10 | 246.1 | 73.1 | 20 | |
| 22 | Oleic acid, trimethylsilyl ester | 19.24 | 232.2 | 73.1 | 22 | 262.3 | 82.1 | 12 | |
| 23 | Myristic acid, 2,3-bis(trimethylsiloxy)propyl ester | 19.94 | 231.1 | 116.1 | 8 | 231.1 | 132.2 | 8 | |
| 24 | D-(+)-Trehalose, octakis(trimethylsilyl) ether | 20.84 | 204.8 | 117.1 | 10 | 204.8 | 147.2 | 10 | |
| 25 | D-Turanose, heptakis(trimethylsilyl)- Isomer 1 | 21.59 | 299.2 | 147.2 | 20 | 299.2 | 225.2 | 10 | |
| 26 | α-D-Galactopyranoside, methyl 2,3,4,6-tetrakis-O-(trimethylsilyl)- | 21.81 | 155.1 | 45 | 28 | 156 | 45.1 | 28 | |
| 27 | 1,2-Propanediol-1-phosphate, tris(trimethylsilyl)- isomer 1 | 22.41 | 437.4 | 191.2 | 10 | 437.4 | 257.2 | 10 | |
| 28 | 1,2-Propanediol-1-phosphate, tris(trimethylsilyl)- isomer 2 | 22.69 | 273.1 | 67.1 | 10 | 273.1 | 183.2 | 10 | |
| 29 | Silane, [[(3β,24R)-ergost-5-en-3-yl]oxy]trimethyl- | 23.3 | 285.2 | 95.2 | 12 | 285.2 | 131.1 | 12 | |
| 30 | β-Sitosterol trimethylsilyl ether | 22.86 | 518.5 | 188.3 | 12 | 518.5 | 428.3 | 12 | |
| 31 | D-Glucose, 4-O-[2,3,4,6-tetrakis-O-(trimethylsilyl) -β-D-galactopyranosyl]-2,3,5,6-tetrakis-O-(trimethylsilyl)- | 24.34 | 191.1 | 45.1 | 30 | 191.1 | 147.2 | 10 | |
| 32 | 2-O-Glycerol-α-d-galactopyranoside, hexa-TMS isomer 1 | 25.02 | 218.1 | 73.1 | 8 | 218.1 | 100.1 | 8 | |
| 33 | D-Turanose, heptakis(trimethylsilyl)- isomer 3 | 27.31 | 73.1 | 45.1 | 12 | 73.1 | 58 | 22 | |
| 34 | 9,19-Cyclolanostan-3-ol, 24-methylene-, (3β)- | 28.15 | 305.2 | 147.2 | 8 | 305.2 | 217.2 | 8 | |
| 35 | 2-Methyl-2(p-methoxy)mandelate, bis(trimethylsilyl)- | 30.27 | 202.1 | 45.1 | 32 | 202.1 | 58 | 32 | |
| 36 | α-D-Galactopyranoside, methyl 2,3,4,6-tetrakis-O-(trimethylsilyl)Isomer1 | 30.7 | 81 | 79.1 | 10 | 337.2 | 131.1 | 10 | |
| 37 | β-D-Xylopyranose,1,2,3,4-tetrakis-O-(trimethylsilyl)- | 30.82 | 339.5 | 75.1 | 20 | 339.5 | 131.1 | 10 | |
| 38 | β-D-Xylopyranose, 1,2,3,4-tetrakis-O-(trimethylsilyl) | 32.65 | 146.9 | 45 | 30 | 146.9 | 131.2 | 10 | |
| 39 | D-Turanose, heptakis(trimethylsilyl)- Isomer 2 | 32.78 | 85 | 43.9 | 12 | 98.1 | 55.1 | 12 | |
| 40 | 1,2-Propanediol-1-phosphate, tris(trimethylsilyl)- | 34.4 | 211.2 | 71.1 | 10 | 343.5 | 95.1 | 10 | |
| 41 | β-D-Xylopyranose, 1,2,3,4-tetrakis-O-(trimethylsilyl)- | 34.64 | 318.4 | 147.2 | 20 | 318.4 | 215.2 | 10 | |
| 42 | 9-Octadecenoic acid, 2-[(trimethylsilyl)oxy]-1-[[(trimethylsilyl)oxy]methyl]ethyl ester | 34.73 | 225.1 | 79.1 | 10 | 225.1 | 93.1 | 10 | |
| 43 | Sucrose, octakis(trimethylsilyl) ether | 35.27 | 117 | 47.1 | 20 | 117 | 75.1 | 10 | |
| 44 | Sucrose, octakis(trimethylsilyl) ether | 38 | 147.1 | 45.1 | 32 | 147.1 | 131.1 | 12 | |
| 45 | β-D-Galactopyranoside, methyl 2,3-bis-O-(trimethylsilyl)-, cyclic methylboronate | 39.09 | 352.7 | 169.2 | 10 | 362.7 | 170.2 | 10 | |
| 46 | Myo-Inositol, pentakis-O-(trimethylsilyl)-, bis(trimethylsilyl) phosphate | 40.81 | 361.4 | 169.1 | 10 | 361.4 | 243.3 | 10 | |
| 47 | 2-Pyrrolidinone, 1-(9-octadecenyl)- | 41 | 103 | 45.1 | 20 | 103 | 58.1 | 30 | |
| 48 | α-L-Galactopyranose, 6-deoxy-1,2,3,4-tetrakis-O-imethylsilyl)- | 41.54 | 399.3 | 81.1 | 10 | 399.3 | 95.5 | 10 | |
| 49 | 9,12-Octadecadienoic acid (Z,Z)-, trimethylsilyl ester | 42.68 | 298.9 | 147.1 | 20 | 298.9 | 225.2 | 10 | |
| 50 | Tryptophan, bis(trimethylsilyl)- | 42.81 | 373.4 | 167.1 | 12 | 373.4 | 211.2 | 12 | |
| 51 | Hexadecanoic acid, trimethylsilyl ester | 43 | 361.1 | 169.2 | 10 | 361.1 | 243.3 | 10 | |
| 52 | N,O,O'-Tris-(trimethylsilyl)tyrosine | 43.67 | 147.2 | 45.1 | 32 | 147.2 | 131.1 | 12 | |
| 53 | Mannose, 2,3,4,5,6-pentakis-O-(trimethylsilyl)-, D- | 44.71 | 204 | 73.1 | 10 | 204 | 189.2 | 10 | |
| 54 | β-Hydroxypyruvic acid, trimethylsilyl ether, trimethylsilyl ester | 44.81 | 147.3 | 131.1 | 10 | 129.3 | 45 | 20 | |
| 55 | Tetradecanoic acid, trimethylsilyl ester | 45.01 | 488.7 | 222.8 | 20 | 488.7 | 223.5 | 10 | |
| 56 | 1,2,3-Propanetricarboxylic acid, 2-[(trimethylsilyl)oxy]-, tris(trimethylsilyl) ester | 45.85 | 357.1 | 225.1 | 18 | 357.1 | 341.2 | 8 | |
| 57 | l-Glutamine, tris(trimethylsilyl) deriv. | 46.57 | 205.2 | 45.3 | 30 | 205.2 | 190.1 | 10 | |
| 58 | Phosphoric acid, bis(trimethylsilyl) 2,3-bis[(trimethylsilyl)oxy]propyl ester | 46.81 | 222 | 45.1 | 32 | 222 | 194.1 | 12 | |
| 59 | Arabinitol, pentakis-O-(trimethylsilyl)- | 48.44 | 211 | 115.1 | 30 | 211 | 133.1 | 10 | |
| 60 | L-Asparagine, N,N2-bis(trimethylsilyl)-, trimethylsilyl ester | 48.84 | 343.2 | 95.2 | 20 | 434.2 | 121.1 | 10 | |
| 61 | Glutamic acid, N-(trimethylsilyl)-, bis(trimethylsilyl) ester, L- | 50.16 | 357.3 | 95.1 | 20 | 357.3 | 107.1 | 20 | |
| 62 | L-Aspartic acid, N-(trimethylsilyl)-, bis(trimethylsilyl) ester | 51.24 | 362.5 | 169.2 | 12 | 362.5 | 170.2 | 12 | |
| 63 | Acetic acid, iodo-, trimethylsilyl ester | 52.13 | 147 | 105.1 | 10 | 147 | 119.1 | 10 | |
| 64 | α-D-Glucopyranoside, 1,3,4,6-tetrakis-O-(trimethylsilyl)-β-D-fructofuranosyl 2,3,4,6-tetrakis-O-(trimethylsilyl)- | 53.49 | 217.2 | 45.1 | 28 | 361.2 | 169.2 | 8 | |
| 65 | D-Glucose, 4-O-[2,3,4,6-tetrakis-O-(trimethylsilyl)-β-D-galactopyranosyl]-2,3,5,6-tetrakis-O-(trimethylsilyl)- | 54.95 | 217.1 | 45.1 | 32 | 217.1 | 143.1 | 12 | |
| 66 | α-D-Glucopyranoside, 1,3,4,6-tetrakis-O-(trimethylsilyl)-β-D-fructofuranosyl 2,3,4,6-tetrakis-O-(trimethylsilyl)- | 59.21 | 204.1 | 45.1 | 30 | 204.1 | 189.2 | 10 | |

Fig. 2. Selected reaction monitoring mode comparison heat map for 66 compounds in 12 tested rice cultivars.The colors in the heat map represent the relative concentration with green and red corresponding to 0 and 100%, respectively. The heat map allows direct visual comparison of the relative amounts of these compounds in the 12 tested samples.
| [1] | Bimb H P, Sah R P, Karn N L.2010. Isozyme variations in fine and aromatic rice genotypes.Nepal Agric Res J, 5: 59-66. |
| [2] | Daviss B.2005. Growing pains for metabolomics.Scientist, 19(8): 25-28. |
| [3] | Fukushima A, Kusano M.2014. A network perspective on nitrogen metabolism from model to crop plants using integrated ‘omics’ approaches.J Exp Bot, 65: 5619-5630. |
| [4] | Galal O A, Abo-Youssef M I, Abdelaziz M, Gharib A T, Dora S A.2015. Assessment of genetic purity of some hybrid rice parental lines using protein profile and fertility restorer gene linked markers. Inter J Biotechnol Res, 2(6): 75-88. |
| [5] | Glaszman J C, de Los Reyes B G, Khush G S.1988. Electrophoretic variation of isozymes in plumules of rice (Oryza sativa L.) a key to the identification of 76 alleles at 24 loci. IRRI Res Paper Ser, 134: 1-3. |
| [6] | Gong L, Chen W, Gao Y Q, Liu X Q, Zhang H Y, Xu C G, Yu S B, Zhang Q F, Luo J.2013. Genetic analysis of the metabolome exemplified using a rice population.Proc Natl Acad Sci USA, 110: 20320-20325. |
| [7] | Hussain A, Scanlon M G, Juliano B O, Bushu K W.1989. Discrimination of rice cultivars by PAGE and HPLC.Cereal Chem, 66(4): 353-356. |
| [8] | Kandikattu H K, Venuprasad M P, Pal A, Khanum F.2014. Phytochemical analysis and exercise enhancing effects of hydroalcoholic extract of Celastrus paniculatus Willd. Ind Crops Prod, 55: 217-224. |
| [9] | Kandikattu H K, Rachitha P, Krupashree K, Jayashree G V, Abhishek V, Khanum F.2015. LC-ESI-MS/MS analysis of total oligomeric flavonoid fraction of Cyperus rotundus and its antioxidant, macromolecule damage protective and antihemolytic effects. Pathophysiol, 22(4): 165-173. |
| [10] | Kim H K, Choi Y H, Verpoorte R.2011. NMR-based plant metabolomics: Where do we stand, where do we go?Trends Biotechnol, 29(6): 267-275. |
| [11] | Kusano M, Yang Z, Okazaki Y, Nakabayashi R, Fukushima A, Saito K.2015. Using metabolomic approaches to explore chemical diversity in rice.Mol Plant, 8(1): 58-67. |
| [12] | Laemmli U K.1970. Cleavage of structural proteins during the assembly of the head of bacteriophage.Nature, 227: 680-685. |
| [13] | Nam M H, Bang E, Kwon T Y, Kim Y, Kim E H, Cho K, Park W J, Kim B G, Yoon I S.2015. Metabolite profiling of diverse rice germplasm and identification of conserved metabolic markers of rice roots in response to long-term mild salinity stress.Inter J Mol Sci, 16: 21959-21974. |
| [14] | Nerkar M, Rao G N.1993. Studies on isozyme variability of rice (Oryza sativa L.). Crop Res, 19(3): 477-480. |
| [15] | Nethra N.2007 Studies on varietal characterization based on morphological, biochemical and molecular markers in rice . [Master Thesis]. Bangalore, India: University Agriculture Science of Bangalore. |
| [16] | Pushpa C, Gowda R, Nethra N, Nataraj K, Uma Rani K, Gangaraju N.2014. Identification of parental lines and rice hybrid (KRH-4) using protein and isozyme electrophoresis.Am Inter J Res Sci Tech Eng Math: 2328-3491. |
| [17] | Rao P, Bharathi M, Reddy K, Keshavulu K, Rao L, Neeraja C.2012. Varietal identification in rice (Oryza sativa) through chemical tests and gel electrophoresis of soluble seed proteins. Ind J Agric Sci, 82(4): 304-311. |
| [18] | Rohini D D.2000. Studies on the characterization of varieties based on morphological and biochemical traits in rice (Oryza sativa L.). [Master Thesis]. Bangalore, India: University Agriculture Science of Bangalore. |
| [19] | Shion B B, Morishma H, Oka H I.1994. A survey of variations in peroxidase, acid phosphatase and esterase isozymes of wild and cultivated Oryza sp. Jpn J Genet, 44(5): 303-319. |
| [20] | Venuprasad M P, Kandikattu H K, Razack S, Khanum F.2014. Phytochemical analysis of Ocimum gratissimum by LC-ESI-MS/MS and its antioxidant and anxiolytic effects. S Afr J Bot, 92: 151-158. |
| [21] | Weckwerth W, Wenzel K, Fiehn O.2004. Process for the integrated extraction identification, and quantification of metabolites, proteins and RNA to reveal their co-regulation in biochemical networks.Proteomics, 4(1): 78-83. |
| [22] | Yu W J, Wang X.1989. Classification of sickle-shaped rice (Oryza sativa L.) based on esterase isoenzymes. Zuowupinzhongziyuan, 3: 4-7. (in Chinese) |
| [23] | Zhou J, Ma C F, Xu H L, Yuan K L, Lu X, Zhu Z, Wu Y N, Xu G W.2009. Metabolic profiling of transgenic rice with cryIAc and sck genes: An evaluation of unintended effects at metabolic level by using GC-FID and GC-MS. J Chromatogr, 877: 725-732. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||