
Rice Science ›› 2021, Vol. 28 ›› Issue (5): 457-465.DOI: 10.1016/j.rsci.2021.07.006
• Research Paper • Previous Articles Next Articles
Junjie Yu1, Mina Yu1, Tianqiao Song1, Huijuan Cao1, Mingli Yong1, Xiayan Pan1, Zhongqiang Qi1, Yan Du1, Rongsheng Zhang1, Xiaole Yin1, Dong Liang1, Yongfeng Liu1,2( )
)
Received:2020-09-14
Accepted:2021-01-12
Online:2021-09-28
Published:2021-09-28
Junjie Yu, Mina Yu, Tianqiao Song, Huijuan Cao, Mingli Yong, Xiayan Pan, Zhongqiang Qi, Yan Du, Rongsheng Zhang, Xiaole Yin, Dong Liang, Yongfeng Liu. UvSMEK1, a Suppressor of MEK Null, Regulates Pathogenicity, Conidiation and Conidial Germination in Rice False Smut Fungus Ustilaginoidea virens[J]. Rice Science, 2021, 28(5): 457-465.
Add to citation manager EndNote|Ris|BibTeX
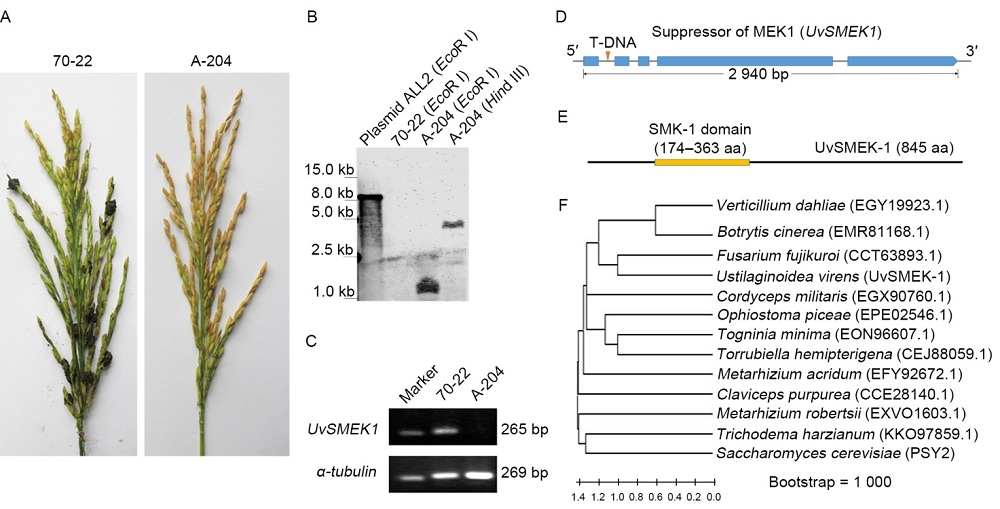
Fig. 1. Identification of mutated gene in U. virens T-DNA insertion mutant A-204.A, Rice false smuts on rice spikelets inoculated with wild-type strain 70-22 and T-DNA insertion mutant A-204. B, Detection of copy number of T-DNA inserted in the genome of A-204 by Southern blotting. C, Detection of UvSMEK1 expression via reversed-transcription PCR, and α-tubulin was employed as a reference gene. D, Inserted site of T-DNA in the coding region of UvSMEK1 in U. virens mutant A-204. E, Conserved SMK-1 domain in UvSMEK1. F, Phylogenetic analysis of SMEK-1 homologs to UvSMEK-1 in fungi.
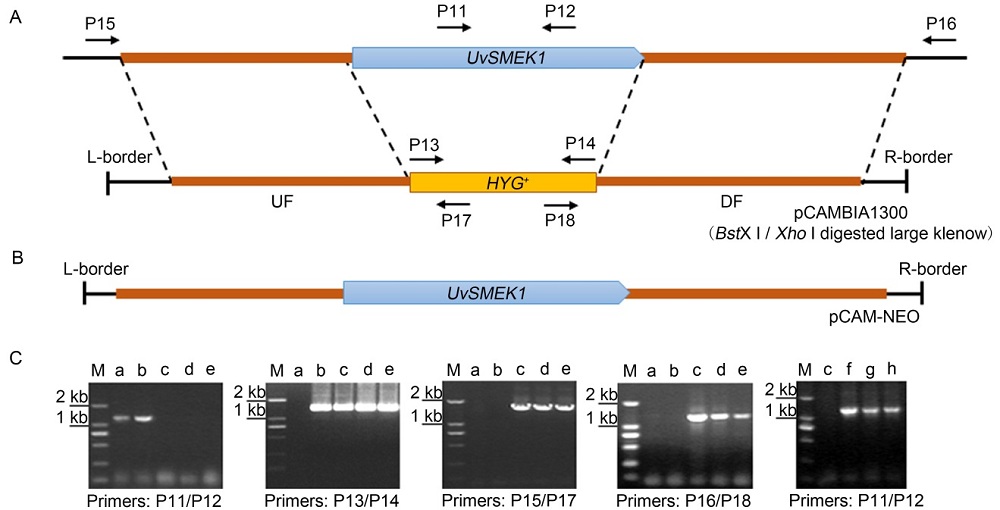
Fig. 2. Deletion of UvSMEK1 gene in U. virens.A, Illustration of targeted deletion of UvSMEK1 by Agrobacterium tumefaciens-mediated transformation (ATMT) and homologous replacement. P11 to P18 represent the primers. UF, Upstream flank of UvSMEK1; HYG+, Hygromycin resistant gene; DF, Downstream flank of UvSMEK1.B, Illustration of complementation cassette of UvSMEK1 gene by ATMT transformation. C, Reversed-transcription PCR analysis of UvSMEK1 deletion mutants and UvSMEK1 complement mutants. The wild type strain P-1 and transformant Trans-9 with ectopically inserted UF-HYG+-RF cassette were included as controls. M, Marker; a, P-1; b, Trans-9; c, ΔUvSMEK1-51; d, ΔUvSMEK1-71; e, ΔUvSMEK1-177; f, CUvSMEK1-1; g, CUvSMEK1-2; h, CUvSMEK1-6.
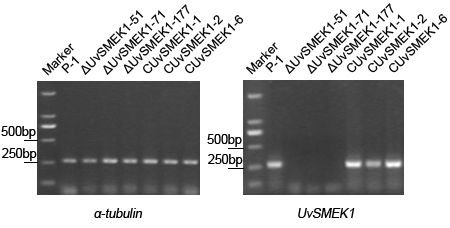
Fig. S1. Detection of UvSMEK1 expression via reversed-transcription PCR (RT-PCR).The expression of UvSMEK1 gene in the wild-type strain P-1, UvSMEK1 deletion mutants ΔUvSMEK1-51, ΔUvSMEK1-71, ΔUvSMEK1-177, and UvSMEK1 completentary mutants CUvSMEK1-1, CUvSMEK1-2, CUvSMEK1-6 was detected. House keeping α-tubulin gene was employed as a reference gene.
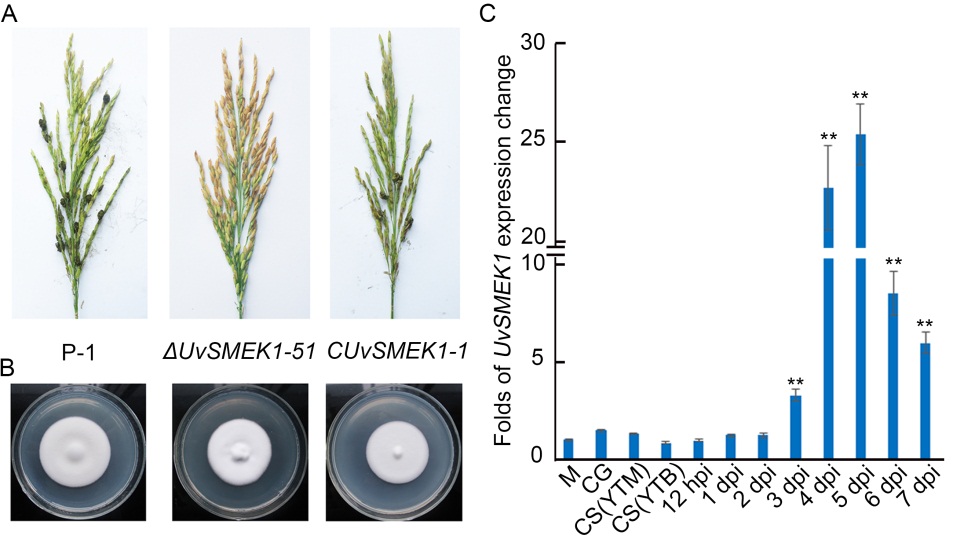
Fig. 3. Characterization of UvSMEK1 deletion and complement mutants in U. virens.A, Rice false smut balls on rice spikelets inoculated with wild type strain P-1, UvSMEK-1 deletion mutant ΔUvSMEK1-51 and complement mutant CUvSMEK1-1. B, Colonies of P-1, ΔUvSMEK1-51 and CUvSMEK1-1 on potato sucrose agar at 28 ºC after 12 d. C, Expression pattern of UvSMEK1 was determined by quantitative PCR. House-keeping gene α-tubulin was employed as a reference gene. Data are Mean ± SD (n = 3). **, Significant difference at the 0.01 level (t-test). M, Mycelia; CG, Conidial germination; CS(YTM), Conidial sporulation on yeast extract tryptone (YT) media; CS(YTB), Conidial sporulation in YT broth; hpi, Hours post inoculation; dpi, Days post inoculation.
| Strain | Mycelial growth on PSA (mm) a | Pathogenicity b | Concentration of conidia in YT broth (Log10 of spores/mL) c | Percentage of conidial gemination on YT medium (%) d | Percentage of micro-cycle conidiation on YT medium (%) d |
|---|---|---|---|---|---|
| P-1 | 49.00 ± 2.16 Aa | 13.7 ± 2.7 Aa | 6.51 ± 0.03 BCc | 90.75 ± 1.71 Aa | 19.75 ± 4.27 Aa |
| ΔUvSMEK1-51 | 48.75 ± 0.96 Aa | 0 Cc | 6.89 ± 0.03 Aa | 89.25 ± 3.59 Aa | 0 Bb |
| ΔUvSMEK1-71 | 48.75 ± 1.71 Aa | 0 Cc | 6.90 ± 0.02 Aa | 88.50 ± 2.38 Aa | 0 Bb |
| ΔUvSMEK1-177 | 47.75 ± 1.71 Aa | 0 Cc | 6.90 ± 0.03 Aa | 89.00 ± 2.71 Aa | 0 Bb |
| CUvSMEK1-1 | 47.75 ± 2.22 Aa | 3.7 ± 2.2 Bb | 6.58 ± 0.03 Bb | 89.50 ± 2.89 Aa | 15.75 ± 2.87 Aa |
| CUvSMEK1-2 | 46.75 ± 1.71 Aa | 3.2 ± 1.6 Bb | 6.50 ± 0.03 Cc | 90.00 ± 2.94 Aa | 18.75 ± 2.22 Aa |
| CUvSMEK1-6 | 48.25 ± 2.06 Aa | 4.1 ± 1.9 Bb | 6.57 ± 0.03 BCb | 89.00 ± 2.58 Aa | 14.25 ± 2.99 Aa |
Table 1 Pathogenicity, conidiation and conidial germination in UvSMEK1 deletion and complement mutants.
| Strain | Mycelial growth on PSA (mm) a | Pathogenicity b | Concentration of conidia in YT broth (Log10 of spores/mL) c | Percentage of conidial gemination on YT medium (%) d | Percentage of micro-cycle conidiation on YT medium (%) d |
|---|---|---|---|---|---|
| P-1 | 49.00 ± 2.16 Aa | 13.7 ± 2.7 Aa | 6.51 ± 0.03 BCc | 90.75 ± 1.71 Aa | 19.75 ± 4.27 Aa |
| ΔUvSMEK1-51 | 48.75 ± 0.96 Aa | 0 Cc | 6.89 ± 0.03 Aa | 89.25 ± 3.59 Aa | 0 Bb |
| ΔUvSMEK1-71 | 48.75 ± 1.71 Aa | 0 Cc | 6.90 ± 0.02 Aa | 88.50 ± 2.38 Aa | 0 Bb |
| ΔUvSMEK1-177 | 47.75 ± 1.71 Aa | 0 Cc | 6.90 ± 0.03 Aa | 89.00 ± 2.71 Aa | 0 Bb |
| CUvSMEK1-1 | 47.75 ± 2.22 Aa | 3.7 ± 2.2 Bb | 6.58 ± 0.03 Bb | 89.50 ± 2.89 Aa | 15.75 ± 2.87 Aa |
| CUvSMEK1-2 | 46.75 ± 1.71 Aa | 3.2 ± 1.6 Bb | 6.50 ± 0.03 Cc | 90.00 ± 2.94 Aa | 18.75 ± 2.22 Aa |
| CUvSMEK1-6 | 48.25 ± 2.06 Aa | 4.1 ± 1.9 Bb | 6.57 ± 0.03 BCb | 89.00 ± 2.58 Aa | 14.25 ± 2.99 Aa |
| Strain | Conidiaa | Abnormal germinating conidiab | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Lengthc (μm) | Widthc (μm) | Length-width ratio | Width of germ tube | Length (μm) | Widthd (μm) | Length-width ratio | Ratio of abnormal germination (%) | Width of germ tube | ||
| P-1 | 7.83 ± 1.05 Aa | 4.74 ± 0.72 Aa | 1.67 ± 0.31 Aa | 3.24 ± 0.47 Aa | - | - | - | - | - | |
| ΔUvSMEK1-51 | 7.74 ± 0.98 Aa | 4.54 ± 0.67 Aa | 1.71 ± 0.29 Aa | - | 20.47 ± 1.63 Aa | 13.63 ± 1.20 Aa | 1.50 ± 0.08 Aa | 75.25 ± 6.18 Aa | 5.72 ± 1.05 Aa | |
| ΔUvSMEK1-71 | 7.67 ± 1.11 Aa | 4.52 ± 0.62 Aa | 1.69 ± 0.37 Aa | - | 19.90 ± 1.93 Aa | 12.89 ± 1.68 Aa | 1.56 ± 0.11 Aa | 74.75 ± 8.01 Aa | 5.93 ± 0.98 Aa | |
| ΔUvSMEK1-177 | 7.85 ± 1.24 Aa | 4.66 ± 0.58 Aa | 1.68 ± 0.36 Aa | - | 20.89 ± 1.84 Aa | 13.99 ± 1.77 Aa | 1.49 ± 0.13 Aa | 72.75 ± 7.18 Aa | 5.29 ± 1.12 Aa | |
| CUvSMEK1-1 | 7.49 ± 1.01 Aa | 4.51 ± 0.70 Aa | 1.67 ± 0.40 Aa | 3.07 ± 0.39 Aa | - | - | - | - | - | |
| CUvSMEK1-2 | 7.68 ± 1.14 Aa | 4.64 ± 0.68 Aa | 1.66 ± 0.31 Aa | 3.11 ± 0.31 Aa | - | - | - | - | - | |
| CUvSMEK1-6 | 7.84 ± 1.27 Aa | 4.71 ± 0.59 Aa | 1.67 ± 0.38 Aa | 3.18 ± 0.43 Aa | - | - | - | - | - | |
Table S1 Size and length-width ratio of UvSMEK1 deletion/complement U. virens mutants in YT broth.
| Strain | Conidiaa | Abnormal germinating conidiab | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Lengthc (μm) | Widthc (μm) | Length-width ratio | Width of germ tube | Length (μm) | Widthd (μm) | Length-width ratio | Ratio of abnormal germination (%) | Width of germ tube | ||
| P-1 | 7.83 ± 1.05 Aa | 4.74 ± 0.72 Aa | 1.67 ± 0.31 Aa | 3.24 ± 0.47 Aa | - | - | - | - | - | |
| ΔUvSMEK1-51 | 7.74 ± 0.98 Aa | 4.54 ± 0.67 Aa | 1.71 ± 0.29 Aa | - | 20.47 ± 1.63 Aa | 13.63 ± 1.20 Aa | 1.50 ± 0.08 Aa | 75.25 ± 6.18 Aa | 5.72 ± 1.05 Aa | |
| ΔUvSMEK1-71 | 7.67 ± 1.11 Aa | 4.52 ± 0.62 Aa | 1.69 ± 0.37 Aa | - | 19.90 ± 1.93 Aa | 12.89 ± 1.68 Aa | 1.56 ± 0.11 Aa | 74.75 ± 8.01 Aa | 5.93 ± 0.98 Aa | |
| ΔUvSMEK1-177 | 7.85 ± 1.24 Aa | 4.66 ± 0.58 Aa | 1.68 ± 0.36 Aa | - | 20.89 ± 1.84 Aa | 13.99 ± 1.77 Aa | 1.49 ± 0.13 Aa | 72.75 ± 7.18 Aa | 5.29 ± 1.12 Aa | |
| CUvSMEK1-1 | 7.49 ± 1.01 Aa | 4.51 ± 0.70 Aa | 1.67 ± 0.40 Aa | 3.07 ± 0.39 Aa | - | - | - | - | - | |
| CUvSMEK1-2 | 7.68 ± 1.14 Aa | 4.64 ± 0.68 Aa | 1.66 ± 0.31 Aa | 3.11 ± 0.31 Aa | - | - | - | - | - | |
| CUvSMEK1-6 | 7.84 ± 1.27 Aa | 4.71 ± 0.59 Aa | 1.67 ± 0.38 Aa | 3.18 ± 0.43 Aa | - | - | - | - | - | |
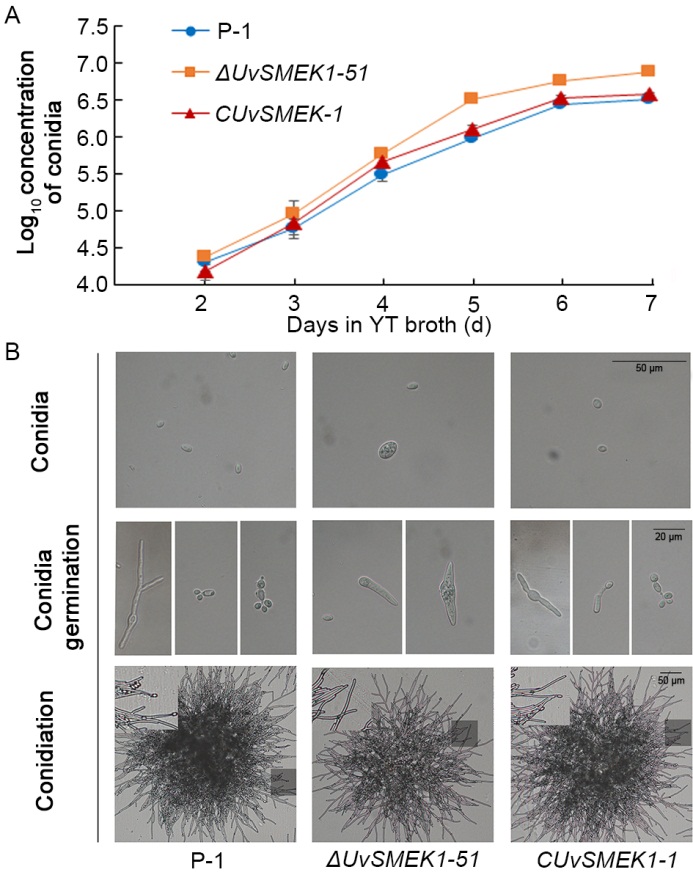
Fig. 4. Conidiation, conidial germination and hyphal branching of UvSMEK-1 deletion mutant and complement mutant.A, Log10 concentration of conidia produced in yeast extract tryptone (YT) broth in 7 d.B, Conidia, germination of conidia and conidiation on YT medium of UvSMEK-1 deletion mutant ΔUvSMEK1-51 and complement mutant CUvSMEK1-1 after 3-day culture.
| Strain | Log10 (concentration of conidia)a | |||||
|---|---|---|---|---|---|---|
| 2 dpi | 3 dpi | 4 dpi | 5 dpi | 6 dpi | 7 dpi | |
| P-1 | 4.33 ± 0.13 | 4.77 ± 0.13 | 5.49 ± 0.09 | 5.99 ± 0.03 | 6.45 ± 0.02 | 6.51 ± 0.02 |
| ΔUvSMEK1-51 | 4.39 ± 0.10 | 4.97 ± 0.17 | 5.78 ± 0.05 | 6.52 ± 0.02 | 6.76 ± 0.02 | 6.89 ± 0.03 |
| ΔUvSMEK1-71 | 4.36 ± 0.11 | 4.85 ± 0.15 | 5.57 ± 0.07 | 6.50 ± 0.03 | 6.79 ± 0.01 | 6.90 ± 0.02 |
| ΔUvSMEK1-177 | 4.16 ± 0.11 | 4.75 ± 0.15 | 5.51 ± 0.03 | 6.45 ± 0.04 | 6.75 ± 0.02 | 6.90 ± 0.02 |
| CUvSMEK1-1 | 4.19 ± 0.12 | 4.85 ± 0.15 | 5.68 ± 0.05 | 6.11 ± 0.05 | 6.54 ± 0.03 | 6.58 ± 0.02 |
| CUvSMEK1-2 | 4.22 ± 0.15 | 4.77 ± 0.13 | 5.68 ± 0.05 | 6.03 ± 0.04 | 6.49 ± 0.02 | 6.51 ± 0.02 |
| CUvSMEK1-6 | 4.19 ± 0.12 | 4.85 ± 0.15 | 5.68 ± 0.05 | 6.09 ± 0.06 | 6.54 ± 0.01 | 6.57 ± 0.02 |
| a Conidia of U. virens strain/mutants were produced in YT broth(28 °C, 150 r/min)for 6 d. | ||||||
Table S2 Concentration of conidia produced by UvSMEK1 deletion/complement U. virens mutants in YT broth.
| Strain | Log10 (concentration of conidia)a | |||||
|---|---|---|---|---|---|---|
| 2 dpi | 3 dpi | 4 dpi | 5 dpi | 6 dpi | 7 dpi | |
| P-1 | 4.33 ± 0.13 | 4.77 ± 0.13 | 5.49 ± 0.09 | 5.99 ± 0.03 | 6.45 ± 0.02 | 6.51 ± 0.02 |
| ΔUvSMEK1-51 | 4.39 ± 0.10 | 4.97 ± 0.17 | 5.78 ± 0.05 | 6.52 ± 0.02 | 6.76 ± 0.02 | 6.89 ± 0.03 |
| ΔUvSMEK1-71 | 4.36 ± 0.11 | 4.85 ± 0.15 | 5.57 ± 0.07 | 6.50 ± 0.03 | 6.79 ± 0.01 | 6.90 ± 0.02 |
| ΔUvSMEK1-177 | 4.16 ± 0.11 | 4.75 ± 0.15 | 5.51 ± 0.03 | 6.45 ± 0.04 | 6.75 ± 0.02 | 6.90 ± 0.02 |
| CUvSMEK1-1 | 4.19 ± 0.12 | 4.85 ± 0.15 | 5.68 ± 0.05 | 6.11 ± 0.05 | 6.54 ± 0.03 | 6.58 ± 0.02 |
| CUvSMEK1-2 | 4.22 ± 0.15 | 4.77 ± 0.13 | 5.68 ± 0.05 | 6.03 ± 0.04 | 6.49 ± 0.02 | 6.51 ± 0.02 |
| CUvSMEK1-6 | 4.19 ± 0.12 | 4.85 ± 0.15 | 5.68 ± 0.05 | 6.09 ± 0.06 | 6.54 ± 0.01 | 6.57 ± 0.02 |
| a Conidia of U. virens strain/mutants were produced in YT broth(28 °C, 150 r/min)for 6 d. | ||||||
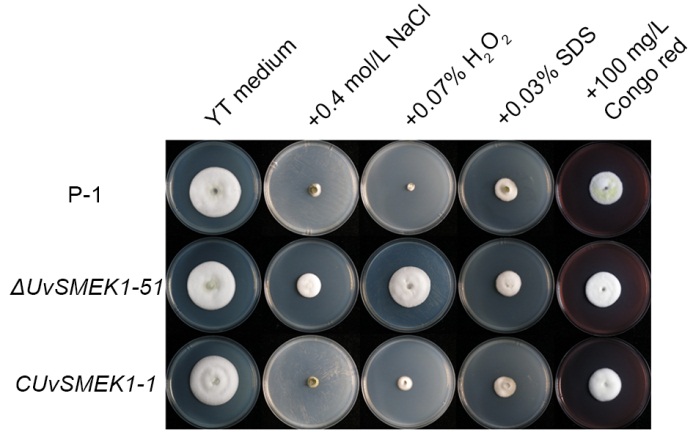

Fig. 5. Growth of UvSMEK1 deletion and complement mutants in presence of different biotic stresses.Wild type strain P-1, UvSMEK1 deletion mutant ΔUvSMEK1-51 and UvSMEK1 complement mutant CUvSMEK1-1 were cultured on plain yeast extract tryptone (YT) medium or amended with 0.4 mol/L NaCl, 0.07% H2O2, 0.03% sodium dodecyl sulfate (SDS) and 100 mg/L Congo red at 28 ºC for 15 d.
| Strain | YT medium | YT (+0.4 mol/L NaCl) | YT (+0.07% H2O2) | YT (+0.03% SDS) | YT (+100 mg/L Congo red) |
|---|---|---|---|---|---|
| P-1 | 44.17 ± 1.76 Aa | 8.00 ± 0.50 Bc | 0 Cc | 18.67 ± 0.76 Aa | 29.83 ± 0.76 Aa |
| ΔUvSMEK1-51 | 45.00 ± 1.00 Aa | 18.33 ± 0.29 Ab | 33.67 ± 0.76 Aa | 18.17 ± 0.76 Aa | 29.83 ± 1.04 Aa |
| ΔUvSMEK1-71 | 42.50 ± 1.80 Aa | 19.50 ± 0.50 Aa | 33.00 ± 1.32 Aa | 17.00 ± 1.00 Aa | 27.67 ± 0.76 Aa |
| ΔUvSMEK1-177 | 44.00 ± 1.50 Aa | 18.22 ± 0.29 Ab | 32.00 ± 0.50 Aa | 17.67 ± 1.04 Aa | 27.67 ± 1.04 Aa |
| CUvSMEK1-1 | 43.83 ± 2.02 Aa | 7.00 ± 0.50 Bcd | 8.67 ± 0.76 Bb | 18.50 ± 0.50 Aa | 27.00 ± 0.86 Aa |
| CUvSMEK1-2 | 42.83 ± 1.26 Aa | 6.83 ± 0.29 Bcd | 7.50 ± 0.50 Bb | 16.50 ± 1.32 Aa | 26.50 ± 1.32 Aa |
| CUvSMEK1-6 | 43.83 ± 1.26 Aa | 7.33 ± 0.29 Bcd | 8.17 ± 1.04 Bb | 16.57 ± 1.44 Aa | 26.83 ± 1.04 Aa |
Table 2 Responses of mycelial growth of UvSMEK1 deletion and complement mutants to abiotic stress. mm
| Strain | YT medium | YT (+0.4 mol/L NaCl) | YT (+0.07% H2O2) | YT (+0.03% SDS) | YT (+100 mg/L Congo red) |
|---|---|---|---|---|---|
| P-1 | 44.17 ± 1.76 Aa | 8.00 ± 0.50 Bc | 0 Cc | 18.67 ± 0.76 Aa | 29.83 ± 0.76 Aa |
| ΔUvSMEK1-51 | 45.00 ± 1.00 Aa | 18.33 ± 0.29 Ab | 33.67 ± 0.76 Aa | 18.17 ± 0.76 Aa | 29.83 ± 1.04 Aa |
| ΔUvSMEK1-71 | 42.50 ± 1.80 Aa | 19.50 ± 0.50 Aa | 33.00 ± 1.32 Aa | 17.00 ± 1.00 Aa | 27.67 ± 0.76 Aa |
| ΔUvSMEK1-177 | 44.00 ± 1.50 Aa | 18.22 ± 0.29 Ab | 32.00 ± 0.50 Aa | 17.67 ± 1.04 Aa | 27.67 ± 1.04 Aa |
| CUvSMEK1-1 | 43.83 ± 2.02 Aa | 7.00 ± 0.50 Bcd | 8.67 ± 0.76 Bb | 18.50 ± 0.50 Aa | 27.00 ± 0.86 Aa |
| CUvSMEK1-2 | 42.83 ± 1.26 Aa | 6.83 ± 0.29 Bcd | 7.50 ± 0.50 Bb | 16.50 ± 1.32 Aa | 26.50 ± 1.32 Aa |
| CUvSMEK1-6 | 43.83 ± 1.26 Aa | 7.33 ± 0.29 Bcd | 8.17 ± 1.04 Bb | 16.57 ± 1.44 Aa | 26.83 ± 1.04 Aa |
| Strain/plasmid | Description in brief | Reference |
|---|---|---|
| P-1 | Wild-type strain of U. virens | Yu et al, 2015 |
| A-204 | T-DNA insertional mutant | Yu et al, 2015 |
| ΔUvSMEK-51 | UvSMEK1 deletion mutant | This study |
| ΔUvSMEK-71 | UvSMEK1 deletion mutant | This study |
| ΔUvSMEK-177 | UvSMEK1 deletion mutant | This study |
| CUvSMEK1-1 | UvSMEK1 complement mutant | This study |
| CUvSMEK1-2 | UvSMEK1 complement mutant | This study |
| CUvSMEK1-6 | UvSMEK1 complement mutant | This study |
| pCAMBIA1300 | Binary vector for Agrobacterium tumefaciens mediated transformation | A stock in lab |
| pCAM-NEO | Binary vector containing geneticin 418 resistance | A stock in lab |
| pD-UvSMEK | Binary vector for UvSMEK1 deletion | This study |
| pC-UvSMEK | Binary vector for UvSMEK1 complementation | This study |
| AGL-1 | Agrobacterium tumefaciens strain | A stock in lab |
Table S3 Strains and vectors used in this study.
| Strain/plasmid | Description in brief | Reference |
|---|---|---|
| P-1 | Wild-type strain of U. virens | Yu et al, 2015 |
| A-204 | T-DNA insertional mutant | Yu et al, 2015 |
| ΔUvSMEK-51 | UvSMEK1 deletion mutant | This study |
| ΔUvSMEK-71 | UvSMEK1 deletion mutant | This study |
| ΔUvSMEK-177 | UvSMEK1 deletion mutant | This study |
| CUvSMEK1-1 | UvSMEK1 complement mutant | This study |
| CUvSMEK1-2 | UvSMEK1 complement mutant | This study |
| CUvSMEK1-6 | UvSMEK1 complement mutant | This study |
| pCAMBIA1300 | Binary vector for Agrobacterium tumefaciens mediated transformation | A stock in lab |
| pCAM-NEO | Binary vector containing geneticin 418 resistance | A stock in lab |
| pD-UvSMEK | Binary vector for UvSMEK1 deletion | This study |
| pC-UvSMEK | Binary vector for UvSMEK1 complementation | This study |
| AGL-1 | Agrobacterium tumefaciens strain | A stock in lab |
| Primer | Sequences (5′ to 3′) | Usage |
|---|---|---|
| P1 | CACACCACTCTCGAGCACTC | For qPCR or RT-PCR detection of α-tubulin expression |
| P2 | GCCTTGGCAGCCGAGATGAC | |
| P3 | CGGACATCTTGGTCTCCATC | For qPCR or RT-PCR detection of UvSMEK1 expression |
| P4 | CGAGGATCGGATGCAGGT | |
| P5 | CACACATTATTATGGAGAAACTCGAGGATGTTCCCGAGATCCAGTG | For construction of UvSMEK1 deletion vector (XhoI BSTXi) |
| P6 | GCTCCTTCAATATCATCTTCTGTAGGTCGGGAGCAATGGCAAG | |
| P7 | CGAGGGCAAAGGAATAGAGTATGGAGCCAAGCGTCACCAAC | |
| P8 | TTGGCTAGAGCAGCTTGCCAACATGGACAGACACCGTTCCAACGAC | |
| P9 | CAGCTATGACCATGATTACGAATTCGCCGCTTGTCCACTCCACCAC | For construction of UvSMEK1 complement vector(EcorI SalI) |
| P10 | CCAAGCTTGCATGCCTGCAGGTCGACACCACGCGAACGGGAAGAAC | |
| P11 | TGCAGCAGACGTTTCCCAAC | For PCR detection of UvSMEK1 DNA sequence |
| P12 | AGACAGGAGTTCGACGAGAG | |
| P13 | ACAGAAGATGATATTGAAGGAGC | For PCR detection of hygromycin-resistant cassette in U. virens mutants |
| P14 | TACTCTATTCCTTTGCCCTCG | |
| P15 | TGTCCTCGTTCCTGTCTGCTA | |
| P16 | TGCGACGCAATCGTCCGATC | |
| P17 | ACGGAGGAGCCTCGCAC | |
| P18 | CGCTTCCGCACCATCGAC |
Table S4 Primers used in this study.
| Primer | Sequences (5′ to 3′) | Usage |
|---|---|---|
| P1 | CACACCACTCTCGAGCACTC | For qPCR or RT-PCR detection of α-tubulin expression |
| P2 | GCCTTGGCAGCCGAGATGAC | |
| P3 | CGGACATCTTGGTCTCCATC | For qPCR or RT-PCR detection of UvSMEK1 expression |
| P4 | CGAGGATCGGATGCAGGT | |
| P5 | CACACATTATTATGGAGAAACTCGAGGATGTTCCCGAGATCCAGTG | For construction of UvSMEK1 deletion vector (XhoI BSTXi) |
| P6 | GCTCCTTCAATATCATCTTCTGTAGGTCGGGAGCAATGGCAAG | |
| P7 | CGAGGGCAAAGGAATAGAGTATGGAGCCAAGCGTCACCAAC | |
| P8 | TTGGCTAGAGCAGCTTGCCAACATGGACAGACACCGTTCCAACGAC | |
| P9 | CAGCTATGACCATGATTACGAATTCGCCGCTTGTCCACTCCACCAC | For construction of UvSMEK1 complement vector(EcorI SalI) |
| P10 | CCAAGCTTGCATGCCTGCAGGTCGACACCACGCGAACGGGAAGAAC | |
| P11 | TGCAGCAGACGTTTCCCAAC | For PCR detection of UvSMEK1 DNA sequence |
| P12 | AGACAGGAGTTCGACGAGAG | |
| P13 | ACAGAAGATGATATTGAAGGAGC | For PCR detection of hygromycin-resistant cassette in U. virens mutants |
| P14 | TACTCTATTCCTTTGCCCTCG | |
| P15 | TGTCCTCGTTCCTGTCTGCTA | |
| P16 | TGCGACGCAATCGTCCGATC | |
| P17 | ACGGAGGAGCCTCGCAC | |
| P18 | CGCTTCCGCACCATCGAC |
| [1] | Byun H J, Kim B R, Yoo R, Park S Y, Rho S B. 2012. sMEK1 enhances gemcitabine anti-cancer activity through inhibition of phosphorylation of Akt/mTOR. Apoptosis, 17(10): 1095-1103. |
| [2] | Chen X Y, Hai D, Tang J T, Liu H, Huang J B, Luo C X, Hsiang T, Zheng L. 2020. UvCom1 is an important regulator required for development and infection in the rice false smut fungus Ustilaginoidea virens. Phytopathology, 110(2): 483-493. |
| [3] | Fan J, Guo X Y, Li L, Huang F, Sun W X, Li Y, Huang Y Y, Xu Y J, Shi J, Lei Y, Zheng A P, Wang W M. 2015. Infection of Ustilaginoidea virens intercepts rice seed formation but activates grain-filling-related genes. J Integr Plant Biol, 57(6): 577-590. |
| [4] | Fan J, Yang J, Wang Y Q, Li G B, Li Y, Huang F, Wang W M. 2016. Current understanding on Villosiclava virens, a unique flower- infecting fungus causing rice false smut disease. Mol Plant Pathol, 17(9): 1321‒1330. |
| [5] | Fan J, Liu J, Gong Z Y, Xu P Z, Hu X H, Wu J L, Li G B, Yang J, Wang Y Q, Zhou Y F, Li S C, Wang L, Chen X Q, He M, Zhao J Q, Li Y, Huang Y Y, Hu D W, Wu X J, Li P, Wang W M. 2020. The false smut pathogen Ustilaginoidea virens requires rice stamens for false smut ball formation. Environ Microbiol, 22(2): 646-659. |
| [6] | Fang A F, Gao H, Zhang N, Zheng X H, Qiu S S, Li Y J, Zhou S, Cui F H, Sun W X. 2019. A novel effector gene SCRE2 contributes to full virulence of Ustilaginoidea virens to rice. Front Microbiol, 10: 845. |
| [7] | Gingras A C, Caballero M, Zarske M, Sanchez A, Hazbun T R, Fields S, Sonenberg N, Hafen E, Raught B, Aebersold R. 2005. A novel, evolutionarily conserved protein phosphatase complex involved in cisplatin sensitivity. Mol Cell Proteomics, 4(11): 1725-1740. |
| [8] | Guest G M, Lin X R, Momany M. 2004. Aspergillus nidulans RhoA is involved in polar growth, branching, and cell wall synthesis. Fungal Genet Biol, 41(1): 13-22. |
| [9] | Guo W W, Gao Y X, Yu Z M, Xiao Y H, Zhang Z G, Zhang H F. 2019. The adenylate cyclase UvAc1 and phosphodiesterase UvPdeH control the intracellular cAMP level, development, and pathogenicity of the rice false smut fungus Ustilaginoidea virens. Fungal Genet Biol, 129: 65-73. |
| [10] | Hustedt N, Seeber A, Sack R, Tsai-Pflugfelder M, Bhullar B, Vlaming H, van Leeuwen F, Guénolé A, van Attikum H, Srivas R, Ideker R, Shimada K, Gasser S M. 2015. Yeast PP4 interacts with ATR homolog Ddc2-Mec1 and regulates checkpoint signaling. Mol Cell, 57(2): 273-289. |
| [11] | Kim B R, Seo S H, Park M S, Lee S H, Kwon Y, Rho S B. 2015. sMEK1 inhibits endothelial cell proliferation by attenuating VEGFR-2-dependent-Akt/eNOS/HIF-1 alpha signaling pathways. Oncotarget, 6(31): 31830-31843. |
| [12] | Kim B R, Kwon Y, Rho S B. 2017. BMI-1 interacts with sMEK1 and inactivates sMEK1-induced apoptotic cell death. Oncol Rep, 37(1): 579-586. |
| [13] | Knechtle P, Dietrich F, Philippsen P. 2003. Maximal polar growth potential depends on the polarisome component AgSpa2 in the filamentous fungus Ashbya gossypii. Mol Biol Cell, 14(10): 4140-4154. |
| [14] | Koiso Y, Li Y, Iwasaki S, Hanaka K, Kobayashi T, Sonoda R, Fujita Y, Yaegashi H, Sato Z. 1994. Ustiloxins, antimitotic cyclic peptides from false smut balls on rice panicles caused by Ustilaginoidea virens. J Antibiot, 47(7): 765-773. |
| [15] | Li Y, Koiso Y, Kobayashi H, Hashimoto Y, Iwasaki S. 1995. Ustiloxins, new antimitotic cyclic peptides: Interaction with porcine brain tubulin. Biochem Pharmacol, 49(10): 1367-1372. |
| [16] | Liang Y F, Han Y, Wang C F, Jiang C, Xu J R. 2018. Targeted deletion of the USTA and UvSLT2 genes efficiently in Ustilaginoidea virens with the CRISPR-Cas9 system. Front Plant Sci, 9: 699. |
| [17] | Liu Z Y, Liu N, Jiang H X, Yan L Y, Ma Z H, Yin Y N. 2018. The activators of type 2A phosphatases (PP2A) regulate multiple cellular processes via PP2A-dependent and -independent mechanisms in Fusarium graminearum. Mol Plant Microbe, 31(11): 1121-1133. |
| [18] | Lü B, Zheng L, Liu H, Tang J T, Hsiang T, Huang J B. 2016. Use of random T-DNA mutagenesis in identification of gene UvPRO1, a regulator of conidiation, stress response, and virulence in Ustilaginoidea virens. Front Microbiol, 7: 2086. |
| [19] | Lyu J, Kim H R, Yamamoto V, Choi S H, Wei Z, Joo C K, Lu W. 2013. Protein phosphatase 4 and Smek complex negatively regulate Par3 and promote neuronal differentiation of neural stem/progenitor cells. Cell Rep, 5(3): 593-600. |
| [20] | Ma H, Han B K, Guaderrama M, Aslanian A, Yates III J R, Hunter T, Wittenberg C. 2014. Psy2 targets the PP4 family phosphatase Pph3 to dephosphorylate Mth1 and repress glucose transporter gene expression. Mol Cell Biol, 34(3): 452-463. |
| [21] | Mendoza M C, Du F, Iranfar N, Tang N, Ma H, Loomis W F, Firtel R A. 2005. Loss of SMEK, a novel, conserved protein, suppresses mek1 null cell polarity, chemotaxis, and gene expression defects. Mol Cell Biol, 25(17): 7839-7853. |
| [22] | Mendoza M C, Booth E O, Shaulsky G, Firtel R A. 2007. MEK1 and protein phosphatase 4 coordinate dictyostelium development and chemotaxis. Mol Cell Biol, 27(10): 3817-3827. |
| [23] | Meng J J, Sun W B, Mao Z L, Xu D, Wang X H, Lu S Q, Lai D W, Liu Y, Zhou L G, Zhang G Z. 2015. Main ustilaginoidins and their distribution in rice false smut balls. Toxins, 7(10): 4023-4034. |
| [24] | Omidi K, Hooshyar M, Jessulat M, Samanfar B, Sanders M, Burnside D, Pitre S, Schoenrock A, Xu J, Babu M, Golshani A. 2014. Phosphatase complex Pph3/Psy2 is involved in regulation of efficient non-homologous end-joining pathway in the yeast Saccharomyces cerevisiae. PLoS One, 9(1): e87248. |
| [25] | Qiu J H, Meng S, Deng Y Z, Huang S W, Kou Y J. 2019. Ustilaginoidea virens: A fungus infects rice flower and threats world rice production. Rice Sci, 26(4): 199-206. |
| [26] | Sun L L, Li W J, Wang H T, Chen J, Deng P, Wang Y, Sang J L. 2011. Protein phosphatase Pph3 and its regulatory subunit Psy2 regulate Rad53 dephosphorylation and cell morphogenesis during recovery from DNA damage in Candida albicans. Eukaryot Cell, 10(11): 1565-1573. |
| [27] | Sun X Y, Kang S, Zhang Y J, Tan X Q, Yu Y F, He H Y, Zhang X Y, Liu Y F, Wang S, Sun W X, Cai L, Li S J. 2013. Genetic diversity and population structure of rice pathogen Ustilaginoidea virens in China. PLoS One, 8(9): e76879. |
| [28] | Takano Y, Takayanagi N, Hori H, Ikeuchi Y, Suzuki T, Kimura A, Okuno T. 2006. A gene involved in modifying transfer RNA is required for fungal pathogenicity and stress tolerance of Colletotrichum lagenarium. Mol Microbiol, 60(1): 81-92. |
| [29] | Tang J T, Bai J, Chen X Y, Zheng L, Liu H, Huang J B. 2020. Two protein kinases UvPmk1 and UvCDC2 with significant functions in conidiation, stress response and pathogenicity of rice false smut fungus Ustilaginoidea virens. Curr Genet, 66(2): 409-420. |
| [30] | Tang Y X, Jin J, Hu D W, Yong M L, Xu Y, He L P. 2013. Elucidation of the infection process of Ustilaginoidea virens (teleomorph: Villosiclava virens) in rice spikelets. Plant Pathol, 62(1): 1‒8. |
| [31] | Tuorto F, Herbst F, Alerasool N, Bender S, Popp O, Federico G, Reitter S, Liebers R, Stoecklin G, Gröne H J, Dittmar G, Glimm H, Lyko F. 2015. The tRNA methyltransferase Dnmt2 is required for accurate polypeptide synthesis during haematopoiesis. EMBO J, 34(18): 2350-2362. |
| [32] | Wolff S, Ma H, Burch D, Maciel G A, Hunter T, Dillin A. 2006. SMK-1, an essential regulator of DAF-16-mediated longevity. Cell, 124(5): 1039-1053. |
| [33] | Xie S L, Wang Y F, Wei W, Li C Y, Liu Y, Qu J S, Meng Q H, Lin Y, Yin W X, Yang Y N, Luo C X. 2019. The Bax inhibitor UvBI-1, a negative regulator of mycelial growth and conidiation, mediates stress response and is critical for pathogenicity of the rice false smut fungus Ustilaginoidea virens. Curr Genet, 65(5): 1185-1197. |
| [34] | Xu P, Wang H, Tu R R, Liu Q N, Wu W X, Fu X M, Cao L Y, Shen X H. 2019. Orientation improvement of blast resistance in rice via CRISPR/Cas9 system. Chin J Rice Sci, 33(4): 313‒322. (in Chinese with English abstract) |
| [35] | Yong M L, Fan L L, Li D Y, Liu Y J, Cheng F M, Xu Y, Wang Z Y, Hu D W. 2016. Villosiclava virens infects specifically rice and barley stamen filaments due to the unique host cell walls. Microsc Res Tech, 79(9): 838-844. |
| [36] | Yoon Y S, Lee M W, Ryu D, Kim J H, Ma H, Seo W Y, Kim Y N, Kim S S, Lee C H, Hunter T, Choi C S, Montminy M R, Koo S H. 2010. Suppressor of MEK null (SMEK)/protein phosphatase 4 catalytic subunit (PP4C) is a key regulator of hepatic gluconeogenesis. Proc Natl Acad Sci USA, 107(41): 17704-17709. |
| [37] | Yu J J, Sun W X, Yu M N, Yin X L, Meng X K, Zhao J, Huang L, Huang L, Liu Y F. 2015. Characterization of mating-type loci in rice false smut fungus Villosiclava virens. FEMS Microbiol Lett, 362(9): fnv014. |
| [38] | Yu J J, Yu M N, Song T Q, Cao H J, Pan X Y, Yong M L, Qi Z Q, Du Y, Zhang R S, Yin X L, Liu Y F. 2019. A homeobox transcription factor UvHOX2 regulates chlamydospore formation, conidiogenesis, and pathogenicity in Ustilaginoidea virens. Front Microbiol, 10: 1071. |
| [39] | Yu M N, Yu J J, Hu J K, Huang L, Wang Y H, Yin X L, Nie Y F, Meng X K, Wang W D, Liu Y F. 2015. Identification of pathogenicity-related genes in the rice pathogen Ustilaginoidea virens through random insertional mutagenesis. Fungal Genet Biol, 76: 10-19. |
| [40] | Zhang N, Yang J Y, Fang A F, Wang J Y, Li D Y, Li Y J, Wang S Z, Cui F H, Yu J J, Liu Y F, Peng Y L, Sun W X. 2020. The essential effector SCRE1 in Ustilaginoidea virens suppresses rice immunity via a small peptide region. Mol Plant Pathol, 21(4): 445-459. |
| [41] | Zhang Y, Zhang K, Fang A F, Han Y Q, Yang J, Xue M F, Bao J D, Hu D W, Zhou B, Sun X Y, Li S J, Wen M, Yao N, Ma L J, Liu Y F, Zhang M, Huang F, Luo C X, Zhou L, Li J Q, Chen Z Y, Miao J K, Wang S, Lai J S, Xu J R, Hsiang T, Peng Y L, Sun W X. 2014. Specific adaptation of Ustilaginoidea virens in occupying host florets revealed by comparative and functional genomics. Nat Commun, 5: 3849. |
| [42] | Zheng D W, Wang Y, Han Y, Xu J R, Wang C F. 2016. UvHOG1 is important for hyphal growth and stress responses in the rice false smut fungus Ustilaginoidea virens. Sci Rep, 6: 24824. |
| [43] | Zheng M T, Ding H, Huang L, Wang Y H, Yu M N, Zheng R, Yu J J, Liu Y F. 2017. Low-affinity iron transport protein Uvt3277 is important for pathogenesis in the rice false smut fungus Ustilaginoidea virens. Curr Genet, 63(1): 131-144. |
| [1] | Wei Qinghui, Cui Daizong, Zheng Baojiang, Zhao Min. In Vitro Antifungal Activity of Dihydrochelerythrine and Proteomic Analysis in Ustilaginoidea virens [J]. Rice Science, 2023, 30(3): 257-266. |
| [2] | Liu Yueran, Qu Jinsong, Wang Yufu, Yin Weixiao, Luo Chaoxi. bZIP Transcription Factor UvATF21 Mediates Vegetative Growth, Conidiation, Stress Tolerance and Is Required for Full Virulence of Rice False Smut Fungus Ustilaginoidea virens [J]. Rice Science, 2023, 30(1): 50-57. |
| [3] | Wu Zhongling, Qiu Jiehua, Shi Huanbin, Lin Chuyu, Yue Jiangnan, Liu Zhiquan, Xie Wei, Naweed I. Naqvi, Kou Yanjun, Tao Zeng. Polycomb Repressive Complex 2-Mediated H3K27 Trimethylation Is Required for Pathogenicity in Magnaporthe oryzae [J]. Rice Science, 2022, 29(4): 363-374. |
| [4] | Meng Shuai, Qiu Jiehua, Xiong Meng, Liu Zhiquan, Jane Sadhna Jagernath, Lin Fucheng, Shi Huanbin, Kou Yanjun. UvWhi2 Is Required for Stress Response and Pathogenicity in Ustilaginoidea virens [J]. Rice Science, 2022, 29(1): 47-54. |
| [5] | Tianqiao Song, Xiong Zhang, You Zhang, Dong Liang, Jiaoling Yan, Junjie Yu, Mina Yu, Huijuan Cao, Mingli Yong, Xiayan Pan, Zhongqiang Qi, Yan Du, Rongsheng Zhang, Yongfeng Liu. Genome-Wide Identification of Zn2Cys6 Class Fungal-Specific Transcription Factors (ZnFTFs) and Functional Analysis of UvZnFTF1 in Ustilaginoidea virens [J]. Rice Science, 2021, 28(6): 567-578. |
| [6] | Meng Xiong, Shuai Meng, Jiehua Qiu, Huanbin Shi, Xiangling Shen, Yanjun Kou. Putative Phosphatase UvPsr1 Is Required for Mycelial Growth, Conidiation, Stress Response and Pathogenicity in Ustilaginonidea virens [J]. Rice Science, 2020, 27(6): 529-536. |
| [7] | Jiehua Qiu, Shuai Meng, Yizhen Deng, Shiwen Huang, Yanjun Kou. Ustilaginoidea virens: A Fungus Infects Rice Flower and Threats World Rice Production [J]. Rice Science, 2019, 26(4): 199-206. |
| [8] | WANG Jiao-yu1, WANG Xiao-yan1, 2, LI Ling1, ZHANG Xin1, WANG Yan-li1, CHAI Rong-yao1, SUN Guo-chang1. Pathogenicity of Rice Blast Fungus Magnaporthe oryzae on Brachypodium distachyon [J]. RICE SCIENCE, 2012, 19(3): 252-258. |
| [9] | XIAO Yong, LIU Ming-wei, LI Gang, ZHOU Er-xun, WANG Ling-xia, TANG Jie, TAN Fu-rong, ZHENG Ai-ping, LI Ping . Genetic Diversity and Pathogenicity Variation in Rhizoctonia solani Isolates from Rice in Sichuan Province, China [J]. RICE SCIENCE, 2008, 15(2): 137-144 . |
| [10] | ZHU Bo, LOU Miao-miao, HUAI Yan, XIE Guan-lin, LUO Jin-yan, XU Li-hui. Isolation and Identification of Burkholderia glumae from Symptomless Rice Seeds [J]. RICE SCIENCE, 2008, 15(2): 145-149 . |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||