
Rice Science ›› 2020, Vol. 27 ›› Issue (5): 384-395.DOI: 10.1016/j.rsci.2020.03.003
• Orginal Article • Previous Articles Next Articles
Caixia Gao, Xiuwen Zheng, Hubo Li, Ali Mlekwa Ussi, Yu Gao, Jie Xiong( )
)
Received:2019-10-12
Accepted:2020-03-20
Online:2020-09-28
Published:2020-09-28
Caixia Gao, Xiuwen Zheng, Hubo Li, Ali Mlekwa Ussi, Yu Gao, Jie Xiong. Roles of lncRNAs in Rice: Advances and Challenges[J]. Rice Science, 2020, 27(5): 384-395.
Add to citation manager EndNote|Ris|BibTeX
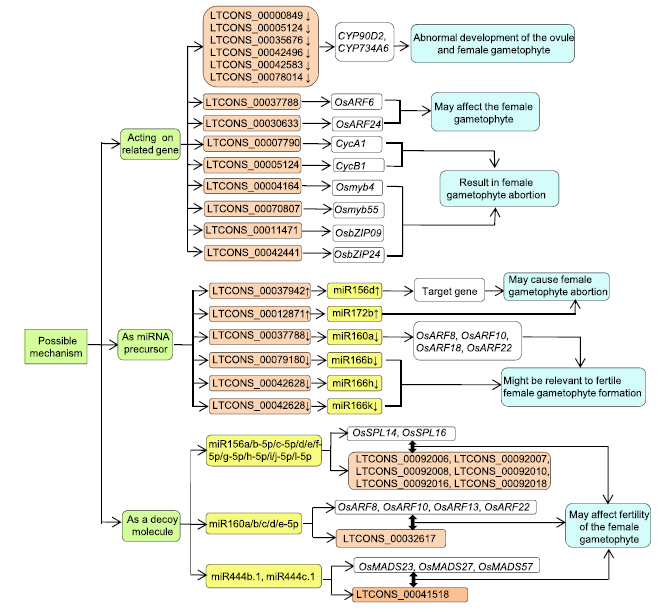
Fig. 4. Possible mechanisms of lncRNAs involved in ovule development and female ligand abortion.Light green boxes, Possible mechanisms; Light pink boxes, lncRNAs; Light yellow boxes, miRNAs; White boxes, Target genes; Light blue boxes, Possible biological functions; Black double arrow, Compete; ↑, Upregulation; ↓, Downregulation.
| [1] | Amor B B, Wirth S, Merchan F, Laporte P, d’Aubenton-Carafa Y, Hirsch J, Maizel A, Mallory A, Lucas A, Deragon J M, Vaucheret H, Thermes C, Crespi M.2009. Novel long non- protein coding RNAs involved in Arabidopsis differentiation and stress responses. Genom Res, 19(1): 57-69. |
| [2] | Ariel F, Jegu T, Latrasse D, Romero-Barrios N, Christ A, Benhamed M, Crespi M.2014. Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin- driven chromatin loop. Mol Cell, 55(3): 383-396. |
| [3] | Baker C C, Sieber P, Wellmer F, Meyerowitz E M.2005. The early extra petals1 mutant uncovers a role for microRNA miR164c in regulating petal number in Arabidopsis. Curr Biol, 15(4): 303-315. |
| [4] | Bardou F, Ariel F, Simpson C G, Romero-Barrios N, Laporte P, Balzergue S, Brown J W S, Crespi M.2014. Long noncoding RNA modulates alternative splicing regulators in Arabidopsis. Dev Cell, 30(2): 166-176. |
| [5] | Berry S, Dean C.2015. Environmental perception and epigenetic memory: Mechanistic insight through FLC. Plant J, 83(1): 133-148. |
| [6] | Campalans A, Kondorosi A, Crespi M.2004. Enod40, a short open reading frame-containing mRNA, induces cytoplasmic localization of a nuclear RNA binding protein in Medicago truncatula. Plant Cell, 16(4): 1047-1059. |
| [7] | Chen L, Shi S L, Jiang N F, Khanzada H, Wassan G M, Zhu C L, Peng X S, Xu J, Chen Y J, Yu Q Y, He X P, Fu J R, Chen X R, Hu L F, Ouyang L J, Sun X T, He H H, Bian J M.2018. Genome-wide analysis of long non-coding RNAs affecting roots development at an early stage in the rice response to cadmium stress. BMC Genom, 19(1): 460. |
| [8] | Csorba T, Questa J I, Sun Q W, Dean C.2014. Antisense COOLAIR mediates the coordinated switching of chromatin states at FLC during vernalization. Proc Natl Acad Sci USA, 111: 16160-16165. |
| [9] | Cui J, Luan Y S, Jiang N, Bao H, Meng J.2017. Comparative transcriptome analysis between resistant and susceptible tomato allows the identification of lncRNA16397 conferring resistance to Phytophthora infestans by co-expressing glutaredoxin. Plant J, 89(3): 577-589. |
| [10] | de Lucia F, Dean C.2011. Long non-coding RNAs and chromatin regulation. Curr Opin Plant Biol, 14(2): 168-173. |
| [11] | Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles D G, Lagarde J, Veeravalli L, Ruan X A, Ruan Y J, Lassmann T, Carninci P, Brown J B, Lipovich L, Gonzalez J M, Thomas M, Davis C A, Shiekhattar R, Gingeras T R, Hubbard T J, Notredame C, Harrow J, Guigó R.2012. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genom Res, 22(9): 1775-1789. |
| [12] | Di C, Yuan J P, Wu Y, Li J R, Lin H X, Hu L, Zhang T, Qi Y J, Gerstein M B, Guo Y, Lu Z J.2014. Characterization of stress-responsive lncRNAs in Arabidopsis thaliana by integrating expression, epigenetic and structural features. Plant J, 80(5): 848-861. |
| [13] | Ding J H, Lu Q, Ouyang Y D, Mao H L, Zhang P B, Yao J L, Xu C G, Li X H, Xiao J H, Zhang Q F.2012a. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc Natl Acad Sci USA, 109(7): 2654-2659. |
| [14] | Ding J H, Shen J Q, Mao H L, Xie W B, Li X H, Zhang Q F.2012b. RNA-directed DNA methylation is involved in regulating photoperiod-sensitive male sterility in rice. Mol Plant, 5(6): 1210-1216. |
| [15] | Dykes I M, Emanueli C.2017. Transcriptional and post-transcriptional gene regulation by long non-coding RNA. Genom Proteom Bioinf, 15(3): 177-186. |
| [16] | Fan Y R, Yang J Y, Mathioni S M, Yu J S, Shen J Q, Yang X F, Wang L, Zhang Q H, Cai Z X, Xu C G, Li X H, Xiao J H, Meyers B C, Zhang Q F.2016. PMS1T, producing phased small-interfering RNAs, regulates photoperiod-sensitive male sterility in rice. Proc Natl Acad Sci USA, 113(52): 15144-15149. |
| [17] | Fang S S, Zhang L L, Guo J C, Niu Y W, Wu Y, Li H, Zhao L H, Li X Y, Teng X Y, Sun X H, Sun L, Zhang M Q, Chen R S, Zhao Y.2018. NONCODEV5: A comprehensive annotation database for long noncoding RNAs. Nucl Acids Res, 46: 308-314. |
| [18] | Fok E T, Scholefield J, Fanucchi S, Mhlanga M M.2017. The emerging molecular biology toolbox for the study of long noncoding RNA biology. Epigenomics, 9(10): 1317-1327. |
| [19] | Franco-Zorrilla J M, Valli A, Todesco M, Mateos I, Puga M I, Rubio-Somoza I, Leyva A, Weigel D, García J A, Paz-Ares J.2007. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat Genet, 39(8): 1033-1037. |
| [20] | Geisler S, Coller J.2013. RNA in unexpected places: Long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol, 14(11): 699-712. |
| [21] | Gerstein M B, Bruce C, Rozowsky J S, Zheng D Y, Du J, Korbel J O, Emanuelsson O, Zhang Z D, Weissman S, Snyder M.2007. What is a gene, post-ENCODE? History and updated definition. Genome Res, 17(6): 669-681. |
| [22] | Gloss B S, Dinger M E.2016. The specificity of long noncoding RNA expression. Biochim Biophys Acta, 1859(1): 16-22. |
| [23] | Guo J X, Liu Y G.2017. Long non-coding RNAs play an important role in regulating photoperiod- and temperature-sensitive male sterility in rice. Sci China Life Sci, 60(4): 443-444. |
| [24] | Haudry A, Platts A E, Vello E, Hoen D R, Leclercq M, Williamson R J, Forczek E, Joly-Lopez Z, Steffen J G, Hazzouri K M, Dewar K, Stinchcombe J R, Schoen D J, Wang X W, Schmutz J, Town C D, Edger P P, Pires J C, Schumaker K S, Jarvis D E, Mandáková T, Lysak M A, van den Bergh E, Schranz M E, Harrison P M, Moses A M, Bureau T E, Wright S I, Blanchette M.2013. An atlas of over 90,000 conserved noncoding sequences provides insight into crucifer regulatory regions. Nat Genet, 45(8): 891-898. |
| [25] | Heo J B, Sung S.2011. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science, 331: 76-79. |
| [26] | Hirayama T, Shinozaki K.2010. Research on plant abiotic stress responses in the post-genome era: Past, present and future. Plant J, 61(6): 1041-1052. |
| [27] | Hirsch J, Lefort V, Vankersschaver M, Boualem A, Lucas A, Thermes C, d’Aubenton-Carafa Y, Crespi M.2006. Characterization of 43 non-protein-coding mRNA genes in Arabidopsis, including the MIR162a-derived transcripts. Plant Physiol, 140(4): 1192-1204. |
| [28] | Huang T, López-Giráldez F, Townsend J P, Irish V F.2012. RBE controls microRNA164 expression to effect floral organogenesis. Development, 139(12): 2161-2169. |
| [29] | Jain P, Sharma V, Dubey H, Singh P K, Kapoor R, Kumari M, Singh J, Pawar D V, Bisht D, Solanke A U, Mondal T K, Sharma T R.2017. Identification of long non-coding RNA in rice lines resistant to rice blast pathogen Maganaporthe oryzae. Bioinformation, 13(8): 249-255. |
| [30] | Johnsson P, Lipovich L, Grandér D, Morris K V.2014. Evolutionary conservation of long non-coding RNAs: Sequence, structure, function. Biochim Biophys Acta, 1840(3): 1063-1071. |
| [31] | Joshi R K, Megha S, Basu U, Rahman M H, Kav N N.2016. Genome wide identification and functional prediction of long non-coding RNAs responsive to Sclerotinia sclerotiorum infection in Brassica napus. PLoS One, 11(7): e0158784. |
| [32] | Karlik E, Ari S, Gozukirmizi N.2019. LncRNAs: Genetic and epigenetic effects in plants. Biotechnol Biotechnol Equip, 33(1): 429-439. |
| [33] | Komiya R, Ohyanagi H, Niihama M, Watanabe T, Nakano M, Kurata N, Nonomura K.2014. Rice germline-specific argonaute MEL1 protein binds to phasiRNAs generated from more than 700 lincRNAs. Plant J, 78(3): 385-397. |
| [34] | Kornienko A E, Dotter C P, Guenzl P M, Gisslinger H, Gisslinger B, Cleary C, Kralovics R, Pauler F M, Barlow D P.2016. Long noncoding RNAs display higher natural expression variation than protein-coding genes in healthy humans. Genome Biol, 17: 14. |
| [35] | Lai F, Orom U A, Cesaroni M, Beringer M, Taatjes D J, Blobel G A, Shiekhattar R.2013. Activating RNAs associate with mediator to enhance chromatin architecture and transcription. Nature, 494: 497-501. |
| [36] | Li P, Yang H, Wang L, Liu H J, Huo H Q, Zhang C J, Liu A Z, Zhu A D, Hu J Y, Lin Y J, Liu L.2019. Physiological and transcriptome analyses reveal short-term responses and formation of memory under drought stress in rice. Front Genet, 10: 55. |
| [37] | Liu F Q, Marquardt S, Lister C, Swiezewski S, Dean C.2010. Targeted 3' processing of antisense transcripts triggers Arabidopsis FLC chromatin silencing. Science, 327: 94-97. |
| [38] | Liu H L, Wang R H, Mao B G, Zhao B R, Wang J B.2019. Identification of lncRNAs involved in rice ovule development and female gametophyte abortion by genome-wide screening and functional analysis. BMC Genom, 20(1): 90. |
| [39] | Liu X, Hao L L, Li D Y, Zhu L H, Hu S N.2015. Long non-coding RNAs and their biological roles in plants. Genom Proteom Bioinf, 13(3): 137-147. |
| [40] | Liu X, Li D Y, Zhang D L, Yin D D, Zhao Y, Ji C J, Zhao X F, Li X B, He Q, Chen R S, Hu S N, Zhu L H.2018. A novel antisense long noncoding RNA, TWISTED LEAF, maintains leaf blade flattening by regulating its associated sense R2R3-MYB gene in rice. New Phytol, 218(2): 774-788. |
| [41] | Liu X D, Huang J, Wang Y, Khanna K, Xie Z X, Owen H A, Zhao D Z.2010. The role of floral organs in carpels, an Arabidopsis loss-of-function mutation in microRNA160a, in organogenesis and the mechanism regulating its expression. Plant J, 62(3): 416-428. |
| [42] | Luan X, Liu S C, Ke S W, Dai H, Xie X M, Hsieh T F, Zhang X Q.2019. Epigenetic modification of ESP, encoding a putative long noncoding RNA, affects panicle architecture in rice. Rice, 12(1): 20. |
| [43] | Ma L, Bajic V B, Zhang Z.2013. On the classification of long non-coding RNAs. RNA Biol, 10(6): 925-933. |
| [44] | Magistri M, Faghihi M A, St Laurent G, Wahlestedt C.2012. Regulation of chromatin structure by long noncoding RNAs: Focus on natural antisense transcripts. Trends Genet, 28(8): 389-396. |
| [45] | Mattick J S, Rinn J L.2015. Discovery and annotation of long noncoding RNAs. Nat Struct Mol Biol, 22(1): 5-7. |
| [46] | Nejat N, Mantri N.2017. Emerging roles of long non-coding RNAs in plant response to biotic and abiotic stresses. Crit Rev Biotechnol, 38(1): 93-105. |
| [47] | Nelson A D, Forsythe E S, Devisetty U K, Clausen D S, Haug- Batzell A K, Meldrum A M, Frank M R, Lyons E, Beilstein M A.2016. A genomic analysis of factors driving lincRNA diversification: Lessons from plants. G3: Gene Genom Genet, 6(9): 2881-2891. |
| [48] | Niazi F, Valadkhan S.2012. Computational analysis of functional long noncoding RNAs reveals lack of peptide-coding capacity and parallels with 3' UTRs. RNA, 18(4): 825-843. |
| [49] | Nitsche A, Rose D, Fasold M, Reiche K, Stadler P F.2015. Comparison of splice sites reveals that long noncoding RNAs are evolutionarily well conserved. RNA, 21(5): 801-812. |
| [50] | Nonomura K, Morohoshi A, Nakano M, Eiguchi M, Miyao A, Hirochika H, Kurata N.2007. A germ cell specific gene of the ARGONAUTE family is essential for the progression of premeiotic mitosis and meiosis during sporogenesis in rice. Plant Cell, 19(8): 2583-2594. |
| [51] | Ponjavic J, Ponting C P, Lunter G.2007. Functionality or transcriptional noise? Evidence for selection within long noncoding RNAs. Genom Res, 17(5): 556-565. |
| [52] | Ponting C P, Oliver P L, Reik W.2009. Evolution and functions of long noncoding RNAs. Cell, 136(4): 629-641. |
| [53] | Qin T, Zhao H Y, Cui P, Albesher N, Xiong L M.2017. A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance. Plant Physiol, 175(3): 1321-1336. |
| [54] | Quan M Y, Chen J H, Zhang D Q.2015. Exploring the secrets of long noncoding RNAs. Int J Mol Sci, 16(3): 5467-5496. |
| [55] | Sanchita, Trivedi P K, Asif M H.2019. Updates on plant long non-coding RNAs (lncRNAs): The regulatory components. Plant Cell Tiss Organ Cult, 140: 259-269. |
| [56] | Shin H, Shin H S, Chen R J, Harrison M J.2006. Loss of At4 function impacts phosphate distribution between the roots and the shoots during phosphate starvation. Plant J, 45(5): 712-726. |
| [57] | Shin S Y, Jeong J S, Lim J Y, Kim T, Park J H, Kim J K, Shin C.2018. Transcriptomic analyses of rice (Oryza sativa) genes and non-coding RNAs under nitrogen starvation using multiple omics technologies. BMC Genom, 19(1): 532. |
| [58] | Shuai P, Liang D, Tang S, Zhang Z J, Ye C Y, Su Y Y, Xia X L, Yin W L.2014. Genome-wide identification and functional prediction of novel and drought-responsive lincRNAs in Populus trichocarpa. J Exp Bot, 65(17): 4975-4983. |
| [59] | Singh U, Khemka N, Rajkumar M S, Garg R, Jain M.2017. PLncPRO for prediction of long non-coding RNAs (lncRNAs) in plants and its application for discovery of abiotic stress- responsive lncRNAs in rice and chickpea. Nucl Acids Res, 45(22): e183. |
| [60] | Spitale R C, Tsai M C, Chang H Y.2011. RNA templating the epigenome: Long noncoding RNAs as molecular scaffolds. Epigenetics, 6(5): 539-543. |
| [61] | Tang Z H, Xu M, Ito H, Cai J H, Ma X X, Qin J P, Yu D L, Meng Y J.2019. Deciphering the non-coding RNA-level response to arsenic stress in rice (Oryza sativa). Plant Signal Behav, 14(9): 1629268. |
| [62] | Tsai M C, Manor O, Wan Y, Mosammaparast N, Wang J K, Lan F, Shi Y, Segal E, Chang H Y.2010. Long noncoding RNA as modular scaffold of histone modification complexes. Science, 329: 689-693. |
| [63] | Vance K W, Ponting C P.2014. Transcriptional regulatory functions of nuclear long noncoding RNAs. Trends Genet, 30(8): 348-355. |
| [64] | van Werven F J, Neuert G, Hendrick N, Lardenois A, Buratowski S, van Oudenaarden A, Primig M, Amon A.2012. Transcription of two long noncoding RNAs mediates mating-type control of gametogenesis in budding yeast. Cell, 150(6): 1170-1181. |
| [65] | Vrbsky J, Akimcheva S, Watson J M, Turner T L, Daxinger L, Vyskot B, Aufsatz W, Riha K.2010. siRNA-mediated methylation of Arabidopsis telomeres. PLoS Genet, 6(6): e1000986. |
| [66] | Wang H, Chung P J, Liu J, Jang I C, Kean M J, Xu J, Chua N H.2014. Genome-wide identification of long noncoding natural antisense transcripts and their responses to light in Arabidopsis. Genome Res, 24(3): 444-453. |
| [67] | Wang H, Niu Q W, Wu H W, Liu J, Ye J, Yu N, Chua N H.2015. Analysis of non-coding transcriptome in rice and maize uncovers roles of conserved lncRNAs associated with agriculture traits. Plant J, 84(2): 404-416. |
| [68] | Wang J Y, Yu W G, Yang Y W, Li X, Chen T Z, Liu T L, Ma N, Yang X, Liu R Y, Zhang B L.2015. Genome-wide analysis of tomato long non-coding RNAs and identification as endogenous target mimic for microRNA in response to TYLCV infection. Sci Rep, 5: 16946. |
| [69] | Wang K C, Chang H Y.2011. Molecular mechanisms of long noncoding RNAs. Mol Cell, 43(6): 904-914. |
| [70] | Wang Y, Luo X J, Sun F, Hu J H, Zha X J, Su W, Yang J S.2018. Overexpressing lncRNA LAIR increases grain yield and regulates neighbouring gene cluster expression in rice. Nat Commun, 9(1): 3516. |
| [71] | Wang Y Q, Fan X D, Lin F, He G M, Terzaghi W, Zhu D M, Deng X W.2014. Arabidopsis noncoding RNA mediates control of photomorphogenesis by red light. Proc Natl Acad Sci USA, 111(28): 10359-10364. |
| [72] | Wierzbicki A T.2012. The role of long non-coding RNA in transcriptional gene silencing. Curr Opin Plant Biol, 15(5): 517-522. |
| [73] | Wilusz J E, Sunwoo H, Spector D L.2009. Long noncoding RNAs: Functional surprises from the RNA world. Genes Dev, 23(13): 1494-1504. |
| [74] | Wu H J, Wang Z M, Wang M, Wang X J.2013. Widespread long noncoding RNAs as endogenous target mimics for microRNAs in plants. Plant Physiol, 161(4): 1875-1884. |
| [75] | Xin M M, Wang Y, Yao Y Y, Song N, Hu Z R, Qin D D, Xie C J, Peng H R, Ni Z F, Sun Q X.2011. Identification and characterization of wheat long non-protein coding RNAs responsive to powdery mildew infection and heat stress by using microarray analysis and SBS sequencing. BMC Plant Biol, 11: 61. |
| [76] | Xu X W, Zhou X H, Wang R R, Peng W L, An Y, Chen L L.2016. Functional analysis of long intergenic non-coding RNAs in phosphate-starved rice using competing endogenous RNA network. Sci Rep, 6: 20715. |
| [77] | Yang Y F, Wen L W, Zhu H L.2015. Unveiling the hidden function of long non-coding RNA by identifying its major partner-protein. Cell Biosci, 5: 59. |
| [78] | Ye C Y, Chen L, Liu C, Zhu Q H, Fan L J.2015. Widespread noncoding circular RNAs in plants. New Phytol, 208(1): 88-95. |
| [79] | Yuan J P, Li J R, Yang Y, Tan C, Zhu Y M, Hu L, Qi Y J, Lu Z J.2018. Stress-responsive regulation of long non-coding RNA polyadenylation in Oryza sativa. Plant J, 93(5): 814-827. |
| [80] | Zhang F T, Luo Yuan, Zhang M, Zhou Y, Chen H P, Hu B L, Xie J K.2018. Identification and characterization of drought stress- responsive novel microRNAs in Dongxiang wild rice. Rice Sci, 25(4): 175-184. |
| [81] | Zhang W, Han Z X, Guo Q L, Liu Y, Zheng Y X, Wu F L, Jin W B.2014. Identification of maize long non-coding RNAs responsive to drought stress. PLoS One, 9(6): e98958. |
| [82] | Zhang Y C, Liao J Y, Li Z Y, Yu Y, Zhang J P, Li Q F, Qu L H, Shu W S, Chen Y Q.2014. Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genom Biol, 15(12): 512. |
| [83] | Zhu Q H, Stephen S, Taylor J, Helliwell C A, Wang M B.2014. Long noncoding RNAs responsive to Fusarium oxysporum infection in Arabidopsis thaliana. New Phytol, 201(2): 574-584. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||