
Rice Science ›› 2020, Vol. 27 ›› Issue (5): 368-383.DOI: 10.1016/j.rsci.2020.03.002
• Orginal Article • Previous Articles Next Articles
Bhatt Tarun, Sharma Aditi, Puri Sanjeev, Priya Minhas Anu( )
)
Received:2019-11-11
Accepted:2020-03-19
Online:2020-09-28
Published:2020-09-28
Bhatt Tarun, Sharma Aditi, Puri Sanjeev, Priya Minhas Anu. Salt Tolerance Mechanisms and Approaches: Future Scope of Halotolerant Genes and Rice Landraces[J]. Rice Science, 2020, 27(5): 368-383.
Add to citation manager EndNote|Ris|BibTeX
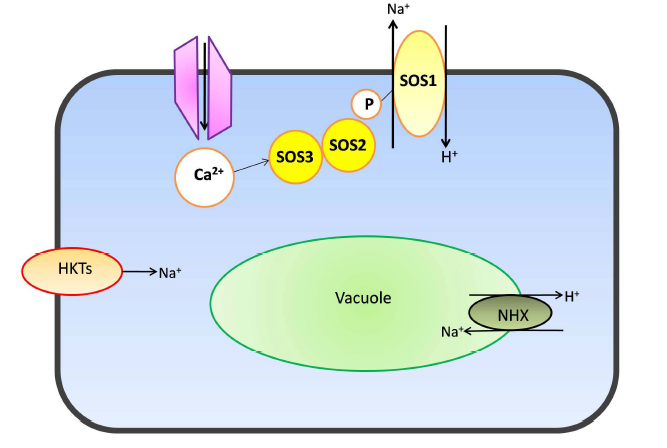
Fig. 1. Diagramatic representation of Na+ transport across cellular and vacuolar membranes through various cation channels.SOS1, Na+/H+ antiporter; SOS2, Serine/Threonine protein kinase; SOS3, Calcium binding protein; NHX1, Sodium/hydrogen; HKT1, High affinity potassium transporter.

Fig. 2. Amino acid alignment of OsHAL3 (accession no Q69K55) to homologs using Clustal X method (Larkin et al, 2007).ZmHAL3, Homolog from Zea mays (Schnable et al, 2009); GmHAL3a and GmHAL3b, Homologs from Glycine max (Guo et al, 2016); PmHAL3, Homolog from Panicum miliaceum (Zou et al, 2019); AtHAL3a and AtHAL3b, Homologs from Arabidopsis thaliana (Espinosa-Ruiz et al, 1999); TaHAL3, Homolog from Triticum aestivum (Choulet et al, 2014); SlHAL3, Homolog from Solanum lycopersicum (Aoki et al, 2010), NtHAL3, Homolog from Nicotiana tabacum (Yonamine et al, 2004); CsHAL3, Homolog from Cucumis sativus (Huang et al, 2009).Sequences in the upper and lower boxes indicate FMN binding motif (PLASANT) and substrate recognition motif (PXMNXXMW) sequences, respectively. Conserved H90 residue is shown in bold letter.
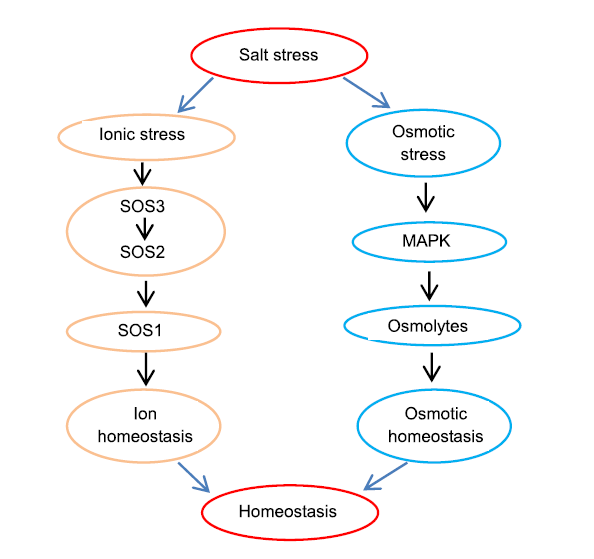
Fig. 3. A model depicting interplay of ionic and osmotic stress induced pathways to maintain cellular homeostasis in response to salt stress.SOS1, Na+/H+ antiporter; SOS2, Serine/Threonine protein kinase; SOS3, Calcium binding protein; MAPK, Mitogen-activated protein kinase.
| [1] | Abrie J A, González A, Strauss E, Ariño J.2012. Functional mapping of the disparate activities of the yeast moonlighting protein Hal3. Biochem J, 442(2): 357-368. |
| [2] | Albert A, Yenush L, Gil-Mascarell M R, Rodriguez P L, Patel S, Martı́nez-Ripoll M, Blundell T L, Serrano R.2000. X-ray structure of yeast hal2p, a major target of lithium and sodium toxicity, and identification of framework interactions determining cation sensitivity. J Mol Biol, 295(4): 927-938. |
| [3] | Ali S, Mannan A, El Oirdi M, Waheed A, Mirza B.2012. Agrobacterium-mediated transformation of rough lemon (Citrus jambhiri Lush) with yeast HAL2 gene. BMC Res Notes, 5: 285. |
| [4] | Anoop N, Gupta A K.2003. Transgenic indica rice cv IR-50 over-expressing Vigna aconitifolia Δ1-pyrroline-5-carboxylate synthetase cDNA shows tolerance to high salt. J Plant Biochem Biotechnol, 12: 109-116. |
| [5] | Aoki K, Yano K, Suzuki A, Kawamura S, Sakurai N, Suda K, Kurabayashi A, Suzuki T, Tsugane T, Watanabe M, Ooga K, Torii M, Narita T, Shin-i T, Kohara Y, Yamamoto N, Takahashi H, Watanabe Y, Egusa M, Kodama M, Ichinose Y, Kikuchi M, Fukushima S, Okabe A, Arie T, Sato Y, Yazawa K, Satoh S, Omura T, Ezura H, Shibata D.2010. Large-scale analysis of full-length cDNAs from the tomato (Solanum lycopersicum) cultivar Micro-Tom, a reference system for the Solanaceae genomics. BMC Genom, 11: 210. |
| [6] | Arrillaga I, Gil-Mascarell R, Gisbert C, Sales E, Montesinos C, Serrano R, Moreno V.1998. Expression of the yeast HAL2 gene in tomato increases the in vitro salt tolerance of transgenic progenies. Plant Sci, 136(2): 219-226. |
| [7] | Bassil E, Coku A, Blumwald E.2012. Cellular ion homeostasis: Emerging roles of intracellular Nhx Na+/H+ antiporters in plant growth and development. J Exp Bot, 63(16): 5727-5740. |
| [8] | Beer S C, Goffreda J, Phillips T D, Murphy J P, Sorrells M E.1993. Assessment of genetic variation in Avena sterilis using morphological traits, isozymes, and RFLPs. Crop Sci, 33(6): 1386-1393. |
| [9] | Binzel M L, Hasegawa P M, Rhodes D, Handa S, Handa A K, Bressan R A.1987. Solute accumulation in tobacco cells adapted to NaCl. Plant Physiol, 84(4): 1408-1415. |
| [10] | Blaesse M, Kupke T, Huber R, Steinbacher S.2000. Crystal structure of the peptidyl-cysteine decarboxylase EpiD complexed with a pentapeptide substrate. EMBO J, 19(23): 6299-6310. |
| [11] | Bohnert H J, Jensen R G.1996. Strategies for engineering water- stress tolerance in plants. Trends Biotechnol, 14(3): 89-97. |
| [12] | Bordas M, Montesinos C, Dabauza M, Salvador A, Roig L A, Serrano R, Moreno V.1997. Transfer of the yeast salt tolerance gene HAL1 to Cucumis melo L. cultivars and in vitro evaluation of salt tolerance. Transg Res, 6: 41-50. |
| [13] | Breseghello F, Coelho A S G.2013. Traditional and modern plant breeding methods with examples in rice (Oryza sativa L.). J Agric Food Chem, 61: 8277-8286. |
| [14] | Cabot C, Sibole J V, Barceló J, Poschenrieder C.2009. Abscisic acid decreases leaf Na+ exclusion in salt-treated Phaseolus vulgaris L. J Plant Growth Regul, 28: 187-192. |
| [15] | Cervera M, Ortega C, Navarro A, Navarro L, Peña L.2000. Generation of transgenic citrus plants with the tolerance-to- salinity gene HAL2 from yeast. J Hort Sci Biotechnol, 75: 26-30. |
| [16] | Chanroj S, Wang G Y, Venema K, Zhang M W, Delwiche C F, Sze H.2012. Conserved and diversified gene families of monovalent cation/H+ antiporters from algae to flowering plants. Front Plant Sci, 3: 25. |
| [17] | Chen J Q, Meng X P, Zhang Y, Xia M, Wang X P.2008. Over-expression of OsDREB genes lead to enhanced drought tolerance in rice. Biotechnol Lett, 30: 2191-2198. |
| [18] | Chen Y, Li F, Ma Y, Chong K, Xu Y Y.2013. Overexpression of OrbHLH001, a putative helix-loop-helix transcription factor, causes increased expression of AKT1 and maintains ionic balance under salt stress in rice. J Plant Physiol, 170(1): 93-100. |
| [19] | Choe Y H, Kim Y S, Kim I S, Bae M J, Lee E J, Kim Y H, Park H M, Yoon H S.2013. Homologous expression of γ-glutamylcysteine synthetase increases grain yield and tolerance of transgenic rice plants to environmental stresses. J Plant Physiol, 170(6): 610-618. |
| [20] | Choi W G, Toyota M, Kim S H, Hilleary R, Gilroy S.2014. Salt stress-induced Ca2+ waves are associated with rapid, long- distance root-to-shoot signaling in plants. Proc Natl Acad Sci USA, 111(17): 6497-6502. |
| [21] | Choulet F, Alberti A, Theil S, Glover N, Barbe V, Daron J, Pingault L, Sourdille P, Couloux A, Paux E, Leroy P, Mangenot S, Guilhot N, Le Gouis J, Balfourier F, Alaux M, Jamilloux V, Poulain J, Durand C, Bellec A, Gaspin C, Safar J, Dolezel J, Rogers J, Vandepoele K, Aury J M, Mayer K, Berges H, Quesneville H, Wincker P, Feuillet C.2014. Structural and functional partitioning of bread wheat chromosome 3B. Science, 345: 1249721. |
| [22] | Chowdhury A D, Haritha G, Sunitha T, Krishnamurthy S L, Divya B, Padmavathi G, Ram T, Sarla N.2016. Haplotyping of rice genotypes using simple sequence repeat markers associated with salt tolerance. Rice Sci, 23(6): 317-325. |
| [23] | Cutler S R, Rodriguez P L, Finkelstein R R, Abrams S R.2010. Abscisic acid: Emergence of a core signaling network. Annu Rev Plant Biol, 61: 651-679. |
| [24] | Danquah A, de Zelicourt A, Colcombet J, Hirt H.2014. The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol Adv, 32(1): 40-52. |
| [25] | Das B, Sengupta S, Parida S K, Roy B, Ghosh M, Prasad M, Ghose T K.2013. Genetic diversity and population structure of rice landraces from eastern and northeastern states of India. BMC Genet, 14: 71. |
| [26] | Davierwala A P, Chowdari K V, Kumar S, Reddy A P K, Ranjekar P K, Gupta V S.2000. Use of three different marker systems to estimate genetic diversity of Indian elite rice varieties. Genetica, 108: 269-284. |
| [27] | de Nadal E, Clotet J, Posas F, Serrano R, Gomez N, Ariño J.1998. The yeast halotolerance determinant Hal3p is an inhibitory subunit of the Ppz1p Ser/Thr protein phosphatase. Proc Natl Acad Sci USA, 95(13): 7357-7362. |
| [28] | Delauney A J, Verma D P S.1993. Proline biosynthesis and osmoregulation in plants. Plant J, 4(2): 215-223. |
| [29] | Dey A, Samanta M K, Gayen S, Maiti M K.2016. The sucrose non-fermenting 1-related kinase 2 gene SAPK9 improves drought tolerance and grain yield in rice by modulating cellular osmotic potential, stomatal closure and stress-responsive gene expression. BMC Plant Biol, 16: 158. |
| [30] | Dhindsa R S, Plumb-Dhindsa P L, Reid D M.1982. Leaf senescence and lipid peroxidation: Effects of some phytohormones, and scavengers of free radicals and singlet oxygen. Physiol Plant, 56(4): 453-457. |
| [31] | di Como C J, Bose R, Arndt K T.1995. Overexpression of SIS2, which contains an extremely acidic region, increases the expression of SWI4, CLN1 and CLN2 in sit4 mutants. Genetics, 139(1): 95-107. |
| [32] | Dichtl B, Stevens A, Tollervey D.1997. Lithium toxicity in yeast is due to the inhibition of RNA processing enzymes. EMBO J, 16(23): 7184-7195. |
| [33] | Diédhiou C J, Popova O V, Dietz K J, Golldack D.2008. The SNF1-type serine-threonine protein kinase SAPK4 regulates stress-responsive gene expression in rice. BMC Plant Biol, 8: 49. |
| [34] | Duan J Z, Zhang M H, Zhang H L, Xiong H Y, Liu P L, Ali J, Li J J, Li Z C.2012. OsMIOX, a myo-inositol oxygenase gene, improves drought tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). Plant Sci, 196: 143-151. |
| [35] | Dubouzet J G, Sakuma Y, Ito Y, Kasuga M, Dubouzet E G, Miura S, Seki M, Shinozaki K, Yamaguchi-Shinozaki K.2003. OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought-, high-salt- and cold- responsive gene expression. Plant J, 33(4): 751-763. |
| [36] | Espinosa-Ruiz A, Bellés J M, Serrano R, Culiáñez-Macià F A.1999. Arabidopsis thaliana AtHAL3: A flavoprotein related to salt and osmotic tolerance and plant growth. Plant J, 20(5): 529-539. |
| [37] | Ferrando A, Kron S J, Rios G, Fink G R, Serrano R.1995. Regulation of cation transport in Saccharomyces cerevisiae by the salt tolerance gene HAL3. Mol Cell Biol, 15(10): 5470-5481. |
| [38] | Flowers T J.2004. Improving crop salt tolerance. J Exp Bot, 55: 307-319. |
| [39] | Forment J, Mulet J M, Vicente O, Serrano R.2002. The yeast SR protein kinase Sky1p modulates salt tolerance, membrane potential and the Trk1,2 potassium transporter. Biochim Biophys Acta, 1565(1): 36-40. |
| [40] | Fukuda A, Nakamura A, Tagiri A, Tanaka H, Miyao A, Hirochika H, Tanaka Y.2004. Function, intracellular localization and the importance in salt tolerance of a vacuolar Na+/H+ antiporter from rice. Plant Cell Physiol, 45: 146-159. |
| [41] | Ganie S A, Molla K A, Henry R J, Bhat K V, Mondal T K.2019. Advances in understanding salt tolerance in rice. Theor Appl Genet, 132: 851-870. |
| [42] | Gaxiola R, de Larrinoa I F, Villalba J M, Serrano R.1992. A novel and conserved salt-induced protein is an important determinant of salt tolerance in yeast. EMBO J, 11(9): 3157-3164. |
| [43] | Gaxiola R A, Rao R, Sherman A, Grisafi P, Alper S L, Fink G R, 1999. The Arabidopsis thaliana proton transporters, AtNhx1 and Avp1, can function in cation detoxification in yeast. Proc Natl Acad Sci USA, 96(4): 1480-1485. |
| [44] | Genard H, Le Saos J, Billard J P, Tremolieres A, Boucaud J.1991. Effect of salinity on lipid composition, glycine betaine content and photosynthetic activity in chloroplasts of Suaeda maritima. Plant Physiol Biochem, 29(5): 421-427. |
| [45] | Gil-Mascarell R, López-Coronado J M, Bellés J M, Serrano R, Rodríguez P L.1999. The Arabidopsis HAL2-like gene family includes a novel sodium-sensitive phosphatase. Plant J, 17(4): 373-383. |
| [46] | Gisbert C, Rus A M, Boları́n M C, López-Coronado J M, Arrillaga I, Montesinos C, Caro M, Serrano R, Moreno V.2000. The yeast HAL1 gene improves salt tolerance of transgenic tomato. Plant Physiol, 123: 393-402. |
| [47] | Gläser H U, Thomas D, Gaxiola R, Montrichard F, Surdin-Kerjan Y, Serrano R.1993. Salt tolerance and methionine biosynthesis in Saccharomyces cerevisiae involve a putative phosphatase gene. EMBO J, 12(8): 3105-3110. |
| [48] | Grattan S R, Zeng L H, Shannon M C, Roberts S R.2002. Rice is more sensitive to salinity than previously thought. Calif Agric, 56: 189-198. |
| [49] | Gumi A M, Guha P K, Mazumder A, Jayaswal P, Mondal T K.2018. Characterization of OglDREB2A gene from African rice (Oryza glaberrima), comparative analysis and its transcriptional regulation under salinity stress. 3 Biotech, 8: 91. |
| [50] | Guo N, Wang M X, Xue C C, Xue D, Xu J Y, Wang H T, Gai J Y, Xing H, Zhao J M.2016. Over-expression of GmHAL3 modulates salt stresses tolerance in transgenic Arabidopsis. J Plant Biol, 59: 444-455. |
| [51] | Guo X L, Wu Y R, Wang Y Q, Chen Y M, Chu C C.2009. OsMSRA4.1 and OsMSRB1.1, two rice plastidial methionine sulfoxide reductases, are involved in abiotic stress responses. Planta, 230: 227-238. |
| [52] | Gupta B, Huang B R. 2014. Mechanism of salinity tolerance in plants: Physiological, biochemical, and molecular characterization. Int J Genom, 2014: 701596. |
| [53] | Gustin M C, Zhou X L, Martinac B, Kung C.1988. A mechanosensitive ion channel in the yeast plasma membrane. Science, 242: 762-765. |
| [54] | Hare P D, Cress W A, van Staden J.2002. Disruptive effects of exogenous proline on chloroplast and mitochondrial ultrastructure in Arabidopsis leaves. South Afr J Bot, 68(3): 393-396. |
| [55] | Harinasut P, Tsutsui K, Takabe T, Nomura M, Takabe T, Kishitani S.1996. Exogenous glycinebetaine accumulation and increased salt-tolerance in rice seedlings. Biosci Biotechnol Biochem, 60: 366-368. |
| [56] | Hasthanasombut S, Paisarnwipatpong N, Triwitayakorn K, Kirdmanee C, Supaibulwatana K.2011. Expression of OsBADH1 gene in indica rice (Oryza sativa L.) in correlation with salt, plasmolysis, temperature and light stresses. Plant Omics, 4(7): 400-407. |
| [57] | Hmida-Sayari A, Gargouri-Bouzid R, Bidani A, Jaoua L, Savouré A, Jaoua S.2005. Overexpression of Δ1-pyrroline-5-carboxylate synthetase increases proline production and confers salt tolerance in transgenic potato plants. Plant Sci, 169(4): 746-752. |
| [58] | Hong C Y, Hsu Y T, Tsai Y C, Kao C H.2007. Expression of ascorbate peroxidase 8 in roots of rice (Oryza sativa L.) seedlings in response to NaCl. J Exp Bot, 58(12): 3273-3283. |
| [59] | Hong C Y, Chao Y Y, Yang M Y, Cheng S Y, Cho S C, Kao C H.2009. NaCl-induced expression of glutathione reductase in roots of rice (Oryza sativa L.) seedlings is mediated through hydrogen peroxide but not abscisic acid. Plant Soil, 320: 103-115. |
| [60] | Hong Z L, Lakkineni K, Zhang Z M, Verma D P S.2000. Removal of feedback inhibition of Δ1-pyrroline-5-carboxylate synthetase results in increased proline accumulation and protection of plants from osmotic stress. Plant Physiol, 122(4): 1129-1136. |
| [61] | Horie T, Sugawara M, Okada T, Taira K, Kaothien Nakayama P, Katsuhara M, Shinmyo A, Nakayama H.2011. Rice sodium- insensitive potassium transporter, OsHAK5, confers increased salt tolerance in tobacco BY2 cells. J Biosci Bioeng, 111: 346-356. |
| [62] | Hu H H, Dai M Q, Yao J L, Xiao B Z, Li X H, Zhang Q F, Xiong L Z.2006. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA, 103: 12987-12992. |
| [63] | Hu H H, You J, Fang Y J, Zhu X Y, Qi Z Y, Xiong L Z.2008. Characterization of transcription factor gene SNAC2 conferring cold and salt tolerance in rice. Plant Mol Biol, 67: 169-181. |
| [64] | Huang S W, Li R Q, Zhang Z H, Li L, Gu X F, Fan W, Lucas W J, Wang X W, Xie B Y, Ni P X, Ren Y Y, Zhu H M, Li J, Lin K, Jin W W, Fei Z J, Li G C, Staub J, Kilian A, van der Vossen E A G, Wu Y, Guo J, He J, Jia Z Q, Ren Y, Tian G, Lu Y, Ruan J, Qian W B, Wang M W, Huang Q F, Li B, Xuan Z L, Cao J J, Asan Wu Z G, Zhang J B, Cai Q L, Bai Y Q, Zhao B W, Han Y H, Li Y, Li X F, Wang S H, Shi Q X, Liu S Q, Cho W K, Kim J Y, Xu Y, Heller-Uszynska K, Miao H, Cheng Z C, Zhang S P, Wu J, Yang Y H, Kang H X, Li M, Liang H Q, Ren X L, Shi Z B, Wen M, Jian M, Yang H L, Zhang G J, Yang Z T, Chen R, Liu S F, Li J W, Ma L J, Liu H, Zhou Y, Zhao J, Fang X D, Li G Q, Fang L, Li Y R, Liu D Y, Zheng H K, Zhang Y, Qin N, Li Z, Yang G H, Yang S, Bolund L, Kristiansen K, Zheng H C, Li S C, Zhang X Q, Yang H M, Wang J, Sun R F, Zhang B X, Jiang S Z, Wang J, Du Y C, Li S G.2009. The genome of the cucumber, Cucumis sativus L. Nat Genet, 41: 1275-1281. |
| [65] | Huang X Y, Chao D Y, Gao J P, Zhu M Z, Shi M, Lin H X.2009. A previously unknown zinc finger protein, DST, regulates drought and salt tolerance in rice via stomatal aperture control. Genes Dev, 23(15): 1805-1817. |
| [66] | Jamil A, Riaz S, Ashraf M, Foolad M R.2011. Gene expression profiling of plants under salt stress. Crit Rev Plant Sci, 30: 435-458. |
| [67] | Jan A, Maruyama K, Todaka D, Kidokoro S, Abo M, Yoshimura E, Shinozaki K, Nakashima K, Yamaguchi-Shinozaki K.2013. OsTZF1, a CCCH-tandem zinc finger protein, confers delayed senescence and stress tolerance in rice by regulating stress- related genes. Plant Physiol, 161(3): 1202-1216. |
| [68] | Jeong M J, Lee S K, Kim B G, Kwon T R, Cho W S, Park Y T, Lee J O, Kwon H B, Byun M O, Park S C.2006. A rice (Oryza sativa L.) MAP kinase gene, OsMAPK44, is involved in response to abiotic stresses. Plant Cell Tissue Organ Cult, 85: 151-160. |
| [69] | Jiang Z H, Zhou X P, Tao M, Yuan F, Liu L L, Wu F H, Wu X M, Xiang Y, Niu Y, Liu F, Li C J, Ye R, Byeon B, Xue Y, Zhao H Y, Wang H N, Crawford B M, Johnson D M, Hu C X, Pei C, Zhou W M, Swift G B, Zhang H, Vo-Dinh T, Hu Z L, Siedow J N, Pei Z M.2019. Plant cell-surface GIPC sphingolipids sense salt to trigger Ca2+ influx. Nature, 572: 341-346. |
| [70] | Jones-Rhoades M W, Bartel D P.2004. Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell, 14(6): 787-799. |
| [71] | Jones R G W, Gorham J. 1983. Osmoregulation BT: Physiological Plant Ecology III: Responses to the chemical and biological environment. In: Lange O L, Nobel P S, Osmond C B, Ziegler H. Physiological Plant Ecology III. Springer Berlin Heidelberg, Berlin, Heidelberg: 35-58. |
| [72] | Kanawapee N, Sanitchon J, Srihaban P, Theerakulpisut P.2011. Genetic diversity analysis of rice cultivars (Oryza sativa L.) differing in salinity tolerance based on RAPD and SSR markers. Electron J Biotechnol, 14: 6. |
| [73] | Kang D J, Seo Y J, Lee J D, Ishii R, Kim K U, Shin D H, Park S K, Jang S W, Lee I J.2005. Jasmonic acid differentially affects growth, ion uptake and abscisic acid concentration in salt- tolerant and salt-sensitive rice cultivars. J Agron Crop Sci, 191(4): 273-282. |
| [74] | Karthikeyan A, Pandian S K, Ramesh M.2011. Transgenic indica rice cv. ADT43 expressing a Δ1-pyrroline-5-carboxylate synthetase (P5CS) gene from Vigna aconitifolia demonstrates salt tolerance. Plant Cell Tissue Organ Cult, 107: 383-395. |
| [75] | Kim B G, Waadt R, Cheong Y H, Pandey G K, Dominguez-Solis J R, Schültke S, Lee S C, Kudla J, Luan S.2007. The calcium sensor CBL10 mediates salt tolerance by regulating ion homeostasis in Arabidopsis. Plant J, 52(3): 473-484. |
| [76] | Kishor P B K, Hong Z, Miao G H, Hu C A A, Verma D P S.1995. Overexpression of Δ1-pyrroline-5-carboxylate synthetase increases proline production and confers osmotolerance in transgenic plants. Plant Physiol, 108: 1387-1394. |
| [77] | Kishor P B K, Sangam S, Amrutha R N, Laxmi P S, Naidu K R, Rao K R S S, Rao S, Reddy K J, Theriappan P, Sreenivasulu N.2005. Regulation of proline biosynthesis, degradation, uptake and transport in higher plants: Its implications in plant growth and abiotic stress tolerance. Curr Sci, 88(3): 424-438. |
| [78] | Kobayashi Y, Yamamoto S, Minami H, Kagaya Y, Hattori T.2004. Differential activation of the rice sucrose nonfermenting1-related protein kinase2 family by hyperosmotic stress and abscisic acid. Plant Cell, 16(5): 1163-1177. |
| [79] | Komis G, Šamajová O, Ovečka M, Šamaj J.2018. Cell and developmental biology of plant mitogen-activated protein kinases. Annu Rev Plant Biol, 69: 237-265. |
| [80] | Kumar G, Kushwaha H R, Purty R S, Kumari S, Singla Pareek S L, Pareek A.2012. Cloning, structural and expression analysis of OsSOS2 in contrasting cultivars of rice under salinity stress. Genes Genom Genom, 6: 34-41. |
| [81] | Kumar S, Dhingra A, Daniell H.2004. Plastid-expressed betaine aldehyde dehydrogenase gene in carrot cultured cells, roots, and leaves confers enhanced salt tolerance. Plant Physiol, 136(1): 2843-2854. |
| [82] | Kumar S K, Sivanesan I, Murugesan K, Jeong B R, Hwang S J.2014. Enhancing salt tolerance in eggplant by introduction of foreign halotolerance gene, HAL1 isolated from yeast. Hort Environ Biotechnol, 55: 222-229. |
| [83] | Kumar V, Shriram V, Kavi Kishor P B, Jawali N, Shitole M G.2010. Enhanced proline accumulation and salt stress tolerance of transgenic indica rice by over-expressing P5CSF129A gene. Plant Biotechnol Rep, 4: 37-48. |
| [84] | Kwon H M, Handler J S.1995. Cell volume regulated transporters of compatible osmolytes. Curr Opin Cell Biol, 7(4): 465-471. |
| [85] | Larkin M A, Blackshields G, Brown N P, Chenna R, McGettgan P A, McWilliam H, Valentin F, Wallace I M, Wilm A, Lopez R, Thompson J D, Gibson T J, Higgins D G.2007. Clustal W and Clustal X version 2.0. Bioinformatics, 23(21): 2947-2948. |
| [86] | Lee S K, Kim B G, Kwon T R, Jeong M J, Park S R, Lee J W, Byun M O, Kwon H B, Matthews B F, Hong C B, Park S C.2011. Overexpression of the mitogen-activated protein kinase gene OsMAPK33 enhances sensitivity to salt stress in rice (Oryza sativa L.). J Biosci, 36: 139-151. |
| [87] | Li M F, Guo S J, Xu Y, Meng Q W, Li G, Yang X H.2014. Glycine betaine-mediated potentiation of HSP gene expression involves calcium signaling pathways in tobacco exposed to NaCl stress. Physiol Plant, 150(1): 63-75. |
| [88] | Lisa L A, Seraj Z I, Fazle Elahi C M, Das K C, Biswas K, Islam M R, Salam M A, Gomosta A R.2004. Genetic variation in microsatellite DNA, physiology and morphology of coastal saline rice (Oryza sativa L.) landraces of Bangladesh. Plant Soil, 263: 213-228. |
| [89] | Lisar S Y S, Motafakkerazad R, Hossain M M, Rahman I M M. 2012. Water stress in plants: Causes, effects and responses. In: Rahman I M M, Hasegawa H. Water Stress. Rijeka, Croatia: Intech: 1-14. |
| [90] | Liu C T, Mao B G, Ou S J, Wang W, Liu L C, Wu Y B, Chu C C, Wang X P.2014. OsbZIP71, a bZIP transcription factor, confers salinity and drought tolerance in rice. Plant Mol Biol, 84: 19-36. |
| [91] | Liu K M, Wang L, Xu Y Y, Chen N, Ma Q B, Li F, Chong K.2007. Overexpression of OsCOIN, a putative cold inducible zinc finger protein, increased tolerance to chilling, salt and drought, and enhanced proline level in rice. Planta, 226: 1007-1016. |
| [92] | Liu S P, Zheng L Q, Xue Y H, Zhang Q, Wang L, Shou H X.2010. Overexpression of OsVP1 and OsNHX1 increases tolerance to drought and salinity in rice. J Plant Biol, 53: 444-452. |
| [93] | Lu Z Q, Liu D L, Liu S K.2007. Two rice cytosolic ascorbate peroxidases differentially improve salt tolerance in transgenic Arabidopsis. Plant Cell Rep, 26: 1909-1917. |
| [94] | Luo D, Niu X L, Yu J D, Yan J, Gou X J, Lu B R, Liu Y S.2012. Rice choline monooxygenase (OsCMO) protein functions in enhancing glycine betaine biosynthesis in transgenic tobacco but does not accumulate in rice (Oryza sativa L. ssp. japonica). Plant Cell Rep, 31: 1625-1635. |
| [95] | Lutts S.2000. Exogenous glycinebetaine reduces sodium accumulation in salt-stressed rice plants. Int Rice Res Notes, 25: 39-40. |
| [96] | Lutts S, Majerus V, Kinet J M.1999. NaCl effects on proline metabolism in rice (Oryza sativa) seedlings. Physiol Plant, 105(3): 450-458. |
| [97] | Macovei A, Tuteja N.2012. microRNAs targeting DEAD-box helicases are involved in salinity stress response in rice (Oryza sativa L.). BMC Plant Biol, 12: 183. |
| [98] | Mäkelä P, Munns R, Colmer T D, Condon A G, Peltonen-Sainio P.1998. Effect of foliar applications of glycinebetaine on stomatal conductance, abscisic acid and solute concentrations in leaves of salt- or drought-stressed tomato. Funct Plant Biol, 25(6): 655-663. |
| [99] | Mallikarjuna G, Mallikarjuna K, Reddy M K, Kaul T.2011. Expression of OsDREB2A transcription factor confers enhanced dehydration and salt stress tolerance in rice (Oryza sativa L.). Biotechnol Lett, 33: 1689-1697. |
| [100] | Mansour M M F.2000. Nitrogen containing compounds and adaptation of plants to salinity stress. Biol Plant, 43(4): 491-500. |
| [101] | Martínez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu J K, Pardo J M, Quintero F J.2007. Conservation of the salt overly sensitive pathway in rice. Plant Physiol, 143: 1001-1012. |
| [102] | McNeil S D, Rhodes D, Russell B L, Nuccio M L, Shachar-Hill Y, Hanson A D.2000. Metabolic modeling identifies key constraints on an engineered glycine betaine synthesis pathway in tobacco. Plant Physiol, 124(1): 153-162. |
| [103] | Minhas A, Sharma A, Kaur H, Rawal Y, Ganesan K, Mondal A K.2012. Conserved Ser/Arg-rich motif in PPZ orthologs from fungi is important for its role in cation tolerance. J Biol Chem, 287(10): 7301-7312. |
| [104] | Mukherjee K, Choudhury A R, Gupta B, Gupta S, Sengupta D N.2006. An ABRE-binding factor, OSBZ8, is highly expressed in salt tolerant cultivars than in salt sensitive cultivars of indica rice. BMC Plant Biol, 6: 18. |
| [105] | Mukherjee R, Mukherjee A, Bandyopadhyay S, Mukherjee S, Sengupta S, Ray S, Majumder A L.2019. Selective manipulation of the inositol metabolic pathway for induction of salt-tolerance in indica rice variety. Sci Rep, 9: 5358. |
| [106] | Mulet J M, Leube M P, Kron S J, Rios G, Fink G R, Serrano R.1999. A novel mechanism of ion homeostasis and salt tolerance in yeast: The Hal4 and Hal5 protein kinases modulate the Trk1-Trk2 potassium transporter. Mol Cell Biol, 19(5): 3328-3337. |
| [107] | Munns R, Tester M.2008. Mechanisms of salinity tolerance. Annu Rev Plant Biol, 59: 651-681. |
| [108] | Munson A M, Haydon D H, Love S L, Fell G L, Palanivel V R, Rosenwald A G.2004. Yeast ARL1 encodes a regulator of K+ influx. J Cell Sci, 117: 2309-2320. |
| [109] | Mustilli A C, Merlot S, Vavasseur A, Fenzi F, Giraudat J.2002. Arabidopsis OST1 protein kinase mediates the regulation of stomatal aperture by abscisic acid and acts upstream of reactive oxygen species production. Plant Cell, 14(12): 3089-3099. |
| [110] | Mutum R D, Balyan S C, Kansal S, Agarwal P, Kumar S, Kumar M, Raghuvanshi S.2013. Evolution of variety-specific regulatory schema for expression of osa-miR408 in indica rice varieties under drought stress. FEBS J, 280(7): 1717-1730. |
| [111] | Nakashima K, Tran L S P, Van Nguyen D, Fujita M, Maruyama K, Todaka D, Ito Y, Hayashi N, Shinozaki K, Yamaguchi- Shinozaki K.2007. Functional analysis of a NAC-type transcription factor OsNAC6 involved in abiotic and biotic stress-responsive gene expression in rice. Plant J, 51(4): 617-630. |
| [112] | Nam M H, Huh S M, Kim K M, Park W J, Seo J B, Cho K, Kim D Y, Kim B G, Yoon I S.2012. Comparative proteomic analysis of early salt stress-responsive proteins in roots of SnRK2 transgenic rice. Proteom Sci, 10: 25. |
| [113] | Nuccio M L, Russell B L, Nolte K D, Rathinasabapathi B, Gage D A, Hanson A D.1998. The endogenous choline supply limits glycine betaine synthesis in transgenic tobacco expressing choline monooxygenase. Plant J, 16(4): 487-496. |
| [114] | Obata T, Kitamoto H K, Nakamura A, Fukuda A, Tanaka Y.2007. Rice shaker potassium channel OsKAT1 confers tolerance to salinity stress on yeast and rice cells. Plant Physiol, 144: 1978-1985. |
| [115] | Ouyang S Q, He S J, Liu P, Zhang W K, Zhang J S, Chen S Y.2011. The role of tocopherol cyclase in salt stress tolerance of rice (Oryza sativa). Sci China Life Sci, 54: 181-188. |
| [116] | Park J M, Park C J, Lee S B, Ham B K, Shin R, Paek K H.2001. Overexpression of the tobacco TSI1 gene encoding an EREBP/ AP2-type transcription factor enhances resistance against pathogen attack and osmotic stress in tobacco. Plant Cell, 13(5): 1035-1046. |
| [117] | Patel B B, Bharat B, Dave R S.2011. Studies on infiltration of saline-alkali soils of several parts of Mehsana and Patan districts of North Gujarat. J Appl Technol Environ Sanit, 1: 87-92. |
| [118] | Peng Z, Verma D P.1995. A rice HAL2-like gene encodes a Ca2+-sensitive 3′(2′),5′-diphosphonucleoside 3′(2′)-phosphohydrolase and complements yeast met22 and Escherichia coli cysQ mutations. J Biol Chem, 270: 29105-29110. |
| [119] | Pérez-Valle J, Jenkins H, Merchan S, Montiel V, Ramos J, Sharma S, Serrano R, Yenush L.2007. Key role for intracellular K+ and protein kinases Sat4/Hal4 and Hal5 in the plasma membrane stabilization of yeast nutrient transporters. Mol Cell Biol, 27(16): 5725-5736. |
| [120] | Posas F, Camps M, Ario J.1995. The PPZ protein phosphatases are important determinants of salt tolerance in yeast cells. J Biol Chem, 270: 13036-13041. |
| [121] | Prasad K V S K, Sharmila P, Kumar P A, Saradhi P P.2000. Transformation of Brassica juncea L. Czern with bacterial codA gene enhances its tolerance to salt stress. Mol Breeding, 6: 489-499. |
| [122] | Qiu D Y, Xiao J, Xie W B, Liu H B, Li X H, Xiong L Z, Wang S P.2008. Rice gene network inferred from expression profiling of plants overexpressing OsWRKY13, a positive regulator of disease resistance. Mol Plant, 1(3): 538-551. |
| [123] | Qiu D Y, Xiao J, Xie W B, Cheng H T, Li X H, Wang S P.2009. Exploring transcriptional signaling mediated by OsWRKY13, a potential regulator of multiple physiological processes in rice. BMC Plant Biol, 9: 74. |
| [124] | Quintero F J, Garciadeblás B, Rodríguez-Navarro A.1996. The SAL1 gene of Arabidopsis, encoding an enzyme with 3'(2'), 5'-bisphosphate nucleotidase and inositol polyphosphate 1-phosphatase activities, increases salt tolerance in yeast. Plant Cell, 8(3): 529-537. |
| [125] | Ranganayakulu G S, Veeranagamallaiah G, Sudhakar C.2013. Effect of salt stress on osmolyte accumulation in two groundnut cultivars (Arachis hypogaea L.) with contrasting salt tolerance. Afr J Plant Sci, 7: 586-592. |
| [126] | Reddy I N B L, Kim B K, Yoon I S, Kim K H, Kwon T R.2017. Salt tolerance in rice: Focus on mechanisms and approaches. Rice Sci, 24(3): 123-144. |
| [127] | Rhodes D, Hanson A D.1993. Quaternary ammonium and aertiary sulfonium compounds in higher plants. Annu Rev Plant Physiol Plant Mol Biol, 44: 357-384. |
| [128] | Rodriguez P L, Ali R, Serrano R.1996. CtCdc55p and CtHal3p: Two putative regulatory proteins from Candida tropicalis with long acidic domains. Yeast, 12(13): 1321-1329. |
| [129] | Roy D, Basu N, Bhunia A, Banerjee S K.1993. Counteraction of exogenous L-proline with NaCl in salt-sensitive cultivar of rice. Biol Plant, 35: 69. |
| [130] | Šamajová O, Komis G, Šamaj J.2013a. Emerging topics in the cell biology of mitogen-activated protein kinases. Trends Plant Sci, 18(3): 140-148. |
| [131] | Šamajová O, Plíhal O, Al-Yousif M, Hirt H, Šamaj J.2013b. Improvement of stress tolerance in plants by genetic manipulation of mitogen-activated protein kinases. Biotechnol Adv, 31(1): 118-128. |
| [132] | Schmidt R, Mieulet D, Hubberten H M, Obata T, Hoefgen R, Fernie A R, Fisahn J, San Segundo B, Guiderdoni E, Schippers J H M, Mueller-Roeber B.2013. SALT-RESPONSIVE ERF1 regulates reactive oxygen species dependent signaling during the initial response to salt stress in rice. Plant Cell, 25: 2115-2131. |
| [133] | Schnable P S, Ware D, Fulton R S, Stein J C, Wei F S, Pasternak S, Liang C Z, Zhang J W, Fulton L, Graves TA, Minx P, Reily A D, Courtney L, Kruchowski S S, Tomlinson C, Strong C, Delehaunty K, Fronick C, Courtney B, Rock S M, Belter E, Du F Y, Kim K, Abbott R M, Cotton M, Levy A, Marchetto P, Ochoa K, Jackson S M, Gillam B, Chen W Z, Yan L, Higginbotham J, Cardenas M, Waligorski J, Applebaum E, Phelps L, Falcone J, Kanchi K, Thane T, Scimone A, Thane N, Henke J, Wang T, Ruppert J, Shah N, Rotter K, Hodges J, Ingenthron E, Cordes M, Kohlberg S, Sgro J, Delgado B, Mead K, Chinwalla A, Leonard S, Crouse K, Collura K, Kudrna D, Currie J, He R F, Angelova A, Rajasekar S, Mueller T, Lomeli R, Scara G, Ko A, Delaney K, Wissotski M, Lopez G, Campos D, Braidotti M, Ashley E, Golser W, Kim H, Lee S, Lin J K, Dujmic Z, Kim W, Talag J, Zuccolo A, Fan C Z, Sebastian A, Kramer M, Spiegel L, Nascimento L, Zutavern T, Miller B, Ambroise C, Muller S, Spooner W, Narechania A, Ren L Y, Wei S, Kumari S, Faga B, Levy M J, McMahan L, Van Buren P, Vaughn M W, Ying K, Yeh C T, Emrich S J, Jia Y, Kalyanaraman A, Hsia A P, Barbazuk W B, Baucom R S, Brutnell T P, Carpita N C, Chaparro C, Chia J M, Deragon J M, Estill J C, Fu Y, Jeddeloh J A, Han Y J, Lee H, Li P H, Lisch D R, Liu S Z, Liu Z J, Nagel D H, McCann M C, SanMiguel P, Myers A M, Nettleton D, Nguyen J, Penning B W, Ponnala L, Schneider K L, Schwartz D C, Sharma A, Soderlund C, Springer N M, Sun Q, Wang H, Waterman M, Westerman R, Wolfgruber T K, Yang L X, Yu Y, Zhang L F, Zhou S G, Zhu Q H, Bennetzen J L, Dawe R K, Jiang J M, Jiang N, Presting G G, Wessler S R, Aluru S, Martienssen R A, Clifton S W, McCombie W R, Wing R A, Wilson R K.2009. The B73 maize genome: Complexity, diversity, and dynamics. Science, 326: 1112-1115. |
| [134] | Serraj R, Sinclair T R.2002. Osmolyte accumulation: Can it really help increase crop yield under drought conditions? Plant Cell Environ, 25(2): 333-341. |
| [135] | Serrano R.1996. Salt tolerance in plants and microorganisms: Toxicity targets and defense responses. Int Rev Cytol, 165: 1-52. |
| [136] | Serrano R, Gaxiola R.1994. Microbial models and salt stress tolerance in plants. Crit Rev Plant Sci, 13: 121-138. |
| [137] | Shahbaz M, Ashraf M.2013. Improving salinity tolerance in cereals. Crit Rev Plant Sci, 32: 237-249. |
| [138] | Sharma N, Tripathi A, Sanan-Mishra N.2015. Profiling the expression domains of a rice-specific microRNA under stress. Front Plant Sci, 6: 333. |
| [139] | Shiu S H, Karlowski W M, Pan R S, Tzeng Y H, Mayer K F X, Li W H.2004. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell, 16(5): 1220-1234. |
| [140] | Shrivastava P, Kumar R.2015. Soil salinity: A serious environmental issue and plant growth promoting bacteria as one of the tools for its alleviation. Saud J Biol Sci, 22(2): 123-131. |
| [141] | Si-Ammour A, Windels D, Arn-Bouldoires E, Kutter C, Ailhas J, Meins F, Vazquez F.2011. miR393 and secondary siRNAs regulate expression of the TIR1/AFB2 auxin receptor clade and auxin-related development of Arabidopsis leaves. Plant Physiol, 157(2): 683-691. |
| [142] | Singh R K, Mishra B, Gregorio G B.2006. CSR23: A new salt- tolerant rice variety for India. Int Rice Res Notes, 31(1): 16-18. |
| [143] | Su J, Wu R.2004. Stress-inducible synthesis of proline in transgenic rice confers faster growth under stress conditions than that with constitutive synthesis. Plant Sci, 166(4): 941-948. |
| [144] | Sun M Z, Yang J K, Cai X X, Shen Y, Cui N, Zhu Y M, Jia B W, Sun X L.2018. The opposite roles of OsmiR408 in cold and drought stress responses in Oryza sativa. Mol Breeding, 38: 120. |
| [145] | Sun S Y, Chao D Y, Li X M, Shi M, Gao J P, Zhu M Z, Yang H Q, Luan S, Lin H X.2009. OsHAL3 mediates a new pathway in the light-regulated growth of rice. Nat Cell Biol, 11: 845-851. |
| [146] | Takasaki H, Maruyama K, Kidokoro S, Ito Y, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K, Nakashima K.2010. The abiotic stress-responsive NAC-type transcription factor OsNAC5 regulates stress-inducible genes and stress tolerance in rice. Mol Genet Genom, 284: 173-183. |
| [147] | Tang W, Sun J Q, Liu J, Liu F F, Yan J, Gou X J, Lu B R, Liu Y S.2014. RNAi-directed downregulation of betaine aldehyde dehydrogenase 1 (OsBADH1) results in decreased stress tolerance and increased oxidative markers without affecting glycine betaine biosynthesis in rice (Oryza sativa). Plant Mol Biol, 86: 443-454. |
| [148] | Teixeira F K, Menezes-Benavente L, Galvão V C, Margis R, Margis-Pinheiro M.2006. Rice ascorbate peroxidase gene family encodes functionally diverse isoforms localized in different subcellular compartments. Planta, 224: 300-314. |
| [149] | Teng X X, Cao W L, Lan H X, Tang H J, Bao Y M, Zhang H S.2017. OsNHX2, an Na+/H+ antiporter gene, can enhance salt tolerance in rice plants through a more effective accumulation of toxic Na+ in leaf mesophyll and bundle sheath cells. Acta Physiol Plant, 39: 113. |
| [150] | Tran L S P, Nakashima K, Sakuma Y, Simpson S D, Fujita Y, Maruyama K, Fujita M, Seki M, Shinozaki K, Yamaguchi- Shinozaki K.2004. Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell, 16(9): 2481-2498. |
| [151] | Uddin M I, Qi Y, Yamada S, Shibuya I, Deng X P, Kwak S S, Kaminaka H, Tanaka K.2008. Overexpression of a new rice vacuolar antiporter regulating protein OsARP improves salt tolerance in tobacco. Plant Cell Physiol, 49: 880-890. |
| [152] | Umezawa T, Yoshida R, Maruyama K, Yamaguchi-Shinozaki K, Shinozaki K.2004. SRK2C, a SNF1-related protein kinase 2, improves drought tolerance by controlling stress-responsive gene expression in Arabidopsis thaliana. Proc Natl Acad Sci USA, 101: 17306-17311. |
| [153] | Vaidyanathan H, Sivakumar P, Chakrabarty R, Thomas G.2003. Scavenging of reactive oxygen species in NaCl-stressed rice (Oryza sativa L.) differential response in salt-tolerant and sensitive varieties. Plant Sci, 165(6): 1411-1418. |
| [154] | Venkatesan A, Chellappan K P.1998. Accumulation of proline and glycine betaine in Ipomoea pes-caprae induced by NaCl. Biol Plant, 41: 271-276. |
| [155] | Vij S, Giri J, Dansana P K, Kapoor S, Tyagi A K.2008. The receptor-like cytoplasmic kinase (OsRLCK) gene family in rice: Organization, phylogenetic relationship, and expression during development and stress. Mol Plant, 1(5): 732-750. |
| [156] | Wang F B, Liu J C, Zhou L J, Pan G, Li Z W, Zaidi S H R, Cheng F M.2016. Senescence-specific change in ROS scavenging enzyme activities and regulation of various SOD isozymes to ROS levels in psf mutant rice. Plant Physiol Biochem, 109: 248-261. |
| [157] | Wang M C, Zhao X, Xiao Z, Yin X H, Xing T, Xia G M.2016. A wheat superoxide dismutase gene TaSOD2 enhances salt resistance through modulating redox homeostasis by promoting NADPH oxidase activity. Plant Mol Biol, 91: 115-130. |
| [158] | Wei D D, Zhang W, Wang C C, Meng Q W, Li G, Chen T H H, Yang X H.2017. Genetic engineering of the biosynthesis of glycinebetaine leads to alleviate salt-induced potassium efflux and enhances salt tolerance in tomato plants. Plant Sci, 257: 74-83. |
| [159] | Winicov I.2000. Alfin1 transcription factor overexpression enhances plant root growth under normal and saline conditions and improves salt tolerance in alfalfa. Planta, 210: 416-422. |
| [160] | Winicov I, Bastola D R.1999. Transgenic overexpression of the transcription factor alfin1 enhances expression of the endogenous MsPRP2 gene in alfalfa and improves salinity tolerance of the plants. Plant Physiol, 120(2): 473-480. |
| [161] | Xia K F, Wang R, Ou X J, Fang Z M, Tian C G, Duan J, Wang Y Q, Zhang M Y.2012. OsTIR1 and OsAFB2 downregulation via OsmiR393 overexpression leads to more tillers, early flowering and less tolerance to salt and drought in rice. PLoS One, 7(1): e30039. |
| [162] | Xiong L Z, Yang Y N.2003. Disease resistance and abiotic stress tolerance in rice are inversely modulated by an abscisic acid- inducible mitogen-activated protein kinase. Plant Cell, 15(3): 745-759. |
| [163] | Xu D P, Duan X L, Wang B Y, Hong B M, Ho T H D, Wu R.1996. Expression of a late embryogenesis abundant protein gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol, 110: 249-257. |
| [164] | Yancey P H, Clark M E, Hand S C, Bowlus R D, Somero G N.1982. Living with water stress: Evolution of osmolyte systems. Science, 217: 1214-1222. |
| [165] | Yang A, Dai X Y, Zhang W H.2012. A R2R3-type MYB gene, OsMYB2, is involved in salt, cold, and dehydration tolerance in rice. J Exp Bot, 63(7): 2541-2556. |
| [166] | Yang M F, Sun F, Wang S Y, Qi W W, Wang Q J, Dong X X, Yang J S, Luo X J.2013. Down-regulation of OsPDCD5, a homolog of the mammalian PDCD5, increases rice tolerance to salt stress. Mol Breeding, 31: 333-346. |
| [167] | Yang S X, Zhao Y X, Zhang Q, He Y K, Zhang H, Luo D.2001. HAL1 mediate salt adaptation in Arabidopsis thaliana. Cell Res, 11(2): 142-148. |
| [168] | Yang W J, Rich P J, Axtell J D, Wood K V, Bonham C C, Ejeta G, Mickelbart M V, Rhodes D.2003. Genotypic variation for glycinebetaine in sorghum. Crop Sci, 43: 162-169. |
| [169] | Yang X H, Lu C M.2005. Photosynthesis is improved by exogenous glycinebetaine in salt-stressed maize plants. Physiol Plant, 124(3): 343-352. |
| [170] | Yenush L, Mulet J M, Ariño J, Serrano R.2002. The Ppz protein phosphatases are key regulators of K+ and pH homeostasis: Implications for salt tolerance, cell wall integrity and cell cycle progression. EMBO J, 21(5): 920-929. |
| [171] | Yesmin N, Elias S M, Rahman M S, Haque T, Mahbub Hasan A K M, Seraj Z I. 2014. Unique genotypic differences discovered among indigenous Bangladeshi rice landraces. Int J Genom, 2014: 210328. |
| [172] | Yonamine I, Yoshida K, Kido K, Nakagawa A, Nakayama H, Shinmyo A.2004. Overexpression of NtHAL3 genes confers increased levels of proline biosynthesis and the enhancement of salt tolerance in cultured tobacco cells. J Exp Bot, 55: 387-395. |
| [173] | Yoshida R, Hobo T, Ichimura K, Mizoguchi T, Takahashi F, Aronso J, Ecker J R, Shinozaki K.2002. ABA-activated SnRK2 protein kinase is required for dehydration stress signaling in Arabidopsis. Plant Cell Physiol, 43(12): 1473-1483. |
| [174] | You J, Hu H H, Xiong L Z.2012a. An ornithine δ-aminotransferase gene OsOAT confers drought and oxidative stress tolerance in rice. Plant Sci, 197: 59-69. |
| [175] | You J, Zong W, Li X K, Ning J, Hu H H, Li X H, Xiao J H, Xiong L Z.2012b. The SNAC1-targeted gene OsSRO1c modulates stomatal closure and oxidative stress tolerance by regulating hydrogen peroxide in rice. J Exp Bot, 64: 569-583. |
| [176] | Yuan S R, Li Z G, Li D Y, Yuan N, Hu Q, Luo H.2015. Constitutive expression of rice microRNA528 alters plant development and enhances tolerance to salinity stress and nitrogen starvation in creeping bentgrass. Plant Physiol, 169(1): 576-593. |
| [177] | Zhang B H, Pan X P, Cannon C H, Cobb G P, Anderson T A.2006. Conservation and divergence of plant microRNA genes. Plant J, 46(2): 243-259. |
| [178] | Zhang B H, Wang Q L, Pan X P.2007. MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol, 210(2): 279-289. |
| [179] | Zhang N, Wang X C, Chen J.2009. Role of OsHAL3 protein, a putative 4-phosphopantothenoylcysteine decarboxylase in rice. Biochemistry (Mose), 74: 61-67. |
| [180] | Zhang S X, Haider I, Kohlen W, Jiang L, Bouwmeester H, Meijer A H, Schluepmann H, Liu C M, Ouwerkerk P B F.2012. Function of the HD-Zip I gene Oshox22 in ABA-mediated drought and salt tolerances in rice. Plant Mol Biol, 80: 571-585. |
| [181] | Zhao B T, Liang R Q, Ge L F, Li W, Xiao H S, Lin H X, Ruan K C, Jin Y X.2007. Identification of drought-induced microRNAs in rice. Biochem Biophys Res Commun, 354: 585-590. |
| [182] | Zhou M, Li D Y, Li Z G, Hu Q, Yang C H, Zhu L H, Luo H.2013. Constitutive expression of a miR319 gene alters plant development and enhances salt and drought tolerance in transgenic creeping bentgrass. Plant Physiol, 161(3): 1375-1391. |
| [183] | Zhou Y B, Liu C, Tang D Y, Yan L, Wang D, Yang Y Z, Gui J S, Zhao X Y, Li L G, Tang X D, Yu F, Li J L, Liu L L, Zhu Y H, Lin J Z, Liu X M.2018. The receptor-like cytoplasmic kinase STRK1 phosphorylates and activates CatC, thereby regulating H2O2 homeostasis and improving salt tolerance in rice. Plant Cell, 30(5): 1100-1118. |
| [184] | Zou C S, Li L T, Miki D, Li D L, Tang Q M, Xiao L H, Rajput S, Deng P, Peng L, Jia W, Huang R, Zhang M L, Sun Y D, Hu J M, Fu X, Schnable P S, Chang Y X, Li F, Zhang Hui, Feng B L, Zhu X G, Liu R Y, Schnable J C, Zhu J K, Zhang H.2019. The genome of broomcorn millet. Nat Commun, 10: 436. |
| [185] | Zou M J, Guan Y C, Ren H B, Zhang F, Chen F.2008. A bZIP transcription factor, Osabi5, is involved in rice fertility and stress tolerance. Plant Mol Biol, 66: 675-683. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||