
Rice Science ›› 2021, Vol. 28 ›› Issue (5): 511-520.DOI: 10.1016/j.rsci.2021.07.011
• Research Paper • Previous Articles
Qina Huang1, Yinliang Wu2, Guosheng Shao1( )
)
Received:2020-08-02
Accepted:2020-11-17
Online:2021-09-28
Published:2021-09-28
Qina Huang, Yinliang Wu, Guosheng Shao. Root Aeration Promotes Cadmium Accumulation in Rice by Regulating Iron Uptake-Associated System[J]. Rice Science, 2021, 28(5): 511-520.
Add to citation manager EndNote|Ris|BibTeX
| Genotype | Cd (µmol/L) | Aeration | Plant height (cm) | Root length (cm) | Shoot dry weight (g) | Root dry weight (g) | Root vigor (mg/g) |
| EJN1 | 0 | Non | 64.70 ± 2.74 ab | 27.64 ± 0.96 bc | 7.43 ± 1.49 a | 0.89 ± 0.16 b | 384.08 ± 43.07 b |
| O2 | 67.66 ± 1.29 a | 31.98 ± 3.42 a | 7.96 ± 0.65 a | 1.16 ± 0.18 a | 505.69 ± 37.38 a | ||
| 1.0 | Non | 62.86 ± 3.64 bc | 22.34 ± 1.54 d | 6.64 ± 0.30 a | 0.86 ± 0.02 bc | 318.64 ± 14.27 c | |
| O2 | 55.64 ± 2.16 d | 28.58 ± 4.01 b | 4.51 ± 0.37 b | 0.68 ± 0.12 cd | 384.39 ± 23.91 b | ||
| FP36 | 0 | Non | 59.64 ± 2.57 c | 21.60 ± 0.54 d | 4.75 ± 0.94 b | 0.74 ± 0.11 bcd | 179.41 ± 1.04 e |
| O2 | 59.80 ± 3.58 c | 28.12 ± 3.04 b | 4.23 ± 0.23 bc | 0.69 ± 0.03 cd | 302.02 ± 19.81 c | ||
| 1.0 | Non | 47.90 ± 3.45 e | 21.50 ± 0.72 d | 3.88 ± 0.35 bc | 0.63 ± 0.09 d | 123.55 ± 8.66 f | |
| O2 | 43.40 ± 1.45 f | 24.60 ± 2.44 cd | 3.13 ± 0.25 c | 0.63 ± 0.06 d | 246.54 ± 27.85 d |
Table 1 Agronomic characteristics of Erjiunan 1 (EJN1) and Fupin 36 (FP36) at 20 d after two Cd treatments under aeration (O2) and nonaeration (Non) conditions.
| Genotype | Cd (µmol/L) | Aeration | Plant height (cm) | Root length (cm) | Shoot dry weight (g) | Root dry weight (g) | Root vigor (mg/g) |
| EJN1 | 0 | Non | 64.70 ± 2.74 ab | 27.64 ± 0.96 bc | 7.43 ± 1.49 a | 0.89 ± 0.16 b | 384.08 ± 43.07 b |
| O2 | 67.66 ± 1.29 a | 31.98 ± 3.42 a | 7.96 ± 0.65 a | 1.16 ± 0.18 a | 505.69 ± 37.38 a | ||
| 1.0 | Non | 62.86 ± 3.64 bc | 22.34 ± 1.54 d | 6.64 ± 0.30 a | 0.86 ± 0.02 bc | 318.64 ± 14.27 c | |
| O2 | 55.64 ± 2.16 d | 28.58 ± 4.01 b | 4.51 ± 0.37 b | 0.68 ± 0.12 cd | 384.39 ± 23.91 b | ||
| FP36 | 0 | Non | 59.64 ± 2.57 c | 21.60 ± 0.54 d | 4.75 ± 0.94 b | 0.74 ± 0.11 bcd | 179.41 ± 1.04 e |
| O2 | 59.80 ± 3.58 c | 28.12 ± 3.04 b | 4.23 ± 0.23 bc | 0.69 ± 0.03 cd | 302.02 ± 19.81 c | ||
| 1.0 | Non | 47.90 ± 3.45 e | 21.50 ± 0.72 d | 3.88 ± 0.35 bc | 0.63 ± 0.09 d | 123.55 ± 8.66 f | |
| O2 | 43.40 ± 1.45 f | 24.60 ± 2.44 cd | 3.13 ± 0.25 c | 0.63 ± 0.06 d | 246.54 ± 27.85 d |
| Genotype | Cd (µmol/L) | Aeration | Plant height (cm) | Root length (cm) | Shoot dry weight (g) | Root dry weight (g) | Root vigor [mg/(g·h)] |
|---|---|---|---|---|---|---|---|
| EJN1 | 0 | Non | 54.67 ± 2.52 b | 11.03 ± 0.97 d | 4.18 ± 0.36 a | 0.44 ± 0.04 cd | 315.73 ± 20.70 cd |
| O2 | 58.58 ± 1.28 a | 15.38 ± 1.62 b | 3.47 ± 0.35 b | 0.58 ± 0.02 b | 513.25 ± 36.27 a | ||
| 1.0 | Non | 52.28 ± 2.21 c | 10.77 ± 1.53 d | 3.86 ± 0.10 ab | 0.48 ± 0.04 c | 271.24 ± 15.50 e | |
| O2 | 50.23 ± 1.30 | 12.15 ± 1.13 cd | 3.95 ± 0.34 a | 0.65 ± 0.06 a | 407.12 ± 7.44 b | ||
| FP36 | 0 | Non | 47.12 ± 3.09 d | 16.48 ± 0.76 b | 2.88 ± 0.29 c | 0.45 ± 0.03 cd | 155.62 ± 14.50 f |
| O2 | 51.93 ± 0.90 c | 18.13 ± 1.12 a | 2.08 ± 0.26 d | 0.39 ± 0.04 de | 323.80 ± 15.45 c | ||
| 1.0 | Non | 44.48 ± 1.53 e | 13.48 ± 1.46 c | 2.74 ± 0.22 c | 0.37 ± 0.04 e | 124.95 ± 18.85 f | |
| O2 | 39.17 ± 1.28 f | 16.87 ± 0.98 ab | 2.55 ± 0.19 c | 0.36 ± 0.04 e | 281.82 ± 18.85 de |
Table S1 Agronomic characteristics of Erjiunan 1 (EJN1) and Fupin 36 (FP36) after 10 d of different Cd stress and aeration treatments.
| Genotype | Cd (µmol/L) | Aeration | Plant height (cm) | Root length (cm) | Shoot dry weight (g) | Root dry weight (g) | Root vigor [mg/(g·h)] |
|---|---|---|---|---|---|---|---|
| EJN1 | 0 | Non | 54.67 ± 2.52 b | 11.03 ± 0.97 d | 4.18 ± 0.36 a | 0.44 ± 0.04 cd | 315.73 ± 20.70 cd |
| O2 | 58.58 ± 1.28 a | 15.38 ± 1.62 b | 3.47 ± 0.35 b | 0.58 ± 0.02 b | 513.25 ± 36.27 a | ||
| 1.0 | Non | 52.28 ± 2.21 c | 10.77 ± 1.53 d | 3.86 ± 0.10 ab | 0.48 ± 0.04 c | 271.24 ± 15.50 e | |
| O2 | 50.23 ± 1.30 | 12.15 ± 1.13 cd | 3.95 ± 0.34 a | 0.65 ± 0.06 a | 407.12 ± 7.44 b | ||
| FP36 | 0 | Non | 47.12 ± 3.09 d | 16.48 ± 0.76 b | 2.88 ± 0.29 c | 0.45 ± 0.03 cd | 155.62 ± 14.50 f |
| O2 | 51.93 ± 0.90 c | 18.13 ± 1.12 a | 2.08 ± 0.26 d | 0.39 ± 0.04 de | 323.80 ± 15.45 c | ||
| 1.0 | Non | 44.48 ± 1.53 e | 13.48 ± 1.46 c | 2.74 ± 0.22 c | 0.37 ± 0.04 e | 124.95 ± 18.85 f | |
| O2 | 39.17 ± 1.28 f | 16.87 ± 0.98 ab | 2.55 ± 0.19 c | 0.36 ± 0.04 e | 281.82 ± 18.85 de |
| Genotype | Cd (µmol/L) | Aeration | 10 d | 20 d | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Chlorophyll a | Chlorophyll b | Carotenoid | Chl a/b | Chlorophyll a | Chlorophyll b | Carotenoid | Chl a/b | |||
| EJN1 | 0 | Non | 2.44 ± 0.58 b | 0.68 ± 0.17 c | 0.50 ± 0.11 b | 3.58 ± 0.03 a | 3.35 ± 0.36 abc | 1.15 ± 0.11 a | 0.60 ± 0.16 ab | 2.95 ± 0.59 a |
| O2 | 2.93 ± 0.21 b | 0.83 ± 0.06 c | 0.59 ± 0.04 b | 3.51 ± 0.01 a | 3.03 ± 0.22 d | 1.05 ± 0.35 a | 0.53 ± 0.15 b | 3.06 ± 0.87 a | ||
| 1.0 | Non | 2.48 ± 0.91 b | 0.71 ± 0.25 c | 0.50 ± 0.17 b | 3.51 ± 0.05 a | 3.64 ± 0.25 a | 1.09 ± 0.08 a | 0.71 ± 0.05 a | 3.34 ± 0.10 a | |
| O2 | 2.57 ± 0.26 b | 0.73 ± 0.05 c | 0.52 ± 0.06 b | 3.52 ± 0.11 a | 3.59 ± 0.35 ab | 1.07 ± 0.19 a | 0.70 ± 0.02 a | 3.38 ± 0.26 a | ||
| FP36 | 0 | Non | 4.02 ± 0.42 a | 1.22 ± 0.17 b | 0.80 ± 0.06 a | 3.31 ± 0.10 b | 3.34 ± 0.21 abcd | 0.95 ± 0.07 a | 0.72 ± 0.03 a | 3.51 ± 0.04 a |
| O2 | 4.05 ± 0.13 a | 1.20 ± 0.05 b | 0.82 ± 0.03 a | 3.39 ± 0.03 b | 3.33 ± 0.28 bcd | 0.98 ± 0.11 a | 0.69 ± 0.03 a | 3.41 ± 0.09 a | ||
| 1.0 | Non | 4.01 ± 0.66 a | 1.23 ± 0.25 b | 0.80 ± 0.11 a | 3.27 ± 0.12 b | 3.19 ± 0.30 cd | 0.92 ± 0.10 a | 0.67 ± 0.04 a | 3.46 ± 0.05 a | |
| O2 | 4.81 ± 0.06 a | 1.57 ± 0.02 a | 0.93 ± 0.03 a | 3.07 ± 0.01 c | 3.04 ± 0.10 cd | 0.87 ± 0.04 a | 0.65 ± 0.02 ab | 3.52 ± 0.06 a | ||
Table S2 Contents of chlorophyll a, chlorophyll b, carotenoid and chlorophyll a/b of Erjiunan 1 (EJN1) and Fupin 36 (FP36) after 10 d and 20 d of different treatments.
| Genotype | Cd (µmol/L) | Aeration | 10 d | 20 d | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Chlorophyll a | Chlorophyll b | Carotenoid | Chl a/b | Chlorophyll a | Chlorophyll b | Carotenoid | Chl a/b | |||
| EJN1 | 0 | Non | 2.44 ± 0.58 b | 0.68 ± 0.17 c | 0.50 ± 0.11 b | 3.58 ± 0.03 a | 3.35 ± 0.36 abc | 1.15 ± 0.11 a | 0.60 ± 0.16 ab | 2.95 ± 0.59 a |
| O2 | 2.93 ± 0.21 b | 0.83 ± 0.06 c | 0.59 ± 0.04 b | 3.51 ± 0.01 a | 3.03 ± 0.22 d | 1.05 ± 0.35 a | 0.53 ± 0.15 b | 3.06 ± 0.87 a | ||
| 1.0 | Non | 2.48 ± 0.91 b | 0.71 ± 0.25 c | 0.50 ± 0.17 b | 3.51 ± 0.05 a | 3.64 ± 0.25 a | 1.09 ± 0.08 a | 0.71 ± 0.05 a | 3.34 ± 0.10 a | |
| O2 | 2.57 ± 0.26 b | 0.73 ± 0.05 c | 0.52 ± 0.06 b | 3.52 ± 0.11 a | 3.59 ± 0.35 ab | 1.07 ± 0.19 a | 0.70 ± 0.02 a | 3.38 ± 0.26 a | ||
| FP36 | 0 | Non | 4.02 ± 0.42 a | 1.22 ± 0.17 b | 0.80 ± 0.06 a | 3.31 ± 0.10 b | 3.34 ± 0.21 abcd | 0.95 ± 0.07 a | 0.72 ± 0.03 a | 3.51 ± 0.04 a |
| O2 | 4.05 ± 0.13 a | 1.20 ± 0.05 b | 0.82 ± 0.03 a | 3.39 ± 0.03 b | 3.33 ± 0.28 bcd | 0.98 ± 0.11 a | 0.69 ± 0.03 a | 3.41 ± 0.09 a | ||
| 1.0 | Non | 4.01 ± 0.66 a | 1.23 ± 0.25 b | 0.80 ± 0.11 a | 3.27 ± 0.12 b | 3.19 ± 0.30 cd | 0.92 ± 0.10 a | 0.67 ± 0.04 a | 3.46 ± 0.05 a | |
| O2 | 4.81 ± 0.06 a | 1.57 ± 0.02 a | 0.93 ± 0.03 a | 3.07 ± 0.01 c | 3.04 ± 0.10 cd | 0.87 ± 0.04 a | 0.65 ± 0.02 ab | 3.52 ± 0.06 a | ||
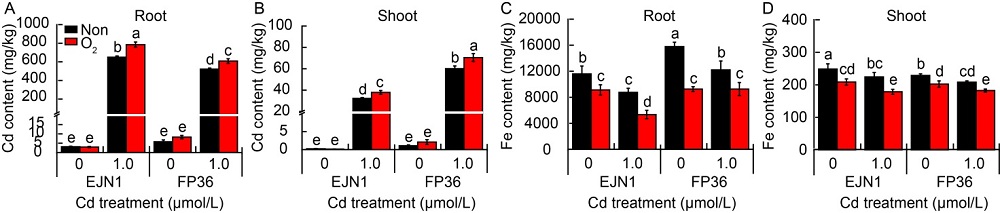
Fig. 1. Cd and Fe contents in roots and shoots of Erjiunan 1 (EJN1) and Fupin 36 (FP36) at 20 d after two Cd treatments (0, 1.0 µmol/L) under aeration (O2) and nonaeration (Non) conditions.Data are Mean ± SE (n = 3). Different lowercase letters above the error bars indicate significant difference (P < 0.05) among different treatments.
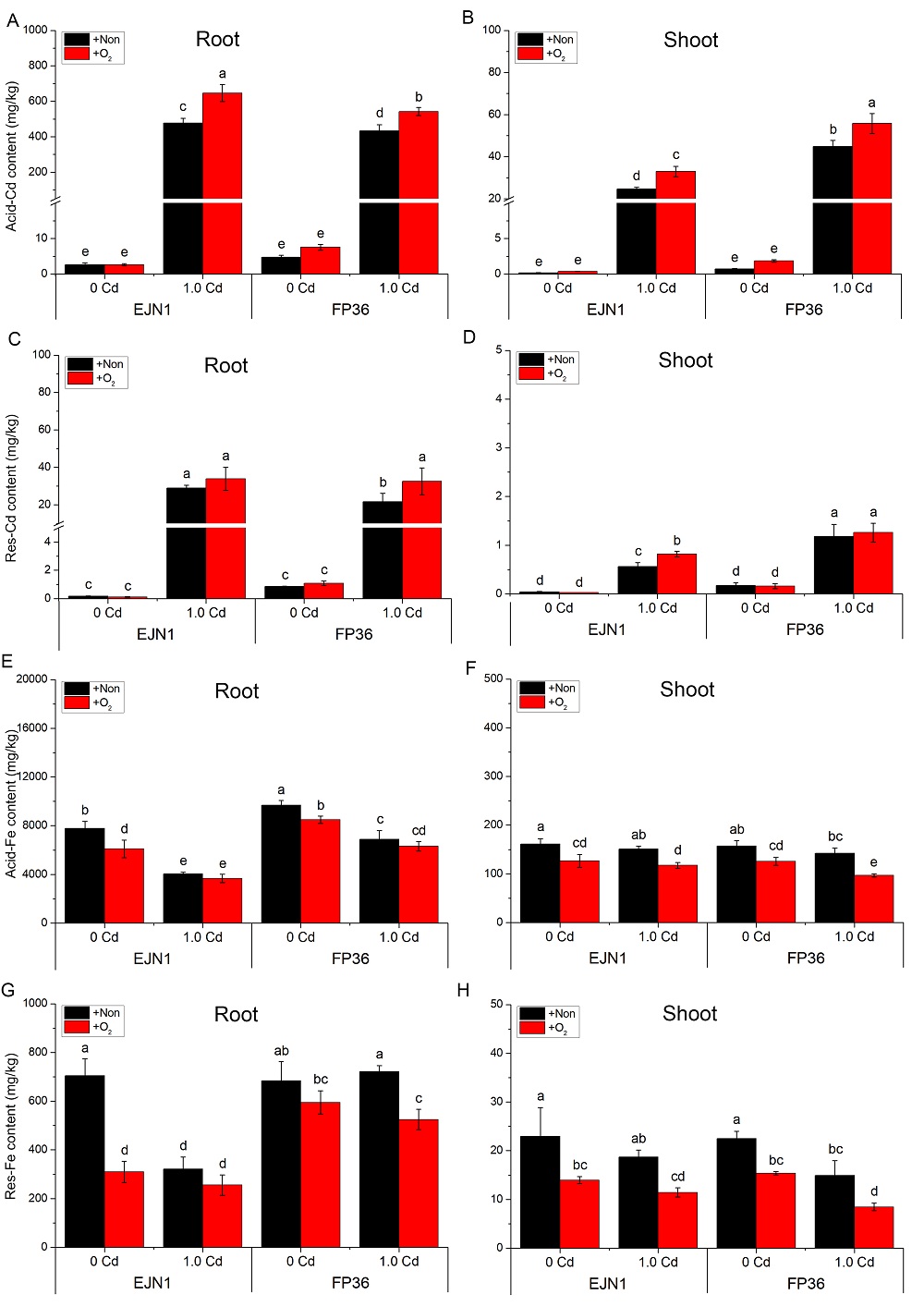
Fig. S2. Acid-/Res-Cd (A-D) and Acid-/Res-Fe (E-H) contents in Erjiunan 1 (EJN1) and Fupin 36 (FP36).With two Cd treatments (0 and 1.0 µmol/L), and aeration (O2) and nonaeration (Non) conditions. Data were Mean ± SE (n = 3). Different letters above the error bars indicated significant difference (P < 0.05) among different treatments.
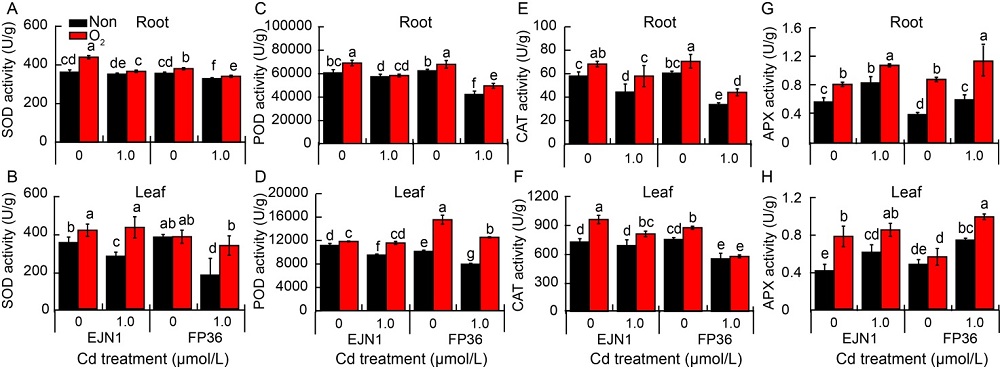
Fig. 2. Activities of antioxidant enzymes in roots and leaves of Erjiunan 1 (EJN1) and Fupin 36 (FP36) at 20 d after two Cd treatments (0, 1.0 µmol/L) under aeration (O2) and nonaeration (Non) conditions. SOD, Superoxide dismutase; POD, Peroxidase; CAT, Catalase; APX, Ascorbate peroxidase.Data are Mean ± SE (n = 3). Different lowercase letters above the error bars indicate significant difference (P < 0.05) among different treatments.
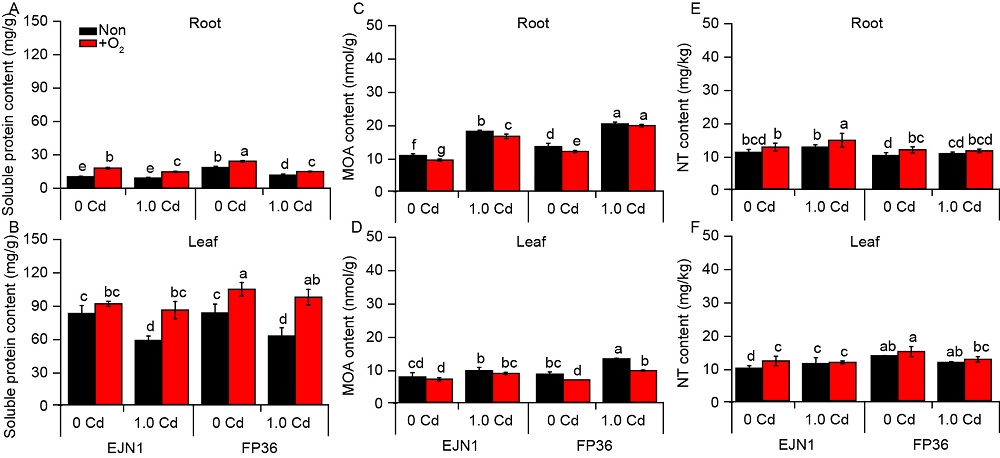
Fig. S3. Contents of soluble protein, malonaldehyde (MOA) and nicotianamine (NT) in roots (A, C and E) and in leaves(B, D and F).With two Cd treatments (0 and 1.0 µmol/L), and aeration (O2) and nonaeration (Non) conditions. Data were Mean ± SE (n = 3). Different letters above the error bars indicated significant difference (P < 0.05) among different treatments.
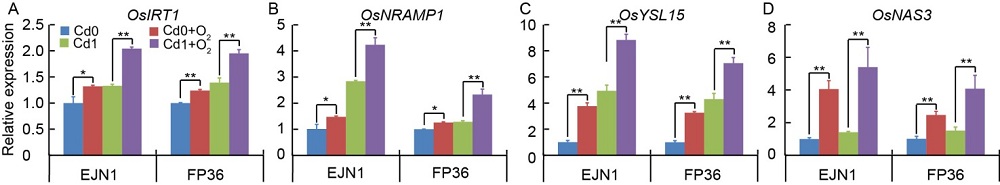
Fig. 3. Expression of Fe-inducible genes (with the housekeeping gene OsActin as the internal control) in roots of Erjiunan 1 (EJN1) and Fupin 36 (FP36) at 20 d after two Cd treatments (Cd0, 0 µmol/L; Cd1, 1.0 µmol/L) under aeration (O2) and nonaeration conditions.The relative expression values were calculated using the 2-ΔΔCt method. Data are Mean ± SE (n = 3). *, P < 0.05; **, P < 0.01.
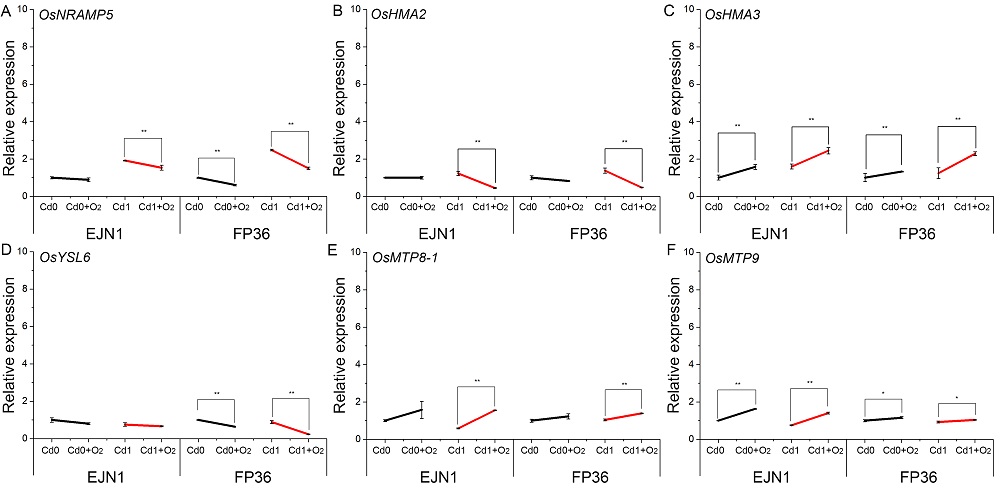
Fig. S4. Expression of genes in the roots of Erjiunan 1 (EJN1) and Fupin 36 (FP36).With two Cd treatments (Cd0, 0 µmol/L; Cd1, 1.0 µmol/L), and aeration (O2) and nonaeration conditions. Data were Mean ± SE (n = 3). *, Significance at P < 0.05 ; **, Highly significance at P < 0.01.
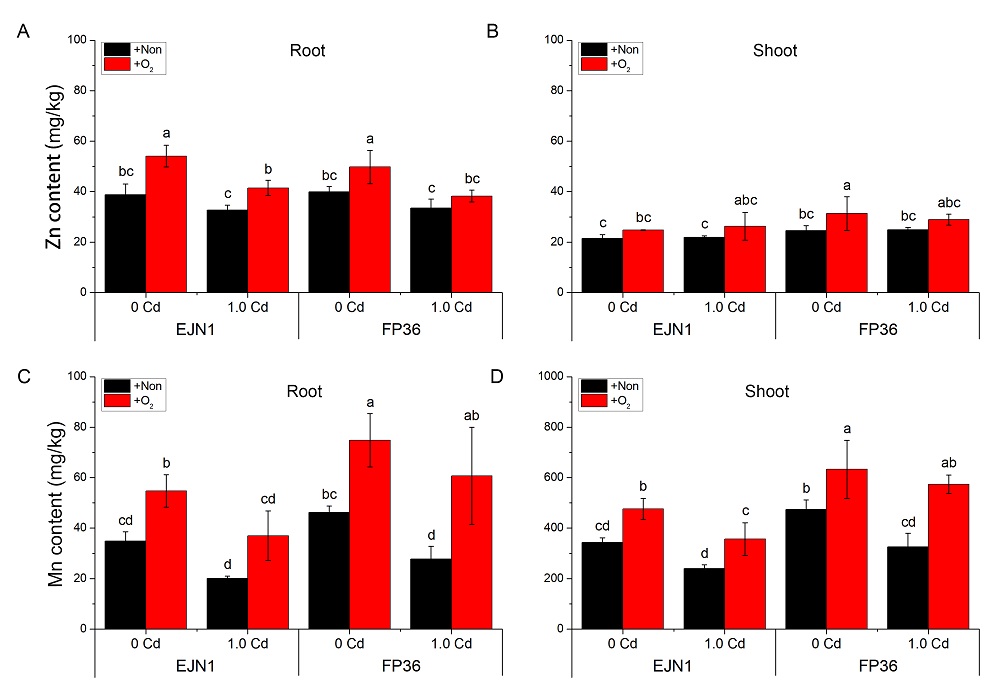
Fig. S5. Contents of Zn and Mn in the roots and shoots of Erjiunan 1 (EJN1) and Fupin 36 (FP36).With two Cd treatments (0 and 1.0 µmol/L, supplied by CdSO4), and aeration (O2) and nonaeration (Non) conditions. Data were Mean ±SE (n = 3). Different letters above the error bars indicated significant difference (P < 0.05) among different treatments.
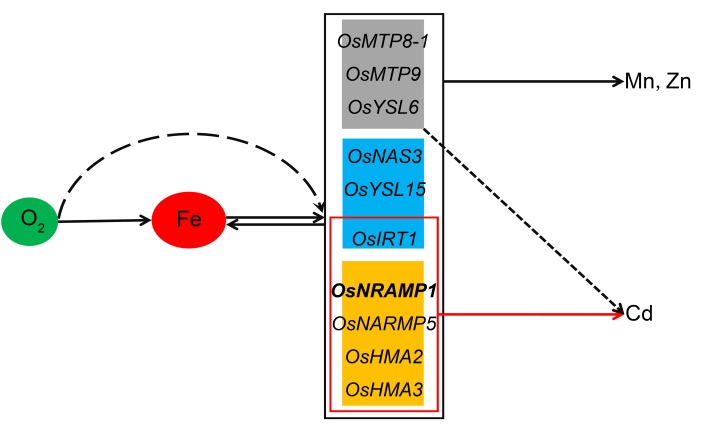
Fig. 4. Model outlining of aeration promotes Cd accumulation due to Fe reduction in rice.The yellow box represents the Cd-absorb/transport genes, the blue box represents the Fe-uptake/transport genes, and the grey box represents the Mn/Zn-transport genes.
| Gene | Former primer (5'-3') | Reverse primer (5'-3') |
|---|---|---|
| OsNRAMP1 | GTGATTGCTTCCGATATTCCA | CAACCTCCAGTTTCCTTACCC |
| OsNRAMP5 | CAGCAGCAGTAAGAGCAAGATGG | GGGAGGTCGTTGTGGATGAGC |
| OsHMA2 | AATCAGGCAAGGTTGGAGGC | CAGCGACACGACCAGAAGAAG |
| OsHMA3 | AGCCTCCAAGAAGCCAACTG | CAACATTTGGCAACGACGG |
| OsIRT1 | CGATGTGCTTCCACCAGATG | GCGTCGTGGTGGAGAAGAAG |
| OsYSL15 | AGCAGGGCTCACGGACATCA | TGCTTCACCAGGGTGCCAAC |
| OsNAS3 | TGGAGCACCTCGGGAAGCAC | GCGCGATGATGACGGAGTTG |
| OsYSL6 | CGATGGCTATGGCTGTCCC | CTCAGATTCCTTGCGGTTCAC |
| OsMTP8-1 | CTGGTAACAAGATAGTCCGTGCC | TGCACCAACTGGGTCAATCC |
| OsMTP9 | GAGGAAGAGTTGAGGCAAATGG | TCCAGTAGGGAGTCCAGTGTCG |
| OsActin | AGGAAGGCTGGAAGAGGACC | CGGGAAATTGTGAGGGACAT |
Table S3 Primer sequences of genes for qRT-PCR.
| Gene | Former primer (5'-3') | Reverse primer (5'-3') |
|---|---|---|
| OsNRAMP1 | GTGATTGCTTCCGATATTCCA | CAACCTCCAGTTTCCTTACCC |
| OsNRAMP5 | CAGCAGCAGTAAGAGCAAGATGG | GGGAGGTCGTTGTGGATGAGC |
| OsHMA2 | AATCAGGCAAGGTTGGAGGC | CAGCGACACGACCAGAAGAAG |
| OsHMA3 | AGCCTCCAAGAAGCCAACTG | CAACATTTGGCAACGACGG |
| OsIRT1 | CGATGTGCTTCCACCAGATG | GCGTCGTGGTGGAGAAGAAG |
| OsYSL15 | AGCAGGGCTCACGGACATCA | TGCTTCACCAGGGTGCCAAC |
| OsNAS3 | TGGAGCACCTCGGGAAGCAC | GCGCGATGATGACGGAGTTG |
| OsYSL6 | CGATGGCTATGGCTGTCCC | CTCAGATTCCTTGCGGTTCAC |
| OsMTP8-1 | CTGGTAACAAGATAGTCCGTGCC | TGCACCAACTGGGTCAATCC |
| OsMTP9 | GAGGAAGAGTTGAGGCAAATGG | TCCAGTAGGGAGTCCAGTGTCG |
| OsActin | AGGAAGGCTGGAAGAGGACC | CGGGAAATTGTGAGGGACAT |
| [1] | Aoyama T, Kobayashi T, Takahashi M, Nagasaka S, Usuda K, Kakei Y, Ishimaru Y, Nakanishi H, Mori S, Nishizawa N K. 2009. OsYSL18 is a rice iron(III)-deoxymugineic acid transporter specifically expressed in reproductive organs and phloem of lamina joints. Plant Mol Biol, 70(6): 681-692. |
| [2] | Bashir K, Nozoye T, Ishimaru Y, Nakanishi H, Nishizawa N K. 2013. Exploiting new tools for iron bio-fortification of rice. Biotechnol Adv, 31(8): 1624-1633. |
| [3] | Chang J D, Huang S, Yamaji N, Zhang W W, Ma J F, Zhao F J. 2020. OsNRAMP1 contributes to cadmium and manganese uptake in rice. Plant Cell Environ, 43(10): 2476-2491. |
| [4] | Chen J M, Yang Y J, Huang Q N, Hu P S, Tang S Q, Wu L Q, Wang J L, Shao G S. 2017. Effects of continuous flooding on cadmium absorption and its regulation mechanisms in rice. Sci Agric Sin, 50(17): 3300-3310. (in Chinese with English abstract) |
| [5] | Clemens S, Ma J F. 2016. Toxic heavy metal and metalloid accumulation in crop plants and foods. Annu Rev Plant Biol, 67: 489-512. |
| [6] | DalCorso G, Farinati S, Maistri S, Furini A. 2008. How plants cope with cadmium: Staking all on metabolism and gene expression. J Integr Plant Biol, 50(10): 1268-1280. |
| [7] | Ding C P, Xu R K. 2011. Oxidation-Reduction Processes of Soils and Their Research Methods. Beijing, China: Beijing Science Press. (in Chinese) |
| [8] | Ding S L, Liu C L, Qian Q, Gao Z Y. 2019. Research advances on molecular genetic mechanism for cadmium absorption and transportation in rice. Chin J Rice Sci, 33(5): 383-390. (in Chinese with English abstract) |
| [9] | Dolferus R, Jacobs M, Peacock W J, Dennis E S. 1994. Differential interactions of promoter elements in stress responses of the Arabidopsis Adh gene. Plant Physiol, 105(4): 1075-1087. |
| [10] | Ekmekci Y, Tanyolac D, Ayhan B. 2008. Effects of cadmium on antioxidant enzyme and photosynthetic activities in leaves of two maize cultivars. J Plant Physiol, 165(6): 600-611. |
| [11] | Ghnaya T, Nouairi I, Slama I, Messedi D, Grignon C, Abdelly C, Ghorbel M H. 2005. Cadmium effects on growth and mineral nutrition of two halophytes: Sesuvium portulacastrum and Mesembryanthemum crystallinum. J Plant Physiol, 162(10): 1133-1140. |
| [12] | Gill S S, Tuteja N. 2010. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem, 48(12): 909-930. |
| [13] | Huang J H, Wang S L, Lin J H, Chen Y M, Wang M K. 2013. Dynamics of cadmium concentration in contaminated rice paddy soils with submerging time. Paddy Water Environ, 11: 483-491. |
| [14] | Huang L, Li W C, Tam N F Y, Ye Z H. 2019. Effects of root morphology and anatomy on cadmium uptake and translocation in rice (Oryza sativa L.). J Environ Sci, 75: 296-306. |
| [15] | Huq E, Hodges T K. 1999. An anaerobically inducible early (ale) gene family from rice. Plant Mol Biol, 40(4): 591-601. |
| [16] | Inoue H, Higuchi K, Takahashi M, Nakanishi H, Mori S, Nishizawa N K. 2003. Three rice nicotianamine synthase genes, OsNAS1, OsNAS2, and OsNAS3 are expressed in cells involved in long-distance transport of iron and differentially regulated by iron. Plant J, 36(3): 366-381. |
| [17] | Inoue H, Takahashi M, Kobayashi T, Suzuki M, Nakanishi H, Mori S, Nishizawa N K. 2008. Identification and localisation of the rice nicotianamine aminotransferase gene OsNAAT1 expression suggests the site of phytosiderophore synthesis in rice. Plant Mol Biol, 66: 193-203. |
| [18] | Inoue H, Kobayashi T, Nozoye T, Takahashi M, Kakei Y, Suzuki K, Nakazono M, Nakanishi H, Mori S, Nishizawa N K. 2009. Rice OsYSL15 is an iron-regulated iron(III)-deoxymugineic acid transporter expressed in the roots and is essential for iron uptake in early growth of the seedlings. J Biol Chem, 284(6): 3470-3479. |
| [19] | Ishikawa S, Abe T, Kuramata M, Yamaguchi M, Ando T, Yamamoto T, Yano M. 2010. A major quantitative trait locus for increasing cadmium-specific concentration in rice grain is located on the short arm of chromosome 7. J Exp Bot, 61(3): 923-934. |
| [20] | Ishimaru Y, Suzuki M, Tsukamoto T, Suzuki K, Nakazono M, Kobayashi T, Wada Y, Watanabe S, Matsuhashi S, Takahashi M, Nakanishi H, Mori S, Nishizawa N K. 2006. Rice plants take up iron as an Fe3+-phytosiderophore and as Fe2+. Plant J, 45(3): 335-346. |
| [21] | Ishimaru Y, Kim S, Tsukamoto T, Oki H, Kobayashi T, Watanabe S, Matsuhashi S, Takahashi M, Nakanishi H, Mori S, Nishizawa N K. 2007. Mutational reconstructed ferric chelate reductase confers enhanced tolerance in rice to iron deficiency in calcareous soil. Proc Natl Acad Sci USA, 104(18): 7373-7378. |
| [22] | Ishimaru Y, Takahashi R, Bashir K, Shimo H, Senoura T, Sugimoto K, Ono K, Yano M, Ishikawa S, Arao T, Nakanishi H, Nishizawa N K. 2012. Characterizing the role of rice NRAMP5 in manganese, iron and cadmium transport. Sci Rep, 2: 286. |
| [23] | Kakei Y, Ishimaru Y, Kobayashi T, Yamakawa T, Nakanishi H, Nishizawa N K. 2012. OsYSL16 plays a role in the allocation of iron. Plant Mol Biol, 79(6): 583-594. |
| [24] | Kobayashi T, Nakanishi Itai R, Nishizawa N K. 2014. Iron deficiency responses in rice roots. Rice, 7(1): 27. |
| [25] | Koike S, Inoue H, Mizuno D, Takahashi M, Nakanishi H, Mori S, Nishizawa N K. 2004. OsYSL2 is a rice metal-nicotianamine transporter that is regulated by iron and expressed in the phloem. Plant J, 39(3): 415-424. |
| [26] | Kuramata M, Masuya S, Takahashi Y, Kitagawa E, Inoue C, Ishikawa S, Youssefian S, Kusano T. 2009. Novel cysteine-rich peptides from Digitaria ciliaris and Oryza sativa enhance tolerance to cadmium by limiting its cellular accumulation. Plant Cell Physiol, 50(1): 106-117. |
| [27] | Lee S, Chiecko J C, Kim S A, Walker E L, Lee Y, Guerinot M L, An G. 2009a. Disruption of OsYSL15 leads to iron inefficiency in rice plants. Plant Physiol, 150(2): 786-800. |
| [28] | Lee S, Jeon U S, Lee S J, Kim Y K, Persson D P, Husted S, Schjørring J K, Kakei Y, Masuda H, Nishizawa N K, An G. 2009b. Iron fortification of rice seeds through activation of the nicotianamine synthase gene. Proc Natl Acad Sci USA, 106: 22014-22019. |
| [29] | Li H B, Zheng X W, Tao L X, Yang Y J, Gao L, Xiong J. 2019. Aeration increases cadmium (Cd) retention by enhancing iron plaque formation and regulating pectin synthesis in the roots of rice (Oryza sativa) seedlings. Rice, 12(1): 28. |
| [30] | Liu S M, Jiang J, Liu Y, Meng J, Xu S L, Tan Y Y, Li Y F, Shu Q Y, Huang J Z. 2019. Characterization and evaluation of OsLCT1 and OsNramp5 mutants generated through CRISPR/Cas9- mediated mutagenesis for breeding low Cd rice. Rice Sci, 26(2): 88-97. |
| [31] | Livak K J, Schmittgen T D. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods, 25(4): 402-408. |
| [32] | Meharg A A, Norton G, Deacon C, Williams P, Adomako E E, Price A, Zhu Y G, Li G, Zhao F J, McGrath S, Villada A, Sommella A, de Silva P M, Brammer H, Dasgupta T, Islam M R. 2013. Variation in rice cadmium related to human exposure. Environ Sci Technol, 47(11): 5613-5618. |
| [33] | Mittler R, Vanderauwers S, Gollery M, van Breusegem F. 2004. Reactive oxygen gene network of plants. Trends Plant Sci, 9(10): 490-498. |
| [34] | Redjala T, Zelko I, Sterckeman T, Legué V, Lux A. 2011. Relationship between root structure and root cadmium uptake in maize. Environ Exp Bot, 71(2): 241-248. |
| [35] | Romero-Puertas M C, Rodriguez-Serrano M, Corpas F J, Gomez M, Del Rio L A, Sandalio L M. 2004. Cadmium-induced subcellular accumulation of O2·̄ and H2O2 in pea leaves. Plant Cell Environ, 27(9): 1122-1134. |
| [36] | Roth U, von Roepenack-Lahaye E, Clemens S. 2006. Proteome changes in Arabidopsis thaliana roots upon exposure to Cd2+. J Exp Bot, 57(15): 4003-4013. |
| [37] | Sasaki A, Yamaji N, Xia J X, Ma J F. 2011. OsYSL6 is involved in the detoxification of excess manganese in rice. Plant Physiol, 157(4): 1832-1840. |
| [38] | Satoh-Nagasawa N, Mori M, Sakurai K, Takahashi H, Watanabe A, Akagi H. 2013. Functional relationship heavy metal P-type ATPases (OsHMA2 and OsHMA3) of rice (Oryza sativa) using RNAi. Plant Biol, 30(5): 511-515. |
| [39] | Schutzendübel A, Schwanz P, Teichmann T, Gross K, Langenfeld- Heyser R, Godbold D L, Polle A. 2001. Cadmium-induced changes in antioxidative systems, H2O2 content and differentiation in pine (Pinus sylvestris) roots. Plant Physiol, 127(3): 887-898. |
| [40] | Shahid M, Dumat C, Khalid S, Niazi N K, Antunes P M C. 2017. Cadmium bioavailability, uptake, toxicity and detoxification in soil-plant system. Rev Environ Contam Toxicol, 241: 73-137. |
| [41] | Shimo H, Ishimaru Y, An G, Yamakawa T, Nakanishi H, Nishizawa N K. 2011. Low cadmium (LCD), a novel gene related to cadmium tolerance and accumulation in rice. J Exp Bot, 62(15): 5727-5734. |
| [42] | Smith E, Kempson I M, Juhasz A L, Weber J, Rofe A, Gancarz D, Naidu R, McLaren R G, Gräfe M. 2011. In vivo-in vitro and XANES spectroscopy assessments of lead bioavailability in contaminated periurban soils. Environ Sci Technol, 45(14): 6145-6152. |
| [43] | Song W Y, Lee H S, Jin S R, Ko D, Martinoia E, Lee Y, An G, Ahn S N. 2015. Rice PCR1 influences grain weight and Zn accumulation in grains. Plant Cell Environ, 38(11): 2327-2339. |
| [44] | Takahashi R, Ishimaru Y, Nakanishi H, Nishizawa N K. 2011. Role of the iron transporter OsNRAMP1 in cadmium uptake and accumulation in rice. Plant Signal Behav, 6(11): 1813-1816. |
| [45] | Tripathi R D, Tripath P, Dwivedi S, Kumar A, Mishra A, Chauhan P S, Norton G J, Nautiyal C S. 2014. Roles for root iron plaque in sequestration and uptake of heavy metals and metalloids in aquatic and wetland plants. Metallomics, 6(10): 1789-1800. |
| [46] | Uraguchi S, Kamiya T, Sakamoto T, Kasai K, Sato Y, Nagamura Y, Yoshida A, Kyozuka J, Ishikawa S, Fujiwara T. 2011. Low- affinity cation transporter (OsLCT1) regulates cadmium transport into rice grains. Proc Natl Acad Sci USA, 108: 20959-20964. |
| [47] | Vert G, Grotz N, Dédaldéchamp F, Gaymard F, Guerinot M L, Briat J F, Curie C. 2002. IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth. Plant Cell, 14(6): 1223-1233. |
| [48] | Wang J, Peng T, Zhang J, Liu J, Chen H, Du Y, Li J, Sun H, Zhao Q. 2016. Effects of different rhizosphere dissolved oxygen mass concentration on rice root growth and antioxidant enzyme activities at middle and late growth stage. J Henan Agric Univ, 50(6): 720-725. (in Chinese with English abstract) |
| [49] | Xin J P, Tang J Y, Liu Y L, Zhang Y, Tian R N. 2019. Pre-aeration of the rhizosphere offers potential for phytoremediation of heavy metal-contaminated wetlands. J Hazard Mater, 374: 437-446. |
| [50] | Yamaguchi H, Uchida R. 2012. Determination of nicotianamine in soy sauce and other plant-based foods by LC-MS/MS. J Agric Food Chem, 60: 10000-10006. |
| [51] | Yoneyama T, Ishikawa S, Fujimaki S. 2015. Route and regulation of zinc, cadmium, and iron transport in rice plants (Oryza sativa L.) during vegetative growth and grain filling: Metal transporters, metal speciation, grain Cd reduction and Zn and Fe biofortification. Int J Mol Sci, 16(8): 19111-19129. |
| [52] | Zhao S J, Shi G A, Dong X C. 2002. Plant Physiology Experiment Guide. Beijing, China: Chinese Agricultural Science and Technology Press: 49-138. (in Chinese) |
| [1] | Prathap V, Suresh KUMAR, Nand Lal MEENA, Chirag MAHESHWARI, Monika DALAL, Aruna TYAGI. Phosphorus Starvation Tolerance in Rice Through a Combined Physiological, Biochemical and Proteome Analysis [J]. Rice Science, 2023, 30(6): 8-. |
| [2] | Serena REGGI, Elisabetta ONELLI, Alessandra MOSCATELLI, Nadia STROPPA, Matteo Dell’ANNO, Kiril PERFANOV, Luciana ROSSI. Seed-Specific Expression of Apolipoprotein A-IMilano Dimer in Rice Engineered Lines [J]. Rice Science, 2023, 30(6): 6-. |
| [3] | Sundus ZAFAR, XU Jianlong. Recent Advances to Enhance Nutritional Quality of Rice [J]. Rice Science, 2023, 30(6): 4-. |
| [4] | Kankunlanach KHAMPUANG, Nanthana CHAIWONG, Atilla YAZICI, Baris DEMIRER, Ismail CAKMAK, Chanakan PROM-U-THAI. Effect of Sulfur Fertilization on Productivity and Grain Zinc Yield of Rice Grown under Low and Adequate Soil Zinc Applications [J]. Rice Science, 2023, 30(6): 9-. |
| [5] | FAN Fengfeng, CAI Meng, LUO Xiong, LIU Manman, YUAN Huanran, CHENG Mingxing, Ayaz AHMAD, LI Nengwu, LI Shaoqing. Novel QTLs from Wild Rice Oryza longistaminata Confer Rice Strong Tolerance to High Temperature at Seedling Stage [J]. Rice Science, 2023, 30(6): 14-. |
| [6] | LIN Shaodan, YAO Yue, LI Jiayi, LI Xiaobin, MA Jie, WENG Haiyong, CHENG Zuxin, YE Dapeng. Application of UAV-Based Imaging and Deep Learning in Assessment of Rice Blast Resistance [J]. Rice Science, 2023, 30(6): 10-. |
| [7] | Md. Forshed DEWAN, Md. AHIDUZZAMAN, Md. Nahidul ISLAM, Habibul Bari SHOZIB. Potential Benefits of Bioactive Compounds of Traditional Rice Grown in South and South-East Asia: A Review [J]. Rice Science, 2023, 30(6): 5-. |
| [8] | Raja CHAKRABORTY, Pratap KALITA, Saikat SEN. Phenolic Profile, Antioxidant, Antihyperlipidemic and Cardiac Risk Preventive Effect of Chakhao Poireiton (A Pigmented Black Rice) in High-Fat High-Sugar induced Rats [J]. Rice Science, 2023, 30(6): 11-. |
| [9] | LI Qianlong, FENG Qi, WANG Heqin, KANG Yunhai, ZHANG Conghe, DU Ming, ZHANG Yunhu, WANG Hui, CHEN Jinjie, HAN Bin, FANG Yu, WANG Ahong. Genome-Wide Dissection of Quan 9311A Breeding Process and Application Advantages [J]. Rice Science, 2023, 30(6): 7-. |
| [10] | JI Dongling, XIAO Wenhui, SUN Zhiwei, LIU Lijun, GU Junfei, ZHANG Hao, Tom Matthew HARRISON, LIU Ke, WANG Zhiqin, WANG Weilu, YANG Jianchang. Translocation and Distribution of Carbon-Nitrogen in Relation to Rice Yield and Grain Quality as Affected by High Temperature at Early Panicle Initiation Stage [J]. Rice Science, 2023, 30(6): 12-. |
| [11] | Nazaratul Ashifa Abdullah Salim, Norlida Mat Daud, Julieta Griboff, Abdul Rahim Harun. Elemental Assessments in Paddy Soil for Geographical Traceability of Rice from Peninsular Malaysia [J]. Rice Science, 2023, 30(5): 486-498. |
| [12] | Monica Ruffini Castiglione, Stefania Bottega, Carlo Sorce, Carmelina SpanÒ. Effects of Zinc Oxide Particles with Different Sizes on Root Development in Oryza sativa [J]. Rice Science, 2023, 30(5): 449-458. |
| [13] | Tan Jingyi, Zhang Xiaobo, Shang Huihui, Li Panpan, Wang Zhonghao, Liao Xinwei, Xu Xia, Yang Shihua, Gong Junyi, Wu Jianli. ORYZA SATIVA SPOTTED-LEAF 41 (OsSPL41) Negatively Regulates Plant Immunity in Rice [J]. Rice Science, 2023, 30(5): 426-436. |
| [14] | Ammara Latif, Sun Ying, Pu Cuixia, Noman Ali. Rice Curled Its Leaves Either Adaxially or Abaxially to Combat Drought Stress [J]. Rice Science, 2023, 30(5): 405-416. |
| [15] | Liu Qiao, Qiu Linlin, Hua Yangguang, Li Jing, Pang Bo, Zhai Yufeng, Wang Dekai. LHD3 Encoding a J-Domain Protein Controls Heading Date in Rice [J]. Rice Science, 2023, 30(5): 437-448. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||