
Rice Science ›› 2019, Vol. 26 ›› Issue (3): 133-146.DOI: 10.1016/j.rsci.2019.04.001
• Review • Next Articles
Mohammed Sulaiman1,2, Abd Samad Azman1, Rahmat Zaidah1( )
)
Received:2018-05-15
Accepted:2018-09-10
Online:2019-05-28
Published:2019-01-25
Mohammed Sulaiman, Abd Samad Azman, Rahmat Zaidah. Agrobacterium-Mediated Transformation of Rice: Constraints and Possible Solutions[J]. Rice Science, 2019, 26(3): 133-146.
Add to citation manager EndNote|Ris|BibTeX
| Cultivar | Sub-species | Medium a | Target explant | Improvement trait | Reference | |
|---|---|---|---|---|---|---|
| Pusa Basmati | indica | MS | Embryogenic callus | Development of fertile transgenic rice | ||
| Sambha Mahsuri, Cotton Sannalu, Pusa Basmati, Taraori Basmati | indica | MS | Embryogenic callus | Abiotic stress (salt) resistant; SOD gene | ||
| IR64 | indica | MS | Shoot apex | Antibiotic resistant plant | ||
| Cempo Ireng | indica | 2N6, MS | Embryogenic callus | Early flowering development | ||
| MR219 | indica | MS | Embryogenic callus | Overexpression of stress related gene, Auxin Binding Protein 57 (Abp 57) | ||
| Sambha Mahsuri, Cotton Sannalu | indica | N6, MS | Embryogenic callus | Improvement of drought tolerance | ||
| Tsukinohikari, Asanohikari, Koshihikari | japonica | N6 | Immature embryo | Hygromycin resistant plant | ||
| Nipponbare | japonica | N6 | Embryogenic callus | Identification of HAP2 genes, 11 HAP3 genes and 7 HAP5 genes binding to the CCAAT-box | ||
| Nipponbare | japonica | MS | Embryogenic callus | Resistance to drought, salinity and pathogens and increasing photosynthesis potential and tiller number | ||
| Nipponbare | japonica | YN | Embryo | Heat tolerance | ||
| Dongjin | japonica | YN | Embryo | Increase tolerance to cold stress, OsCYP19-4 gene | ||
| Dongjin | japonica | 2N6 | Embryogenic callus | Promote flowering in rice at short-day condition | ||
| Taipei 309 | japonica | MS, N6 | Embryogenic callus | Iron vitamin improvement | ||
| Taipei 309 | japonica | MS | Embryogenic callus | Early development of rice inflorescence | ||
| Zhonghua 11 | japonica | N6 | Immature embryo | Floral activator OsELF3 controlling heading date at long-day condition | ||
| Zhonghua 11 | japonica | MS | Embryogenic callus | Increase plant height, grain yield and grain weight | ||
Table 1 Rice genotypes, medium for explant culture and regeneration, and their improvements via Agrobacterium-mediated transformation.
| Cultivar | Sub-species | Medium a | Target explant | Improvement trait | Reference | |
|---|---|---|---|---|---|---|
| Pusa Basmati | indica | MS | Embryogenic callus | Development of fertile transgenic rice | ||
| Sambha Mahsuri, Cotton Sannalu, Pusa Basmati, Taraori Basmati | indica | MS | Embryogenic callus | Abiotic stress (salt) resistant; SOD gene | ||
| IR64 | indica | MS | Shoot apex | Antibiotic resistant plant | ||
| Cempo Ireng | indica | 2N6, MS | Embryogenic callus | Early flowering development | ||
| MR219 | indica | MS | Embryogenic callus | Overexpression of stress related gene, Auxin Binding Protein 57 (Abp 57) | ||
| Sambha Mahsuri, Cotton Sannalu | indica | N6, MS | Embryogenic callus | Improvement of drought tolerance | ||
| Tsukinohikari, Asanohikari, Koshihikari | japonica | N6 | Immature embryo | Hygromycin resistant plant | ||
| Nipponbare | japonica | N6 | Embryogenic callus | Identification of HAP2 genes, 11 HAP3 genes and 7 HAP5 genes binding to the CCAAT-box | ||
| Nipponbare | japonica | MS | Embryogenic callus | Resistance to drought, salinity and pathogens and increasing photosynthesis potential and tiller number | ||
| Nipponbare | japonica | YN | Embryo | Heat tolerance | ||
| Dongjin | japonica | YN | Embryo | Increase tolerance to cold stress, OsCYP19-4 gene | ||
| Dongjin | japonica | 2N6 | Embryogenic callus | Promote flowering in rice at short-day condition | ||
| Taipei 309 | japonica | MS, N6 | Embryogenic callus | Iron vitamin improvement | ||
| Taipei 309 | japonica | MS | Embryogenic callus | Early development of rice inflorescence | ||
| Zhonghua 11 | japonica | N6 | Immature embryo | Floral activator OsELF3 controlling heading date at long-day condition | ||
| Zhonghua 11 | japonica | MS | Embryogenic callus | Increase plant height, grain yield and grain weight | ||
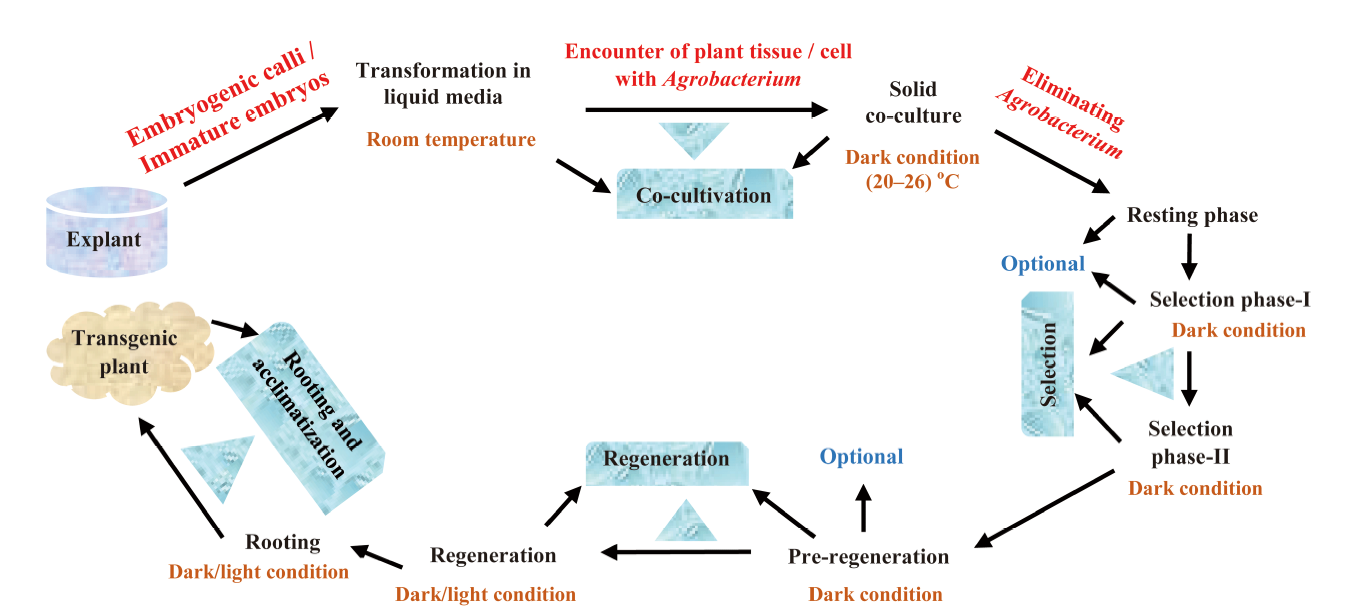
Fig. 1. A complete schematic representation of Agrobacterium-mediated transformation and recovery of transgenic rice adapted from Shrawat and Lörz (2006).
| Variety | Sub-species | Agrobacterium strain | Reporter gene | Selectable marker | Transformation efficiency (%) | Reference |
|---|---|---|---|---|---|---|
| Kasalath | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 66.9 | |
| CaMV35S:sgfp | ||||||
| CaMV35S:luc | ||||||
| Pusa Basmati | indica | EHA105 | CaMV35S:gus | CaMV35S:nptII | 26 | |
| MR219 | indica | LBA4404 | CaMV35S:gus | CaMV35S:hpt | ± 35 | |
| CaMV35S:nptII | ||||||
| Handao 297 | japonica | ALG1 | CaMV35S:gus | CaMV35S:hpt | 20 | |
| Pusa Basmati, Taraori Basmati | indica | LBA4404 | CaMV35S:gus | CaMV35S:hpt | 80 | |
| IR64, CSR10, PB1, Swarna | indica | EHA105, LBA4404 | CaMV35S:gus | CaMV35S:hpt | 45 | |
| BR29, IR68899B | indica | EHA105 | rd29 | rd29:hpt | 12 | |
| Zhenshan 97 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 85.2 | |
| Nipponbare | japonica | EHA105 | CaMV35S:rUbi | CaMV35S:hpt | 1.8 | |
| IR64 | indica | LBA4404 | CaMV35S:SUV | CaMV35S:hpt | 12 | |
| BPT5204 | indica | LBA4404 | CaMV35S:gus | CaMV35S:hpt | 16.7 | |
| CaMV35S:nptII | ||||||
| IR36 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 99.5 | |
| Zhonghua 11 | japonica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 66.7 | |
| Samba Mahsuri (BPT-5204) | indica | LBA4404 | CaMV35S:rd29A | CaMV35S:hpt | ± 30 | |
| MR219 | indica | EHA101, EHA105, LBA4404 | mgfp:gus | mGFP:hptII | 5.8 | |
| JK1044R, JKRH401 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 30 | |
| Nipponbare | japonica | EHA105 | Ubi promoter | Ubi:hpt | 41.2 | |
| Zhonghua 11 | japonica | EHA105 | CaMV35S:gus | CaMV35S:hpt | ± 67.4 | |
| SR1, Jhelum, K332 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 9.3 | |
| CaMV35S:nptII | ||||||
| MR219 | indica | LBA4404 | CaMV35S | CaMV35S:hptII | 26.4 | |
| Sambha Mahsuri, Cotton Sannula | indica | LBA4404 | CaMV35S:gus | CaMV35S:nptII | ± 50 |
Table 2 Transgenic rice generated via Agrobacterium-mediated transformation system (2010-2017), along with the bacterium strain, the vector’s promoter and selectable marker used.
| Variety | Sub-species | Agrobacterium strain | Reporter gene | Selectable marker | Transformation efficiency (%) | Reference |
|---|---|---|---|---|---|---|
| Kasalath | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 66.9 | |
| CaMV35S:sgfp | ||||||
| CaMV35S:luc | ||||||
| Pusa Basmati | indica | EHA105 | CaMV35S:gus | CaMV35S:nptII | 26 | |
| MR219 | indica | LBA4404 | CaMV35S:gus | CaMV35S:hpt | ± 35 | |
| CaMV35S:nptII | ||||||
| Handao 297 | japonica | ALG1 | CaMV35S:gus | CaMV35S:hpt | 20 | |
| Pusa Basmati, Taraori Basmati | indica | LBA4404 | CaMV35S:gus | CaMV35S:hpt | 80 | |
| IR64, CSR10, PB1, Swarna | indica | EHA105, LBA4404 | CaMV35S:gus | CaMV35S:hpt | 45 | |
| BR29, IR68899B | indica | EHA105 | rd29 | rd29:hpt | 12 | |
| Zhenshan 97 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 85.2 | |
| Nipponbare | japonica | EHA105 | CaMV35S:rUbi | CaMV35S:hpt | 1.8 | |
| IR64 | indica | LBA4404 | CaMV35S:SUV | CaMV35S:hpt | 12 | |
| BPT5204 | indica | LBA4404 | CaMV35S:gus | CaMV35S:hpt | 16.7 | |
| CaMV35S:nptII | ||||||
| IR36 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 99.5 | |
| Zhonghua 11 | japonica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 66.7 | |
| Samba Mahsuri (BPT-5204) | indica | LBA4404 | CaMV35S:rd29A | CaMV35S:hpt | ± 30 | |
| MR219 | indica | EHA101, EHA105, LBA4404 | mgfp:gus | mGFP:hptII | 5.8 | |
| JK1044R, JKRH401 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 30 | |
| Nipponbare | japonica | EHA105 | Ubi promoter | Ubi:hpt | 41.2 | |
| Zhonghua 11 | japonica | EHA105 | CaMV35S:gus | CaMV35S:hpt | ± 67.4 | |
| SR1, Jhelum, K332 | indica | EHA105 | CaMV35S:gus | CaMV35S:hpt | 9.3 | |
| CaMV35S:nptII | ||||||
| MR219 | indica | LBA4404 | CaMV35S | CaMV35S:hptII | 26.4 | |
| Sambha Mahsuri, Cotton Sannula | indica | LBA4404 | CaMV35S:gus | CaMV35S:nptII | ± 50 |
| [1] | Alam M M, Tanaka T, Nakamura H, Ichikawa H, Kobayashi K, Yaeno T, Yamaoka N, Shimomoto K, Takayama K, Nishina H, Nishiguchi M.2015. Overexpression of a rice heme activator protein gene (OsHAP2E) confers resistance to pathogens, salinity and drought, and increases photosynthesis and tiller number. Plant Biotechnol J, 13(1): 85-96. |
| [2] | Arntzen C.2015. Plant-made pharmaceuticals: From ‘edible vaccines’ to ebola therapeutics.Plant Biotechnol J, 13(8): 1013-1016. |
| [3] | Ashby A M, Watson M D, Shaw C H.1987. A Ti-plasmid determined function is responsible for chemotaxis of Agrobacterium tumefaciens towards the plant wound product acetosyringone. FEMS Microbiol Lett, 41(2): 189-192. |
| [4] | Azhakanandam K, Silverstone A, Daniell H, Davey M R.2015. Recent Advancements in Gene Expression and Enabling Technologies in Crop Plants. Springer. |
| [5] | Bhatia R, Gallagher J A, Gomez L D, Bosch M.2017. Genetic engineering of grass cell wall polysaccharides for biorefining.Plant Biotechnol J, 15(9): 1071-1092. |
| [6] | Binns A, Campbell A.2001. Agrobacterium tumefaciens-Mediated Transformation of Plant Cells. John Wiley & Sons. doi: 10.1038/npg.els.0001492. |
| [7] | Birch R G, Bower R, Elliott A, Potier B, Franks T, Cordeiro G.1995. Expression of foreign genes in sugarcane. In: Proceedings of the International Society of Sugarcane Technologists XXII Congress. Cartegena, Springer: 368-373. |
| [8] | Birch R G.1997. Plant transformation: Problems and strategies for practical application.Ann Rev Plant Biol, 48: 297-326. |
| [9] | Bourras S, Rouxel T, Meyer M.2015. Agrobacterium tumefaciens gene transfer: How a plant pathogen hacks the nuclei of plant and nonplant organisms. Phytopathology, 105(10): 1288-1301. |
| [10] | Broothaerts W, Mitchell H J, Weir B, Kaines S, Smith L M A, Yang W, Mayer J E, Roa-Rodriguez C, Jefferson R A.2005. Gene transfer to plants by diverse species of bacteria.Nature, 433: 629-633. |
| [11] | Bundock P, Hooykaas P J J.1996. Integration of Agrobacterium tumefaciens T-DNA in the Saccharomyces cerevisiae genome by illegitimate recombination. Proc Natl Acad Sci USA, 93: 15272-15275. |
| [12] | Buttimer C, McAuliffe O, Ross R P, Hill C, O’Mahony J, Coffey A.2017. Bacteriophages and bacterial plant diseases.Front Microb, 8: e33227. |
| [13] | Chakraborty M, Reddy P S, Narasu M L, Krishna G, Rana D.2016. Agrobacterium-mediated genetic transformation of commercially elite rice restorer line using nptII gene as a plant selection marker. Physiol Mol Biol Plants, 22(1): 51-60. |
| [14] | Chandra S.2012. Natural plant genetic engineer Agrobacterium rhizogenes: Role of T-DNA in plant secondary metabolism. Biotechnol Lett, 34(3): 407-415. |
| [15] | Cheng M, Lowe B A, Spencer T M, Ye X D, Armstrong C L.2004. Factors influencing Agrobacterium-mediated transformation of monocotyledonous species. In vitro Cellul Dev Biol Plant, 40(1): 31-45. |
| [16] | Christou P, Ford T L, Kofron M.1991. Production of transgenic rice (Oryza sativa L.) plants from agronomically important indica and japonica varieties via electric discharge particle acceleration of exogenous DNA into immature zygotic embryos. Nat Biotechnol, 9(10): 957-962. |
| [17] | Christou P, McCabe D E.1992. Prediction of germ-line transformation events in chimeric Ro transgenic soybean plantlets using tissue-specific expression patterns. Plant J, 2(3): 283-290. |
| [18] | Clement W K F, Lai K S, Wong M Y, Maziah M.2016. Heat and hydrolytic enzymes treatment improved the Agrobacterium- mediated transformation of recalcitrant indica rice(Oryza sativa L.). Plant Cell, Tiss Organ Cult, 125(1): 183-190. |
| [19] | Danilova S A, da Silva J A T, Kusnetsov V V.2006. Novel approaches for Agrobacterium-mediated transformation of maize and ornamental grasses. Floric, Ornam Plant Biotechnol, 2: 66-69. |
| [20] | Datta K, Baisakh N, Ganguly M, Krishnan S, Yamaguchi Shinozaki K, Datta S K.2012. Overexpression of Arabidopsis and rice stress genes’ inducible transcription factor confers drought and salinity tolerance to rice. Plant Biotechnol J, 10(5): 579-586. |
| [21] | Datta S K, Peterhans A, Datta K, Potrykus I.1990. Genetically engineered fertile indica-rice recovered from protoplasts. Biotechnology, 8(8): 736-740. |
| [22] | Dey M, Bakshi S, Galiba G, Sahoo L, Panda S K.2012. Development of a genotype independent and transformation amenable regeneration system from shoot apex in rice (Oryza sativa spp. indica) using TDZ. 3 Biotech, 2(3): 233-240. |
| [23] | Din A R J M, Ahmad F I, Wagiran A, Samad A A, Rahmat Z, Sarmidi M R.2016. Improvement of efficient in vitro regeneration potential of mature callus induced from Malaysian upland rice seed(Oryza sativa cv. Panderas). Saud J Biol Sci, 23(1): S69-S77. |
| [24] | Endo M, Mikami M, Toki S.2015. Multigene knockout utilizing off-target mutations of the CRISPR/Cas9 system in rice.Plant Cell Physiol, 56(1): 41-47. |
| [25] | Escobar M A, Dandekar A M.2003. Agrobacterium tumefaciens as an agent of disease. Trends Plant Sci, 8(8): 380-386. |
| [26] | Fang R X, Nagy F, Sivasubramaniam S, Chua N H.1989. Multiple cis regulatory elements for maximal expression of the cauliflower mosaic virus 35S promoter in transgenic plants.Plant Cell, 1: 141-150. |
| [27] | Fook C W K, Song L K, Wong Y M, Mahmood M.2015. Efficient regeneration and Agrobacterium-mediated transformation protocol for recalcitrant indica rice(Oryza sativa L.). Em J Food Agric, 27(11): 837-848. |
| [28] | Gelvin S B.2000. Agrobacterium and plant genes involved in T-DNA transfer and integration. Ann Rev Plant Biol, 51(1): 223-256. |
| [29] | Grabber J H.2005. How do lignin composition, structure, and cross-linking affect degradability? A review of cell wall model studies.Crop Sci, 45(3): 820-831. |
| [30] | Han S H, Yoo S C, Lee B D, An G, Paek N C.2015. Rice FLAVIN-BINDING, KELCH REPEAT, F-BOX 1 (OsFKF1) promotes flowering independent of photoperiod. Plant, Cell Environ, 38(12): 2527-2540. |
| [31] | Hansen G, Das A, Chilton M D.1994. Constitutive expression of the virulence genes improves the efficiency of plant transformation by Agrobacterium. Proc Natl Acad Sci USA, 91(16): 7603-7607. |
| [32] | Hansen G, Shillito R D, Chilton M D.1997. T-strand integration in maize protoplasts after codelivery of a T-DNA substrate and virulence genes.Proc Natl Acad Sci USA, 94: 11726-11730. |
| [33] | Hellwig S, Drossard J, Twyman R M, Fischer R.2004. Plant cell cultures for the production of recombinant proteins.Nat Biotechnol, 22(11): 1415-1422. |
| [34] | Hempel F, Bozarth A S, Lindenkamp N, Klingl A, Zauner S, Linne U, Steinbüchel A, Maier U G.2011. Microalgae as bioreactors for bioplastic production.Microb Cell Fact, 10(1): 81. |
| [35] | Hiei Y, Ohta S, Komari T, Kumashiro T.1994. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J, 6(2): 271-282. |
| [36] | Hiei Y, Komari T, Kubo T.1997. Transformation of rice mediated by Agrobacterium tumefaciens. Plant Mol Biol, 35: 205-218. |
| [37] | Hiei Y, Ishida Y, Komari T.2014. Progress of cereal transformation technology mediated by Agrobacterium tumefaciens. Front Plant Sci, 5: 628. |
| [38] | Hoffmann M C, Pfänder Y, Tintel M, Masepohl B.2017. Bacterial PerO permeases transport sulfate and related oxyanions.J Bacteriol, 199(14): 83-117. |
| [39] | Horsch R B, Fraley R T, Rogers S G, Sanders P R, Lloyd A, Hoffmann N.1984. Inheritance of functional foreign genes in plants.Science, 223: 496-498. |
| [40] | Hou C X, Lv T, Zhan Y H, Peng Y Y, Huang Y Y, Jiang D A, Weng X Y.2015. Overexpression of the RIXI xylanase inhibitor improves disease resistance to the fungal pathogen,Magnaporthe oryzae, in rice. Plant Cell, Tiss Organ Cult, 120(1): 167-177. |
| [41] | Isikgor F H, Becer C R.2015. Lignocellulosic biomass: A sustainable platform for the production of bio-based chemicals and polymers.Polym Chem, 6: 4497-4559. |
| [42] | Ithape D M, Maharana M, Tripathy Swapan K.2017. Scope of genetic transformation in Sugarcane: A review.Genom Appl Biol, 8(1): 1-7. |
| [43] | Izawa T, Shimamoto K.1996. Becoming a model plant: The importance of rice to plant science.Trends Plant Sci, 1(3): 95-99. |
| [44] | Jiang H M, Doerge R W, Gelvin S B.2003. Transfer of T-DNA and vir proteins to plant cells by Agrobacterium tumefaciens induces expression of host genes involved in mediating transformation and suppresses host defense gene expression. Plant J, 35(2): 219-236. |
| [45] | Joersbo M, Okkels F T.1996. A novel principle for selection of transgenic plant cells: Positive selection.Plant Cell Rep, 16: 219-221. |
| [46] | Kang J M, Turano F J.2003. The putative glutamate receptor 1.1 (AtGLR1.1) functions as a regulator of carbon and nitrogen metabolism in Arabidopsis thaliana. Proc Natl Acad Sci USA, 100(11): 6872-6877. |
| [47] | Karthikeyan A, Pandian S T K, Ramesh M.2009. High frequency plant regeneration from embryogenic callus of a popular indica rice(Oryza sativa L.). Physiol Mol Biol Plants, 15(4): 371-375. |
| [48] | Koetle M J, Finnie J F, Balázs E, van Staden J.2015. A review on factors affecting the Agrobacterium-mediated genetic transformation in ornamental monocotyledonous geophytes. South Afr J Bot, 98: 37-44. |
| [49] | Koetle M J, Baskaran P, Finnie J F, Soos V, Balázs E, van Staden J.2017. Optimization of transient GUS expression of Agrobacterium- mediated transformation in Dierama erectum Hilliard using sonication and Agrobacterium. South Afr J Bot, 111: 307-312. |
| [50] | Komiya R, Ikegami A, Tamaki S, Yokoi S, Shimamoto K.2008. Hd3a and RFT1 are essential for flowering in rice. Development, 135(4): 767-774. |
| [51] | Komiya R, Yokoi S, Shimamoto K.2009. A gene network for long-day flowering activates RFT1 encoding a mobile flowering signal in rice. Development, 136(20): 3443-3450. |
| [52] | Koncz C, Schell J.1986. The promoter of TL-DNA gene 5 controls the tissue-specific expression of chimaeric genes carried by a novel type of Agrobacterium binary vector. Mol Gener Genet, 204(3): 383-396. |
| [53] | Koziel M G, Beland G L, Bowman C, Carozzi N B, Crenshaw R, Crossland L, Dawson J, Desai N, Hill M, Kadwell S, Launis K, Lewis K, Maddox D, McPherson K, Meghji M R, Merlin E, Rhodes R, Warren G W, Wright M, Evola S V.1993. Field performance of elite transgenic maize plants expressing an insecticidal protein derived from Bacillus thuringiensis. Nat Biotechnol, 11(2): 194-200. |
| [54] | Krishnan S R, Priya A M, Ramesh M.2013. Rapid regeneration and ploidy stability of ‘cv IR36’ indica rice(Oryza sativa L.) confers efficient protocol for in vitro callus organogenesis and Agrobacterium tumefaciens mediated transformation. Bot Stud, 54: 47. |
| [55] | Kumria R, Waie B, Rajam M.2001. Plant regeneration from transformed embryogenic callus of an elite indica rice via Agrobacterium. Plant Cell, Tiss Organ Cult, 67(1): 63-71. |
| [56] | Lacroix B, Zaltsman A, Citovsky V.2011. Host factors involved in genetic transformation of plant cells by Agrobacterium. In: Neal Stewart Jr C, Touraev A, Tzfira T. Plant Transformation Technologies. John Wiley & Sons: 1-29. |
| [57] | Laurenceau R, Péhau-Arnaudet G, Baconnais S, Gault J, Malosse C, Dujeancourt A, Campo N, Chamot-Rooke J, Le Cam E, Claverys J P, Fronzes R.2013. A type IV pilus mediates DNA binding during natural transformation in Streptococcus pneumoniae. PLoS Pathog, 9(6): e1003473. |
| [58] | Lee Y S, An G.2015. OsGI controls flowering time by modulating rhythmic flowering time regulators preferentially under short day in rice. J Plant Biol, 58(2): 137-145. |
| [59] | Li J, Zhu S H, Song X W, Shen Y, Chen H M, Yu J, Yi K K, Liu Y F, Karplus V J, Wu P, Deng X W.2006. A rice glutamate receptor-like gene is critical for the division and survival of individual cells in the root apical meristem. Plant Cell, 18(2): 340-349. |
| [60] | Liang Z, Chen K L, Li T D, Zhang Y, Wang Y P, Zhao Q, Liu J X, Zhang H W, Liu C M, Ran Y D, Gao C X.2017. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes.Nat Commun, 8: 1-5. |
| [61] | Lippincott J A, Lippincott B B.1978. Cell walls of crown-gall tumors and embryonic plant tissues lack Agrobacterium adherence sites. Science, 199: 1075-1078. |
| [62] | Liu J P, Zhang C C, Wei C C, Liu X, Wang M G, Yu F F, Xie Q, Tu J M.2016. The RING finger ubiquitin E3 ligase OsHTAS enhances heat tolerance by promoting H2O2-induced stomatal closure in rice.Plant Physiol, 170(1): 429-443. |
| [63] | Lorenz M G, Wackernagel W.1994. Bacterial gene transfer by natural genetic transformation in the environment.Microb Rev, 58(3): 563-602. |
| [64] | Lörz H, Baker B, Schell J.1985. Gene transfer to cereal cells mediated by protoplast transformation.Mol Gene Genet, 199(2): 178-182. |
| [65] | Lowe K, Wu E, Wang N, Hoerster G, Hastings C, Cho M J, Scelonge C, Lenderts B, Chamberlin M, Cushatt J, Wang L J, Ryan L, Khan T, Chow-Yiu J, Hua W, Yu M, Banh J, Bao Z M, Brink K, Igo E, Rudrappa B, Shamseer P M, Bruce W, Newman L, Shen B, Zheng P Z, Bidney D, Carl Falco S, Register J C, Zhao Z Y, Xu D P, Jones T, Gordon-Kamm W.2016. Morphogenic regulators baby boom and Wuschel improve monocot transformation.Plant Cell, 28(9): 1998-2015. |
| [66] | Lucca P, Hurrell R, Potrykus I.2001. Genetic engineering approaches to improve the bioavailability and the level of iron in rice grains.Theor Appl Genet, 102(2): 392-397. |
| [67] | Ma J K C, Drossard J, Lewis D, Altmann F, Boyle J, Christou P, Cole T, Dale P, van Dolleweerd C J, Isitt V, Katinger D, Lobedan M, Mertens H, Paul M J, Rademacher T, Sack M, Hundleby P A C, Stiegler G, Stoger E, Twyman R M, Vcelar B, Fischer R.2015. Regulatory approval and a first-in-human phase I clinical trial of a monoclonal antibody produced in transgenic tobacco plants.Plant Biotechnol J, 13(8): 1106-1120. |
| [68] | Mafakheri H, Taghavi S M, Banihashemi Z, Osdaghi E, Lamichhane J R.2017. Pathogenicity, host range and phylogenetic position of Agrobacterium species associated with sugar beet crown gall outbreaks in Southern Iran. Eur J Plant Pathol, 147(3): 721-730. |
| [69] | Manimaran P, Kumar G R, Reddy M R, Jain S, Rao T B, Mangrauthia S K, Sundaram R M, Ravichandran S, Balachandran S M.2013. Infection of early and young callus tissues of indica rice BPT5204 enhances regeneration and transformation efficiency. Rice Sci, 20(6): 415-426. |
| [70] | Mann D G J, LaFayette P R, Abercrombie L L, King Z R, Mazarei M, Halter M C, Poovaiah C R, Baxter H, Shen H, Dixon R A, Parrott W A, Neal Stewart Jr C.2012. Gateway-compatible vectors for high-throughput gene functional analysis in switchgrass (Panicum virgatum L.) and other monocot species. Plant Biotechnol J, 10(2): 226-236. |
| [71] | Martins P K, Ribeiro A P, da Cunha B A D B, Kobayashi A K, Molinari H B C.2015. A simple and highly efficient Agrobacterium-mediated transformation protocol for Setaria viridis. Biotechnol Rep, 6: 41-44. |
| [72] | Matzke M A, Matzke A J.1995. How and why do plants inactivate homologous (trans) genes?Plant Physiol, 107(3): 679-685. |
| [73] | Mohanty A, Sarma N P, Tyagi A K.1999. Agrobacterium- mediated high frequency transformation of an elite indica rice variety Pusa Basmati 1 and transmission of the transgenes to R2 progeny. Plant Sci, 147(2): 127-137. |
| [74] | Mulwa R M S, Norton M A, Farrand S K, Skirvin R M.2015. Agrobacterium-mediated transformation and regeneration of transgenic Chancellor wine grape plants expressing the tfd A gene. VITIS J Grap Res, 46(3): 110-115. |
| [75] | Opabode J T.2006. Agrobacterium-mediated transformation of plants: Emerging factors that influence efficiency. Biotechnol Mol Biol Rev, 1(1): 12-20. |
| [76] | Ou-Lee T M, Turgeon R, Wu R.1986. Expression of a foreign gene linked to either a plant-virus or a Drosophila promoter, after electroporation of protoplasts of rice, wheat, and sorghum.Proc Natl Acad Sci USA, 83(18): 6815-6819. |
| [77] | Ozawa K, Wakasa Y, Ogo Y, Matsuo K, Kawahigashi H, Takaiwa F.2012. Development of an efficient Agrobacterium-mediated gene targeting system for rice and analysis of rice knockouts lacking granule-bound starch synthase (Waxy) and β1, 2-xylosyltransferase. Plant Cell Physiol, 53(4): 755-761. |
| [78] | Park S Y, Vaghchhipawala Z, Vasudevan B, Lee L Y, Shen Y, Singer K, Waterworth W M, Zhang Z J, West C E, Mysore K S, Gelvin S B.2015. Agrobacterium T-DNA integration into the plant genome can occur without the activity of key non-homologous end-joining proteins. Plant J, 81(6): 934-946. |
| [79] | Pathi K M, Tula S, Huda K M K, Srivastava V K, Tuteja N.2013. An efficient and rapid regeneration via multiple shoot induction from mature seed derived embryogenic and organogenic callus of Indian maize (Zea mays L.). Plant Signal Behav, 8(10): e25891. |
| [80] | Prasad K, Kushalappa K, Vijayraghavan U.2003. Mechanism underlying regulated expression of RFL, a conserved transcription factor, in the developing rice inflorescence.Mech Dev, 120(4): 491-502. |
| [81] | Purwestri Y A, Sari R D K, Anggraeni L N, Sasongko A B.2015. Agrobacterium tumefaciens mediated transformation of rolC:: Hd3a-GFP in black rice(Oryza sativa L. cv. Cempo Ireng) to promote early flowering. Proc Chem, 14: 469-473. |
| [82] | Rahman Z A, Seman Z A, Basirun N, Julkifle A L, Zainal Z, Subramaniam S.2011. Preliminary investigations of Agrobacterium- mediated transformation in indica rice MR219 embryogenic callus using gusA gene. Afr J Biotechnol, 10: 7805-7813. |
| [83] | Rao K V, Rathore K S, Hodges T K, Fu X, Stoger E, Sudhakar D, Williams S, Christou P, Bharathi M, Bown D P, Powell K S, Spence J, Gatehouse A M R, Gatehouse J A.1998. Expression of snowdrop lectin (GNA) in transgenic rice plants confers resistance to rice brown planthopper.Plant J, 15(4): 469-477. |
| [84] | Rashid B, Tariq M, Khalid A, Shams F, Ali Q, Ashraf F, Ghaffar I, Khan M I, Rehman R, Husnain T.2017. Crop improvement: New approaches and modern techniques.Plant Gene Trait, 8(3): 18-30. |
| [85] | Rathus C, Bower R, Birch R G.1993. Effects of promoter, intron and enhancer elements on transient gene expression in sugar-cane and carrot protoplasts.Plant Mol Biol, 23(3): 613-618. |
| [86] | Ravikumar G, Manimaran P, Voleti S R, Subrahmanyam D, Sundaram R M, Bansal K C, Viraktamath B C, Balachandran S M.2014. Stress-inducible expression of AtDREB1A transcription factor greatly improves drought stress tolerance in transgenic indica rice. Transg Res, 23(3): 421-439. |
| [87] | Reddy S S S, Singh B, Peter A J, Rao T V.2018. Production of transgenic local rice cultivars (Oryza sativa L.) for improved drought tolerance using Agrobacterium mediated transformation. Saud J Biol Sci, 25(8): 1535-1545. |
| [88] | Reddy S S S, Singh B, Peter A J, Rao T V.2019. Genetic transformation of indica rice varieties involving Am-SOD gene for improved abiotic stress tolerance. Saud J Biol Sci, 26(2): 294-300. |
| [89] | Sack M, Hofbauer A, Fischer R, Stoger E.2015. The increasing value of plant-made proteins.Curr Opin Biotechnol, 32: 163-170. |
| [90] | Saharan V, Yadav R C, Yadav N R, Ram K.2004. Studies on improved Agrobacterium-mediated transformation in two indica rice(Oryza sativa L.). Afr J Biotechnol, 3(11): 572-575. |
| [91] | Sahoo K K, Tripathi A K, Pareek A, Sopory S K, Singla-Pareek S L.2011. An improved protocol for efficient transformation and regeneration of diverse indica rice cultivars. Plant Meth, 7: 49. |
| [92] | Sahoo R K, Tuteja N.2012. Development of Agrobacterium- mediated transformation technology for mature seed-derived callus tissues of indica rice cultivar IR64. GM Crops Food, 3(2): 123-128. |
| [93] | Saidu H, Mohammed S.2013. Plant response to abiotic stresses: From the perspectives of gene expression.Res J Biotechnol, 8(9): 81-91. |
| [94] | Saika H, Toki S.2010. Mature seed-derived callus of the model indica rice variety Kasalath is highly competent in Agrobacterium- mediated transformation. Plant Cell Rep, 29(12): 1351-1364. |
| [95] | Sarangi S, Ghosh J, Bora A, Das S, Mandal A B.2011. Agrobacterium-mediated genetic transformation of indica rice varieties involving Am-SOD gene. Ind J Biotechnol, 10(1): 9-18. |
| [96] | Schoonbeek H J, Wang H H, Stefanato F L, Craze M, Bowden S, Wallington E, Zipfel C, Ridout C J.2015. Arabidopsis EF-Tu receptor enhances bacterial disease resistance in transgenic wheat. New Phytol, 206(2): 606-613. |
| [97] | Shi J, Gao H, Wang H, Lafitte H R, Archibald R L, Yang M, Hakimi S M, Mo H, Habben J E.2017. ARGOS8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions.Plant Biotechnol J, 15(2): 207-216. |
| [98] | Shrawat A K, Lörz H.2006. Agrobacterium-mediated transformation of cereals: A promising approach crossing barriers. Plant Biotechnol J, 4(6): 575-603. |
| [99] | Snell K D, Peoples O P.2009. PHA bioplastic: A value-added coproduct for biomass biorefineries.Biotechnol Bioen Bioprod, 3(4): 456-467. |
| [100] | Sood P, Bhattacharya A, Sood A.2011. Problems and possibilities of monocot transformation.Biol Plant, 55(1): 1-15. |
| [101] | Streatfield S J.2007. Approaches to achieve high-level heterologous protein production in plants.Plant Biotechnol J, 5(1): 2-15. |
| [102] | Svitashev S, Young J K, Schwartz C, Gao H R, Falco S C, Cigan A M.2015. Targeted mutagenesis, precise gene editing and site-specific gene insertion in maize using Cas9 and guide RNA.Plant Physiol, 169: 931-945. |
| [103] | Tabashnik B E, Gassmann A J, Crowder D W, Carriére Y.2008. Insect resistance to Bt crops: Evidence versus theory.Nat Biotechnol, 26(2): 199-202. |
| [104] | Tan L W, Rahman Z A, Goh H H, Hwang D J, Ismail I, Zainal Z.2017. Production of transgenic rice (indica cv. MR219) overexpressing Abp57 gene through Agrobacterium-mediated transformation. Sains Malays, 46(5): 703-711. |
| [105] | Thirumurugan T, Ito Y, Kubo T, Serizawa A, Kurata N.2008. Identification, characterization and interaction of HAP family genes in rice.Mol Genet Genom, 279(3): 279-289. |
| [106] | Tie W W, Zhou F, Wang L, Xie W B, Chen H, Li X H, Lin Y J.2012. Reasons for lower transformation efficiency in indica rice using Agrobacterium tumefaciens-mediated transformation: Lessons from transformation assays and genome-wide expression profiling. Plant Mol Biol, 78: 1-18. |
| [107] | Toriyama K, Arimoto Y, Uchimiya H, Hinata K.1988. Transgenic rice plants after direct gene transfer into protoplasts.Nat Biotechnol, 6(9): 1072-1074. |
| [108] | Tripathi R, Bisht H, Singh R.2010. Effect of acetosyringone and callus age on transformation for scutellum-derived callus of rice.Int J Pharm Biol Sci, 1(4): 163-170. |
| [109] | Tzfira T, Li J, Lacroix B, Citovsky V.2004. Agrobacterium T-DNA integration: Molecules and models. Trends Genet, 20(8): 375-383. |
| [110] | Tzfira T, Citovsky V.2006. Agrobacterium-mediated genetic transformation of plants: Biology and biotechnology. Curr Opin Biotechnol, 17(2): 147-154. |
| [111] | Ullah H, Ullah I, Jadoon S A, Rashid H.2007. Tissue culture techniques for callus induction in rice.Sarh J Agric, 23(1): 81-86. |
| [112] | Uzé M, Potrykus I, Sautter C.2000. Factors influencing T-DNA transfer from Agrobacterium to precultured immature wheat embryos(Triticum aestivum L.). Cereal Res Commun, 28(2): 17-23. |
| [113] | Vain P.2007. Thirty years of plant transformation technology development.Plant Biotechnol J, 5(2): 221-229. |
| [114] | van der Graaff E, den Dulk-Ras A, Hooykaas P J J.1996. Deviating T-DNA transfer from Agrobacterium tumefaciens to plants. Plant Mol Biol, 31(3): 677-681. |
| [115] | Veluthambi K, Gupta A K, Sharma A.2003. The current status of plant transformation technologies.Curr Sci, 84(3): 368-380. |
| [116] | Vogel J.2008. Unique aspects of the grass cell wall.Curr Opin Plant Biol, 11(3): 301-307. |
| [117] | Wang C G, Hu Z L, Lei A P, Jin B H.2010. Biosynthesis of poly-3-hydroxybutyrate (PHB) in the transgenic green alga Chlamydomonas reinhardtii. J Phycol, 46(2): 396-402. |
| [118] | Wendt T, Doohan F, Mullins E.2012. Production of Phytophthora infestans-resistant potato(Solanum tuberosum) utilising Ensifer adhaerens OV14. Transg Res, 21(3): 567-578. |
| [119] | Wright M, Dawson J, Dunder E, Suttie J, Reed J, Kramer C, Chang Y, Novitzky R, Wang H, Artim-Moore L.2001. Efficient biolistic transformation of maize (Zea mays L.) and wheat(Triticum aestivum L.) using the phosphomannose isomerase gene, pmi, as the selectable marker. Plant Cell Rep, 20(5): 429-436. |
| [120] | Wright T R, Shan G, Walsh T A, Lira J M, Cui C, Song P, Zhuang M, Arnold N L, Lin G, Yau K, Russell S M, Clcchillo R M, Peterson M A, Simpson D M, Zhou N, Ponsamuel J, Zhang Z.2010. Robust crop resistance to broadleaf and grass herbicides provided by aryloxyalkanoate dioxygenase transgenes.Proc Natl Acad Sci USA, 107: 20240-20245. |
| [121] | Xing S C, Li F, Guo Q F, Liu D R, Zhao X X, Wang W.2009. The involvement of an expansin gene TaEXPB23 from wheat in regulating plant cell growth. Biol Plant, 53(3): 429-434. |
| [122] | Xu R F, Qin R Y, Li H, Li D D, Li L, Wei P C, Yang J B.2017. Generation of targeted mutant rice using a CRISPR-Cpf1 system.Plant Biotechnol J, 15(6): 713-717. |
| [123] | Xue W Y, Xing Y Z, Weng X Y, Zhao Y, Tang W J, Wang L, Zhou H J, Yu S B, Xu C G, Li X H, Zhang Q F.2008. Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet, 40(6): 761-767. |
| [124] | Yang Y, Peng Q, Chen G X, Li X H, Wu C Y.2013. OsELF3 is involved in circadian clock regulation for promoting flowering under long-day conditions in rice. Mol Plant, 6(1): 202-215. |
| [125] | Yao J L, Cohen D, Atkinson R, Richardson K, Morris B.1995. Regeneration of transgenic plants from the commercial apple cultivar Royal Gala.Plant Cell Rep, 14(7): 407-412. |
| [126] | Yaqoob U, Kaul T, Nawchoo I A.2017. Development of an efficient protocol for Agrobacterium mediated transformation of some recalcitrant indica rice varieties. Ind J Plant Physiol, 22(3): 346-353. |
| [127] | Yookongkaew N, Srivatanakul M, Narangajavana J.2007. Development of genotype-independent regeneration system for transformation of rice (Oryza sativa ssp. indica). J Plant Res, 120(2): 237-245. |
| [128] | Yoon D H, Lee S S, Park H J, Lyu J I, Chong W S, Liu J R, Kim B G, Ahn J C, Cho H S.2015. Overexpression of OsCYP19-4 increases tolerance to cold stress and enhances grain yield in rice(Oryza sativa). J Exp Bot, 67(1): 69-82. |
| [129] | Zaidi M A, Narayanan M, Sardana R, Taga I, Postel S, Johns R, McNulty M, Mottiar Y, Mao J, Loit E, Altosaar I.2006. Optimizing tissue culture media for efficient transformation of different indica rice genotypes. Agron Res, 4(2): 563-575. |
| [130] | Zhang H M, Yang H, Rech E L, Golds T J, Davis A S, Mulligan B J, Cocking E C, Davey M R.1988. Transgenic rice plants produced by electroporation-mediated plasmid uptake into protoplasts.Plant Cell Rep, 7(6): 379-384. |
| [131] | Zhang H X, Blumwald E.2001. Transgenic salt-tolerant tomato plants accumulate salt in foliage but not in fruit.Nat Biotechnol, 19(8): 765-768. |
| [132] | Zhao J, Chen H Y, Ren D, Tang H W, Qiu R, Feng J L, Long Y M, Niu B X, Chen D P, Zhong T Y, Liu Y G, Guo J X.2015. Genetic interactions between diverged alleles of early heading date 1 (Ehd1) and heading date 3a (Hd3a)/RICE FLOWERING LOCUS T1 (RFT1) control differential heading and contribute to regional adaptation in rice(Oryza sativa). New Phytol, 208(3): 936-948. |
| [133] | Zhao W N, Zheng S S, Ling H Q.2011. An efficient regeneration system and Agrobacterium-mediated transformation of Chinese upland rice cultivar Handao 297. Plant Cell, Tiss Organ Cult, 106: 475. |
| [134] | Zhu Y J, Fan Y Y, Wang K, Huang D R, Liu W Z, Ying J Z, Zhuang J Y.2017. Rice flowering locus T1 plays an important role in heading date influencing yield traits in rice. Sci Rep, 7: 4918. |
| [135] | Ziemienowicz A.2014. Agrobacterium-mediated plant transformation: Factors, applications and recent advances. Biocat Agric Biotechnol, 3(4): 95-102. |
| [136] | Zuniga-Soto E, Mullins E, Dedicova B.2015. Ensifer-mediated transformation: An efficient non-Agrobacterium protocol for the genetic modification of rice. Spring Plus, 4: 600. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||