
Rice Science ›› 2020, Vol. 27 ›› Issue (3): 184-200.DOI: 10.1016/j.rsci.2020.04.003
• Review • Previous Articles Next Articles
Adijat Ajadi Abolore1,2, Cisse Amara1, Ahmad Shakeel1, Yifeng Wang1, Yazhou Shu1, Shufan Li1, Xixi Liu1, Kazeem Bello Babatunde3, Muhammad Tajo Sani1,4, Xiaohong Tong1, Jian Zhang1( )
)
Received:2019-05-06
Accepted:2019-10-31
Online:2020-05-28
Published:2020-01-17
Adijat Ajadi Abolore, Cisse Amara, Ahmad Shakeel, Yifeng Wang, Yazhou Shu, Shufan Li, Xixi Liu, Kazeem Bello Babatunde, Muhammad Tajo Sani, Xiaohong Tong, Jian Zhang. Protein Phosphorylation and Phosphoproteome: An Overview of Rice[J]. Rice Science, 2020, 27(3): 184-200.
Add to citation manager EndNote|Ris|BibTeX
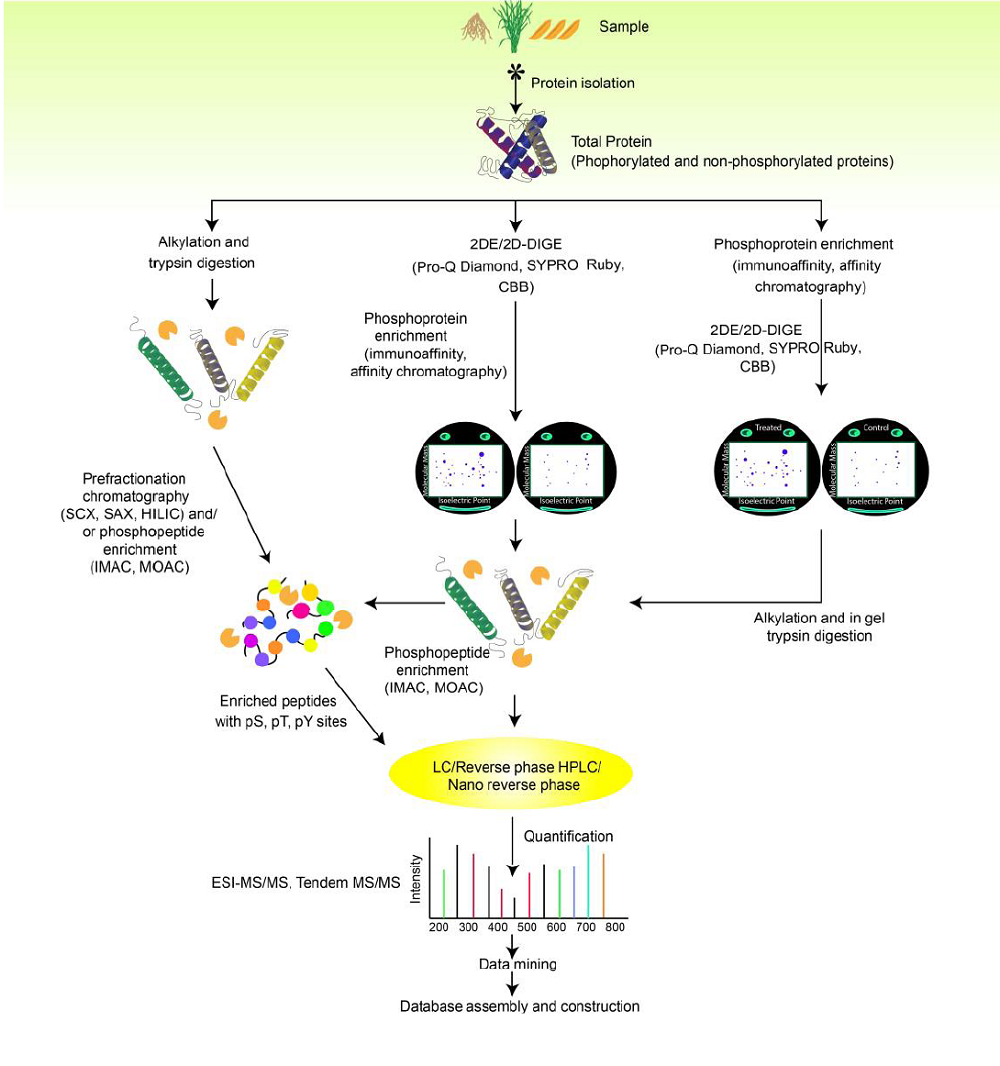
Fig. 2. Overview of common techniques used in identification and quantification of rice phosphoproteome.2DE, Two-dimensional gel electrophoresis; 2D-DIGE, Two-dimensional differential gel electrophoresis; CBB, Coomassie brilliant blue; SCX, Strong cation exchange; SAX, Small-angle X-ray scattering; HILIC, Hydrophilic interaction liquid chromatography; IMAC, Immobilized metal affinity chromatography; MOAC, Metal oxide/hydroxide affinity chromatography; LC, Liquid chromatography; HPLC, High performance liquid chromatography; ESI-MS/MS; Electrospray ionization tandem mass spectrometry.
| Treatment/Sample type | Proteomic approach; Subcellular fraction | Number of phosphoproteins and number of sites identified | Reference |
|---|---|---|---|
| GAs, BL, 2,4-D, ABA, cold or salt treatment of rice seedlings | Total protein extracts from leaf sheaths, leaf blades and roots, in vitro 32P labeling, 2DE; ESI-Q-TOF-MS/MS; MALDI-TOF-MS | 42 phosphoproteins identified, among them 13 were stress/hormonal responsive; 8 sites of phosphorylation in 6 phosphoproteins | Khan et al, 2005 |
| Salt treatment of rice roots | Total protein extracts; 2DE; Pro-Q Diamond, SYPRO Ruby; MALDI-TOF/TOF PMF and MS/MS | 17 differentially up-regulated and 11 down-regulated phosphoproteins for Pro-Q Diamond dye; 26 up-regulated and 5 down-regulated phosphoproteins for SYPRO Ruby | Chitteti and Peng, 2007 |
| Protoplast isolation from rice cell suspension | Nucleus and chromatin isolation; Total protein extracts; 2D-PAGE; Pro-Q Diamond; MALDI-TOF/TOF MS and LC/LC-MS/MS | 972 distinct protein spots and 509 proteins identified, which correspond to 269 unique proteins | Tan et al, 2007 |
| ABA treatment of rice seedlings | Total protein; 2DE; Immunodetection of phospho-Ser and phospho-Thr; MALDI-TOF/TOF MS and MS/MS | 6 ABA responsive phosphoproteins out of 28 proteins | He and Li, 2008 |
| Cd stress in leaves and root of rice seedlings | Total protein extracts; 2DE; Colloidal coomassie blue; MALDI-TOF/TOF PMF and MS/MS | 13 proteins in the roots and 12 proteins in the leaves | Ge et al, 2009 |
| Drought treatment of rice seedlings | Total protein; 2DE; Immunodetection of phospho-Ser and phospho-Thr; MALDI-TOF/TOF MS and MS/MS | 3 drought-responsive differentially expressed proteins and 10 differentially phosphorylated (dp) phosphoproteins | Ke et al, 2009 |
| Unfractionated whole-cell lysates of rice and Arabidopsis cells | Total protein extract; SCX prefractionation; Ti-HAMMOC; Zr-HAMMOC; Fe-IMAC using Phos-select; Nano-LC-MS/MS | 6 919 phosphopeptides from 3 393 proteins | Nakagami et al, 2010 |
| Salinity, cold and drought stress | Genome sequences; Microarray gene expression analysis; Online databases and in silico search tools; RT-PCR | 132 proteins; 46 phosphatases were differentially expressed in 3 abiotic stresses | Singh et al, 2010 |
| Heat (at 42 ºC) of rice seedlings | Total protein; 2DE; Pro-Q Diamond and coomassie, MALDI-TOF/TOF MS and MS/MS; Immunoblotting of ATP-β | 10 dp proteins | Chen et al, 2011 |
| High salinity treatment of A. thaliana and rice plants | Microsomal fractions; Protein digestion; Enrichment using Zr4+-IMAC and TiO2 magnetic beads; Q-TOF Nano LC-MS/MS; Label-free quantification on Q-TOF, LTQ-FT and Q-Trap | 3 membrane phosphoproteins and 4 nonmembrane phosphoproteins; 15 phosphopeptides from rice microsomes | Chang et al, 2012 |
| Cold stress (6 ºC and 15 ºC) of rice seedlings | Total protein extracts; 2DE; Coomassie brilliant blue; Pro-Q Diamond; MALDI-TOF/TOF MS | 12 up-regulated and 1 down-regulated phosphoprotein among 13 phosphoproteins | Chen et al, 2012 |
| ABA treatment of rice inferior spiklets at 20 d after flowering | Total protein extracts; 2DE; Coomassie brilliant blue; Pro-Q Diamond; MALDI-TOF/TOF MS | 16 up-regulated and 15 down-regulated among 31 differentially regulated phosphoproteins | Zhang et al, 2012 |
| Rice embryo germination | Total protein extracts; 2DE; Pro-Q Diamond; Coomassie brilliant blue and MALDI-TOF/TOF MS | About 500 phosphorylated protein spots were detected on each gel | Han et al, 2014a |
| Poor grain-filling of rice inferior spikelets | Total protein extracts; 2DE; Coomassie brilliant blue; Pro-Q Diamond; MALDI-TOF/TOF MS and LC-ESI-MS/MS | 123 proteins in abundance and 43 phosphoproteins | Zhang et al, 2014 |
| Rice pistils | Total protein; Fe3+-IMAC protein enrichment; HILIC and Nano-LC-MS/MS | 2 347 phosphorylation sites and 1 588 phosphoproteins | Wang K et al, 2014 |
| Rice leaves in response to bacterial blight | Total protein; TiO2 MOAC and LC-MS/MS | 2 367 and 2 223 phosphosites on 1 334 and 1 297 proteins in 0 and 24 h after Xoo infection, respectively | Hou et al, 2015 |
| Early stage of seed germination | Isolation of nuclei and extraction of nuclear proteins; Total protein; PolyMAC-Ti and MS/MS | 3 467 phosphopeptides and 102 nuclear phosphorproteins from rice embryos; 115 nuclear phosphoproteins identified after 24 h of imbibition | Li M et al, 2015 |
| Early seed development | Total protein; TiO2 protein enrichment and LC-MS/MS | Pistils and seeds at 3 and 7 d after pollination resulted in 38 854 313 and 4 135 phosphopeptides, respectively. A total of 2487 proteins were dp | Qiu et al, 2015 |
| Magnaporthe oryzae treatment of susceptible and resistant rice cultivars | Total protein extracts; Al(OH)3-MOAC; 2DE; Pro-Q Diamond; MALDI-TOF/TOF PMF and Nano LC-MS/MS | 1 522 unique peptides, of which 1 365 were dp | Li Y F et al, 2015 |
| Developing rice anthers | Total protein; TiO2 MOAC and Nano UHPLC-MS/MS | 4 984 proteins and 3 203 phosphoproteins with 8 973 unique phosphorylation sites | Ye et al, 2015 |
| Brassinosteroid treatment in rice | Total protein; TiO2 MOAC and LC-MS/MS | 4 034 phosphosites on 1 900 phosphoproteins and 1 821 brassinosteroid-responsive proteins | Hou et al, 2017 |
| Callus, leaf, root, shoot meristem, young panicle, and mature panicle | Total protein extracts; Pro-Q diamond; TiO2 enrichment and LC-MS/MS | 7 171 unique phosphosites in 4 792 phosphopeptides from 2 657 identified phosphoproteins | Wang et al, 2017 |
| Cd stress in rice seedlings | Total protein; iTRAQ labeling; TiO2 enrichment and LC-MS/MS | 2 454 phosphosites associated with 1 244 proteins, among them, 482 were dp | Zhong et al, 2017 |
| Response to Cd stress in rice | 32 up-regulated and 1 down-regulated Cd tolerant, and 15 up-regulated and 4 down-regulated Cd sensitive among 53 differentially expressed phosphoproteins | Fang et al, 2019 |
Table 1 Summary of some quantitative studies in rice phosphoproteome study.
| Treatment/Sample type | Proteomic approach; Subcellular fraction | Number of phosphoproteins and number of sites identified | Reference |
|---|---|---|---|
| GAs, BL, 2,4-D, ABA, cold or salt treatment of rice seedlings | Total protein extracts from leaf sheaths, leaf blades and roots, in vitro 32P labeling, 2DE; ESI-Q-TOF-MS/MS; MALDI-TOF-MS | 42 phosphoproteins identified, among them 13 were stress/hormonal responsive; 8 sites of phosphorylation in 6 phosphoproteins | Khan et al, 2005 |
| Salt treatment of rice roots | Total protein extracts; 2DE; Pro-Q Diamond, SYPRO Ruby; MALDI-TOF/TOF PMF and MS/MS | 17 differentially up-regulated and 11 down-regulated phosphoproteins for Pro-Q Diamond dye; 26 up-regulated and 5 down-regulated phosphoproteins for SYPRO Ruby | Chitteti and Peng, 2007 |
| Protoplast isolation from rice cell suspension | Nucleus and chromatin isolation; Total protein extracts; 2D-PAGE; Pro-Q Diamond; MALDI-TOF/TOF MS and LC/LC-MS/MS | 972 distinct protein spots and 509 proteins identified, which correspond to 269 unique proteins | Tan et al, 2007 |
| ABA treatment of rice seedlings | Total protein; 2DE; Immunodetection of phospho-Ser and phospho-Thr; MALDI-TOF/TOF MS and MS/MS | 6 ABA responsive phosphoproteins out of 28 proteins | He and Li, 2008 |
| Cd stress in leaves and root of rice seedlings | Total protein extracts; 2DE; Colloidal coomassie blue; MALDI-TOF/TOF PMF and MS/MS | 13 proteins in the roots and 12 proteins in the leaves | Ge et al, 2009 |
| Drought treatment of rice seedlings | Total protein; 2DE; Immunodetection of phospho-Ser and phospho-Thr; MALDI-TOF/TOF MS and MS/MS | 3 drought-responsive differentially expressed proteins and 10 differentially phosphorylated (dp) phosphoproteins | Ke et al, 2009 |
| Unfractionated whole-cell lysates of rice and Arabidopsis cells | Total protein extract; SCX prefractionation; Ti-HAMMOC; Zr-HAMMOC; Fe-IMAC using Phos-select; Nano-LC-MS/MS | 6 919 phosphopeptides from 3 393 proteins | Nakagami et al, 2010 |
| Salinity, cold and drought stress | Genome sequences; Microarray gene expression analysis; Online databases and in silico search tools; RT-PCR | 132 proteins; 46 phosphatases were differentially expressed in 3 abiotic stresses | Singh et al, 2010 |
| Heat (at 42 ºC) of rice seedlings | Total protein; 2DE; Pro-Q Diamond and coomassie, MALDI-TOF/TOF MS and MS/MS; Immunoblotting of ATP-β | 10 dp proteins | Chen et al, 2011 |
| High salinity treatment of A. thaliana and rice plants | Microsomal fractions; Protein digestion; Enrichment using Zr4+-IMAC and TiO2 magnetic beads; Q-TOF Nano LC-MS/MS; Label-free quantification on Q-TOF, LTQ-FT and Q-Trap | 3 membrane phosphoproteins and 4 nonmembrane phosphoproteins; 15 phosphopeptides from rice microsomes | Chang et al, 2012 |
| Cold stress (6 ºC and 15 ºC) of rice seedlings | Total protein extracts; 2DE; Coomassie brilliant blue; Pro-Q Diamond; MALDI-TOF/TOF MS | 12 up-regulated and 1 down-regulated phosphoprotein among 13 phosphoproteins | Chen et al, 2012 |
| ABA treatment of rice inferior spiklets at 20 d after flowering | Total protein extracts; 2DE; Coomassie brilliant blue; Pro-Q Diamond; MALDI-TOF/TOF MS | 16 up-regulated and 15 down-regulated among 31 differentially regulated phosphoproteins | Zhang et al, 2012 |
| Rice embryo germination | Total protein extracts; 2DE; Pro-Q Diamond; Coomassie brilliant blue and MALDI-TOF/TOF MS | About 500 phosphorylated protein spots were detected on each gel | Han et al, 2014a |
| Poor grain-filling of rice inferior spikelets | Total protein extracts; 2DE; Coomassie brilliant blue; Pro-Q Diamond; MALDI-TOF/TOF MS and LC-ESI-MS/MS | 123 proteins in abundance and 43 phosphoproteins | Zhang et al, 2014 |
| Rice pistils | Total protein; Fe3+-IMAC protein enrichment; HILIC and Nano-LC-MS/MS | 2 347 phosphorylation sites and 1 588 phosphoproteins | Wang K et al, 2014 |
| Rice leaves in response to bacterial blight | Total protein; TiO2 MOAC and LC-MS/MS | 2 367 and 2 223 phosphosites on 1 334 and 1 297 proteins in 0 and 24 h after Xoo infection, respectively | Hou et al, 2015 |
| Early stage of seed germination | Isolation of nuclei and extraction of nuclear proteins; Total protein; PolyMAC-Ti and MS/MS | 3 467 phosphopeptides and 102 nuclear phosphorproteins from rice embryos; 115 nuclear phosphoproteins identified after 24 h of imbibition | Li M et al, 2015 |
| Early seed development | Total protein; TiO2 protein enrichment and LC-MS/MS | Pistils and seeds at 3 and 7 d after pollination resulted in 38 854 313 and 4 135 phosphopeptides, respectively. A total of 2487 proteins were dp | Qiu et al, 2015 |
| Magnaporthe oryzae treatment of susceptible and resistant rice cultivars | Total protein extracts; Al(OH)3-MOAC; 2DE; Pro-Q Diamond; MALDI-TOF/TOF PMF and Nano LC-MS/MS | 1 522 unique peptides, of which 1 365 were dp | Li Y F et al, 2015 |
| Developing rice anthers | Total protein; TiO2 MOAC and Nano UHPLC-MS/MS | 4 984 proteins and 3 203 phosphoproteins with 8 973 unique phosphorylation sites | Ye et al, 2015 |
| Brassinosteroid treatment in rice | Total protein; TiO2 MOAC and LC-MS/MS | 4 034 phosphosites on 1 900 phosphoproteins and 1 821 brassinosteroid-responsive proteins | Hou et al, 2017 |
| Callus, leaf, root, shoot meristem, young panicle, and mature panicle | Total protein extracts; Pro-Q diamond; TiO2 enrichment and LC-MS/MS | 7 171 unique phosphosites in 4 792 phosphopeptides from 2 657 identified phosphoproteins | Wang et al, 2017 |
| Cd stress in rice seedlings | Total protein; iTRAQ labeling; TiO2 enrichment and LC-MS/MS | 2 454 phosphosites associated with 1 244 proteins, among them, 482 were dp | Zhong et al, 2017 |
| Response to Cd stress in rice | 32 up-regulated and 1 down-regulated Cd tolerant, and 15 up-regulated and 4 down-regulated Cd sensitive among 53 differentially expressed phosphoproteins | Fang et al, 2019 |
| Approach | Method | Advantage | Disadvantage |
|---|---|---|---|
| 2D-DIGE | Deliver a visual map of all the proteins and allow the fractionation of thousands of proteins in a complex mixture | Relatively small scale, low sensitivity, poor detection of low-abundance phosphoproteins, and insufficient collected proteins for downstream enrichment methods | |
| Stable isotope labeling quantification | SILAC | Lower experimental error | Allow up to only three samples |
| HILEP, SILIP, SILIA | Applicable to entire plant and other plant tissues | Only be used in auxotrophic cells | |
| iTRAQ and TMT | Allow multiplex up to 8 (iTARQ) and 10 (TMT); Labeling agent doesn’t affect overall peptide behavior thus minimizes experimental error | Reduced identification efficiency as much as 50% in multi-stage activation | |
| Chemical derivatization (DiLeu) | Higher protein coverage and quantification efficiency; Cheaper than iTRAQ and TMT; Provide higher reproducibility in quantitation | ||
| Label-free quantification | Signal intensity and spectral counting | Allow as many samples as possible | Time-consuming; Less degree of reproducibility in quantitation |
| Absolute quantification | AQUA | Allow quantification of isoforms in low abundance and stoichiometry; Achieve accurate and sensitive quantification | |
| AQUIP | Can determine the absolute amount of protein | ||
| 2D-DIGE, Two-dimensional differential gel electrophoresis; SILAC, Stable isotope labeling by amino acids in culture; HILEP, Hydroponic isotope labeling of entire plants; SILIP, Stable isotope labeling in planta; SILIA, Stable isotope labeling in Arabidopsis; iTRAQ, Isobaric tags for relative and absolute quantification; TMT, Tandem mass tags; AQUA, Absolute quantification of proteins; AQUIP, Absolute quantitation of isoforms of post-translationally modified proteins. | |||
Table 2 Different approach, methods, advantages and disadvantages of different phosphoproteome quantitation methods.
| Approach | Method | Advantage | Disadvantage |
|---|---|---|---|
| 2D-DIGE | Deliver a visual map of all the proteins and allow the fractionation of thousands of proteins in a complex mixture | Relatively small scale, low sensitivity, poor detection of low-abundance phosphoproteins, and insufficient collected proteins for downstream enrichment methods | |
| Stable isotope labeling quantification | SILAC | Lower experimental error | Allow up to only three samples |
| HILEP, SILIP, SILIA | Applicable to entire plant and other plant tissues | Only be used in auxotrophic cells | |
| iTRAQ and TMT | Allow multiplex up to 8 (iTARQ) and 10 (TMT); Labeling agent doesn’t affect overall peptide behavior thus minimizes experimental error | Reduced identification efficiency as much as 50% in multi-stage activation | |
| Chemical derivatization (DiLeu) | Higher protein coverage and quantification efficiency; Cheaper than iTRAQ and TMT; Provide higher reproducibility in quantitation | ||
| Label-free quantification | Signal intensity and spectral counting | Allow as many samples as possible | Time-consuming; Less degree of reproducibility in quantitation |
| Absolute quantification | AQUA | Allow quantification of isoforms in low abundance and stoichiometry; Achieve accurate and sensitive quantification | |
| AQUIP | Can determine the absolute amount of protein | ||
| 2D-DIGE, Two-dimensional differential gel electrophoresis; SILAC, Stable isotope labeling by amino acids in culture; HILEP, Hydroponic isotope labeling of entire plants; SILIP, Stable isotope labeling in planta; SILIA, Stable isotope labeling in Arabidopsis; iTRAQ, Isobaric tags for relative and absolute quantification; TMT, Tandem mass tags; AQUA, Absolute quantification of proteins; AQUIP, Absolute quantitation of isoforms of post-translationally modified proteins. | |||
| Rice tissue | Distribution of single (S) : double (D) : triple (T) : multiple (M) phosphorylated protein | Frequency of occurrence ratio (pS : pT : pY) | Reference |
|---|---|---|---|
| Unfractionated whole-cell lysates of rice cells | 84.8 : 12.3 : 2.9 | Nakagami et al, 2010 | |
| Leaves from150 mmol/L NaCl treated rice roots | 75 : 25 : 0 | Shen et al, 2012 | |
| Pistils | 98 : 2 : 0 (S : D : M) | 87 : 12 : 1 | Wang et al, 2014a |
| Magnaporthe oryzae infected seedlings | 86 : 14 : 0 (S : D : M) | Li Y F et al, 2015 | |
| Rice nucleus during the early stage of seed germination | 65 : 30 : 5 (S : D : M) | 64 : 31 : 5 | Li M et al, 2015 |
| Developing rice anthers | 39 : 23 :14 : 24 (S : D : T : M) | 81.33: 17.06: 1.61 | Ye et al, 2015 |
| Rice leaves in response to bacterial blight | (88.3‒89.9) : (11.0‒9.6) : (0.7‒0.5) (S : D : M) | (88.8‒89.2) : (10.6‒10.1) : (0.6‒0.7) | Hou et al, 2015 |
| Pistils and seeds | 95.5 : 4.4 : 0.1 (S : D : M) | 90.93 : 8:84 : 0.23 | Qiu et al, 2015 |
| ABA induced rice seedlings | 93.2 : 6.7 : 0.1 (S : D : M) | 90:93 : 8.84 : 0.23 | Qiu et al, 2017 |
| Brassinosteroid treatment on rice seedlings | (92.9‒93.3) : (6.5‒6.8) : (> 0.3) (S : D : M) | 89.7 : 9.9 : 0.4 | Hou et al, 2017 |
| Callus, leaf, root, shoot meristem, young panicle, and mature panicle | 64.82 : 24.83 : 7.29 : 3.06 (S : D : T : M) | 86.0 : 13.0 : 1.0 | Wang et al, 2017 |
Table 3 Distribution of phosphosites having single, double, triple and multiple sites and frequency of phosphosite in rice phosphoproteome study.
| Rice tissue | Distribution of single (S) : double (D) : triple (T) : multiple (M) phosphorylated protein | Frequency of occurrence ratio (pS : pT : pY) | Reference |
|---|---|---|---|
| Unfractionated whole-cell lysates of rice cells | 84.8 : 12.3 : 2.9 | Nakagami et al, 2010 | |
| Leaves from150 mmol/L NaCl treated rice roots | 75 : 25 : 0 | Shen et al, 2012 | |
| Pistils | 98 : 2 : 0 (S : D : M) | 87 : 12 : 1 | Wang et al, 2014a |
| Magnaporthe oryzae infected seedlings | 86 : 14 : 0 (S : D : M) | Li Y F et al, 2015 | |
| Rice nucleus during the early stage of seed germination | 65 : 30 : 5 (S : D : M) | 64 : 31 : 5 | Li M et al, 2015 |
| Developing rice anthers | 39 : 23 :14 : 24 (S : D : T : M) | 81.33: 17.06: 1.61 | Ye et al, 2015 |
| Rice leaves in response to bacterial blight | (88.3‒89.9) : (11.0‒9.6) : (0.7‒0.5) (S : D : M) | (88.8‒89.2) : (10.6‒10.1) : (0.6‒0.7) | Hou et al, 2015 |
| Pistils and seeds | 95.5 : 4.4 : 0.1 (S : D : M) | 90.93 : 8:84 : 0.23 | Qiu et al, 2015 |
| ABA induced rice seedlings | 93.2 : 6.7 : 0.1 (S : D : M) | 90:93 : 8.84 : 0.23 | Qiu et al, 2017 |
| Brassinosteroid treatment on rice seedlings | (92.9‒93.3) : (6.5‒6.8) : (> 0.3) (S : D : M) | 89.7 : 9.9 : 0.4 | Hou et al, 2017 |
| Callus, leaf, root, shoot meristem, young panicle, and mature panicle | 64.82 : 24.83 : 7.29 : 3.06 (S : D : T : M) | 86.0 : 13.0 : 1.0 | Wang et al, 2017 |
| [1] | Abbasi F M, Komatsu S.2004. A proteomic approach to analyze salt-responsive proteins in rice leaf sheath.Proteomics, 4(7): 2072-2081. |
| [2] | Agrawal G K, Thelen J J J M.2006. Large scale identification and quantitative profiling of phosphoproteins expressed during seed filling in oilseed rape.Mol Cell Proteomics, 5(11): 2044-2059. |
| [3] | Agrawal G K, Rakwal R.2008. Plant proteomics: Technologies, Strategies, and Applications. USA: John Wiley & Sons. |
| [4] | Agrawal G K, Rakwal R.2011. Rice proteomics: A move toward expanded proteome coverage to comparative and functional proteomics uncovers the mysteries of rice and plant biology.Proteomics, 11(9): 1630-1649. |
| [5] | Aryal U K, Krochko J E, Ross A R S.2012. Identification of phosphoproteins inArabidopsis thaliana leaves using polyethylene glycol fractionation, immobilized metal-ion affinity chromatography, two-dimensional gel electrophoresis and mass spectrometry. J Proteom Res, 11: 425-437. |
| [6] | Bajguz A.2007. Metabolism of brassinosteroids in plants.Plant Physiol Biochem, 45: 95-107. |
| [7] | Bantscheff M, Schirle M, Sweetman G, Rick J, Kuster B.2007. Quantitative mass spectrometry in proteomics: A critical review.Anal Bioanal Chem, 389(4): 1017-1031. |
| [8] | Bantscheff M, Lemeer S, Savitski M M, Kuster B.2012. Quantitative mass spectrometry in proteomics: Critical review update from 2007 to the present.Anal Bioanal Chem, 404(4): 939-965. |
| [9] | Batalha I L, Lowe C R, Roque A C.2012. Platforms for enrichment of phosphorylated proteins and peptides in proteomics.Trends Biotechnol, 30(2): 100-110. |
| [10] | Benschop J J, Mohammed S, O’Flaherty M, Heck A J R, Slijper M, Menke F L H.2007. Quantitative phosphoproteomics of early elicitor signaling inArabidopsis. Mol Cell Proteomics, 6: 1198-1214. |
| [11] | Berthet J, Rall T W, Sutherland E W.1957. The relationship of epinephrine and glucagon to liver phosphorylase: IV. Effect of epinephrine and glucagon on the reactivation of phosphorylase in liver homogenates.J Biol Chem, 224(1): 463-475. |
| [12] | Bindschedler L V, Palmblad M, Cramer R.2008. Hydroponic isotope labelling of entire plants (HILEP) for quantitative plant proteomics: An oxidative stress case study.Phytochemistry, 69(10): 1962-1972. |
| [13] | Bindschedler L V, Cramer R.2011. Quantitative plant proteomics.Proteomics, 11(4): 756-775. |
| [14] | Boersema P J, Aye T T, van Veen T A, Heck A J, Mohammed S.2008. Triplex protein quantification based on stable isotope labeling by peptide dimethylation applied to cell and tissue lysates.Proteomics, 8: 4624-4632. |
| [15] | Boex-Fontvieille E, Davanture M, Jossier M, Zivy M, Hodges M, Tcherkez G.2014. Photosynthetic activity influences cellulose biosynthesis and phosphorylation of proteins involved therein inArabidopsis leaves. J Exp Bot, 65: 4997-5010. |
| [16] | Bonhomme L, Valot B, Tardieu F, Zivy M.2012. Phosphoproteome dynamics upon changes in plant water status reveal early events associated with rapid growth adjustment in maize leaves.Mol Cell Proteomics, 11(10): 957-972. |
| [17] | Burnett G, Kennedy E P.1954. The enzymatic phosphorylation of proteins.J Biol Chem, 211(2): 969-980. |
| [18] | Chandrasekhar K, Dileep A, Lebonah D E, Kumari J P.2014. A short review on proteomics and its applications.Int Lett Nat Sci, 17: 77-84. |
| [19] | Chang I F, Hsu J L, Hsu P H, Sheng W A, Lai S J, Lee C, Chen C W, Hsu J C, Wang S Y, Wang L Y, Chen C C.2012. Comparative phosphoproteomic analysis of microsomal fractions ofArabidopsis thaliana and Oryza sativa subjected to high salinity. Plant Sci, 185/186: 131-142. |
| [20] | Chen J H, Tian L, Xu H F, Tian D G, Luo Y M, Ren C M, Yang L M, Shi J S.2012. Cold-induced changes of protein and phosphoprotein expression patterns from rice roots as revealed by multiplex proteomic analysis.Plant Omics, 5(2): 194-199. |
| [21] | Chen S X, Harmon A C.2006. Advances in plant proteomics.Proteomics, 6(20): 5504-5516. |
| [22] | Chen X H, Zhang W F, Zhang B Q, Zhou J C, Wang Y F, Yang Q B, Ke Y Q, He H Q.2011. Phosphoproteins regulated by heat stress in rice leaves.Proteome Sci, 9: 37. |
| [23] | Chitteti B R, Peng Z H.2007. Proteome and phosphoproteome differential expression under salinity stress in rice (Oryza sativa) roots. J Proteome Res, 6(5): 1718-1727. |
| [24] | Cohen P.2002. The origins of protein phosphorylation.Nat Cell Biol, 4(5): E127-E130. |
| [25] | Cori G T, Green A A.1943. Crystalline muscle phosphorylase: II. Prosthetic group.J Biol Chem, 151: 31-38. |
| [26] | Cunningham R, Ma D, Li L J.2012. Mass spectrometry-based proteomics and peptidomics for systems biology and biomarker discovery.Front Biol, 7(4): 313-335. |
| [27] | Dietrich A, Mayer J E, Hahlbrock K.1990. Fungal elicitor triggers rapid, transient, and specific protein phosphorylation in parsley cell suspension cultures.J Biol Chem, 265(11): 6360-6368. |
| [28] | Engelsberger W R, Schulze W X.2012. Nitrate and ammonium lead to distinct global dynamic phosphorylation patterns when resupplied to nitrogen-starvedArabidopsis seedlings. Plant J, 69(6): 978-995. |
| [29] | Fageria N K.2007. Yield physiology of rice.J Plant Nutr, 30(6): 843-879. |
| [30] | Fan S L, Meng Y Y, Song M Z, Pang C Y, Wei H L, Liu J, Zhan X J, Lan J Y, Feng C H, Zhang S X, Yu S X.2014. Quantitative phosphoproteomics analysis of nitric oxide-responsive phospho- proteins in cotton leaf.PLoS One, 9(4): e94261. |
| [31] | Fang Y X, Deng X X, Lu X L, Zheng J J, Jiang H, Rao Y C, Zeng D L, Hu J, Zhang X Q, Xue D W.2019. Differential phosphop- roteome study of the response to cadmium stress in rice.Ecotox Environ Safe, 180: 780-788. |
| [32] | Fíla J, Honys D.2012. Enrichment techniques employed in phosphoproteomics.Amino Acids, 43(3): 1025-1047. |
| [33] | Fischer E H, Krebs E G.1955. Conversion of phosphorylaseb to phosphorylase a in muscle extracts. J Biol Chem, 216(1): 121-132. |
| [34] | Fischer E H.2010. Phosphorylase and the origin of reversible protein phosphorylation.Biol Chem, 391: 131-137. |
| [35] | Fitzgerald M A, McCouch S R, Hall R D.2009. Not just a grain of rice: The quest for quality. Trends Plant Sci, 14(3): 133-139. |
| [36] | Gallego-Bartolome J, Minguet E G, Grau-Enguix F, Abbas M, Locascio A, Thomas S G, Alabadí D, Blazquez M A.2012. Molecular mechanism for the interaction between gibberellin and brassinosteroid signaling pathways inArabidopsis. Proc Natl Acad Sci USA, 109: 13446-13451. |
| [37] | Gaspari M, Cuda G.2011. Nano LC-MS/MS: A robust setup for proteomic analysis.Methods Mol Biol, 790: 115-126. |
| [38] | Gates M B, Tomer K B, Deterding L J.2010. Comparison of metal and metal oxide media for phosphopeptide enrichment prior to mass spectrometric analyses.J Am Soc Mass Spec, 21(10): 1649-1659. |
| [39] | Ge C L, Wang Z G, Wan D Z, Ding Y, Wang Y L, Shang Q, Luo S S.2009. Proteomic study for responses to cadmium stress in rice seedlings.Rice Sci, 16(1): 33-44. |
| [40] | Ghelis T, Bolbach G, Clodic G, Habricot Y, Miginiac E, Sotta B, Jeannette E.2008. Protein tyrosine kinases and protein tyrosine phosphatases are involved in abscisic acid-dependent processes inArabidopsis seeds and suspension cells. Plant Physiol, 148(3): 1668-1680. |
| [41] | Gruhler A, Schulze W X, Matthiesen R, Mann M, Jensen O N.2005. Stable isotope labeling ofArabidopsis thaliana cells and quantitative proteomics by mass spectrometry. Mol Cell Proteomics, 4(11): 1697-1709. |
| [42] | Guo H Q, Li L, Aluru M, Aluru S, Yin Y H.2013. Mechanisms and networks for brassinosteroid regulated gene expression.Curr Opin Plant Biol, 16(5): 545-553. |
| [43] | Gygi S P, Rist B, Gerber S A, Turecek F, Gelb M H, Aebersold R.1999. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags.Nat Biotechnol, 17: 994-999. |
| [44] | Han C, Wang K, Yang P F.2014a. Gel-based comparative phosphoproteomic analysis on rice embryo during germination.Plant Cell Physiol, 55(8): 1376-1394. |
| [45] | Han C, Yang P F, Sakata K, Komatsu S.2014b. Quantitative proteomics reveals the role of protein phosphorylation in rice embryos during early stages of germination.J Proteome Res, 13(3): 1766-1782. |
| [46] | He H Q, Li J X.2008. Proteomic analysis of phosphoproteins regulated by abscisic acid in rice leaves.Biochem Biophl Res Co, 371(4): 883-888. |
| [47] | Hou Y X, Qiu J H, Tong X H, Wei X J, Nallamilli B R, Wu W H, Huang S W, Zhang J.2015. A comprehensive quantitative phosphoproteome analysis of rice in response to bacterial blight.BMC Plant Biol, 15: 163. |
| [48] | Hou Y X, Qiu J H, Wang Y F, Li Z Y, Zhao J, Tong X H, Lin H Y, Zhang J.2017. A quantitative proteomic analysis of brassinosteroid- induced protein phosphorylation in rice (Oryza sativa L.). Front Plant Sci, 8: 514. |
| [49] | Howden A J M, Salek M, Miguet L, Pullen M, Thomas B, Knight M R, Sweetlove L J.2011. The phosphoproteome ofArabidopsis plants lacking the oxidative signal-inducible1 (OXI1) protein kinase. New Phytol, 190(1): 49-56. |
| [50] | Hubbard M J, Cohen P.1993. On target with a new mechanism for the regulation of protein phosphorylation.Trends Biochem Sci, 18(5): 172-177. |
| [51] | Ingebritsen T S, Foulkes J G, Cohen P.1983. The protein phosphatases involved in cellular regulation: 2. Glycogen metabolism.Eur J Biochem, 132(2): 263-274. |
| [52] | Javier G B, Minguet E G, Federico G E, Mohamad A, Locascio A, Thomas S G, Alabadí D, Miguel A B.2012. Molecular mechanism for the interaction between gibberellin and brassinosteroid signaling pathways inArabidopsis. Proc Natl Acad Sci USA, 109: 13446-13451. |
| [53] | Johnson L N, Barford D.1993. The effects of phosphorylation on the structure and function of proteins.Annu Rev Biophys Biomol Struct, 22: 199-232. |
| [54] | Johnson L N, Lewis R J.2001. Structural basis for control by phosphorylation.Chem Rev, 101: 2209-2242. |
| [55] | Jones J D G, Dangl J L.2006. The plant immune system.Nature, 444: 323-329. |
| [56] | Ke Y Q, Han G Q, He H Q, Li J X.2009. Differential regulation of proteins and phosphoproteins in rice under drought stress.Biochem Biophys Res Commun, 379(1): 133-138. |
| [57] | Kersten B, Agrawal G K, Iwahashi H, Rakwal R.2006. Plant phosphoproteomics: A long road ahead.Proteomics, 6(20): 5517-5528. |
| [58] | Kersten B, Agrawal G K, Durek P, Neigenfind J, Schulze W, Walther D, Rakwal R.2009. Plant phosphoproteomics: An update.Proteomics, 9(4): 964-988. |
| [59] | Khan M, Takasaki H, Komatsu S.2005. Comprehensive phosphoproteome analysis in rice and identification of phosphoproteins responsive to different hormones/stresses.J Proteome Res, 4(5): 1592-1599. |
| [60] | Kline J C, De Luca C J.2014. Error reduction in EMG signal decomposition.J Neurophysiol, 112(11): 2718-2728. |
| [61] | Kline-Jonakin K G, Barrett-Wilt G A, Sussman M R.2011. Quantitative plant phosphoproteomics.Curr Opin Plant Biol, 14(5): 507-511. |
| [62] | Krebs E G, Fischer E H.1956. The phosphorylase b to a converting enzyme of rabbit skeletal muscle.Biochimi Biophys Acta, 20: 150-157. |
| [63] | Kresge N, Simoni R D, Hill R L.2011. The Process of reversible phosphorylation: The work of Edmond H. Fischer.J Biol Chem, 286(3): e1-e2. |
| [64] | Kweon H K, Håkansson K.2006. Selective zirconium dioxide- based enrichment of phosphorylated peptides for mass spectrometric analysis.Anal Chem, 78(6): 1743-1749. |
| [65] | Leibler D C.2002. Introduction to Proteomics: Tools for the New Biology. Totowa NJ: Humana press. |
| [66] | Levene P A, Alsberg C L.1906. The cleavage products of vitellin.J Biol Chem, 2: 127-133. |
| [67] | Li M, Yin X J, Sakata K, Yang P F, Komatsu S.2015. Proteomic analysis of phosphoproteins in the rice nucleus during the early stage of seed germination.J Proteome Res, 14(7): 2884-2896. |
| [68] | Li Y F, Ye Z J, Nie Y F, Zhang J, Wang G L, Wang Z Z.2015. Comparative phosphoproteome analysis ofMagnaporthe oryzae- responsive proteins in susceptible and resistant rice cultivars. J Proteomic, 115: 66-80. |
| [69] | Lipmann F A, Levene P A.1932. Serinephosphoric acid obtained on hydrolysis of vitellinic acid.J Biol Chem, 98: 109-114. |
| [70] | Macek B, Mann M, Olsen J V.2009. Global and site-specific quantitative phosphoproteomics: Principles and applications.Annu Rev Pharmacol Toxicol, 49: 199-221. |
| [71] | Magdeldin S, Enany S, Yoshida Y, Xu B, Zhang Y, Zureena Z, Lokamani I, Yaoita E, Yamamoto T.2014. Basics and recent advances of two dimensional-polyacrylamide gel electrophoresis.Clin Proteom, 11(1): 16. |
| [72] | Mandell D J, Chorny I, Groban E S, Wong S E, Levine E, Rapp C S, Jacobson M P.2007. Strengths of hydrogen bonds involving phosphorylated amino acid side chains.J Am Chem Soc, 129: 820-827. |
| [73] | Meleady P.2018. Two-dimensional gel electrophoresis and 2D-DIGE.Methods Mol Biol, 1664: 3-14. |
| [74] | Molina H, Horn D M, Tang N, Mathivanan S, Pandey A.2007. Global proteomic profiling of phosphopeptides using electron transfer dissociation tandem mass spectrometry.Proc Natl Acad Sci USA, 104(7): 2199-2204. |
| [75] | Nakagami H, Sugiyama N, Mochida K, Daudi A, Yoshida Y, Toyoda T, Tomita M, Ishihama Y, Shirasu K.2010. Large-scale comparative phosphoproteomics identifies conserved phosphorylation sites in plants.Plant Physiol, 153(3): 1161-1174. |
| [76] | Neilson K A, Ali N A, Muralidharan S, Mirzaei M, Mariani M, Assadourian G, Lee A, van Sluyter S C, Haynes P A.2011. Less label, more free: Approaches in label-free quantitative mass spectrometry.Proteomics, 11(4): 535-553. |
| [77] | Niittylä T, Fuglsang A T, Palmgren M G, Frommer W B, Schulze W X.2007. Temporal analysis of sucrose-induced phosphorylation changes in plasma membrane proteins ofArabidopsis. Mol Cell Proteomics, 6(10): 1711-1726. |
| [78] | Nilsson C L.2012. Advances in quantitative phosphoproteomics.Anal Chem, 84(2): 735-746. |
| [79] | Nishi H, Shaytan A, Panchenko A R.2014. Physicochemical mechanisms of protein regulation by phosphorylation.Front Genet, 5: 270. |
| [80] | Nühse T S, Stensballe A, Jensen O N, Peck S C.2004. Phospho- proteomics of theArabidopsis plasma membrane and a new phosphorylation site database. Plant Cell, 16(9): 2394-2405. |
| [81] | Oda Y, Huang K, Cross F R, Cowburn D, Chait B T.1999. Accurate quantitation of protein expression and site-specific phosphorylation.Proc Natl Acad Sci USA, 96(12): 6591-6596. |
| [82] | Ong S E, Blagoev B, Kratchmarova I, Kristensen D B, Steen H, Pandey A, Mann M.2002. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics.Mol Cell Proteomics, 1(5): 376-386. |
| [83] | Ong S E, Mann M.2005. Mass spectrometry-based proteomics turns quantitative.Nat Chem Biol, 1: 252-262. |
| [84] | Ozlu N, Akten B, Timm W, Haseley N, Steen H, Steen J.2010. Phosphoproteomics.Wiley Interdiscip Rev: Syst Biol Med, 2(3): 255-276. |
| [85] | Qiu J H, Hou Y X, Tong X H, Wang Y F, Lin H Y, Liu Q, Zhang W, Li Z Y, Nallamilli B R, Zhang J.2015. Quantitative phosphoproteomic analysis of early seed development in rice (Oryza sativa L.). Plant Mol Biol, 90(3): 249-265. |
| [86] | Qiu J H, Hou Y X, Wang Y F, Li Z Y, Zhao J, Tong X H, Lin H Y, Wei X J, Ao H J, Zhang J.2017. A comprehensive proteomic survey of ABA-induced protein phosphorylation in rice (Oryza sativa L.). Int J Mol Sci, 18(1): E60. |
| [87] | Reiland S, Finazzi G, Endler A, Willig A, Baerenfaller K, Grossmann J, Gerrits B, Rutishauser D, Gruissem W, Rochaix J D.2011. Comparative phosphoproteome profiling reveals a function of the STN8 kinase in fine-tuning of cyclic electron flow (CEF).Proc Natl Acad Sci USA, 108: 12955-12960. |
| [88] | Reinders J, Sickmann A.2005. State-of-the-art in phospho- proteomics.Proteomics, 5(16): 4052-4061. |
| [89] | Ross P L, Huang Y N, Marchese J N, Williamson B, Parker K, Hattan S, Khainovski N, Pillai S, Dey S, Daniels S, Purkayastha S, Juhasz P, Martin S, Bartlet-Jones M, He F, Jacobson A, Pappin D J.2004. Multiplexed protein quantitation inSaccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol Cell Proteomics, 3(12): 1154-1169. |
| [90] | Schaff J E, Mbeunkui F, Blackburn K, Bird D M, Goshe M B.2008. SILIP: A novel stable isotope labeling method for in planta quantitative proteomic analysis.Plant J, 56(5): 840-854. |
| [91] | Schmidt S R, Schweikart F, Andersson M E.2007. Current methods for phosphoprotein isolation and enrichment.J Chromatogr B Analyt Technol Biomed Life Sci., 849: 154-162. |
| [92] | Schulze W X, Schneider T, Starck S, Martinoia E, Trentmann O.2012. Cold acclimation induces changes inArabidopsis tonoplast protein abundance and activity and alters phosphorylation of tonoplast monosaccharide transporters. Plant J, 69(3): 529-541. |
| [93] | Shen F, Hu Y, Guan P, Ren X.2012. Ti4+-phosphate functionalized cellulose for phosphopeptides enrichment and its application in rice phosphoproteome analysis.J Chromatogr B Analyt Technol Biomed Life Sci, 902: 108-115. |
| [94] | Silva-Sanchez C, Li H Y, Chen S X.2015. Recent advances and challenges in plant phosphoproteomics.Proteomics, 15: 1127-1141. |
| [95] | Singh A, Giri J, Kapoor S, Tyagi A K, Pandey G K.2010. Protein phosphatase complement in rice: Genome-wide identification and transcriptional analysis under abiotic stress conditions and reproductive development.BMC Genom, 11: 435. |
| [96] | Stokes M P, Farnsworth C L, Moritz A, Silva J C, Jia X Y, Lee K A, Guo A L, Polakiewicz R D, Comb M J.2012. PTMScan direct: Identification and quantification of peptides from critical signaling proteins by immunoaffinity enrichment coupled with LC-MS/MS.Mol Cell Proteomics, 11(5): 187-201. |
| [97] | Sutherland E W, Wosilait W D.1955. Inactivation and activation of liver phosphorylase.Nature, 175: 169-170. |
| [98] | Tan F, Li G S, Chitteti B R, Peng Z H.2007. Proteome and phosphoproteome analysis of chromatin associated proteins in rice (Oryza sativa). Proteomics, 7: 4511-4527. |
| [99] | Trinidad J C, Specht C G, Thalhammer A, Schoepfer R, Burlingame A L.2006. Comprehensive identification of phosphorylation sites in postsynaptic density preparations.Mol Cell Proteomics, 5(5): 914-922. |
| [100] | Unlu M, Morgan M E, Minden J S.1997. Difference gel electrophoresis: A single gel method for detecting changes in protein extracts.Electrophoresis, 18(11): 2071-2077. |
| [101] | Walton A, Stes E, de Smet I, Goormachtig S, Gevaert K.2015. Plant hormone signalling through the eye of the mass spectrometer.Proteomics, 15: 1113-1126. |
| [102] | Wang K, Zhao Y, Li M, Gao F, Yang M K, Wang X, Li S Q, Yang P F.2014. Analysis of phosphoproteome in rice pistil.Proteomics, 14(20): 2319-2334. |
| [103] | Wang P C, Xue L, Batelli G, Lee S, Hou Y J, van Oosten M J, Zhang H M, Tao W A, Zhu J K.2013. Quantitative phosphoproteomics identifies SnRK2 protein kinase substrates and reveals the effectors of abscisic acid action.Proc Natl Acad Sci USA, 110: 11205-11210. |
| [104] | Wang W F, Bai M Y, Wang Z Y.2014. The brassinosteroid signaling network: A paradigm of signal integration.Curr Opin Plant Biol, 21: 147-153. |
| [105] | Wang Y F, Tong X H, Qiu J H, Li Z Y, Zhao J, Hou Y X, Tang L Q, Zhang J.2017. A phosphoproteomic landscape of rice (Oryza sativa) tissues. Physiol Plant, 160(4): 458-475. |
| [106] | Wang Z Y, Bai M Y, Oh E, Zhu J Y.2012. Brassinosteroid signaling network and regulation of photomorphogenesis.Annu Rev Genet, 46: 701-724. |
| [107] | Watson T J, Sparkman D O.2007. Introduction to Mass Spectrometry: Instrumentation, Applications, and Strategies of Data Interpretation. Chichester West Sussex, England: John Wiley & Sons, Ltd. |
| [108] | Wu K J, Shaler T A, Becker C H.1994. Time-of-flight mass spectrometry of underivatized single-stranded DNA oligomers by matrix-assisted laser desorption.Anal Chem, 66(10): 1637-1645. |
| [109] | Xiang F, Ye H, Chen R B, Fu Q, Li L J.2010. N, N-dimethyl leucines as novel isobaric tandem mass tags for quantitative proteomics and peptidomics.Anal Chem, 82(7): 2817-2825. |
| [110] | Yang Z, Guo G Y, Zhang M Y, LiuC Y, Hu Q, Lam H, Cheng H, Xue Y, Li J Y, Li N.2013. Stable isotope metabolic labeling- based quantitative phosphoproteomic analysis ofArabidopsis mutants reveals ethylene-regulated time-dependent phosphoproteins and putative substrates of constitutive triple response 1 kinase. Mol Cell Protemics, 12(12): 3559-3582. |
| [111] | Yao X D, Freas A, Ramirez J, Demirev P A, Fenselau C.2001. Proteolytic 18O labeling for comparative proteomics: Model studies with two serotypes of adenovirus.Anal Chem, 73(13): 2836-2842. |
| [112] | Ye J Y, Zhang Z B, Long H F, Zhang Z M, Hong Y, Zhang X M, You C J, Liang W Q, Ma H, Lu P L.2015. Proteomic and phosphoproteomic analyses reveal extensive phosphorylation of regulatory proteins in developing rice anthers.Plant J, 84: 527-544. |
| [113] | Yi E C, Li X J, Cooke K, Lee H, Raught B, Page A, Aneliunas V, Hieter P, Goodlett D R, Aebersold R.2005. Increased quantitative proteome coverage with 13C/12C-based, acid- cleavable isotope-coded affinity tag reagent and modified data acquisition scheme.Proteomics, 5(2): 380-387. |
| [114] | Yoshida S.1972. Physiological aspects of grain yield.Ann Rev Plant Physiol, 23: 437-464. |
| [115] | Zhang H T, Zhou H J, Berke L, Heck A J R, Mohammed S, Scheres B, Menke F L H.2013. Quantitative phosphoproteomics after auxin-stimulated lateral root induction identifies a SNX1 phosphorylation site required for growth.Mol Cell Proteomics, 12(5): 1158-1169. |
| [116] | Zhang J, Guo D, Chang Y X, You C J, Li X W, Dai X X, Weng Q J, Zhang J W, Chen G X, Li X H, Liu H F, Han B, Zhang Q F, Wu C Y.2007. Non-random distribution of T-DNA insertions at various levels of the genome hierarchy as revealed by analyzing 13804 T-DNA flanking sequences from an enhancer-trap mutant library.Plant J, 49: 947-959. |
| [117] | Zhang Z X, Chen J, Lin S S, Li Z, Cheng R H, Fang C X, Chen H F, Lin W X.2012. Proteomic and phosphoproteomic determination of ABA’s effects on grain-filling ofOryza sativa L. inferior spikelets. Plant Sci, 185/186: 259-273. |
| [118] | Zhang Z X, Zhao H, Tang J, Li Z, Li Z, Chen D M, Lin W X.2014. A proteomic study on molecular mechanism of poor grain-filling of rice (Oryza sativa L.) inferior spikelets. PLoS One, 9(2): e89140. |
| [119] | Zhao Y M, Jensen O N.2009. Modification-specific proteomics: Strategies for characterization of post-translational modifications using enrichment techniques.Proteomics, 9: 4632-4641. |
| [120] | Zhong M, Li S F, Huang F L, Qiu J H, Zhang J, Sheng Z H, Tang S Q, Wei X J, Hu P S.2017. The phosphoproteomic response of rice seedlings to cadmium stress.Int J Mol Sci, 18: E2055. |
| [121] | Zuo J R, Li J Y.2014. Molecular dissection of complex agronomic traits of rice: A team effort by Chinese scientists in recent years. Nat Sci Rev, 1(2): 253-276. |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [13] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||