
Rice Science ›› 2021, Vol. 28 ›› Issue (6): 567-578.DOI: 10.1016/j.rsci.2021.03.001
• Research Paper • Previous Articles Next Articles
Tianqiao Song1,2, Xiong Zhang3, You Zhang1, Dong Liang1, Jiaoling Yan1, Junjie Yu1, Mina Yu1, Huijuan Cao1, Mingli Yong1, Xiayan Pan1, Zhongqiang Qi1, Yan Du1, Rongsheng Zhang1, Yongfeng Liu1,4( )
)
Received:2020-09-14
Accepted:2021-03-01
Online:2021-11-28
Published:2021-11-28
Tianqiao Song, Xiong Zhang, You Zhang, Dong Liang, Jiaoling Yan, Junjie Yu, Mina Yu, Huijuan Cao, Mingli Yong, Xiayan Pan, Zhongqiang Qi, Yan Du, Rongsheng Zhang, Yongfeng Liu. Genome-Wide Identification of Zn2Cys6 Class Fungal-Specific Transcription Factors (ZnFTFs) and Functional Analysis of UvZnFTF1 in Ustilaginoidea virens[J]. Rice Science, 2021, 28(6): 567-578.
Add to citation manager EndNote|Ris|BibTeX
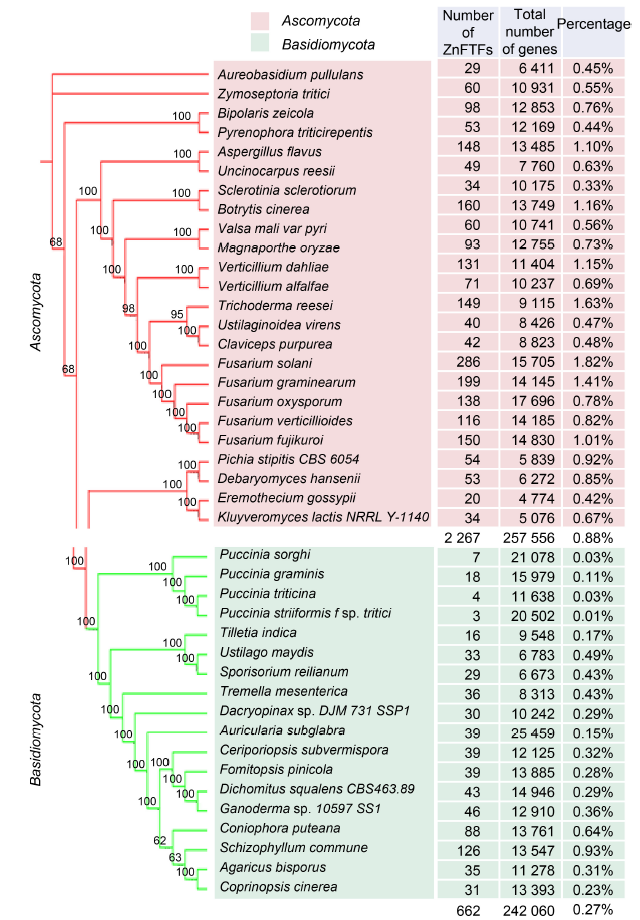
Fig. 1. Distribution of ZnFTFs in fungal species.Phylogenetic relationships of fungal species are shown in the left. The maximum likelihood tree was constructed from a concatenated alignment of core eukaryotic genes identified by the Core Eukaryotic Genes Mapping Approach analysis. The number of ZnFTFs, total number of genes, and percentage of ZnFTFs out of total number of genes in each species are listed in the right.
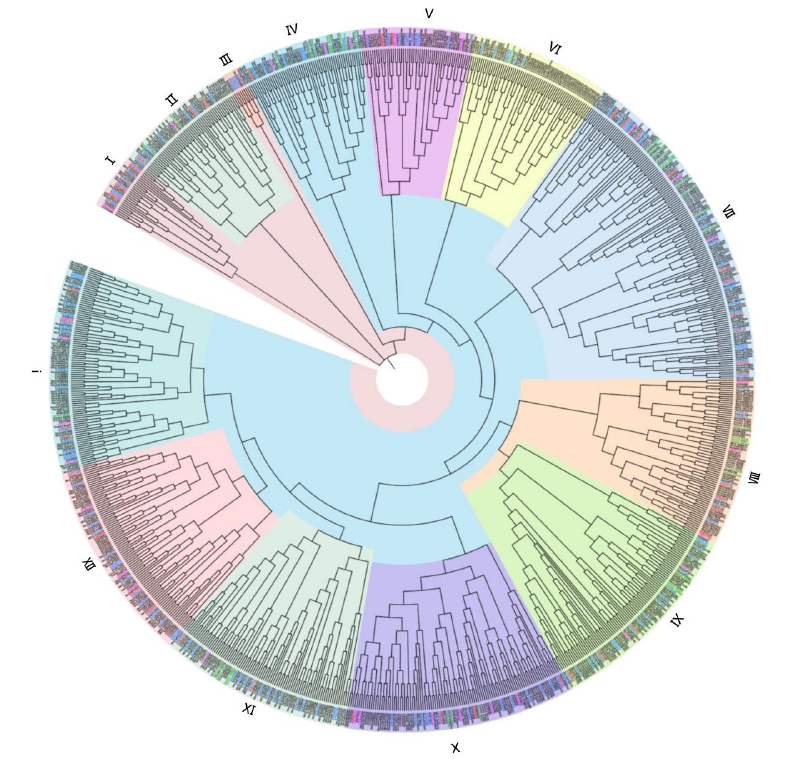
Fig. S1. Phylogenetic analysis of ZnFTFs from diverse fungal species.The genes included in this tree are ZnFTFs from Debaryomyces hansenii (CAG/CAR), Kluyveromyces lactis (CAG/CAH), Sporisorium reilianum (CBQ), Coprinopsis cinerea (CC1G), Claviceps purpurea (CCE), Ceriporiopsis subvermispora (Cersu1), Uncinocarpus reesii (EEP), Trichoderma reesei (EGR), Agaricus bisporus (EKM), Ustilaginoidea virens (KDB), Ustilago maydis (KIS), Magnaporthe oryzae (MGG), Fusarium solani (Necha), Tilletia indica (OAI99796), and Tremella mesenterica (Treme1). The tree was constructed using the neighbor-joining algorithm. These genes can be classified into thirteen groups. Each group is labeled in different background colors and with different Roman numerals. The gene IDs are shown on the outer ring of the circle. Gene IDs from representative pathogenic Ascomycota, non-pathogenic Ascomycota, pathogenic Basidiomycota, and non-pathogenic Basidiomycota species are colored as follows: U. virens, red; T. reesei, blue; S. reilianum, pink; C. subvermispora green. Gene IDs from all other species are shown in black.
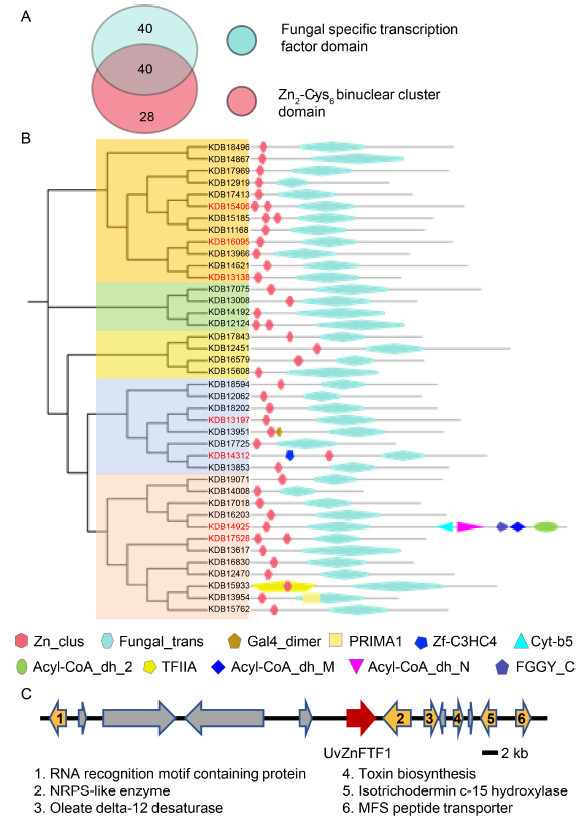
Fig. 2. Characterization of UvZnFTFs in U. virens.A, Venn diagram shows the overlap in proteins, containing the fungal specific transcription factor domain and Zn2-Cys6 binuclear cluster domain.B, Phylogenetic analyses of ZnFTFs in U. virens. The phylogenetic tree was constructed using Geneious R11 by the neighbor-joining method. Domain organization is indicated below.C, UvZnFTF1 is located within a putative secondary metabolite-related gene cluster that encodes proteins possibly involved in toxin biosynthesis.MFS, Major facilitator superfamily; NRPS, Non- ribosomal peptide synthetase.
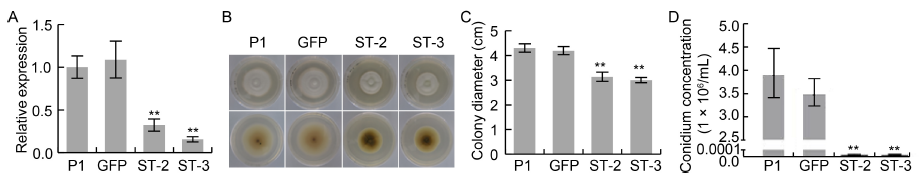
Fig. 3. UvZnFTF1 is involved in vegetative growth, pigment biosynthesis and conidiation of U. virens.A, Relative transcript levels of UvZnFTF1 in the wild type strain P1 and different transgenic U. virens lines (GFP, ST-2 and ST-3). The transcript levels were measured by quantitative real-time PCR using the β-tubulin gene as an internal reference.B, Colony morphology of P1, GFP, ST-2 and ST-3 cultured on potato sucrose agar plates after 14 d of incubation in the dark at 28 ºC. The upper and lower panels represent the front and back of the plates, respectively.C, Colony diameters of P1, GFP, ST-2 and ST-3.D, Statistical analysis of conidium concentration.A U. virens strain expressing GFP was served as a transgenic negative control. Data in A, C and D are Mean ± SE (n = 3). **, P < 0.01 by the Dunnett's test.
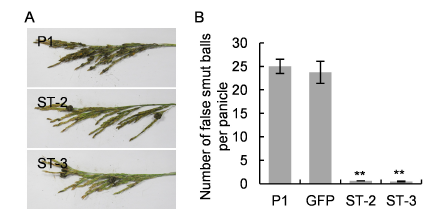
Fig. 4. UvZnFTF1-silenced transformants showed reduced pathogenicity on rice panicles.A, Representative photographs of disease symptoms on rice panicles at 30 d post inoculation with U. virens transformants (ST-2 and ST-3) and the wild type strain P1.B, Quantification of the number of false smut balls per panicle. Data are Mean ± SE (n = 3). **, P < 0.01 by the Dunnett's test.
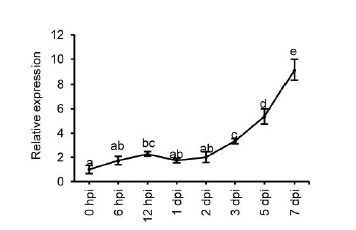
Fig. S2. UvZnFTF1 expression profile during infection.Rice panicles were inoculated with a suspension of U. virens mycelia and conidia and collected at the indicated times post inoculation (hpi, hours post inoculation; dpi, days post inoculation). Values are the means ± SEM of three independent biological replicates. Letters represent significant differences P<0.01 as measured by Duncan’s multiple range test.
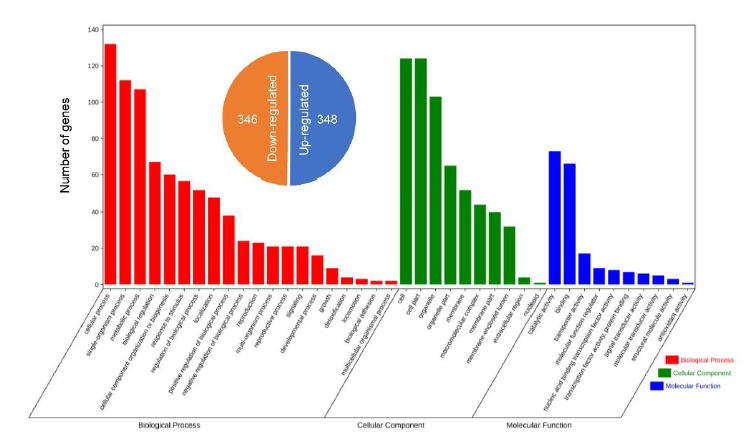
Fig. S3. Gene Ontology annotations of differentially expressed genes in UvZnFTF1-silenced transformants vs. the wild type strain.DEGs are genes with a more than 4-fold change in expression in the UvZNFTF1-silenced transformants compared with the WT strain. The x-axis represents the GO terms. The red bars represent ‘Biological Process’ terms, green bars represent ‘Cellular Component’ terms, and blue bars are the ‘Molecular Function’ terms. The y-axis shows the number of genes for each enriched GO term.
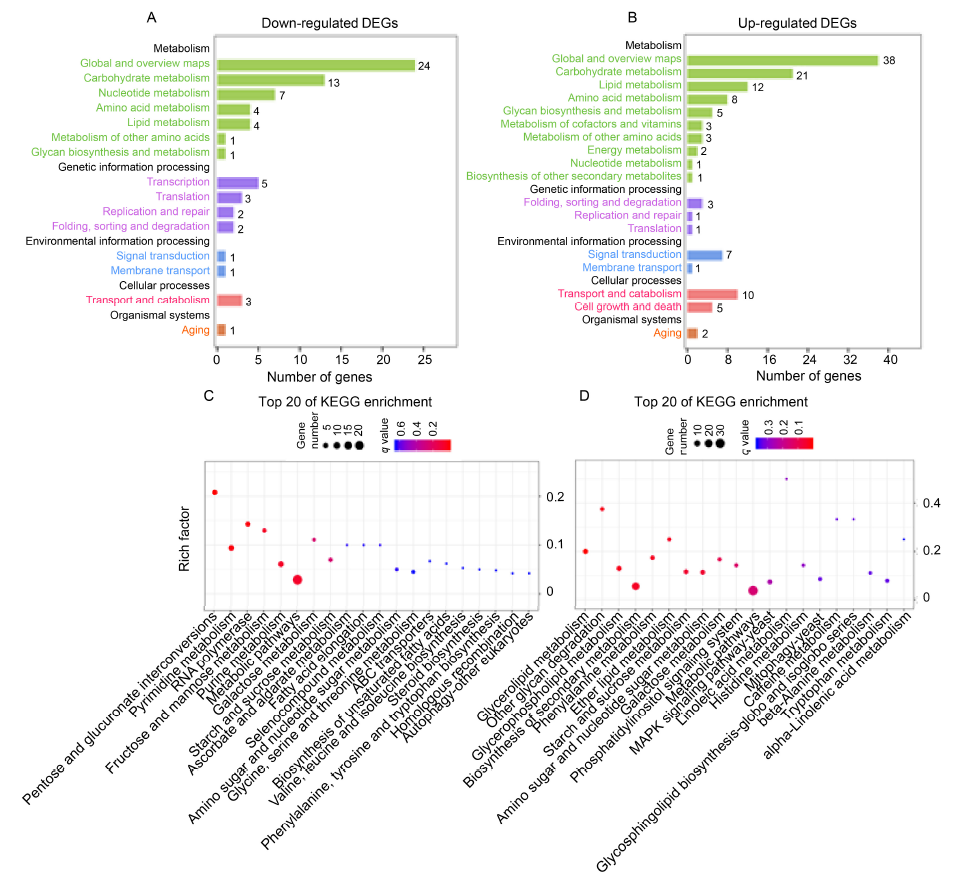
Fig. 5. Kyoto Encyclopedia of Genes and Genomes (KEGG) functional annotations of differentially expressed genes (DEGs) in UvZnFTF1- silenced transformants vs. wild type strain.A and B, KEGG pathway annotations of DEGs. The number of genes related to different KEGG pathways was calculated. The KEGG pathways are listed according to their molecular interactions and reaction networks within ‘Metabolism’ (light green bars), ‘Genetic information processing’ (purple bars), ‘Environmental information processing’ (blue bars), ‘Cellular processes’ (red bars) and ‘Organismal systems’ (orange bar).C and D, Top 20 down-regulated DEGs (C) and up-regulated DEGs (D) enriched KEGG pathways. The color of the bubble represents the corrected q value. The size of the bubble indicates the number of DEGs. The rich factor is the proportion of DEGs in each pathway.
| Gene ID | Biological function | Up-/down-regulated a |
|---|---|---|
| Genes involved in signal transduction | ||
| KDB11856, KDB16287, KDB16312, KDB18181, KDB18666, KDB11669 | G protein-coupled receptor gene | Up |
| KDB12196, KDB13191, KDB12444, KDB17214, KDB11081, KDB16465, KDB12620 | G protein-coupled receptor gene | Down |
| KDB17719, KDB14771, KDB17609, KDB18180, KDB17527, KDB17948, KDB13363, KDB18665, KDB14539, KDB13362, KDB13535, KDB12267 | Transcription factor | Up |
| KDB13663, KDB18728, KDB14016, KDB13664, KDB12105, KDB11763, KDB15876, KDB18718, KDB18880, KDB11956, KDB17862, KDB11405 | Transcription factor | Down |
| KDB14812, KDB14811, KDB11045 | Zn2Cys6 transcription factor | Up |
| KDB11606 | Zn2Cys6 transcription factor | Down |
| KDB13647 | Transcription factor for Polarity Control 1 | Down |
| KDB18369 | Histidine kinase | Up |
| KDB16000, KDB13339, KDB11031 | Protein kinase | Up |
| KDB13650, KDB18230 | Protein kinase | Down |
| Genes involved in transport | ||
| KDB18524 | Putative mitochondrial ornithine transporter | Up |
| KDB16595, KDB15193 | BCAA transporter | Up |
| KDB14098 | BCAA transporter | Down |
| KDB13509, KDB13238 | Copper transporter | Up |
| KDB11479, KDB11766, KDB15819 | Glucose uptake and transport | Up |
| Genes encoding secreted proteins | ||
| KDB15463, KDB11767 | Necrosis inducing protein | Up |
| KDB14361, KDB17416 | Effector protein | Down |
| KDB17641 | LysM effector protein | Down |
| KDB14258, KDB15607 | Small secreted fungal protein | Down |
| KDB12171 | Proteinaceous elicitor | Down |
Table 1 Representative PHI genes regulated by UvZnFTF1.
| Gene ID | Biological function | Up-/down-regulated a |
|---|---|---|
| Genes involved in signal transduction | ||
| KDB11856, KDB16287, KDB16312, KDB18181, KDB18666, KDB11669 | G protein-coupled receptor gene | Up |
| KDB12196, KDB13191, KDB12444, KDB17214, KDB11081, KDB16465, KDB12620 | G protein-coupled receptor gene | Down |
| KDB17719, KDB14771, KDB17609, KDB18180, KDB17527, KDB17948, KDB13363, KDB18665, KDB14539, KDB13362, KDB13535, KDB12267 | Transcription factor | Up |
| KDB13663, KDB18728, KDB14016, KDB13664, KDB12105, KDB11763, KDB15876, KDB18718, KDB18880, KDB11956, KDB17862, KDB11405 | Transcription factor | Down |
| KDB14812, KDB14811, KDB11045 | Zn2Cys6 transcription factor | Up |
| KDB11606 | Zn2Cys6 transcription factor | Down |
| KDB13647 | Transcription factor for Polarity Control 1 | Down |
| KDB18369 | Histidine kinase | Up |
| KDB16000, KDB13339, KDB11031 | Protein kinase | Up |
| KDB13650, KDB18230 | Protein kinase | Down |
| Genes involved in transport | ||
| KDB18524 | Putative mitochondrial ornithine transporter | Up |
| KDB16595, KDB15193 | BCAA transporter | Up |
| KDB14098 | BCAA transporter | Down |
| KDB13509, KDB13238 | Copper transporter | Up |
| KDB11479, KDB11766, KDB15819 | Glucose uptake and transport | Up |
| Genes encoding secreted proteins | ||
| KDB15463, KDB11767 | Necrosis inducing protein | Up |
| KDB14361, KDB17416 | Effector protein | Down |
| KDB17641 | LysM effector protein | Down |
| KDB14258, KDB15607 | Small secreted fungal protein | Down |
| KDB12171 | Proteinaceous elicitor | Down |
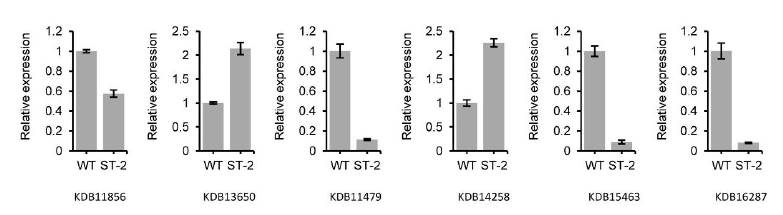
Fig. S4. Quantitative real-time PCR analysis to validate differentially expressed genes.The relative transcript levels of KDB11856 (GPCR gene), KDB13650 (protein kinase), KDB11479 (Glucose uptake and transport), KDB14258 (Small secreted fungal protein), KDB15463 (Necrosis inducing protein), and KDB16287(GPCR gene) in silencing transformants ST-2 was normalized to those in WT line, using the Tubulin gene as an internal reference. Means and standard deviations obtained from three replicates are shown.
| [1] | Brown D W, Villani A, Susca A, Moretti A, Hao G X, Kim H S, Proctor R H, McCormick S P. 2020. Gain and loss of a transcription factor that regulates late trichothecene biosynthetic pathway genes in Fusarium. Fungal Genet Biol, 136: 103317. |
| [2] | Chang P K, Ehrlich K C. 2013. Genome-wide analysis of the Zn(II)2Cys6 zinc cluster-encoding gene family in Aspergillus flavus. Appl Microbiol Biotechnol, 97(10): 4289-4300. |
| [3] | Chen X Y, Hai D, Tang J T, Liu H, Huang J B, Luo C X, Hsiang T, Zheng L. 2020. UvCom1 is an important regulator required for development and infection in the rice false smut fungus Ustilaginoidea virens. Phytopathology, 110(2): 483-493. |
| [4] | Cho Y, Ohm R A, Grigoriev I V, Srivastava A. 2013. Fungal- specific transcription factor AbPf2 activates pathogenicity in Alternaria brassicicola. Plant J, 75(3): 498-514. |
| [5] | Fan J, Yang J, Wang Y Q, Li G B, Li Y, Huang F, Wang W M. 2016. Current understanding on Villosiclava virens, a unique flower- infecting fungus causing rice false smut disease. Mol Plant Pathol, 17(9): 1321-1330. |
| [6] | Fang A F, Han Y Q, Zhang N, Zhang M, Liu L J, Li S, Lu F, Sun W X. 2016. Identification and characterization of plant cell death- inducing secreted proteins from Ustilaginoidea virens. Mol Plant Microbe Interact, 29(5): 405-416. |
| [7] | Flynn P J, Reece R J. 1999. Activation of transcription by metabolic intermediates of the pyrimidine biosynthetic pathway. Mol Cell Biol, 19(1): 882-888. |
| [8] | Hu Z, Zheng L, Huang J B, Zhou L G, Liu C S, Liu H. 2020. Ustiloxin A is produced early in experimental Ustilaginoidea virens infection and affects transcription in rice. Curr Microbiol, 77: 2766-2774. |
| [9] | Jackson J C, Lopes J M. 1996. The yeast UME6 gene is required for both negative and positive transcriptional regulation of phospholipid biosynthetic gene expression. Nucleic Acids Res, 24(7): 1322-1329. |
| [10] | Jones D A B, John E, Rybak K, Phan H T T, Singh K B, Lin S Y, Solomon P S, Oliver R P, Tan K C. 2019. A specific fungal transcription factor controls effector gene expression and orchestrates the establishment of the necrotrophic pathogen lifestyle on wheat. Sci Rep, 9(1): 15884. |
| [11] | Jorgensen T R, Burggraaf A M, Arentshorst M, Schutze T, Lamers G, Niu J, Kwon M J, Park J, Frisvad J C, Nielsen K F, Meyer V, van den Hondel C A M J J, Dyer P S, Ram A F J. 2020. Identification of SclB, a Zn(II)2Cys6 transcription factor involved in sclerotium formation in Aspergillus niger. Fungal Genet Biol, 139: 103377. |
| [12] | Kange A M, Xia A, Si J R, Li B X, Zhang X, Ai G, He F, Dou D L. 2019. The fungal-specific transcription factor VpFSTF1 is required for virulence in Valsa pyri. Front Microbiol, 10: 2945. |
| [13] | Li Y J, Wang M, Liu Z H, Zhang K, Cui F H, Sun W X. 2019. Towards understanding the biosynthetic pathway for ustilaginoidin mycotoxins in Ustilaginoidea virens. Environ Microbiol, 21(8): 2629-2643. |
| [14] | Lu J P, Cao H J, Zhang L L, Huang P Y, Lin F C. 2014. Systematic analysis of Zn2Cys6 transcription factors required for development and pathogenicity by high-throughput gene knockout in the rice blast fungus. PLoS Pathog, 10(10): e1004432. |
| [15] | Luo X M, Mao H Q, Wei Y M, Cai J, Xie C J, Sui A P, Yang X Y, Dong J Y. 2016. The fungal-specific transcription factor Vdpf influences conidia production, melanized microsclerotia formation and pathogenicity in Verticillium dahliae. Mol Plant Pathol, 17(9): 1364-1381. |
| [16] | Lü B, Zheng L, Liu H, Tang J T, Hsiang T, Huang J B. 2016. Use of random T-DNA mutagenesis in identification of gene UvPRO1, a regulator of conidiation, stress response, and virulence in Ustilaginoidea virens. Front Microbiol, 7: 2086. |
| [17] | MacPherson S, Larochelle M, Turcotte B. 2006. A fungal family of transcriptional regulators: The zinc cluster proteins. Microbiol Mol Biol Rev, 70(3): 583-604. |
| [18] | Mullins E D, Chen X, Romaine P, Raina R, Geiser D M, Kang S. 2001. Agrobacterium-mediated transformation of fusarium oxysporum: An efficient tool for insertional mutagenesis and gene transfer. Phytopathology, 91(2): 173-180. |
| [19] | Ness F, Bourot S, Regnacq M, Spagnoli R, Berges T, Karst F. 2001. SUT1 is a putative Zn[II]2Cys6-transcription factor whose upregulation enhances both sterol uptake and synthesis in aerobically growing Saccharomyces cerevisiae cells. Eur J Biochem, 268(6): 1585-1595. |
| [20] | Nishimura H, Kawasaki Y, Kaneko Y, Nosaka K, Iwashima A. 1992. Cloning and characteristics of a positive regulatory gene, THI2 (PHO6), of thiamin biosynthesis in Saccharomyces cerevisiae. FEBS Lett, 297: 155-158. |
| [21] | Qiu J H, Meng S, Deng Y Z, Huang S W, Kou Y J. 2019. Ustilaginoidea virens: A fungus infects rice flower and threats world rice production. Rice Sci, 26(4): 199-206. |
| [22] | Rybak K, See P T, Phan H T T, Syme R A, Moffat C S, Oliver R P, Tan K C. 2017. A functionally conserved Zn2Cys6 binuclear cluster transcription factor class regulates necrotrophic effector gene expression and host-specific virulence of two major Pleosporales fungal pathogens of wheat. Mol Plant Pathol, 18(3): 420-434. |
| [23] | Sengupta P, Cochran B H. 1991. MATα1 can mediate gene activation by α-mating factor. Genes Dev, 5(10): 1924-1934. |
| [24] | Shelest E. 2008. Transcription factors in fungi. FEMS Microbiol Lett, 286(2): 145-151. |
| [25] | Son Y E, Cho H J, Lee M K, Park H S. 2020. Characterizing the role of Zn cluster family transcription factor ZcfA in governing development in two Aspergillus species. PLoS One, 15(2): e0228643. |
| [26] | Song T Q, Zhang Y, Zhang Q, Zhang X, Shen D Y, Yu J J, Yu M N, Pan X Y, Cao H J, Yong M L, Qi Z Q, Du Y, Zhang R S, Yin X L, Qiao J Q, Liu Y Z, Liu W D, Sun W X, Zhang Z G, Wang Y C, Dou D L, Ma Z C, Liu Y F. 2021. The N-terminus of an Ustilaginoidea virens Ser-Thr-rich glycosylphosphatidylinositol- anchored protein elicits plant immunity as a MAMP. Nat Commun, 12(1): 2451. |
| [27] | Turcotte B, Liang X B, Robert F, Soontorngun N. 2010. Transcriptional regulation of nonfermentable carbon utilization in budding yeast. FEMS Yeast Res, 10(1): 2-13. |
| [28] | Urban M, Cuzick A, Rutherford K, Irvine A, Pedro H, Pant R, Sadanadan V, Khamari L, Billal S, Mohanty S, Hammond- Kosack K E. 2017. PHI-base: A new interface and further additions for the multi-species pathogen-host interactions database. Nucleic Acids Res, 45(1): 604-610. |
| [29] | Vallee B L, Coleman J E, Auld D S. 1991. Zinc fingers, zinc clusters, and zinc twists in DNA-binding protein domains. Proc Natl Acad Sci USA, 88(3): 999-1003. |
| [30] | Verma S, Gazara R K, Verma P K. 2017. Transcription factor repertoire of necrotrophic fungal phytopathogen Ascochyta rabiei: Predominance of MYB transcription factors as potential regulators of secretome. Front Plant Sci, 8: 1037. |
| [31] | Vik A, Rine J. 2001. Upc2p and Ecm22p, dual regulators of sterol biosynthesis in Saccharomyces cerevisiae. Mol Cell Biol, 21(19): 6395-6405. |
| [32] | Wang Y, Li J, Xiang S, Zhou J, Peng X, Hai Y, Wang Y, Li S, Wei S. 2020. A putative effector UvHrip1 inhibits BAX-triggered cell death in Nicotiana benthamiana, and infection of Ustilaginoidea virens suppresses defense-related genes expression. PeerJ, 8: e9354. |
| [33] | Wang Z Q, Meng F Z, Zhang M M, Yin L F, Yin W X, Lin Y, Hsiang T, Peng Y L, Wang Z H, Luo C X. 2018. A putative Zn2Cys6 transcription factor is associated with isoprothiolane resistance in Magnaporthe oryzae. Front Microbiol, 9: 2608. |
| [34] | Yin W X, Cui P, Wei W, Lin Y, Luo C X. 2017. Genome-wide identification and analysis of the basic leucine zipper (bZIP) transcription factor gene family in Ustilaginoidea virens. Genome, 60(12): 1051-1059. |
| [35] | Yu M N, Yu J J, Hu J K, Huang L, Wang Y H, Yin X L, Nie Y F, Meng X K, Wang W D, Liu Y F. 2015. Identification of pathogenicity-related genes in the rice pathogen Ustilaginoidea virens through random insertional mutagenesis. Fungal Genet Biol, 76: 10-19. |
| [36] | Yu M N, Yu J J, Cao H J, Yong M L, Liu Y F. 2019. Genome-wide identification and analysis of the GATA transcription factor gene family in Ustilaginoidea virens. Genome, 62(12): 807-816. |
| [37] | Yu J J, Yu M N, Song T Q, Cao H J, Pan X Y, Yong M L, Qi Z Q, Du Y, Zhang R S, Yin X L, Liu Y F. 2019. A homeobox transcription factor UvHOX2 regulates chlamydospore formation, conidiogenesis, and pathogenicity in Ustilaginoidea virens. Front Microbiol, 10: 1071. |
| [38] | Zhang Y, Zhang K, Fang A F, Han Y Q, Yang J, Xue M F, Bao J D, Hu D W, Zhou B, Sun X Y, Li S J, Wen M, Yao N, Ma L J, Liu Y F, Zhang M, Huang F, Luo C X, Zhou L G, Li J Q, Chen Z Y, Miao J K, Wang S, Lai J S, Xu J R, Hsiang T, Peng Y L, Sun W X. 2014. Specific adaptation of Ustilaginoidea virens in occupying host florets revealed by comparative and functional genomics. Nat Commun, 5: 3849. |
| [39] | Zhao Y, Su H, Zhou J, Feng H H, Zhang K Q, Yang J K. 2015. The APSES family proteins in fungi: Characterizations, evolution and functions. Fungal Genet Biol, 81: 271-280. |
| [1] | Wei Qinghui, Cui Daizong, Zheng Baojiang, Zhao Min. In Vitro Antifungal Activity of Dihydrochelerythrine and Proteomic Analysis in Ustilaginoidea virens [J]. Rice Science, 2023, 30(3): 257-266. |
| [2] | Liu Yueran, Qu Jinsong, Wang Yufu, Yin Weixiao, Luo Chaoxi. bZIP Transcription Factor UvATF21 Mediates Vegetative Growth, Conidiation, Stress Tolerance and Is Required for Full Virulence of Rice False Smut Fungus Ustilaginoidea virens [J]. Rice Science, 2023, 30(1): 50-57. |
| [3] | Meng Shuai, Qiu Jiehua, Xiong Meng, Liu Zhiquan, Jane Sadhna Jagernath, Lin Fucheng, Shi Huanbin, Kou Yanjun. UvWhi2 Is Required for Stress Response and Pathogenicity in Ustilaginoidea virens [J]. Rice Science, 2022, 29(1): 47-54. |
| [4] | Junjie Yu, Mina Yu, Tianqiao Song, Huijuan Cao, Mingli Yong, Xiayan Pan, Zhongqiang Qi, Yan Du, Rongsheng Zhang, Xiaole Yin, Dong Liang, Yongfeng Liu. UvSMEK1, a Suppressor of MEK Null, Regulates Pathogenicity, Conidiation and Conidial Germination in Rice False Smut Fungus Ustilaginoidea virens [J]. Rice Science, 2021, 28(5): 457-465. |
| [5] | Meng Xiong, Shuai Meng, Jiehua Qiu, Huanbin Shi, Xiangling Shen, Yanjun Kou. Putative Phosphatase UvPsr1 Is Required for Mycelial Growth, Conidiation, Stress Response and Pathogenicity in Ustilaginonidea virens [J]. Rice Science, 2020, 27(6): 529-536. |
| [6] | Fei Shang, Wenbin Mou, Hao Wu, Furong Xu, Chunyan Xiang, Jianfei Wang. New Allele of HL6 Regulates Trichome Elongation in Rice [J]. Rice Science, 2020, 27(6): 480-492. |
| [7] | Yongqi He, Jia Zhao, Defeng Feng, Zhibo Huang, Jiaming Liang, Yufei Zheng, Jinping Cheng, Jifeng Ying, Zhoufei Wang. RNA-Seq Study Reveals AP2-Domain-Containing Signalling Regulators Involved in Initial Imbibition of Seed Germination in Rice [J]. Rice Science, 2020, 27(4): 302-314. |
| [8] | Jiehua Qiu, Shuai Meng, Yizhen Deng, Shiwen Huang, Yanjun Kou. Ustilaginoidea virens: A Fungus Infects Rice Flower and Threats World Rice Production [J]. Rice Science, 2019, 26(4): 199-206. |
| [9] | Bo Wang, Zhaohui Zhong, Huanhuan Zhang, Xia Wang, Binglin Liu, Lijia Yang, Xiangyan Han, Deshui Yu, Xuelian Zheng, Chunguo Wang, Wenqin Song, Chengbin Chen, Yong Zhang. Targeted Mutagenesis of NAC Transcription Factor Gene, OsNAC041, Leading to Salt Sensitivity in Rice [J]. Rice Science, 2019, 26(2): 98-108. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||